Chapter 21 - Proton Motive Force
1/28
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
29 Terms
Where is [H+] high in the mitochondria? Where is [H+] low?
[H+] is high in the IM space, [H+] is low in the matrix
![<p>[H+] is high in the IM space, [H+] is low in the matrix</p>](https://assets.knowt.com/user-attachments/1a07c0e3-3d07-47a4-bdf9-0306aab4f5bd.png)
What are the two components of the proton-motive force?
A chemical gradient high [H+] → low [H+] and a charge gradient + → —
What links the ETC to ATP synthesis?
The proton-motive force
Does inhibition of ATP synthase by oligomycin block the ETC?
Yes because the [H+] gradient will build up and slow the pumping of protons by the ETC
Does inhibition of the ETC by cyanide block ATP synthase?
Yes because no protons will get pumped and ATP synthase has no gradient to use
What is nonshivering thermogenesis?
The ETC is uncoupled from ATP synthesis, generating heat; this is regulated by uncoupling protein 1 (UCP-1), also called thermogenin, located in the IM membrane
protons flow through UCP1 instead of ATP synthase
present in brown adipose tissue
Is a proton gradient alone enough to drive ATP synthesis?
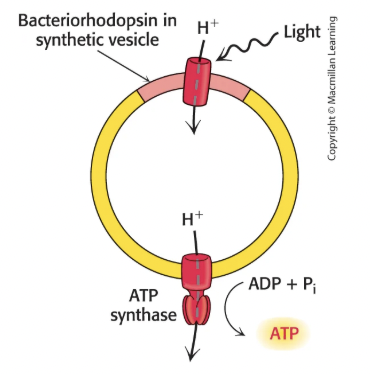
Yes, the experiment with bacteriorhodopsin proved this by making vesicles with just bacteriorhodopsin (light-powered proton pump) and ATP synthase, and they were able to form ATP, showing no chemical intermediates are actually needed
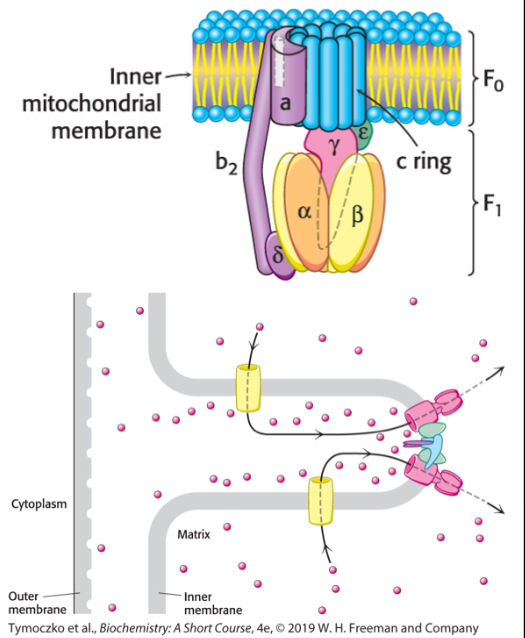
What is the structure of ATP synthase?
Two subunits: F1 and F0
F1 protrudes into the matrix and contains the catalytic active sites in three β subunits
F0 is in the IM membrane and contains the proton channel in the c ring
the γ subunit connects F1 and F0
protons flow through F0 and turn the c ring, which transmits the spinning to the γ subunit, which rotates the F1 subunit, driving conformational changes in the β subunits that drive ATP synthesis
ATP synthases dimerize on the tips of the cristaw
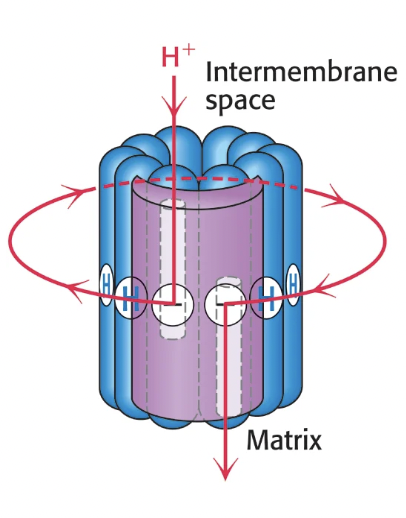
How do protons flow through the F0 subunit?
the a subunit abuts the c ring and contains two offset half-channels, one open to the IM space and one open to the matrix
protons enter the IM half-channel and bind to a glutamate residue on the c ring, causing the ring to rotate one c-subunit position clockwise
the proton is then released when it undergoes a 360 degree rotation around the c ring and reaches the matrix half-channel
c ring rotates one subunit per proton, so the number of c-subunits determines the efficiency of proton flow and thus ATP synthesis
If the c ring of ATP synthase has 10 subunits and each rotation generates 3 ATP, how many protons are required per ATP?
