BSCI331 Exam 3
1/128
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
129 Terms
most common way a cell can receive signals
cell surface receptor
3 phases of cell reception
Reception: binding of ligand to cell surface receptor
Transduction: transfer of information from inside to outside (usually has a signal pathway)
Cellular response
In most cases how are signal molecules created?
they are secreted — released from one cell and diffuses to reach another cell
Secreted molecules that travel locally, affecting cells in the immediate environment is called?
paracrine signals
secreted molecules that act over long distances through the bloodstream?
endocrine signaling
secreted molecules that travel through axons
synaptic signaling — specific and knows who the receptor is, unlike endocrine and paracrine signaling
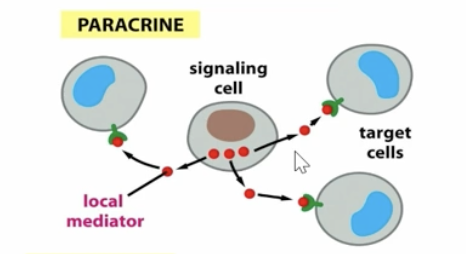
paracrine signaling graphic
paracrine signaling graphic
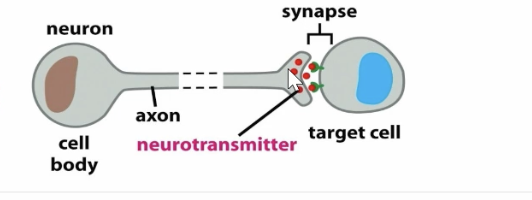
synaptic signaling graphic
can cells secrete a molecule that binds to its own receptor
yes — autocrine signaling
why would we ever have autocrine signaling cells?
Cells are autocrine to self-stimulate, allowing a single cell to regulate its own growth, survival, and differentiation by releasing signaling molecules that bind back to its own receptors. Having this provides instant feedback for rapid responses, immune cell activation (e.g., T-cells), and tissue
why would a signaling molecule stay on the surface?
contact dependent signaling and this happens in vital developments such as embryonic development to ensures precise localized signaling of the moelcule in the active site
Extracellular molecule signalling can work in 2 ways
Rapid or slow
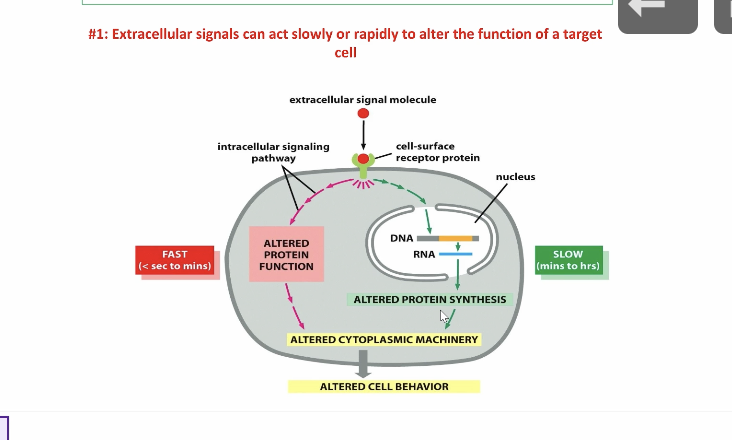
how does rapid extracellular molecule signalling work
it immediately alters the proteins function which then alters the cells behavior
how does slow extracellular molecule signalling work
the extracellular molecule does not immediately alter the protein function, but it send a cascading message inside the nucleus and then goes DNA —> RNA and synthesizes the protein and then alters the cells behavior
causes transcription and the expression of new genes
Rapid and Slow extracellular signalling graphic
how do multiple signal receptors on a cell work?
they all must superscore for a specific response —> alters the protein function —> changes cell behavior
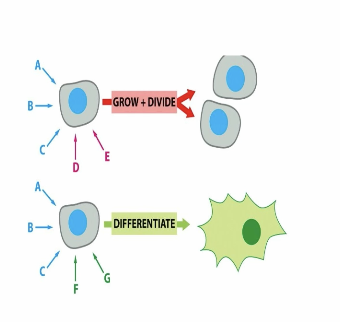
can multicellular organisms decide to die on its own?
no
what is one way in which a cell is destined to survive
receiving the necessary cells to “tell it” to survive, missing these results in deahth
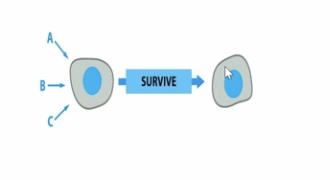
what is the name of the suicide cells commit when they dont receive the necessary signals?
apoptosis
are each survival signalds restricted to the specific environment in the body?
yes
can different receptors bind to the same transport moelcule (ligand)
yes, cardiac cells, mouth cells and skeletal muscle cells all have different receptors that bind to Ach and cause different outcomes — due to different internal machinery
can the same signaling molecule cause different protein alterations and cell responses based on it’s concentration?
yes
cells are specialized to receive and respond to a wide variety of stimuli
mechanical: cells can tell they are in contact with something else when the internal machinery is distorted, detect pressure such as being smushed, sound from people
light: nerves from retina where we can see
heat: feel the fire from the other day
chemical: amino acid signals, nucleotides, small peptides, proteins, steroids, fatty acids, gases, hormones all binding to receptors
Most of these cells are created through exocytosis and secreted
Lipid soluble molecules
signaling molecule that diffuses past the plasma membrane
steroid hormones, cortsol, sex horomones, vitamin D
how do the hydrophic molecules travel to their receptors
lipid soluble/NP signals bind to carrier proteins and bind to their respective intracellular receptors which be cystolic of nuclear
what are the family of intracellular receptors that bind to llipid soluble singnals?
nuclear receptor subfamily
what state is the nuclear receptors in before the ligand binds?
inactive — bound to an inhibitory protein
contains a ligand binding, DNA binding and tracription activating domain
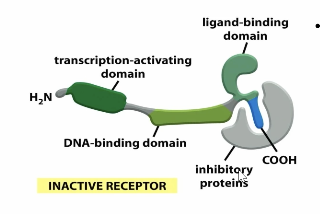
when the ligand binds to the nuclear receptor what happens?
It causes a conformational change and releases the nuclear receptor from the inhibitory protein, allowing it to fold into its correct shape and travel to the DNA, causing transcription
It can bind directly to RNA polymerase or chromatin remodeling (loosens) and activates transcription
what is great about NP/lipid soluble signal molecules?
the receptors are also the downstream molecule
no need of an additional pathway making it fast
what are the majority of the signal molecules?
hydrophilic/lipid insoluble signals
neurotransmitters, peptides, GF, olfactants and tastants
what type of receptors are present for insoluble/polar molecules?
cell-surface receptor proteins that bind to hydrophillic signal moelcules
what are the three main types of cell-surface receptors?
ion-gated channels
enzyme-coupled receptors
G-protein coupled receptors
how do ion gated channels work?
