MASTER CHEM221 FINAL
1/78
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
79 Terms
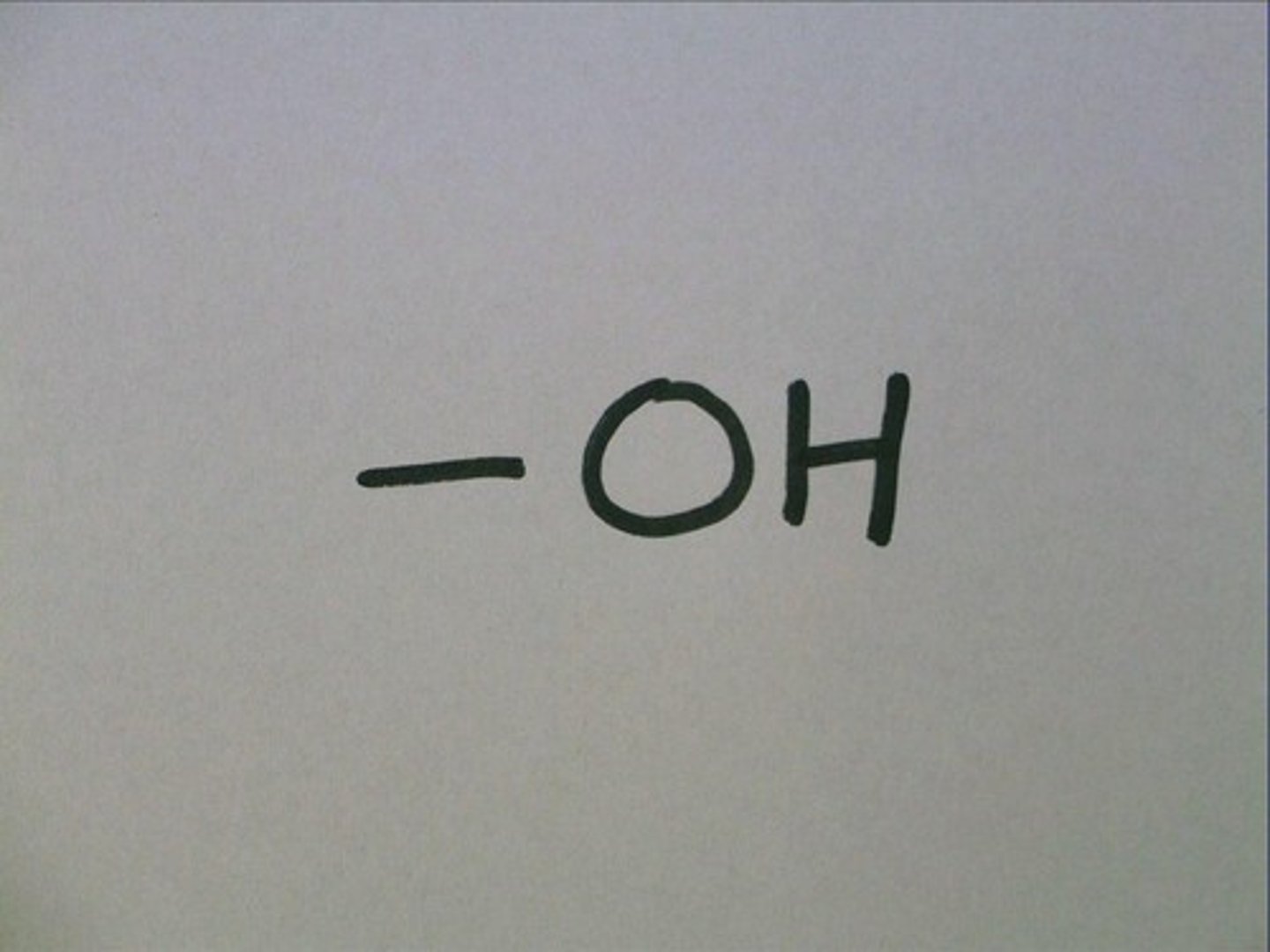
Hydroxyl Group
A functional group characterized by a hydrogen atom bonded to an oxygen atom, commonly found in alcohols.
Amine
A functional group consisting of a nitrogen atom bonded to 3 other atoms (C or H)

Nitrile
carbon triple-bonded to nitrogen
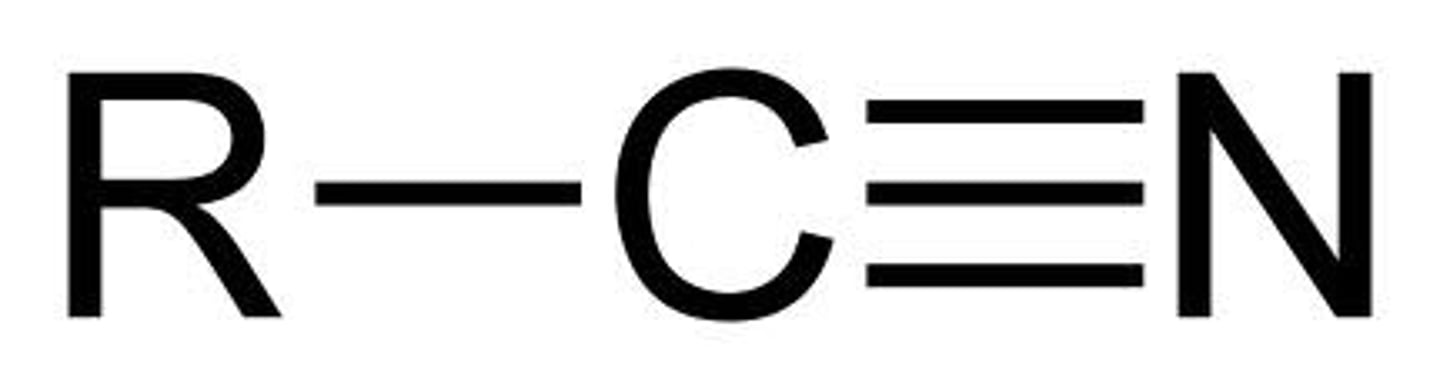
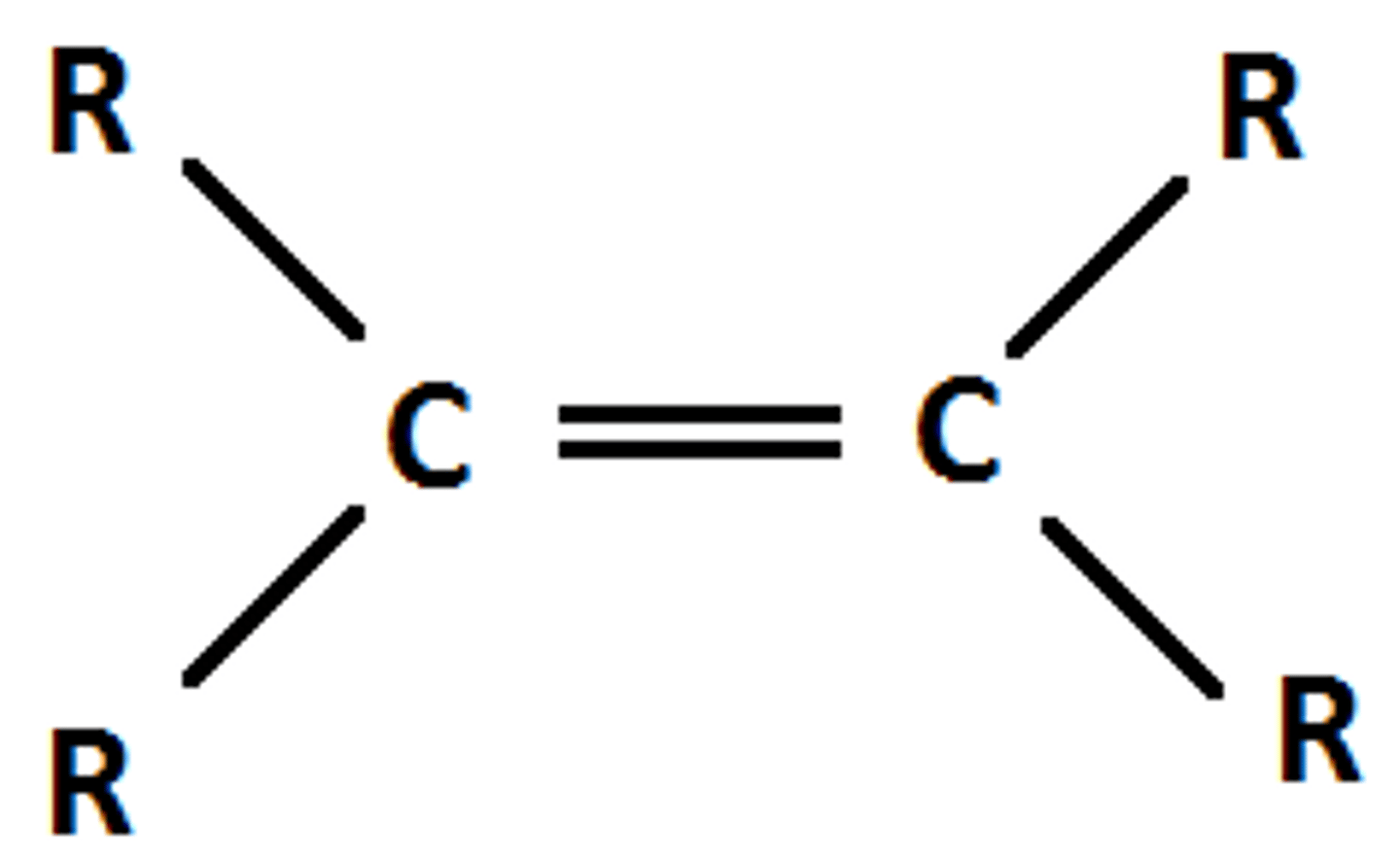
Alkene
A functional group characterized by at least one carbon-carbon double bond
Alkyne
A functional group characterized by at least one carbon-carbon triple bond