10 protons per rotation, so 10/3 = 3.33 protons per ATP
How does the rotation of the c ring generate ATP synthesis?
The binding-change mechanism
the β subunits exist in three distinct conformations at a time:
Open (O) → nucleotides can freely bind/release from subunit
Loose (L) → nucleotides are trapped in the subunit
Tight (T) → ADP + Pi catalyzed to form ATP (binds ATP with high affinity)
when the proton gradient spins the c ring, the γ subunit follows and rotates the F1 subunit, causing the β subunits to change conformation (T → O → L → T), releasing the ATP formed by the T subunit
a full 360 degree rotation produces 3 ATP molecules
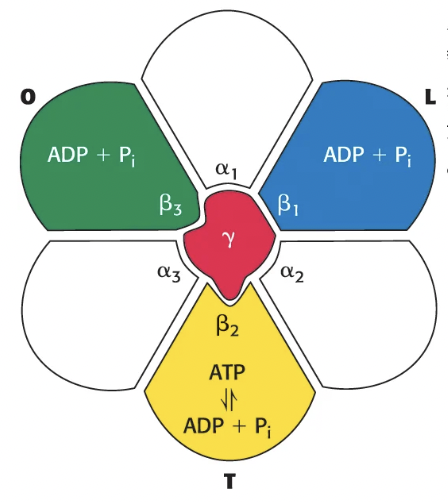
Does the proton gradient provide the energy to form ATP?
No, ATP synthase does not need energy to form ATP, the T binding site does this spontaneously because it binds ATP with such high affinity; rather, the role of the proton gradient is to provide the energy to release the newly formed ATP by rotating the subunits to change their conformation so ATP is not as tightly bound
What does the F1 subunit of ATP synthase do when isolated from the F0 subunit?
It acts as an ATPase and hydrolyzes ATP because without the F0 subunit providing directionality from the proton gradient, the thermodynamically favorable direction is ATP hydrolysis
Will a c ring with more or fewer c ring subunits produce ATP more efficiently?
A c ring with fewer subunits will produce ATP more efficiently because it will require fewer protons to complete a full rotation to produce 3 ATP
Can NADH from glycolysis get into the mitochondrial matrix to participate in the ETC?
No, there are shuttle systems that transfer the electrons from NADH, not the molecule itself
this transport has an energy cost that reduces the final ATP yield
How are electrons shuttled from cytoplasmic NADH to the ETC in muscle/brain?
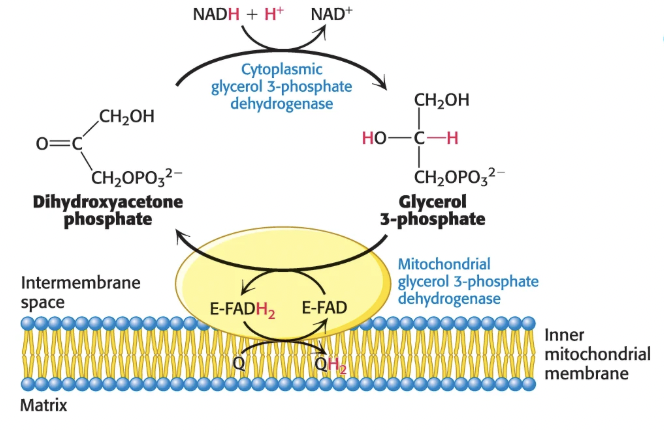
Via the glycerol 3-phosphate shuttle:
cytoplasmic NADH reduces DHAP to glycerol 3-phosphate via glycerol 3-phosphate dehydrogenase
glycerol 3-phosphate moves into the mitochondria where it is reoxidized to DHAP by an IM-membrane bound FAD-linked dehydrogenase, passing the electrons to FAD to form FADH2
FADH2 passes electrons to CoQ, reducing it to QH2 that then enters the ETC at Complex III
Faster, but less efficient because electrons enter ETC as FADH2, not NADH, so 1.5 ATP are formed instead of 2.5
How are electrons shuttled from cytoplasmic NADH to the ETC in liver/heart/kidney?