a protein forms a channel which is closed, when the ligand binds, it opens the protein channel allowing ions to flow across the channel
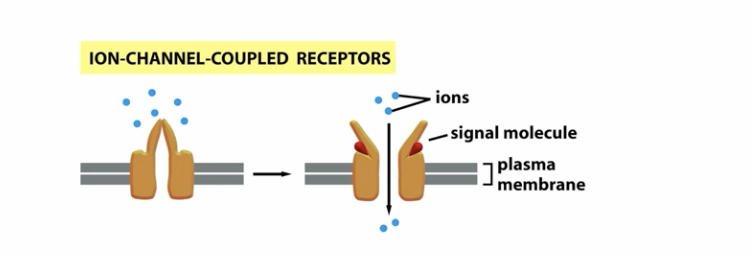
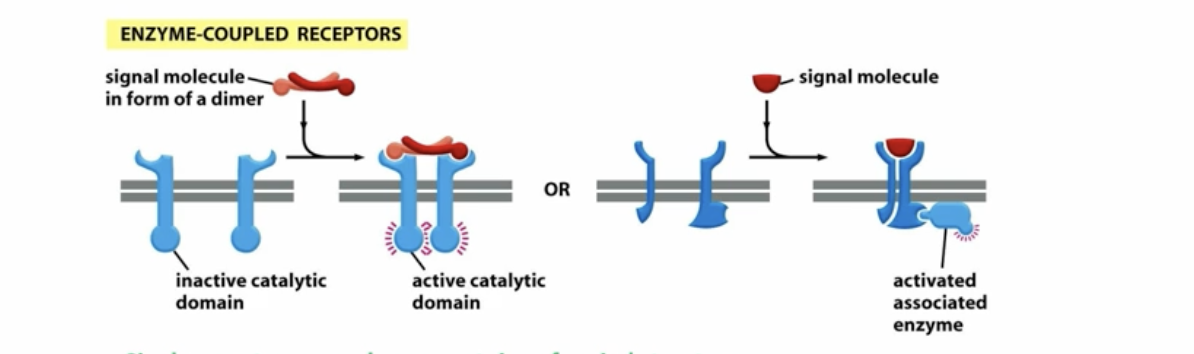
how does enzyme coupled receptors work?
signal molecule binds to the protein receptor, allowing it to form a conformation where the inactive kinase receptor dimerizes allowing one to phosphorylate the other and VV increases the activity of the kinase and phosphorylates the signaling
Keep in mind enzyme-coupled receptors can also work by having the kinase domain act as a separate protein that is tightly associated with the receptor (non receptor tyrosine kinases)
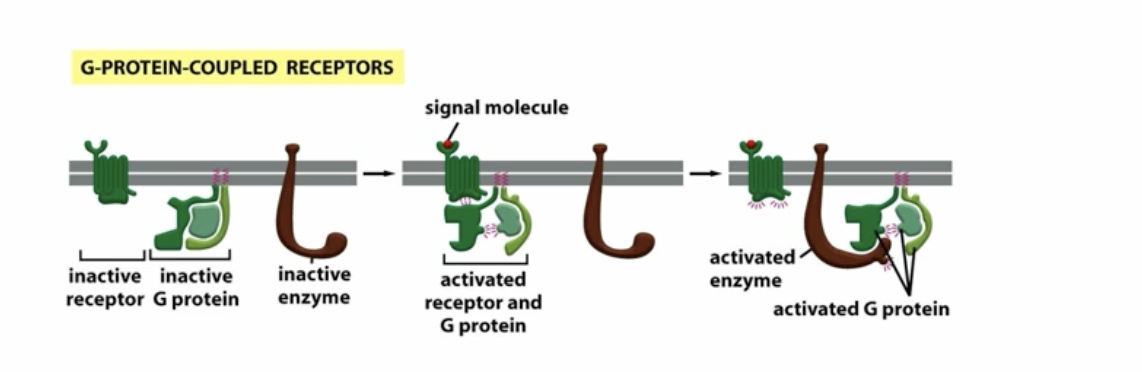
how do G-protein coupled receptors work?
signal molecule binds to the receptor, shifting the G-protein complex over allowing the G-protein to release GDP and bind GTP, putting it in the active state. The alpha chain seperates from the beta and gamma chains, allowing the alpha chain to bind to other enzymes
are extracellular receptors the same as the one-and-done nuclear receptors that act as the downstream effect
NO
what do we use to send the signal in the extracellular signaling proteins?
intracellular signaling proteins that lead to protein alteration and change in cells activity
what do the secondary messengers do?
amplify the signal and spread it throughout the cell
what benefit do signal transduction pathways have over nuclear receptor paths
branch and amplify the signal
many intracellular signaling proteins function as what?
switches — kinases and phosphates
(phosphorylation and dephosphorylation)
what are signaling complexes
They are when multiple proteins assemble together inside cells to rapidly transmit external signals from receptors to intracellular targets
speed
efficient
specific
what are some ways we can assemble intracellular protein signaling complexes
scaffold protein vertically
receptor serves as it’s own scaffold
membrane lipids act as the scaffold
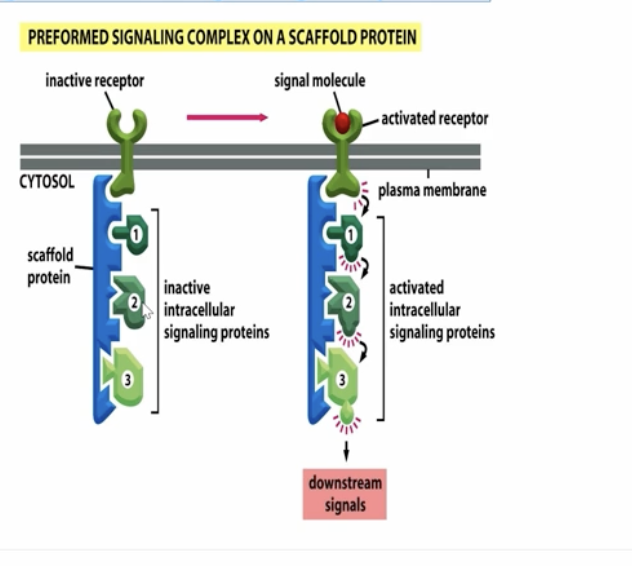
scaffold protein vertically
non enzymatic proteins that act as an adaptor and keep intracellular signaling proteins organized
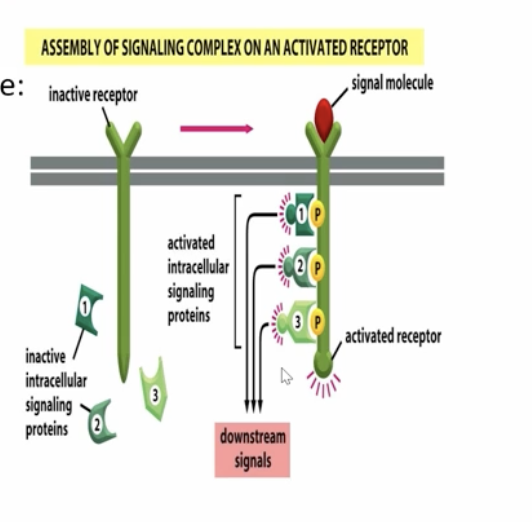
receptor serves as it’s own scaffold
the cytoplasmic portion of the receptor serves as bidning sites for the intracellular signaling proteins
membrane lipids act as the scaffold
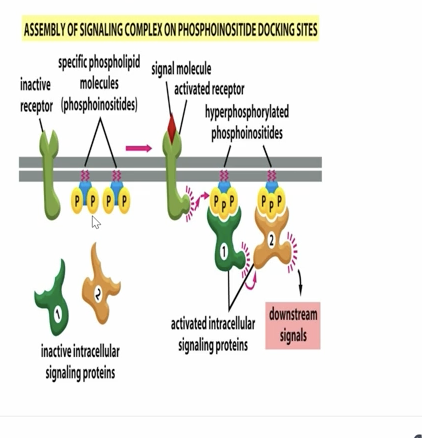
When a ligand binds a receptor tyrosine kinase, the receptor becomes activated and recruits enzymes such as PI3K that phosphorylate membrane phosphoinositides. These phosphorylated lipids serve as docking sites for cytoplasmic signaling proteins, allowing them to assemble at the membrane and generate downstream signaling. Lipid rafts help concentrate receptors and phosphoinositides to make this process more efficient.