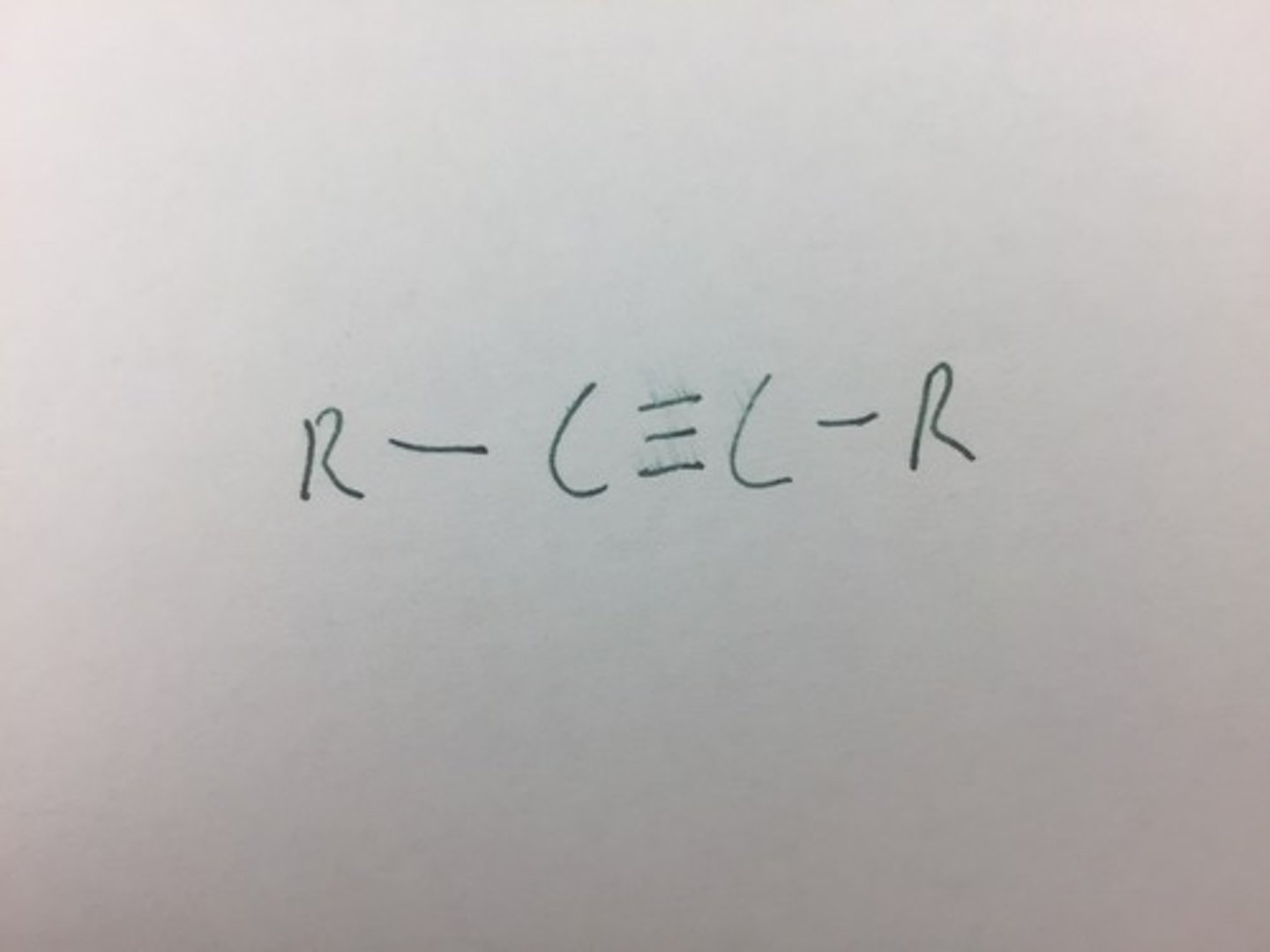
Arene
a functional group that's an aromatic ring with carbon-carbon double bonds
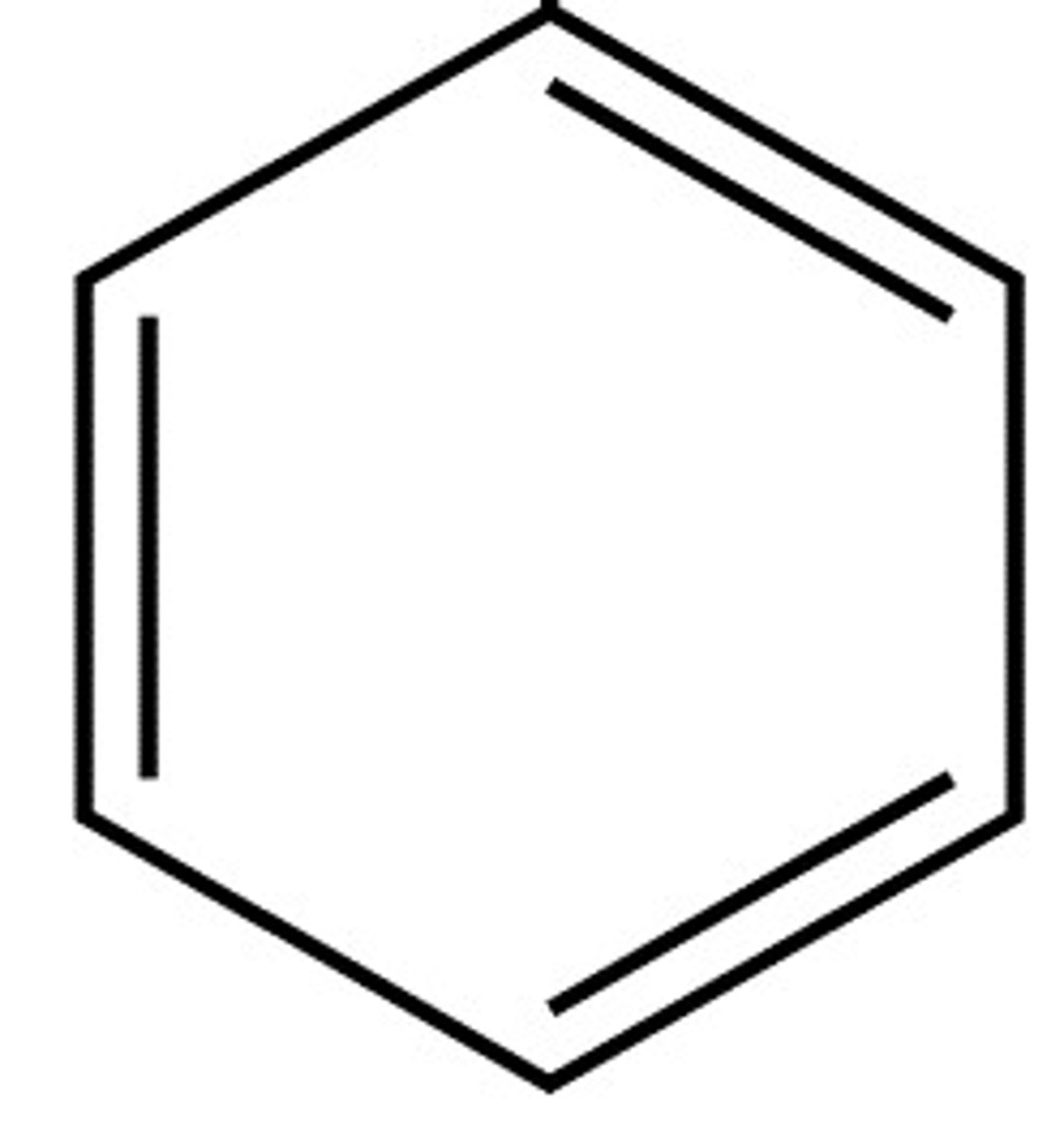
Halide
Carbon bonded to a halogen (Br, Cl, F, I)
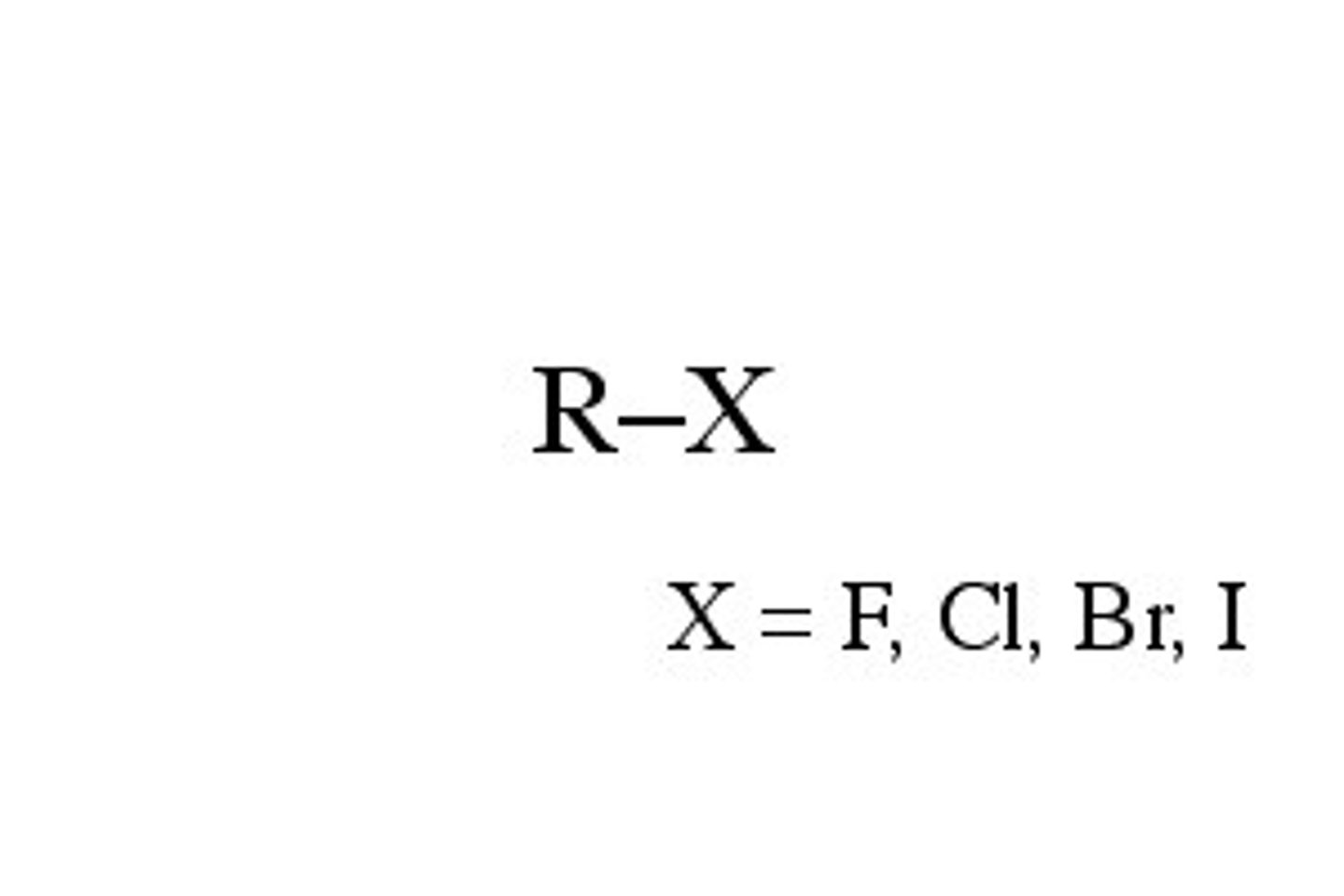
Alcohol
Carbon bonded to an OH (R-OH)
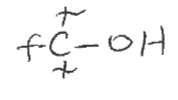
Ether
2 carbons bonded to an O on either side (R-O-R)

Primary carbon
a carbon bonded to only one other carbon

Secondary carbon
carbon bonded to two other carbons
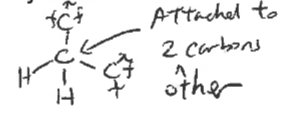
Tertiary carbon
carbon bonded to three other carbons
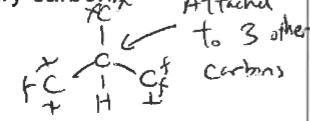