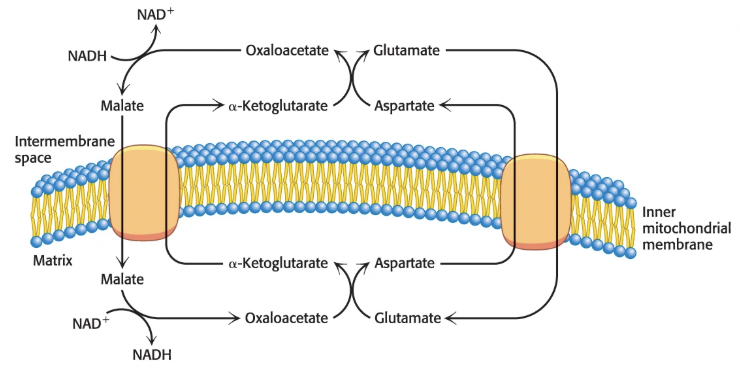
Via the malate-aspartate shuttle:
cytoplasmic NADH reduces OAA to malate, which enters the mitochondria via an a-ketoglutarate antiporter
malate then reoxidized in matrix to OAA, regenerating NADH; this is catalyzed by malate dehydrogenase (citric acid cycle enzyme)
OAA then re-enters cytoplasm as aspartate (transamination) via a transporter
NADH enters ETC at Complex I
Slower but more efficient because preserves electrons in NADH, which produces 2.5 ATP
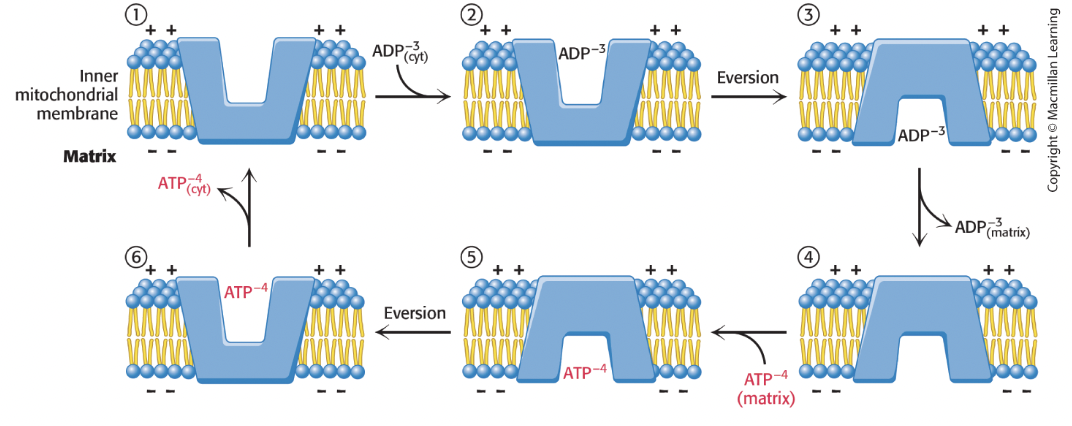
What is the ATP-ADP translocase (ANT)?
Most abundant protein in IM membrane
Swaps one matrix ATP (-4 charge) for one cytoplasmic ADP (-3 charge)
driven by membrane potential since matrix is negative and IM space is positive
inhibition of ANT inhibits cellular respiration
What is the phosphate carrier?
It transports Pi into the matrix coupled to a proton since the proton is moving down its gradient
What is an ATP synthasome?
The ATP-ADP translocase, the phosphate carrier, and ATP synthase associate with one another to form a large complex that increase efficiency
What is the total ATP takeaway from oxidative phosphorylation?
Glycolysis produces 2 ATP and 2 NADH
Pyruvate Dehyrogenase produces 2 NADH
Citric Acid Cycle produces 2 GTP/ATP, 6 NADH, and 2 FADH2
Each NADH produces 2.5 ATP → 25 ATP
Each FADH2 produces 1.5 ATP → 3 ATP
Total is 30-32 ATP per glucose (depending on shuttle system used for glycolysis NADH)
What is respiratory control/acceptor control?
Electrons do not flow through the ETC to O2 unless ADP is available to be phosphorylated to ATP; the rate of oxidative phosphorylation is regulated by the levels of ADP
high [ADP] → fast respiration
low [ADP] → slow respiration
[ADP] affects citric acid cycle too because if ETC is not running, NADH and FADH2 are not being oxidized back to NAD+ and FAD to feed the cycle, so it slows
What enzymes are stimulated when energy charge is low (not a lot of ATP)?
PFK-1, isocitrate dehydrogenase, and pyruvate dehyrogenase
What is inhibitory factor 1 (IF1)?
A mitochondrial protein that inhibits hydrolytic activity of ATP synthase when O2 is low, preventing wasteful hydrolysis of ATP
What are some ETC inhibitors?
Rotenone, antimycin, cyanise
these block electron flow, lowering O2 consumption and ATP synthesis
What is an ATP synthase inhibitor?
oligomycin
proton gradient builds up, back pressure stops the ETC, O2 consumption and ATP synthesis decrease
What are some uncouplers?
UCP1, DNP, FCCP
dissipate the proton gradient, so ATP synthesis drops but ETC runs faster to rebuild gradient, so O2 consumption increases
Inhibitor prediction table