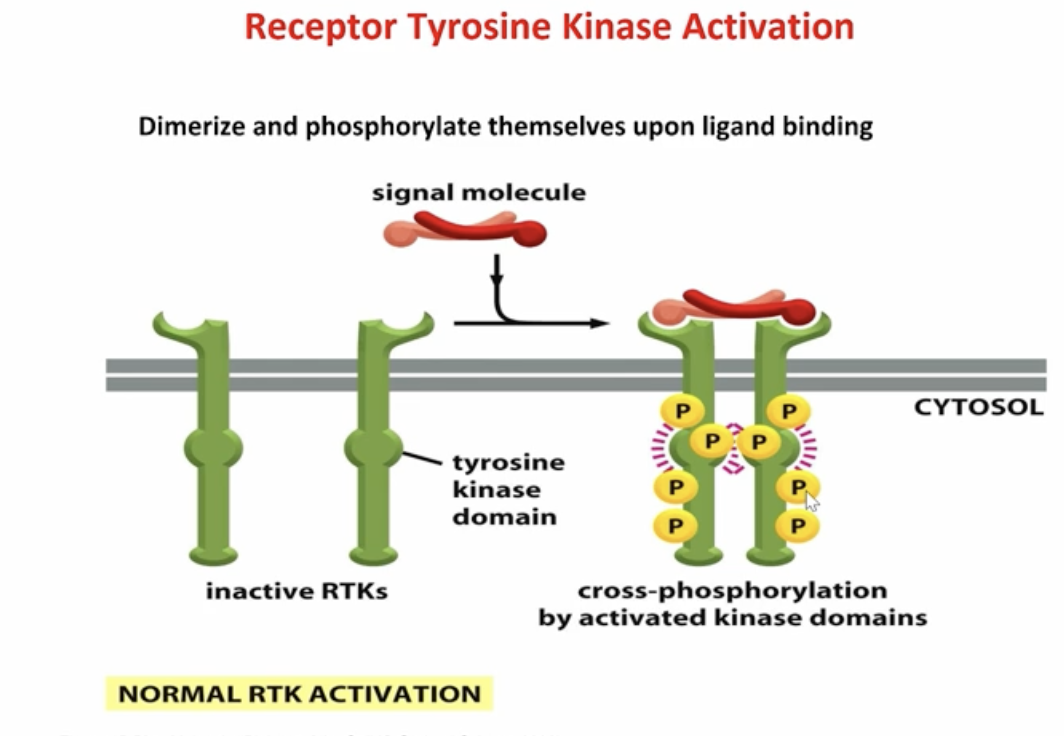
receptor tyrosine kinase activation
Inactive RTKs bind a ligand, which causes dimerization. This brings the kinase domains together, allowing them to activate and cross-phosphorylate each other on tyrosine residues.
what makes RTKs (receptor tyrosine kinases) so damn special
they allow the docking of intracellular singaling proteins to the phosphorylated tyrosines via SH2 domains forming a signaling complex downstream amplifying the lignad binding ti the RTK
Graphic of RTK
ligand binds to the RTK allowing the receptor to dimerize and phosphorylate resulting in intracellular binding proteins binding to the docking sites on the phosphorylated Tyrosines via SH2 domain
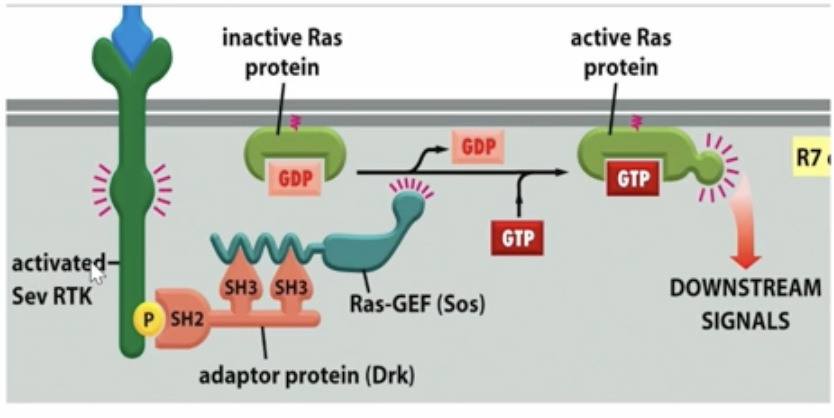
after RTK is phosphorylated what can the signal be passed to?
Ras
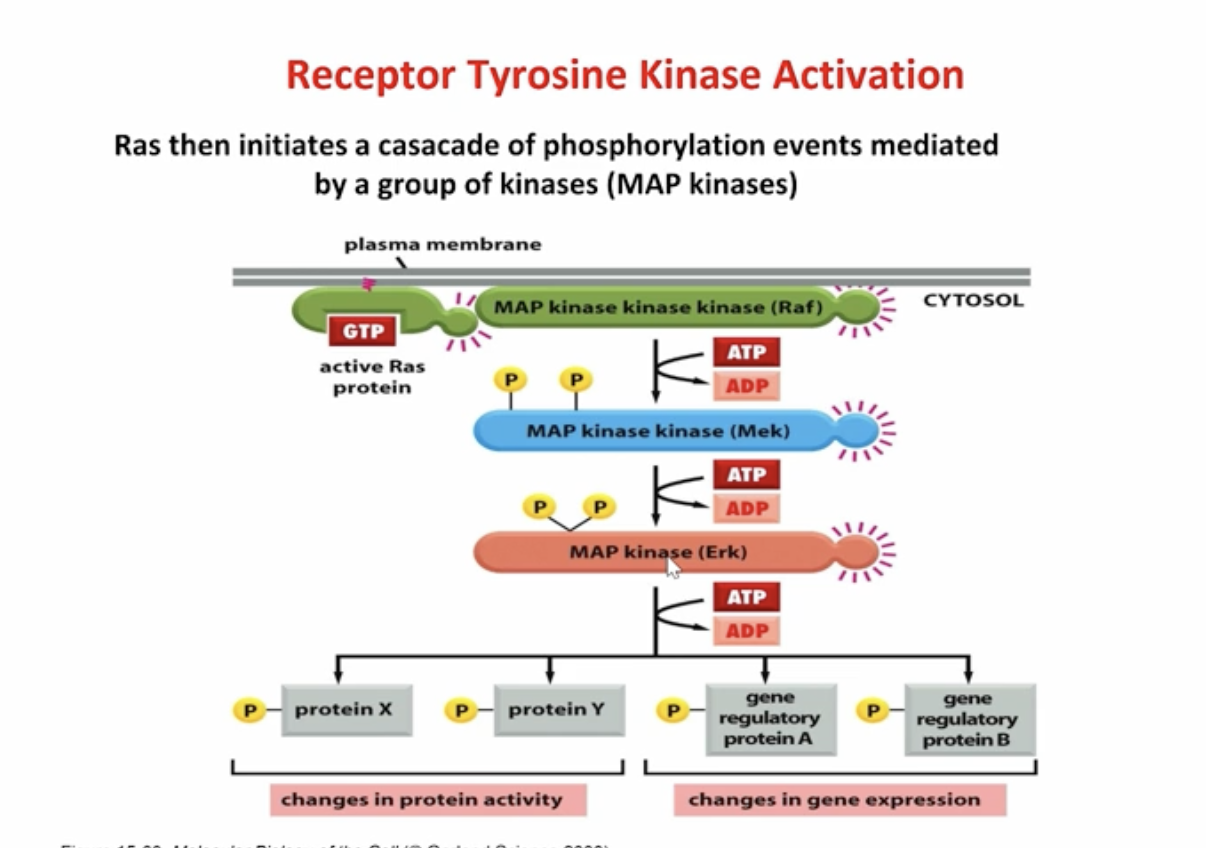
what does Ras do?
it causes a kinase cascade
what type of protein is RAS?