methyl carbon
carbon attached to 3 hydrogens
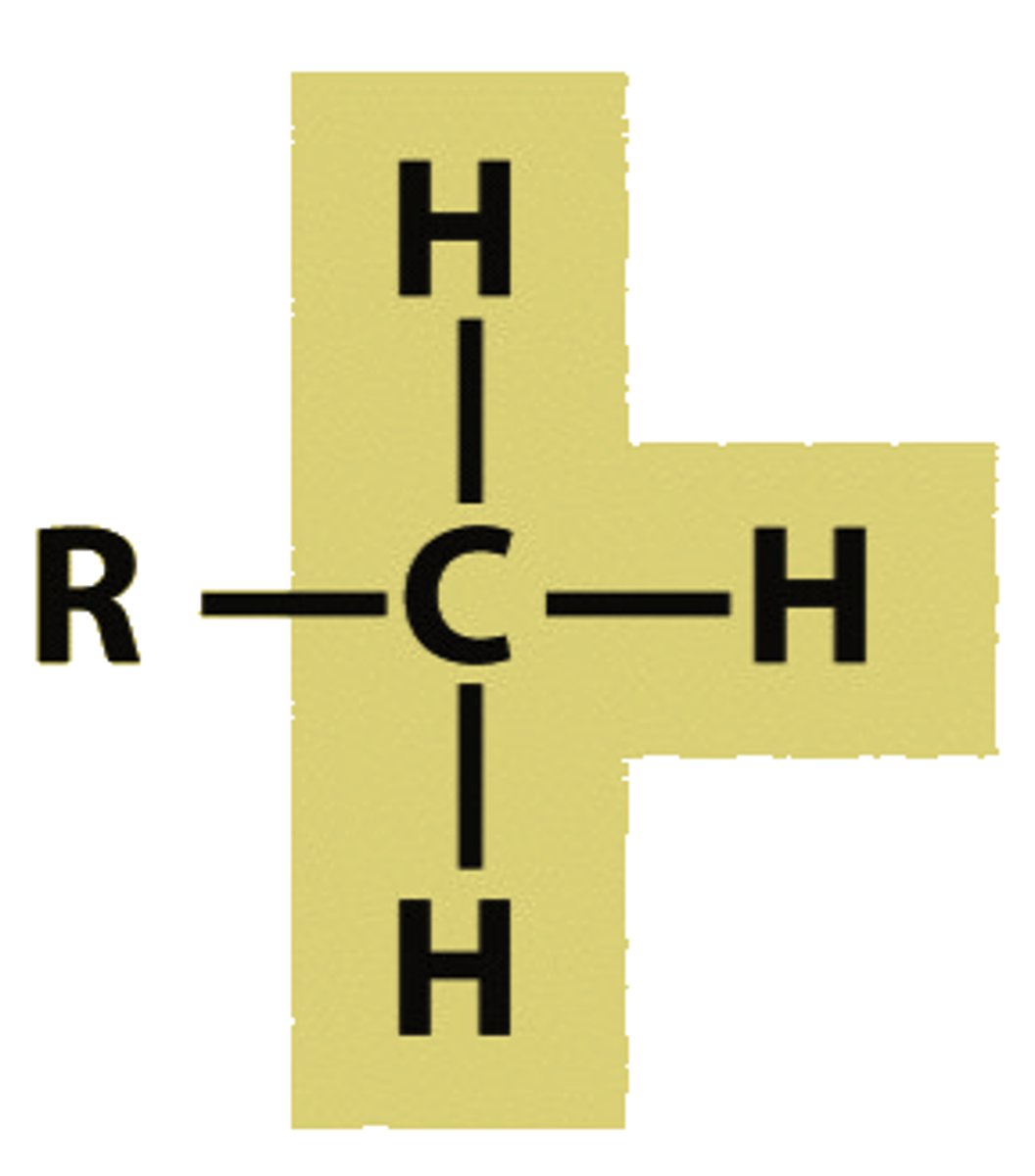
Alkanes
C--C and C--H bonds
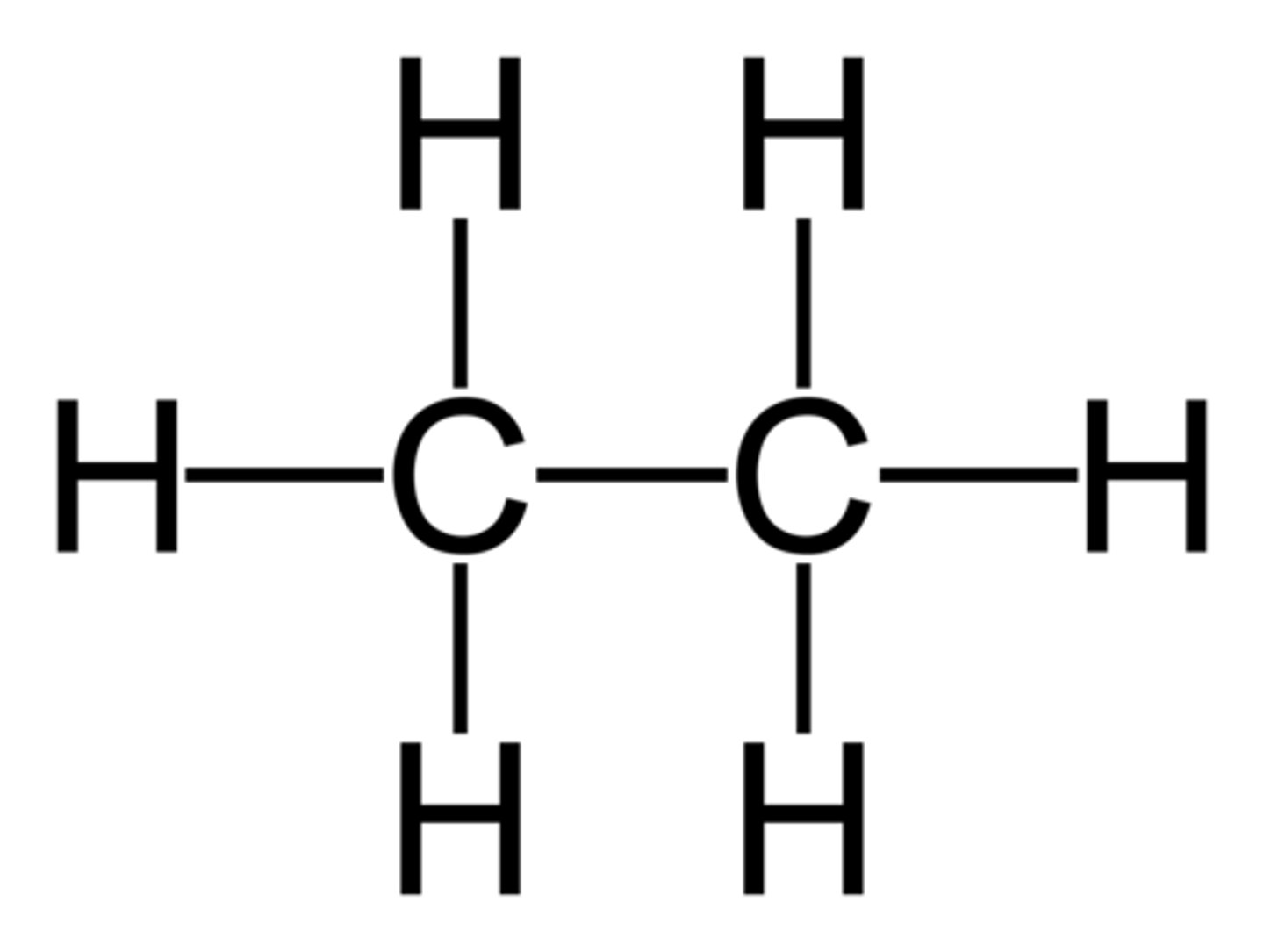
Aldehyde
type of carbonyl FG
C double bonded to O and single bonded to H
Ketone
type of carbonyl FG
3 carbons single bonded to each other, with the middle double bonded to an O
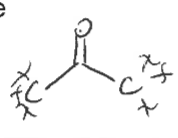
Amide
type of carbonyl FG
Carbon single bonded to N and double bonded to O
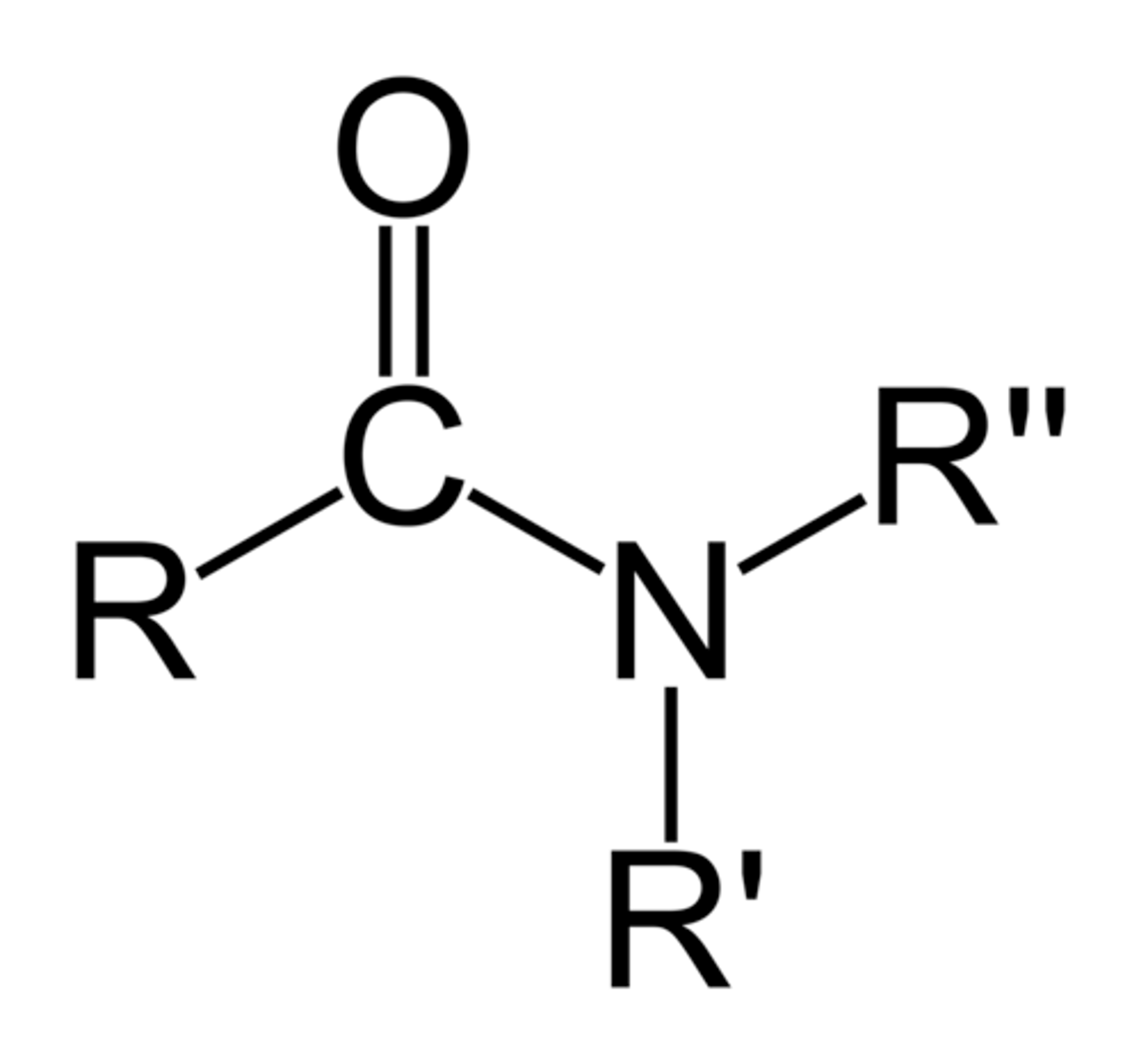
carboxylic acid
type of carbonyl FG
carbon double bonded to O, single bonded to OH
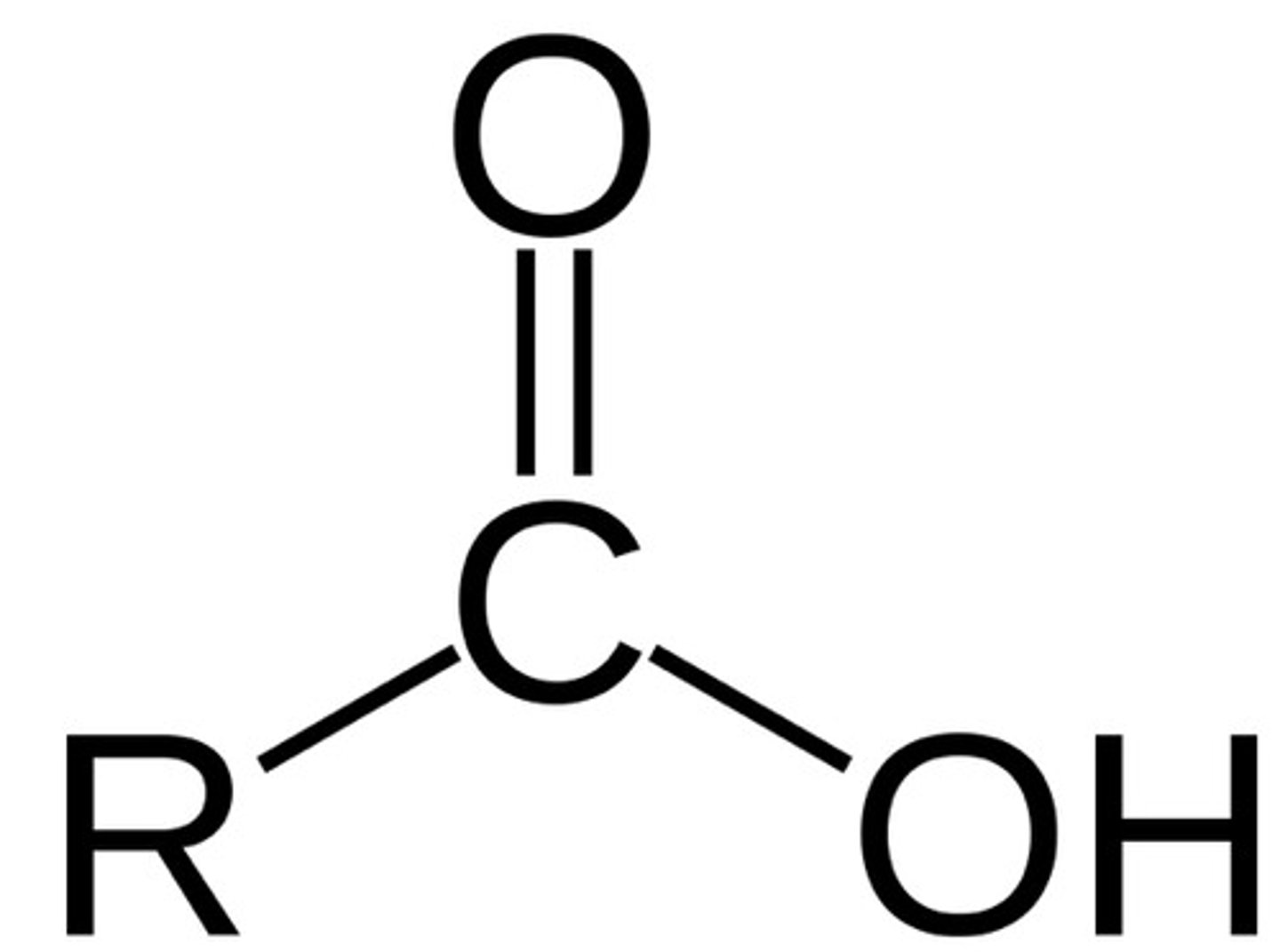