it is a G-protein which is activated when GTP is bound
order of RTK and Ras
RTK gets phosphorylated
Ras becomes activated (GTP on)
Ras activated a cascade: Raf, Mek, Erk (each step gets phosphorylated via MAP kinase)
Erk activated proteins and causes transcriptional factors
graphic of G-protein coupled receptor
G-protein is activated, causing GDP → GTP swapping, allowing the alpha unit to break off and bind to other enzymes and regulating their function
G-protein stimulated adenylyl cyclase, converting ATP to cAMP, a secondary messenger bind and bind to the cAMP responsive protein kinase A where it can diffuse in the cytosol or nucleus
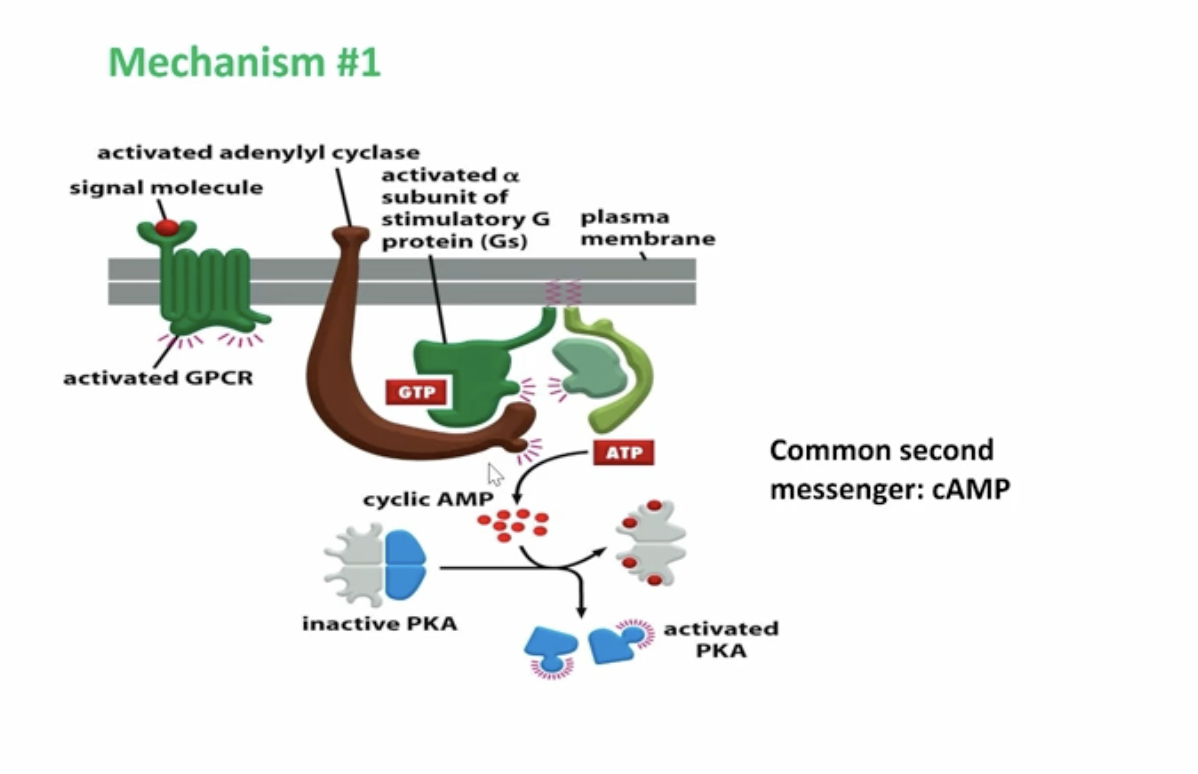
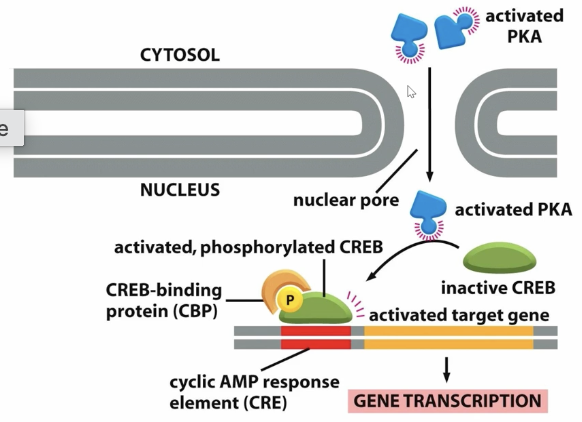
A graphic of protein kinase A diffusing in the nucleus and activating CREB, where it binds to CBP and leads to gene transcription
does the alpha subunit in the G-proteins vary
Yes, they can bind to adenylyl cyclase or any other enzyme causing different behaviors
What is the storyline of G-protein receptors?
Bind → GDP→GTP → split alpha subunit → activate enzyme → make second messenger → response → GTP hydrolysis off
what is the role of Ca2+ ions in intracellular communication
they are second messengers
what activities do Ca2+ ions have a role in?
muscle contraction, cell division, secretion, synaptic transmission
Graphic of gene expression regulation and what points along central dogma we have checkpointst
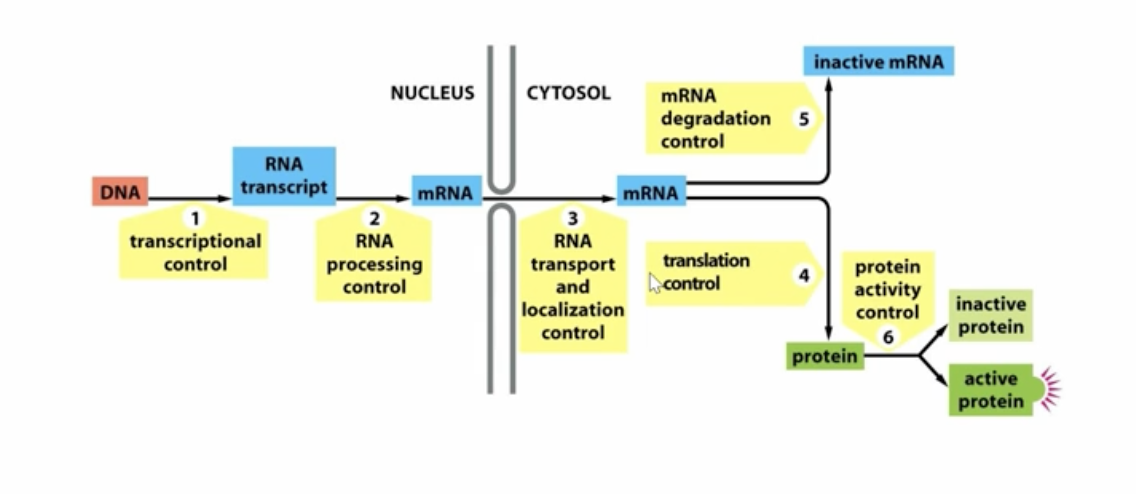
What is the first point of gene regulation?
transcriptional control
for most genes where does the primary regulation occur?
RNA transcription
how can we maximize the central dogma?
create only the amount of RNA we need to transcribe into protein so we don’t waste ATP
where is the location of regulating RNA transcription?
initiation
what are the things that regulate transcription called?
transcription factors
what is the special structure of the transcription factor that aids in regulation
DNA-binding motifs
what do DNA-binding motifs do
they find and bind to specific nucleotide sequences in the DNA through the grooves and increase/decrease RNA polymerase activity
why do transcription factors prefer the major groove?