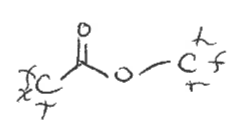
Ester
type of carbonyl FG
C double bonded to O, single bonded to O-C
Alkyl Group
A group of carbon and hydrogen atoms derived from alkanes by removing one hydrogen atom.
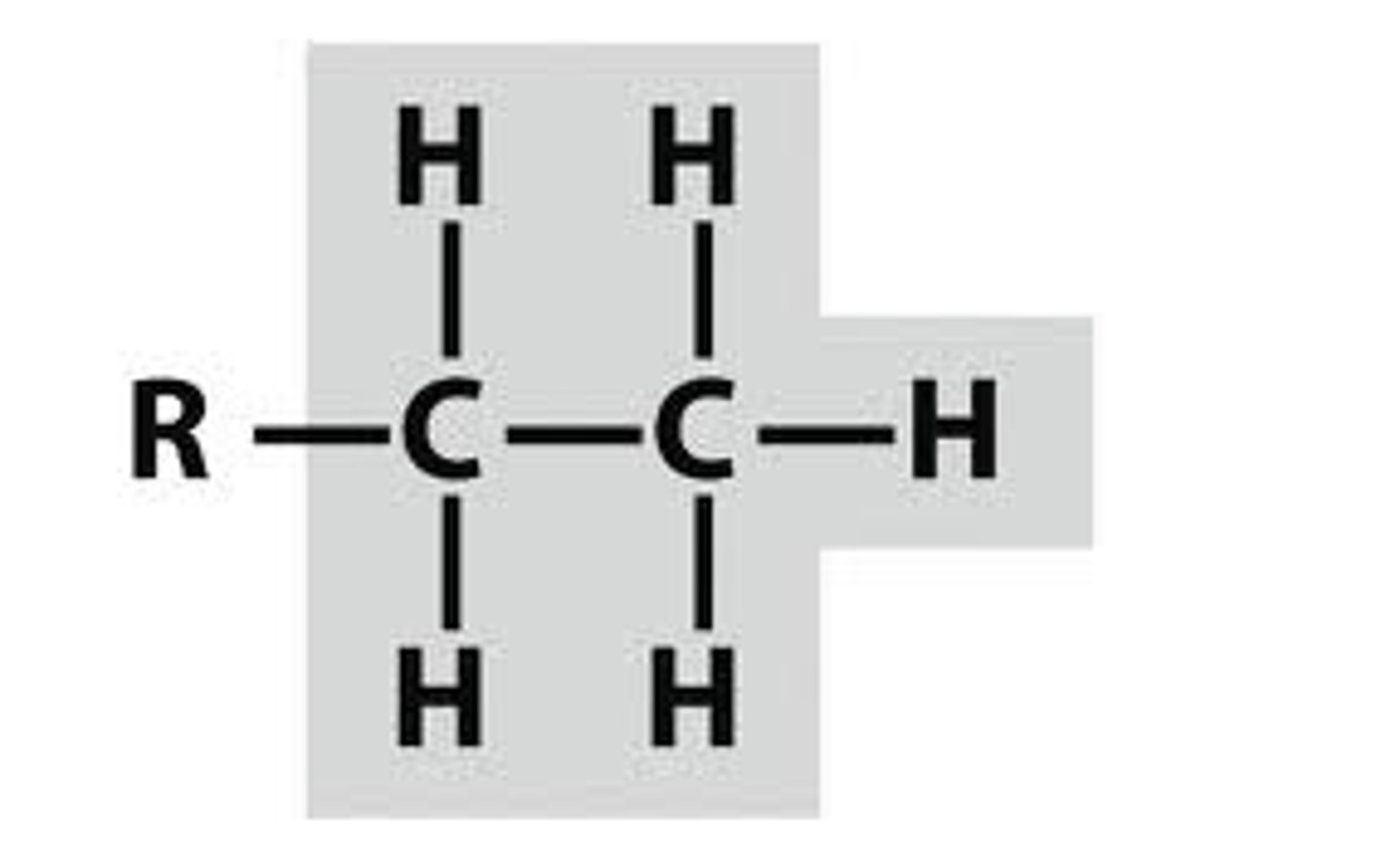
Ethyl Group
An alkyl group containing two carbon atoms and five hydrogen atoms, represented as -C2H5
Et-
Propyl Group
An alkyl group with three carbon atoms and seven hydrogen atoms, represented as -C3H7.
Pr-
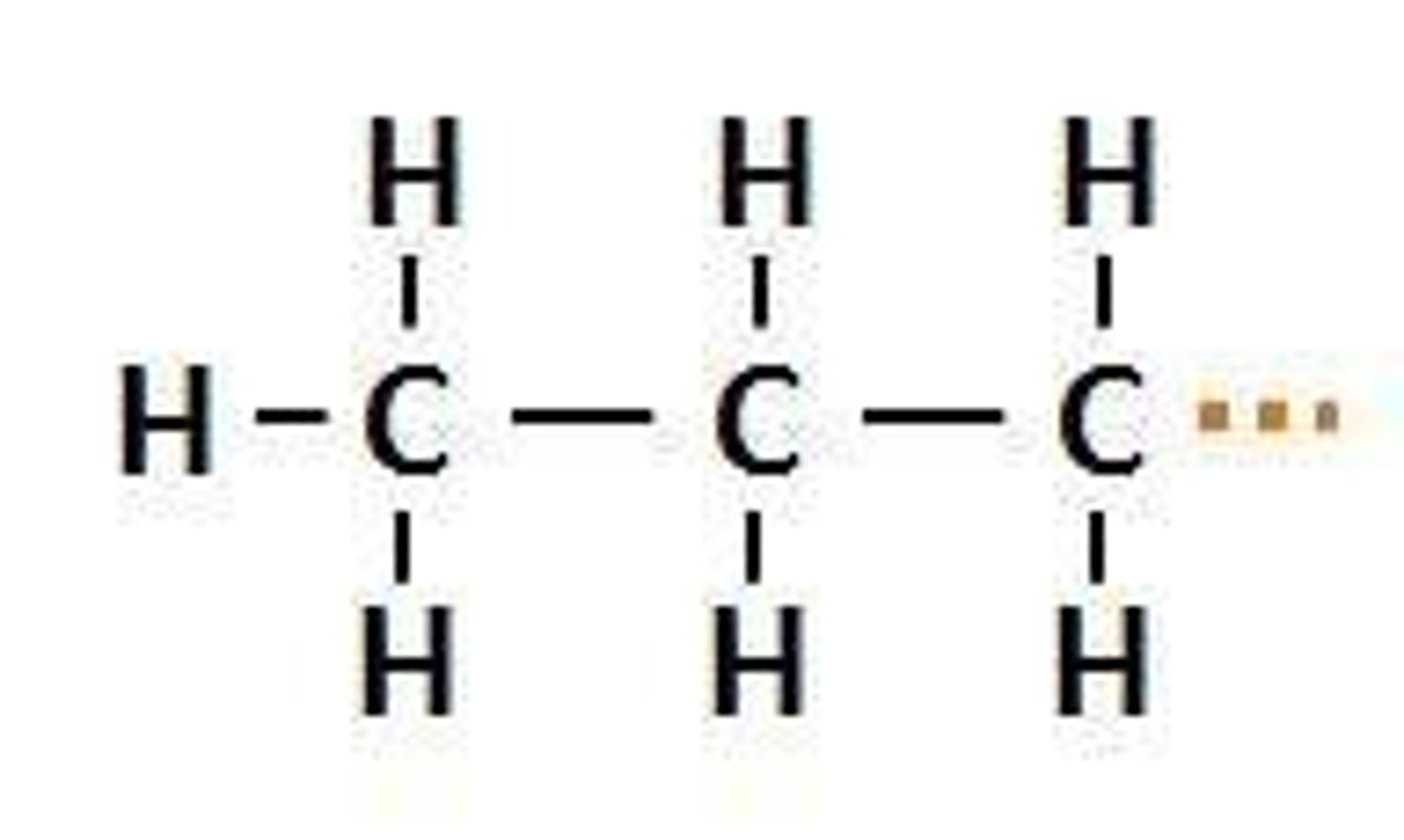
Butyl Group
An alkyl group made up of four carbon atoms and nine hydrogen atoms, represented as -C4H9.
Bu-
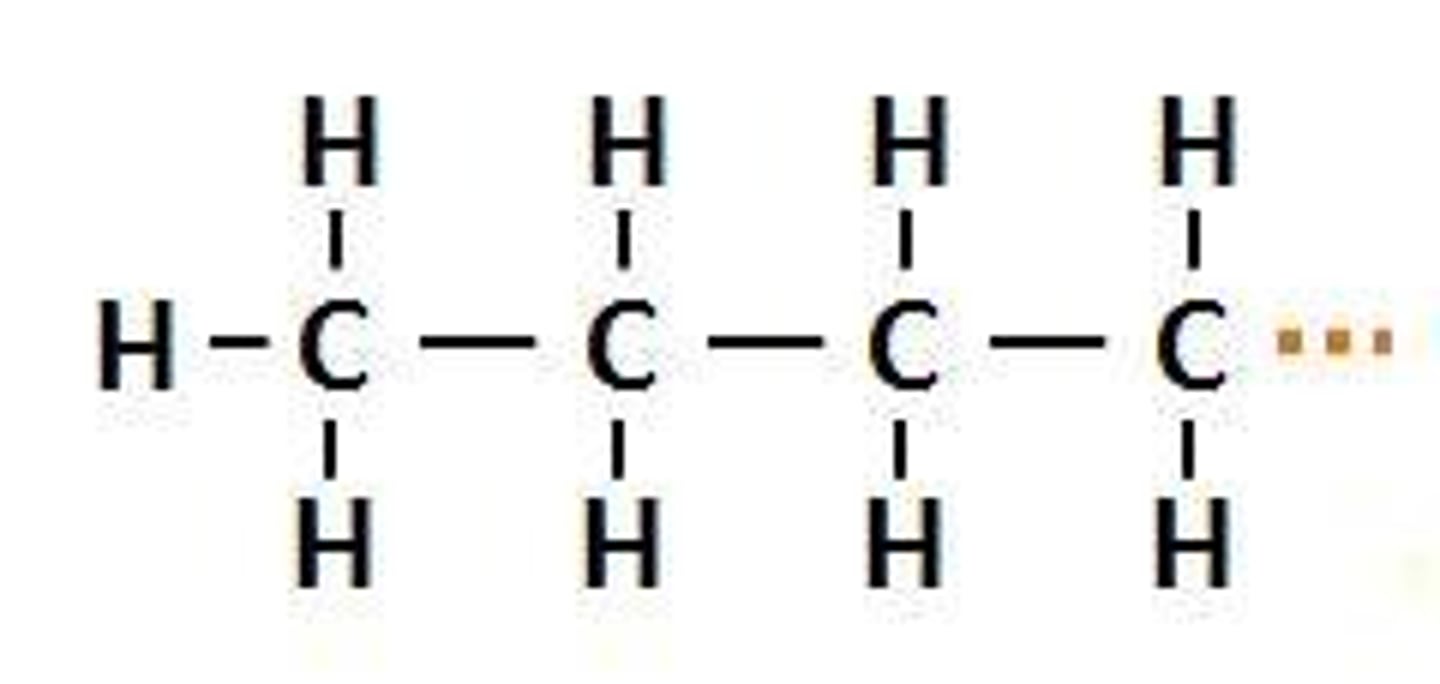
Isopropyl Group
A branched alkyl group derived from propane, with the structure -C3H7, where the central carbon is connected to two other carbons.
iPr-
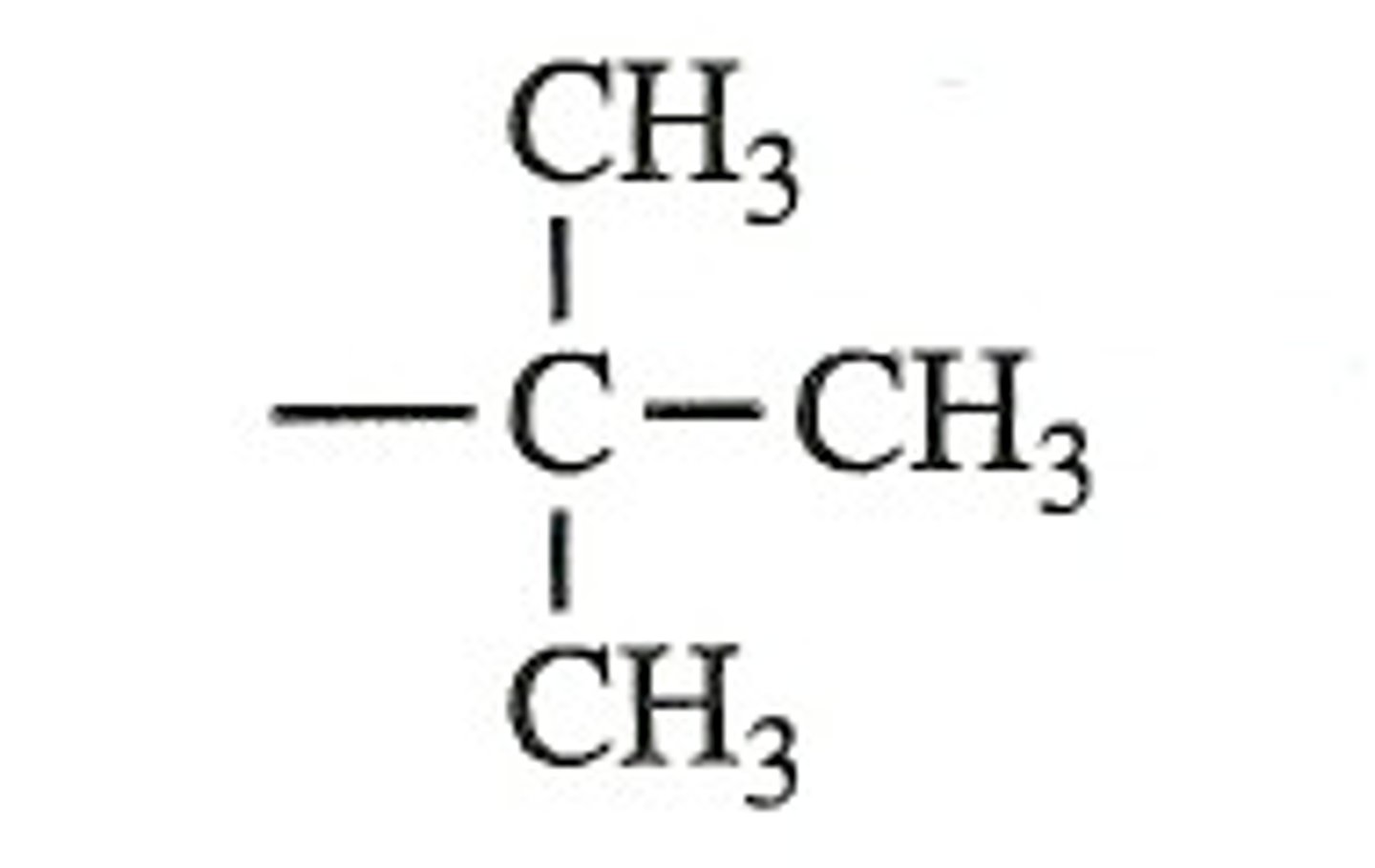
Tert-Butyl Group
A branched alkyl group consisting of four carbon atoms, with the structure -C4H9, where three methyl groups are attached to a central carbon.