they bind to the major groove because it exposes more information about the DNA
How does the formation of transcription factors bind to the DNA
the DNA-binding motifs have H-bond acceptors which bind to the H-bond donors of teh base sequences of DNA to create that “lock” and alter RNA polymerase activity
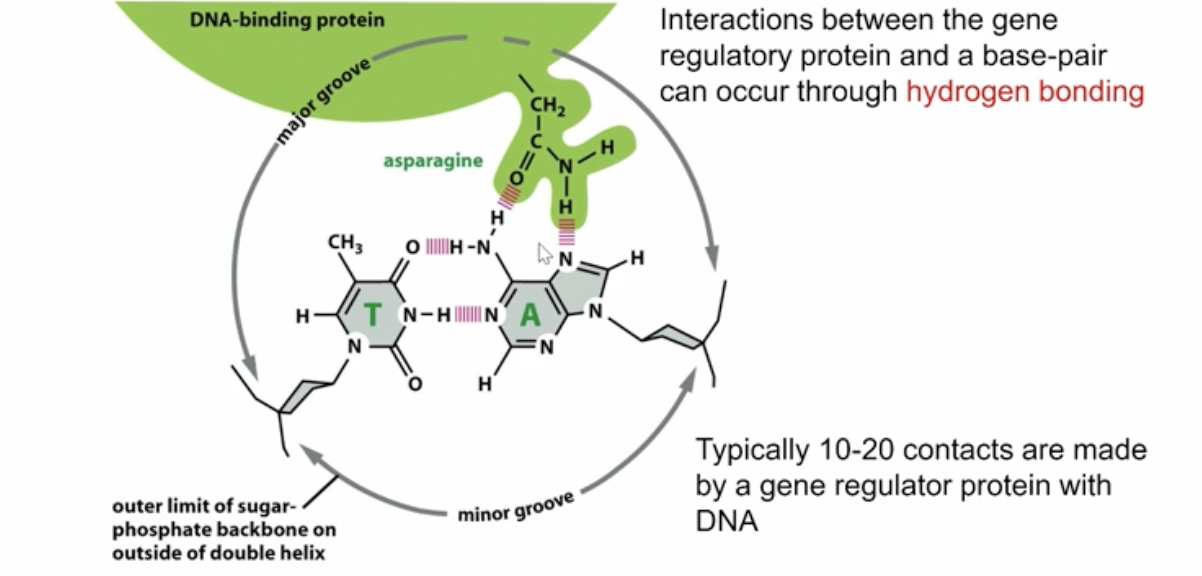
How many DNA-binding motifs are there?
6 different DNA-biding motifs
Helix-turn-helix DNA-binding motif
2 alpha-helices which are conncected by a short unstructured turn with the c-terminus recognition helix makes contact with the major groove of DNA
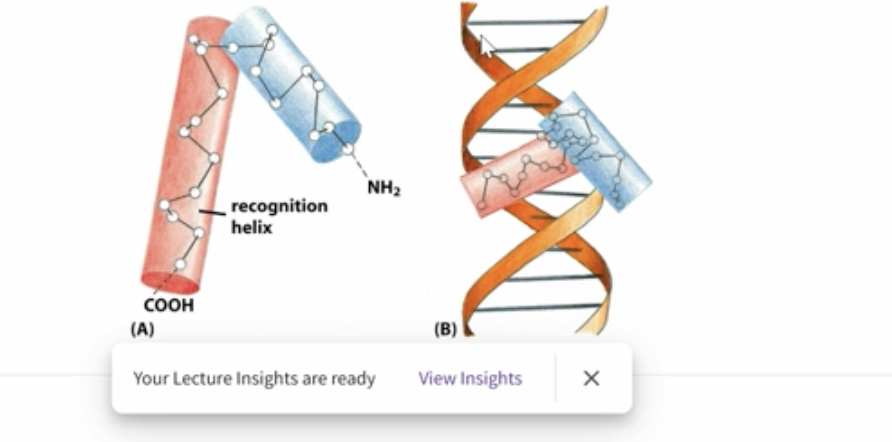
Homeodomain DNA-binding motif
3 alpha turn helices which create bonds with the major groove on the c-terrminus and bonds with the minor groove in the n-terminus
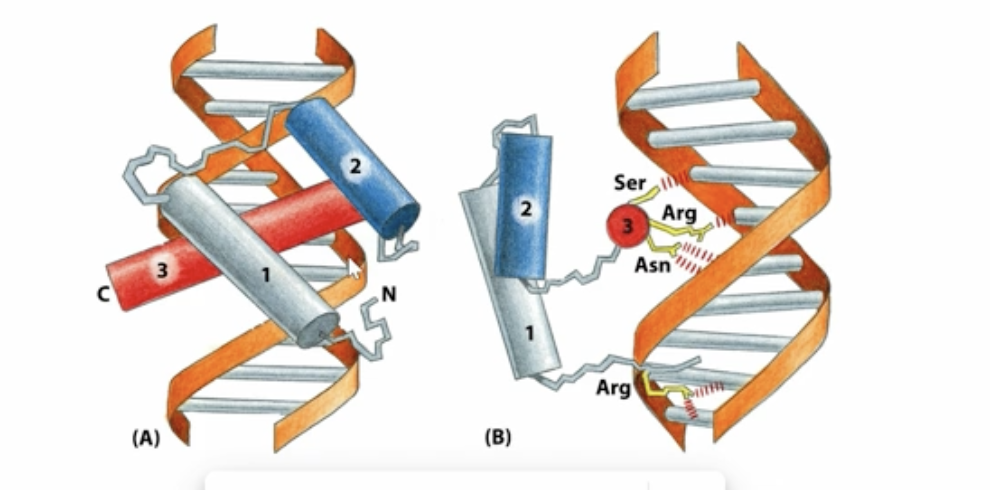
Zinc fingers DNA-binding motif
consisted of an alpha helix and a beta sheet, where the alpha helix binds to the major groove of the DNA while it is stabilized by the Zinc ion
We can alter the specific sequence of amino acids in the alpha helix to create whatever recognition we want
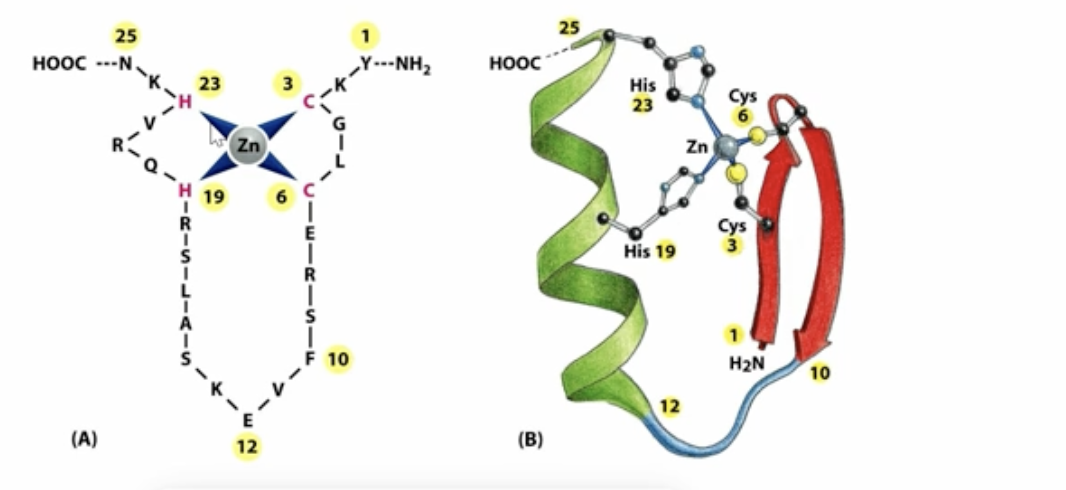
Zinc fingers DNA-binding motif (second one!)