tBu-
Pentyl
alkyl group made up of 5 carbons
Pe-
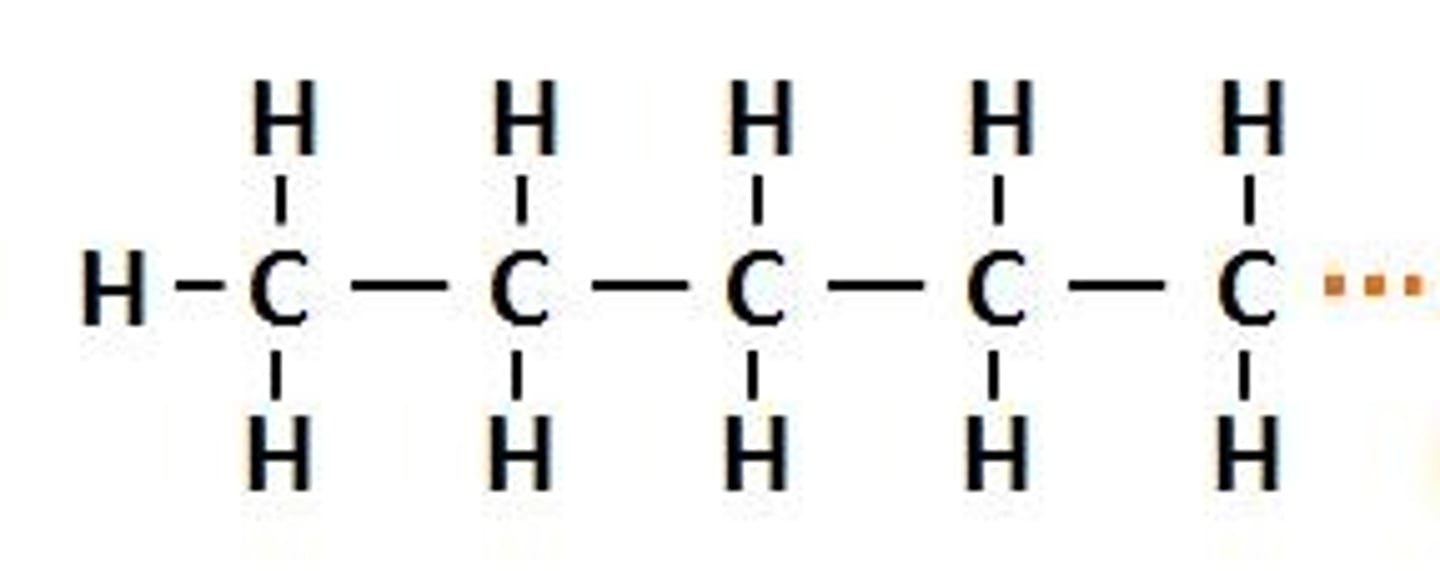
Hexyl
alkyl group made up of 6 carbons
Hex-
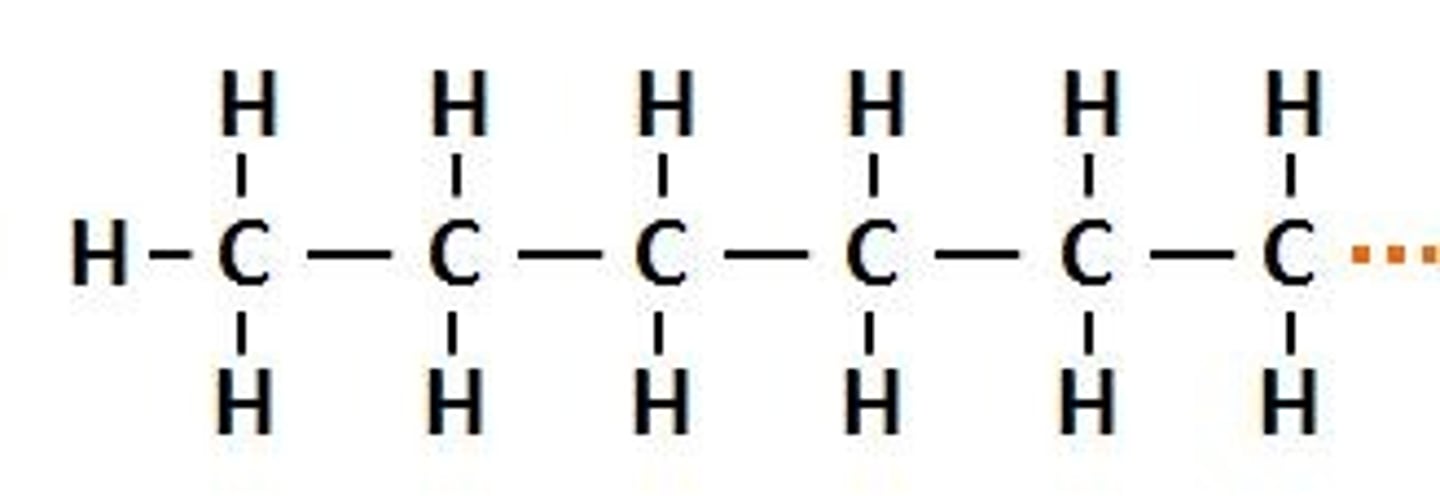
Heptyl
alkyl group made up of 7 carbons
Hep-
Octyl
alkyl group made up of 8 carbons
eclipsed
when the atoms are blocking each other from the frontal view
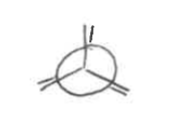
staggered
when all atoms are visible from the frontal view
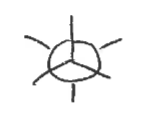
anti
when the atoms are in opposite positions of each other
(the 2 X’s)
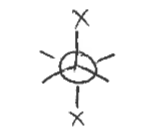
syn
when the atoms are in the same position
(the 2 X’s)
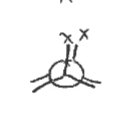
gauche confirmation
This is one of the four conformations seen in a Newman projection. This confirmation occurs when the largest substituents are 60° apart from one another. This is the second-most stable confirmation.
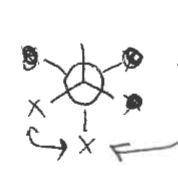
Chair confirmation
Most stable confirmation of cyclohexane
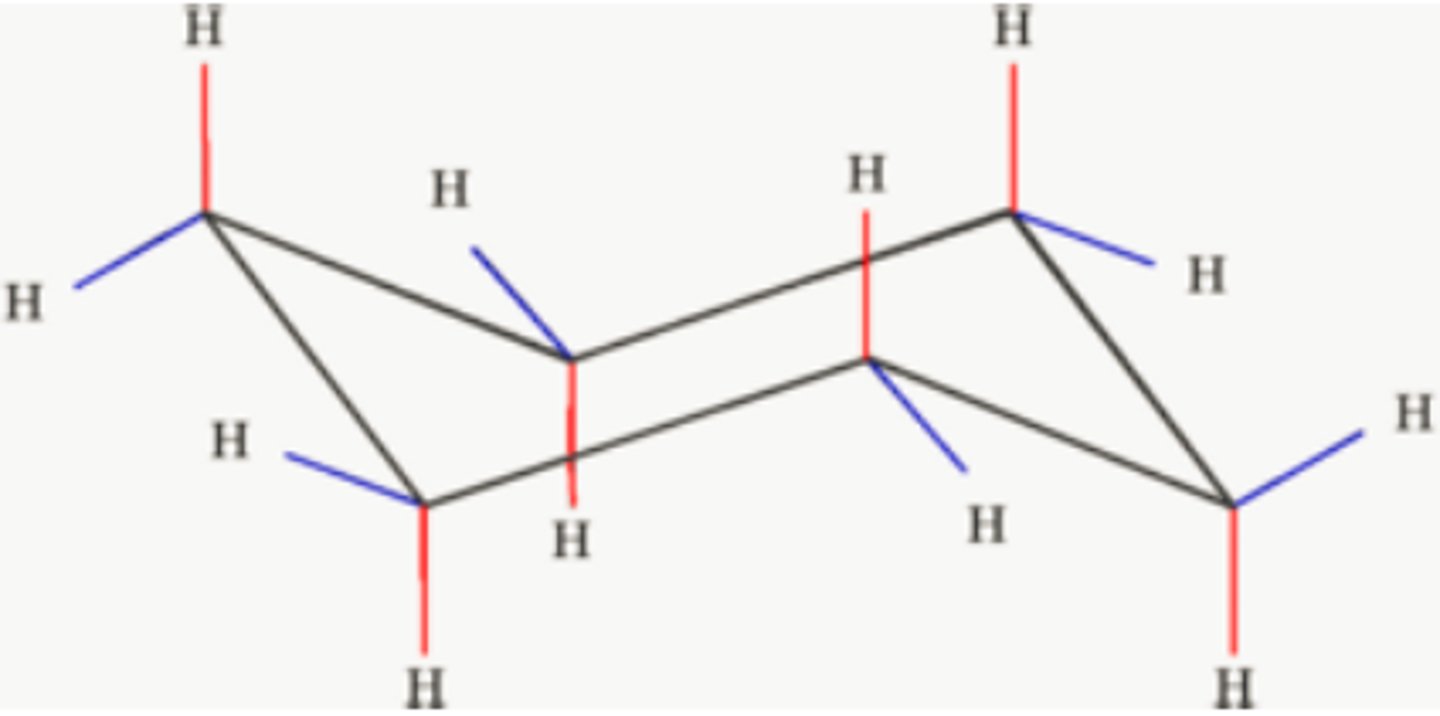
Cis orientation
2 high bonds OR 2 low bonds
(ex. Ha on C1 and He on C2 OR He on C1 and Ha on C2
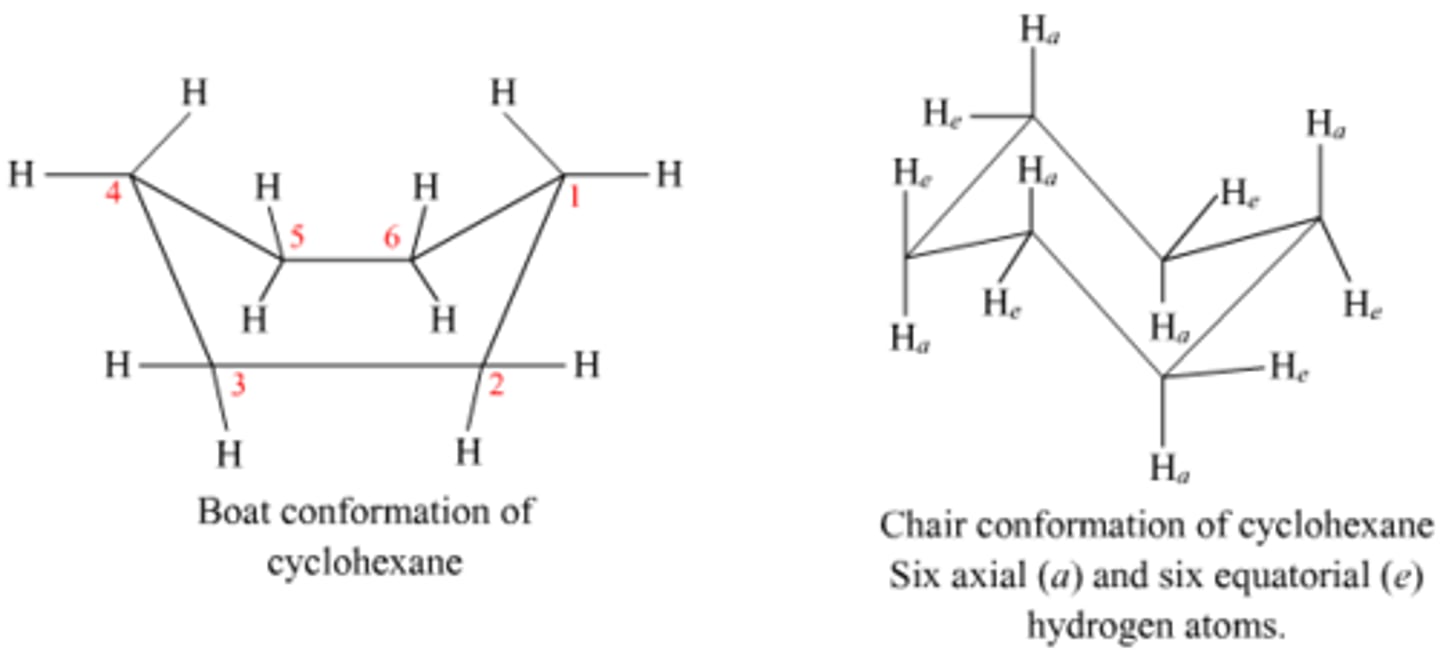
Trans orientation
1 high bond and 1 low bond
(ex. He on C1 and He on C2 OR Ha on C1 and Ha on C2)
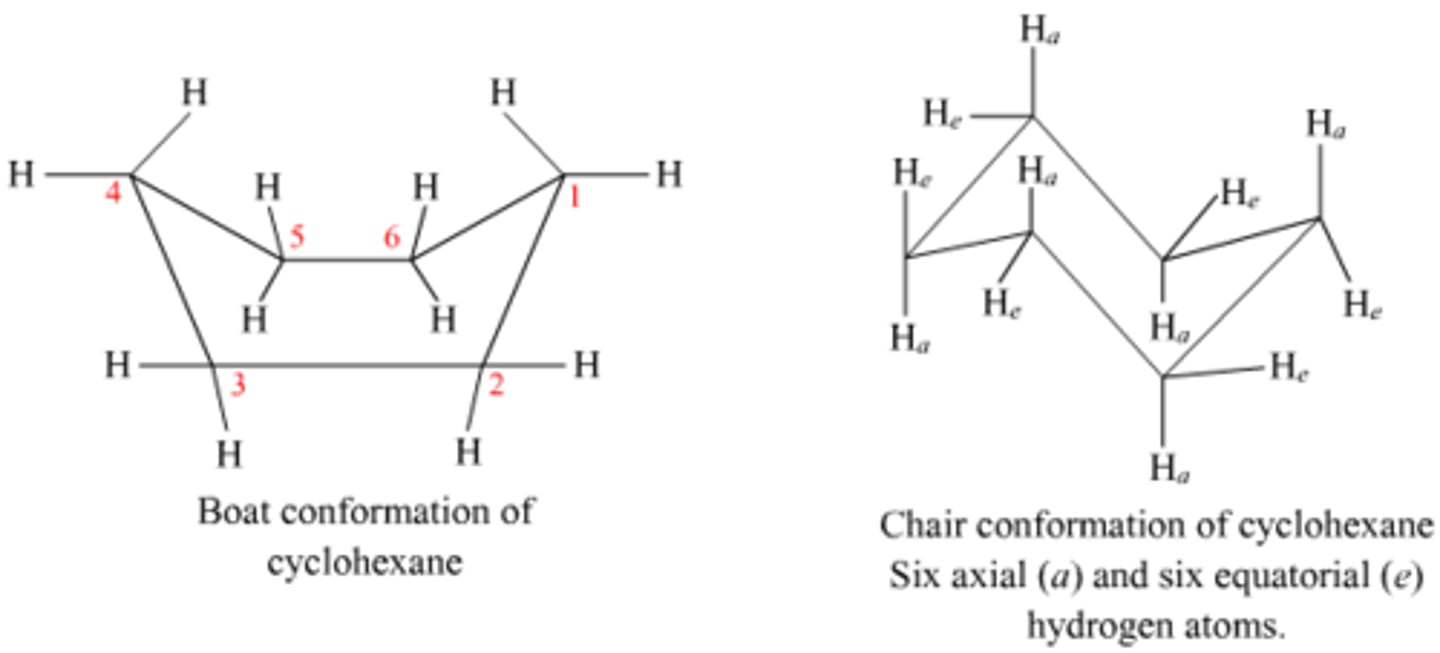
isomer
same formula different structure
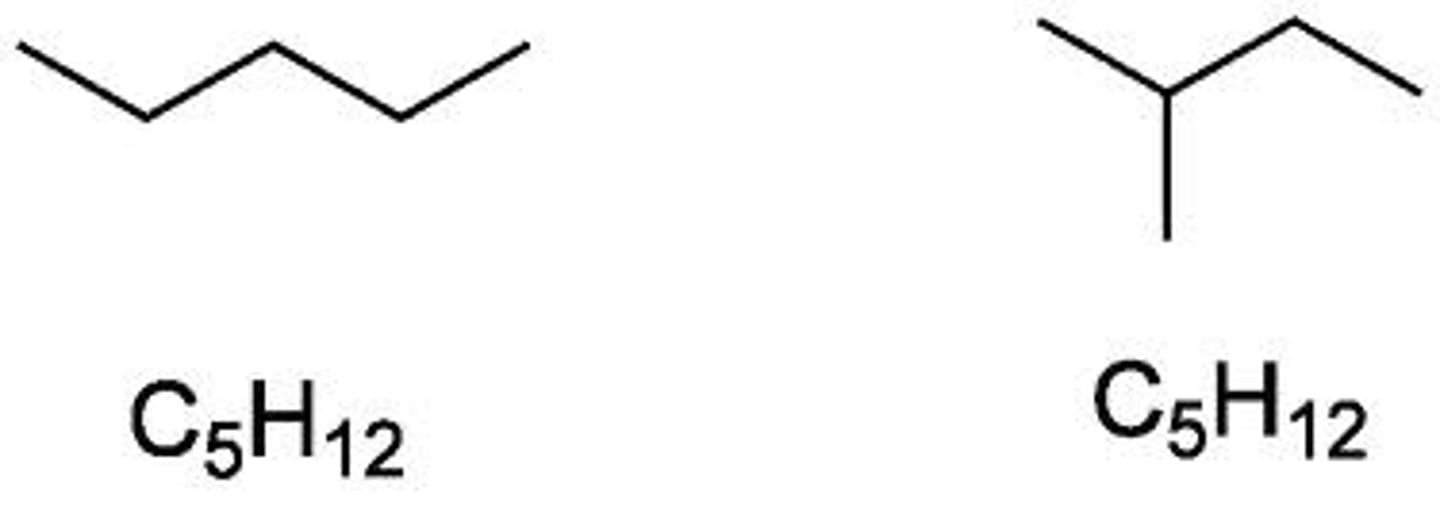
constitutional isomer
differs by atom connectivity
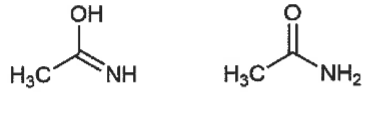
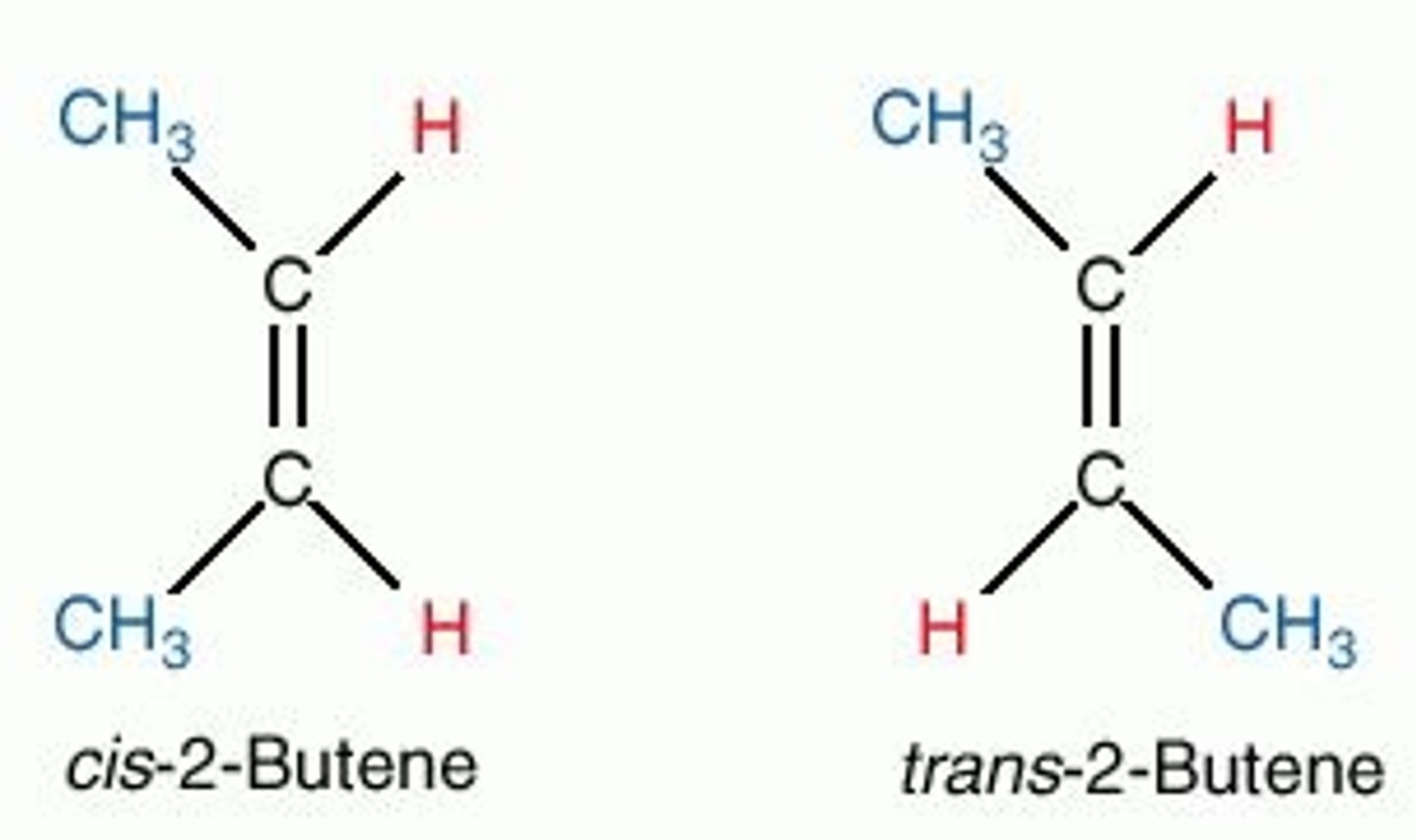
stereoisomer
different in arrangement of atoms in 3D space
superimposable
to align all parts/atoms perfectly overlapping two objects such that they match exactly in three-dimensional space.