This zinc finger model has 2 Zn ions, one is used in the middle of the motif like normal, and the other one is used to stabilize a loop involved in creating DIMERIZATION which bind to DNA as symmetric dimers

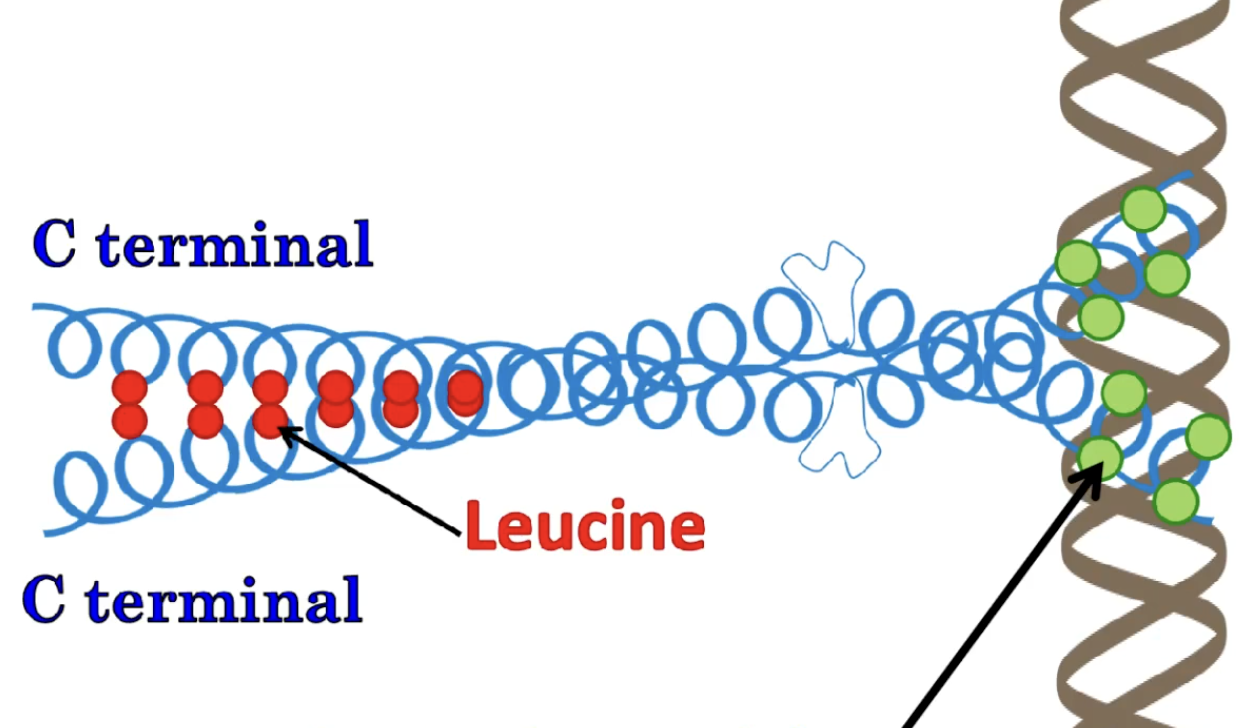
Leucine zipper DNA-binding motif
2 Alpha helices formed dimerized together, with the C-terminus having NP bonds such as Leucine forming a zipper and the other end having Polar bonds (N-terminus) making bonds with the DNA
DIMER
Helix-loop-helix DNA-binding motif
2 alpha helices are held together — one short alpha helix and another long alpha helix held together by a flexible loop
DIMER
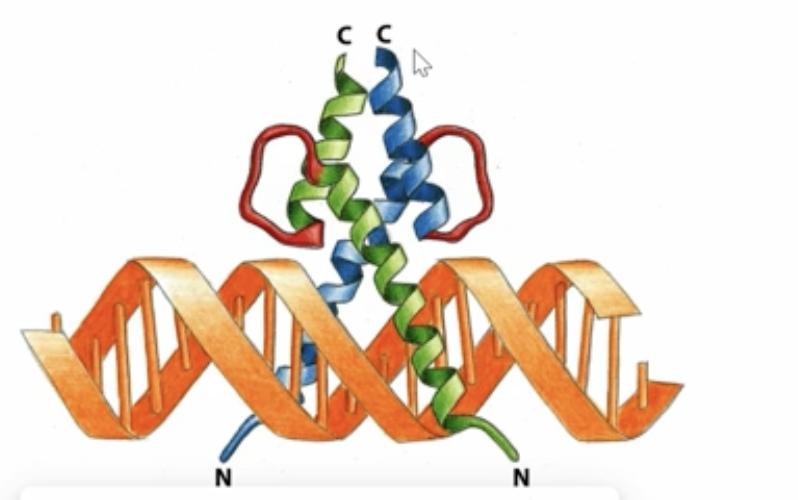
Why do we have these DNA-binding motifs that bind in the major groove using their alpha helices as dimers?
We use dimerized DNA-binding motifs such as leucine zippers, zinc fingers, and helix-loop-helix to enhance the binding and specificity to double the contact area and square the affinity
what are the benefits of heterodimerizing?
increase the range of sequences that can be recognized
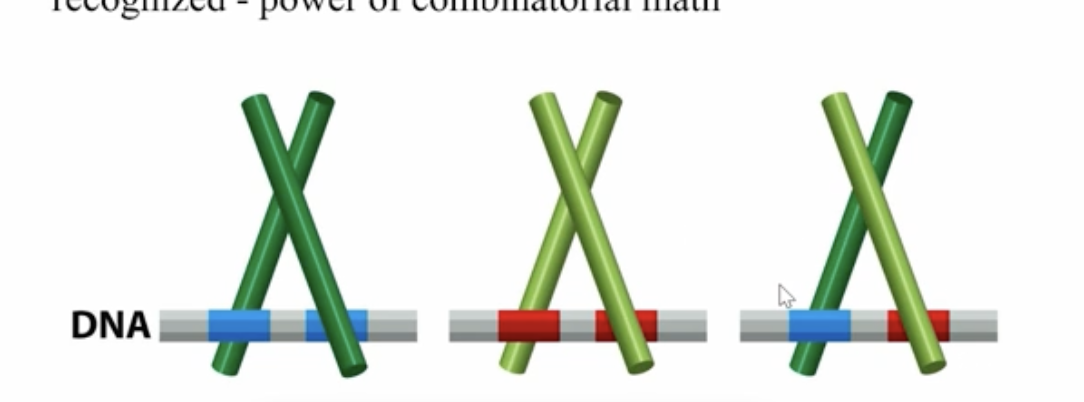
How many different sequences can be recognized by a dimerization of a TF with A, B and C
AA, AB, AC, BC, CC, BB
*AC and CA are the same and result in the same transcription response so that doesn’t count”
Where do transcription factors usually act on in initiation?
Promoters and Enhancers
What is the promotor
Region of the DNA where the RNA polymerase and general transcription factors (essential enzymes used to aid the RNA polymerase in binding to the promoter) assemble
where is the promotor located
just upstream of the 5’ end
can promotors be gene-specific?
yes, they can which means they only work when they are located to their adjacent gene and orientated right
What is an enhancer
independent region outside the promotor which can be very far away which can boost the transcription of a specific gene
why do TFs bind to the enhancer and promoters
they are both used in either aiding in transcription or initiating it
can enhancer work with other promotors? (coworkers)
yes with a heterologous promoter
How can we use the TF to modify the transcription and thus the gene expression?
The TF factors can bind synergistically or antagonistically producing different effects
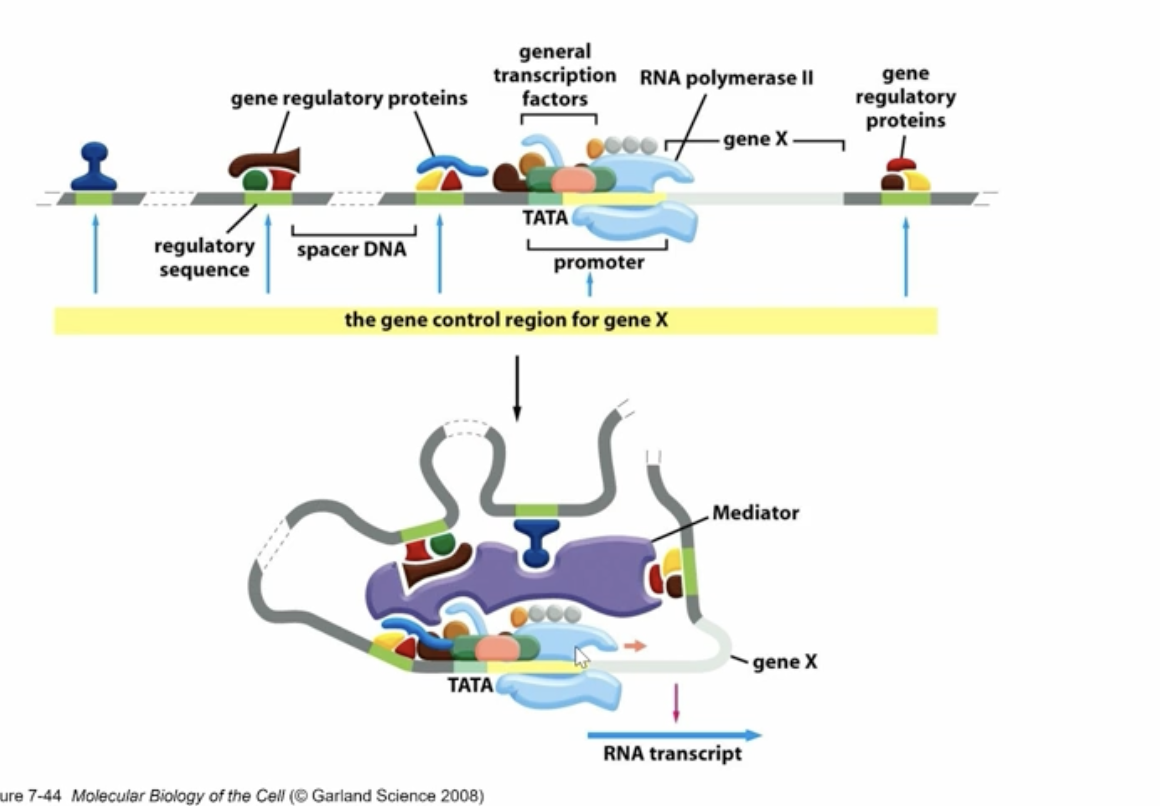
what is going on here?
the TF are binding to the enhancers and regulatory sequences very close to the promotor, then the mediator protein causes the DNA to loop allowing the TF to interact with the promotor region
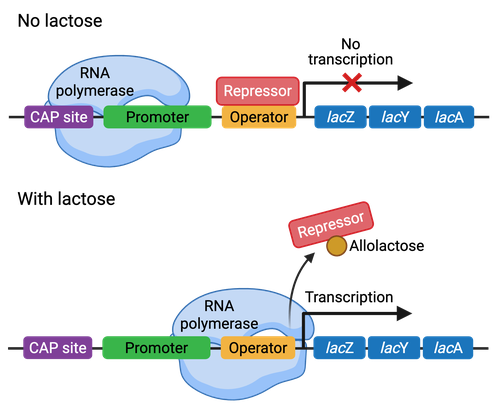
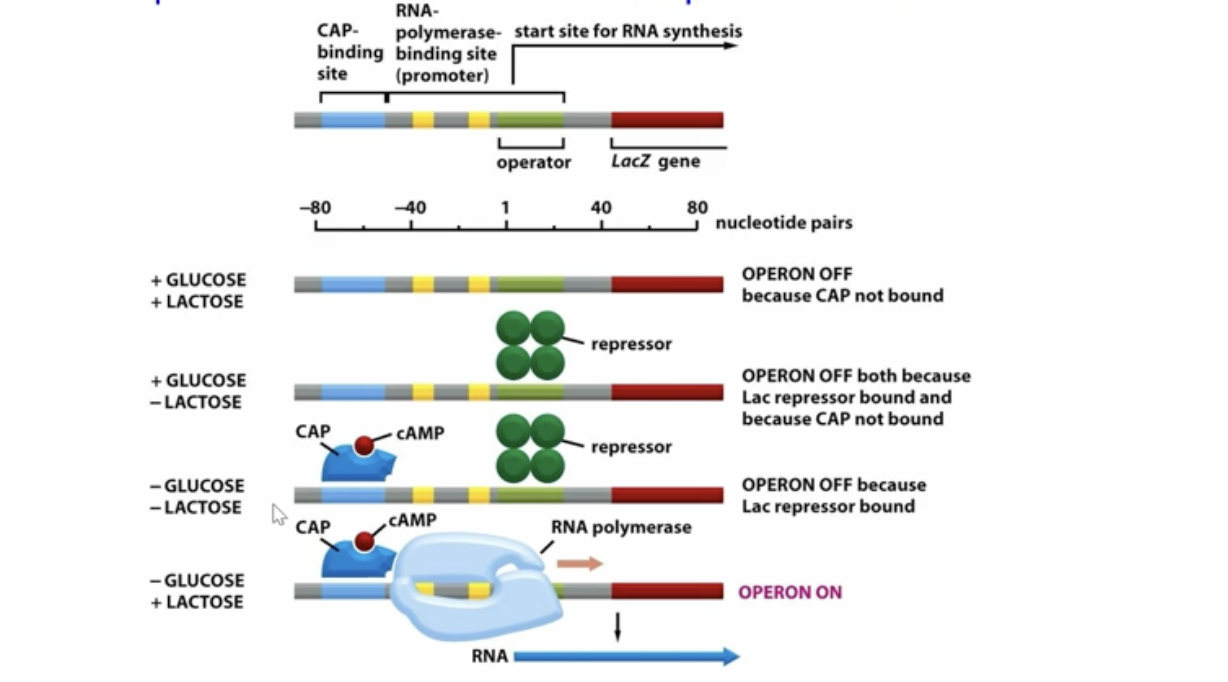
what does the lac operon do?
it creates structural enzymes to bring in and break down lactose when it is available in the environment
how does the lac operon work in the differing environments?
No Lactose: there is no point in running ts so the Lac I regulatory sequence has an active repressor bind to the promotor to prevent RNA Poly from transcribing the gene to make the lactose destroyers
Lactose Present: lactose converts to allolactose allowing it to act as an inducer binding to the repressor moving it away
When we have Low Glucose: we bind a CAP which carries cAMP to the CAP binding site, now keep in mind if lactose is low we still have the repressor but if lactose levels are high and glucose levels are low we will have the CAP/cAMP as well as the repressor not being bound so there is transcription
what mechanisms do TF actually regulate after they bind to regulatory sequences such as promoters and enhancers?
help unpack chromatin making the gene more accessible for RNA polymerase promoting transcription
help recruit more GTF by binding close to the promoter
can regulate the switch from initiation to elongation
recruit histone-modifying enzymes to change the local chromatin structure
can bend the DNA allowing long-distance TF to interact between regulatory sequences
how do we regulate TF?
we can’t have all of the TF on all at once or else there’s no regulation
they’re pretty smart, they bind when they are needed and don’t when they aren’t needed
some TF are for tissue specific expression — TF only bind to liver when needed
some TF are only expressed based on environmental signals — like hunger
some TF are made only during some phases of the cell cycle
why are some TF incactive in the cells?
so the cell can regulate when we want to turn genes on
how can we activate TF
signaling pathways such as RTKs (MAPK occurs downstream), or the kinase enzyme phosphorylating the TF
What is NFAT and NF-κB
they are both TF that are inactive in the cytoplasm until the genes are told to turn it on
NFAT process (triggered by T cells)
inactive TF in the cytoplasm that is phosphorylated, when Ca2+ increases it activates a phosphatase that dephosphorylates the NFAT and allows it to move into the Nucelus to activate genes
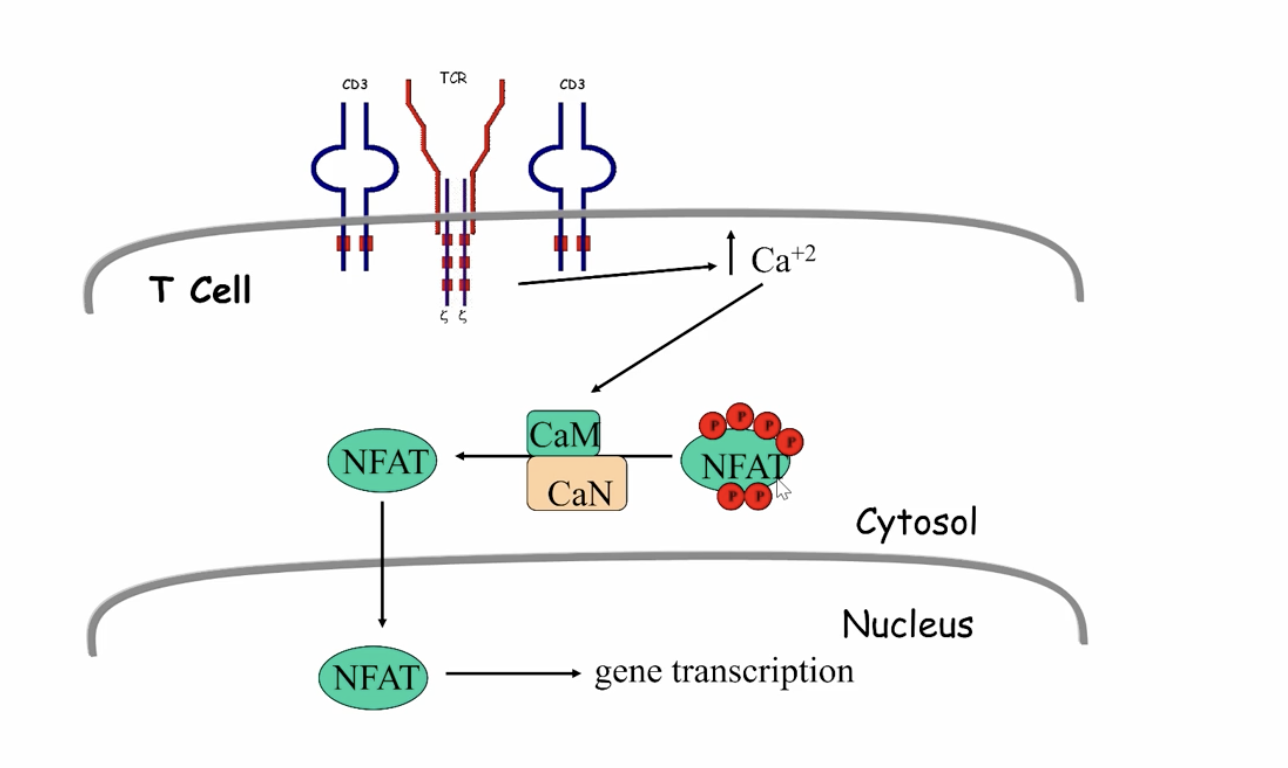
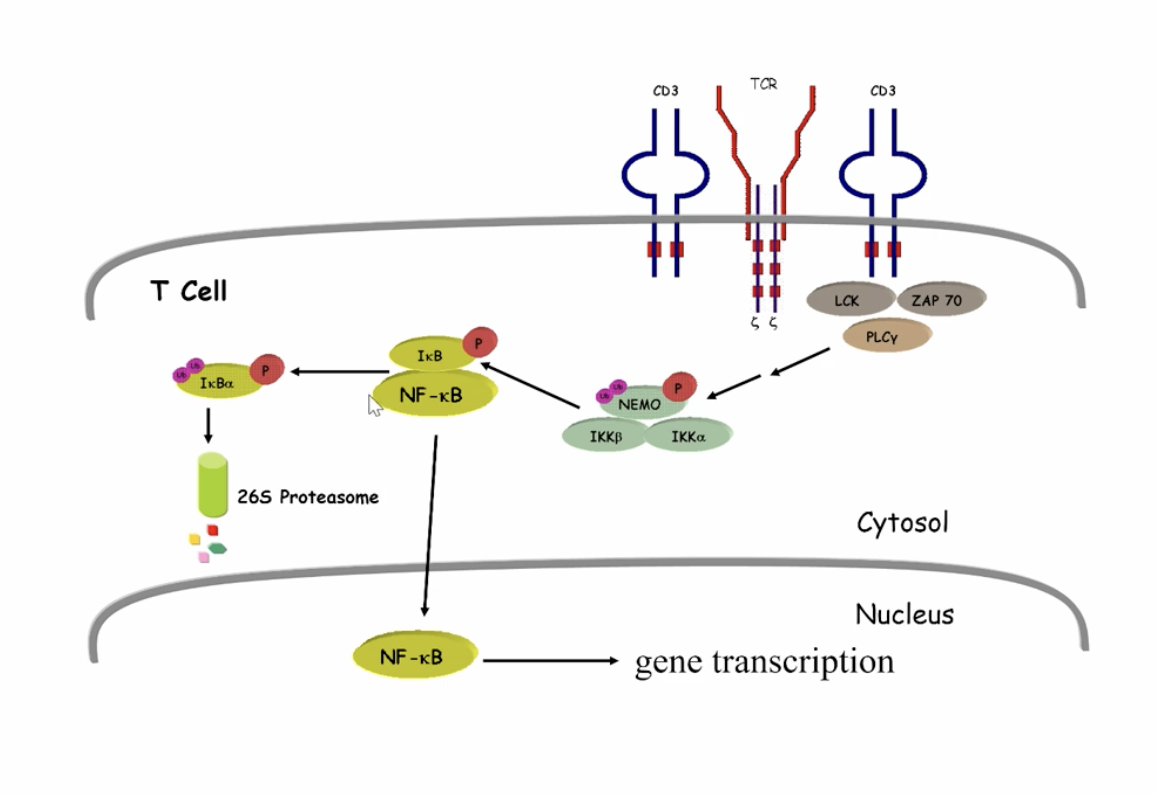
NF-κB process (triggered by T cells)
NF-κB is bound to an iκB inhibitor, when iκB is phosphorylated it becomes ubiquinitized and degreades allowing the NF-κB to move into the cell and result in gene transcription
how is it regulated? NEGATIVE feedback loop: newly synthesized IκB inhibits NF-κB and returns it to the cytoplasm
what is the combinational control of gene expression?
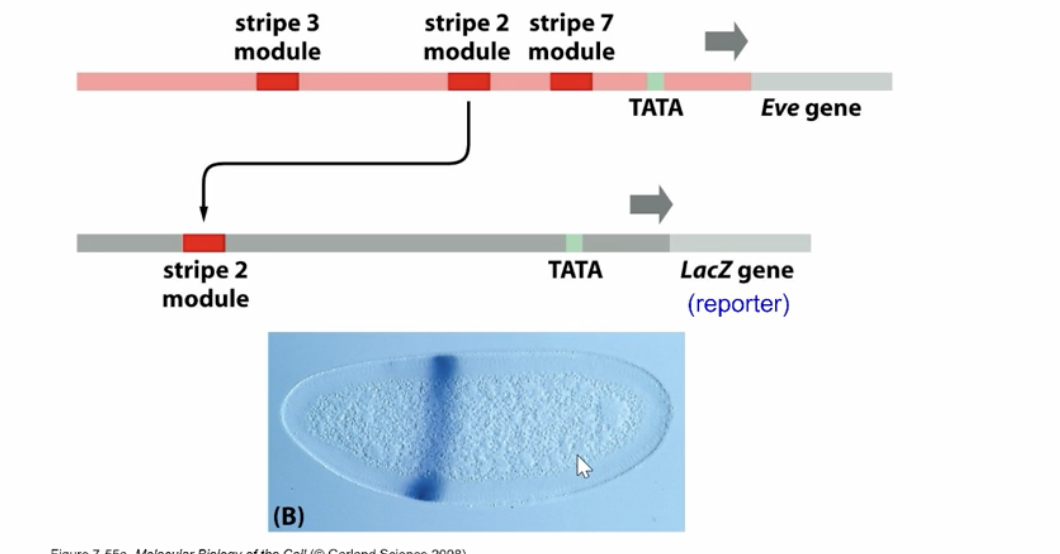
Combinatorial control of TF produces spatial patterns of gene expression, such as the striped expression of the even-skipped gene in embryos
how do we get this on/off sequence of one gene?
what we are used to which is a gradient of TF in the DNA, this is not that case and actually it requires enhancers
how can we express only one of the stripes on it’s own?
Each stripe is controlled by its own enhancer module that integrates inputs from multiple transcription factors to produce expression in a specific region.