IDENTICAL
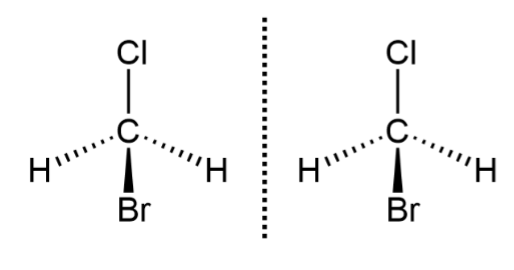
nonsuperimposable
2 mirror image molecules that can’t be placed on top of each other to align perfectly in 3D space
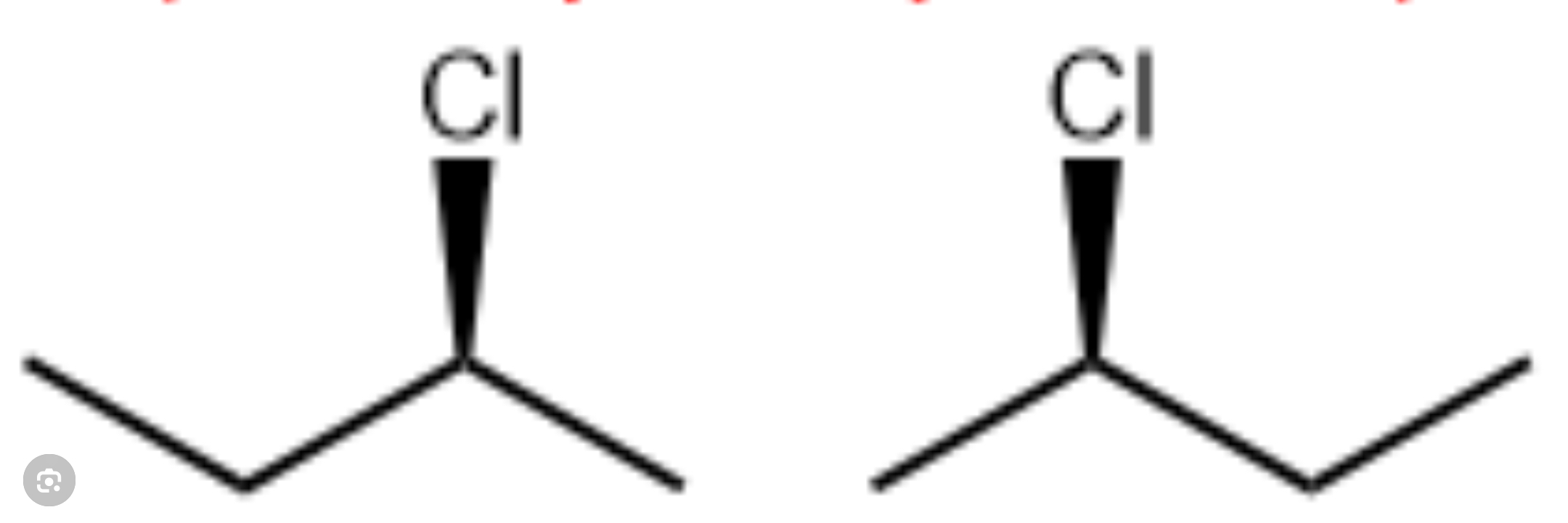
chiralty
hardedness
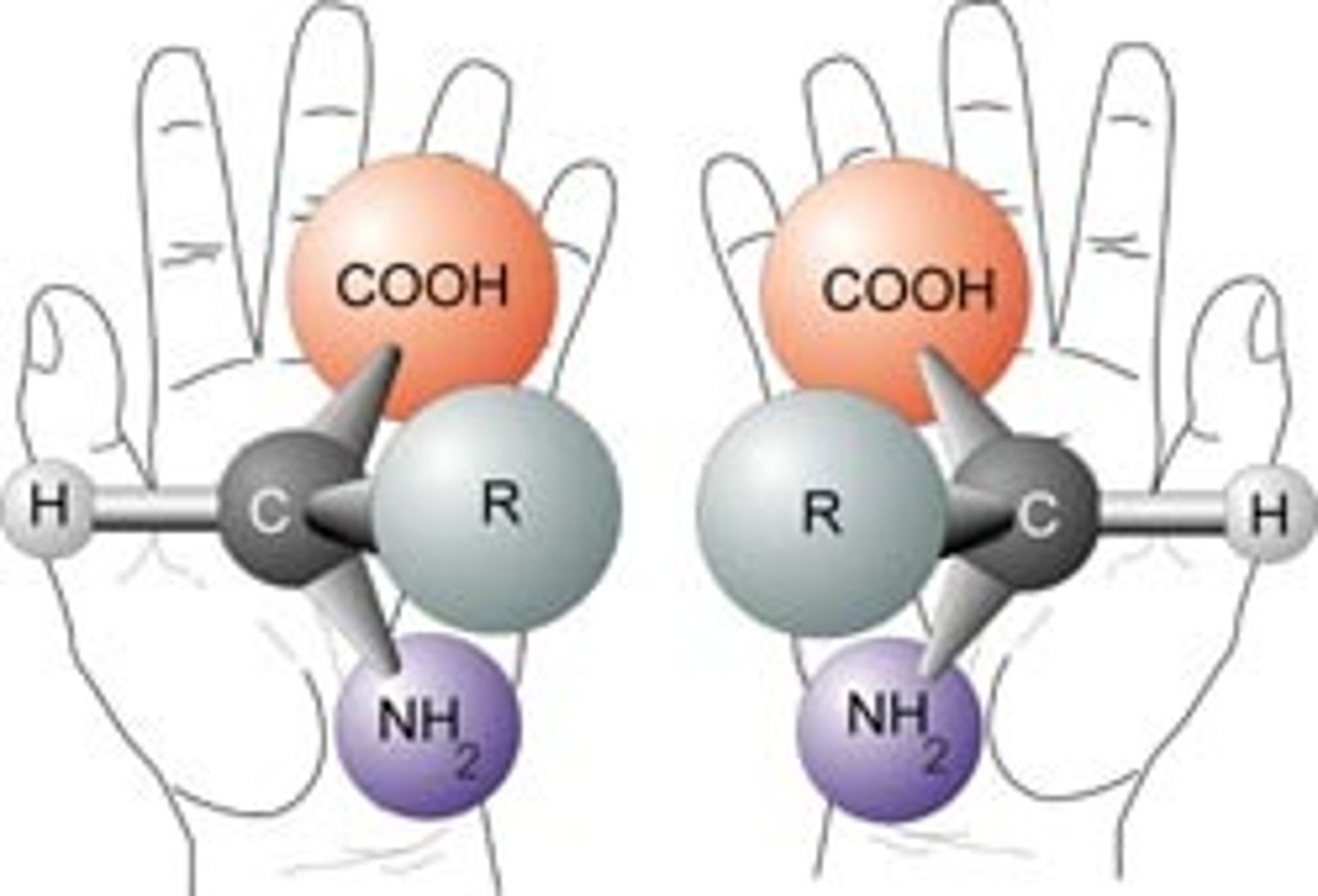
chiral
mirror image is non-superimposable
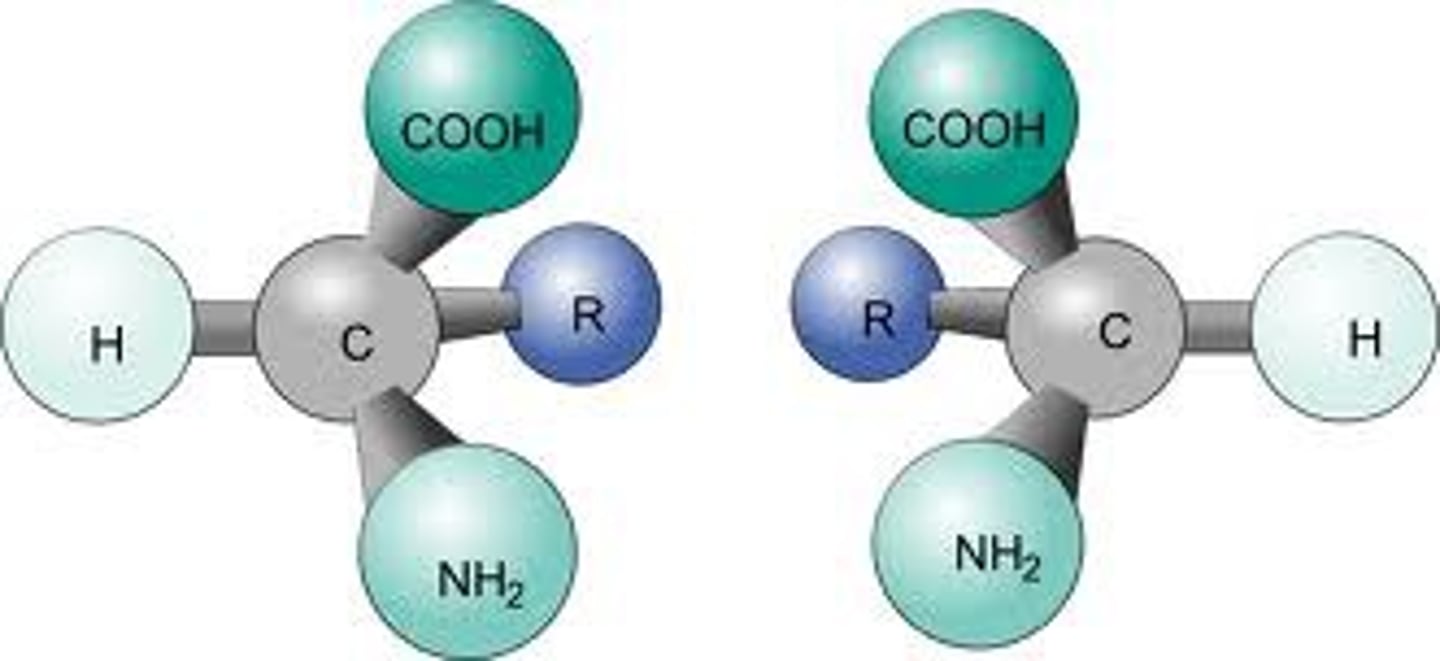
achiral
mirror image is superimposable (bottom part of pic)
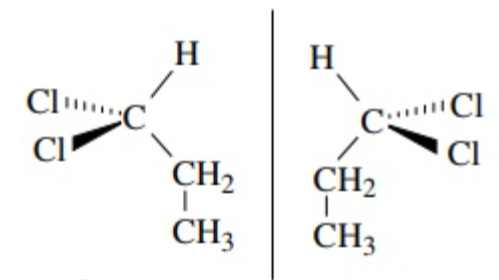
chiral center
a tetrahedral (-like, sp3) atom (C) with 4 different groups attached
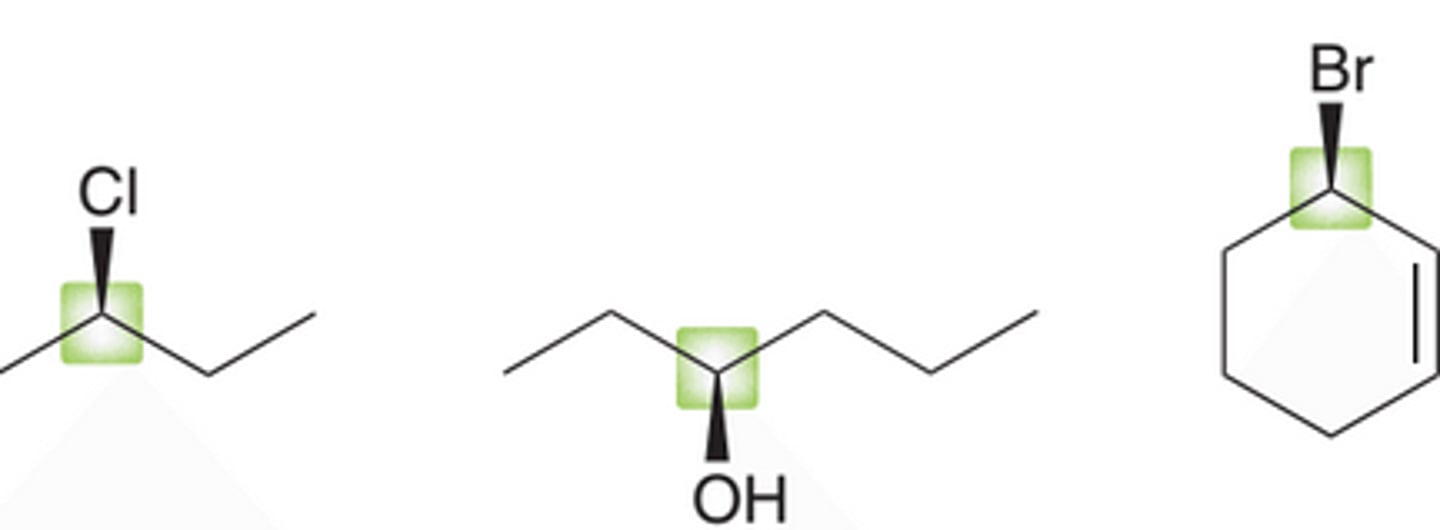
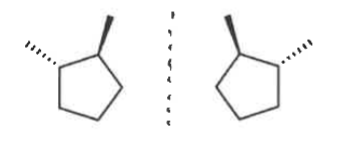
enantiomers
stereoisomers that are non-superimposable mirror images
racemic mixture
1:1 mixture of enantiomer
diastereomer
any stereoisomers that are not an enantiomer (not superimposable, not mirror images
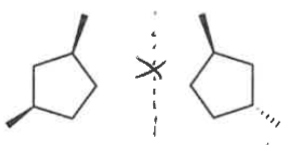
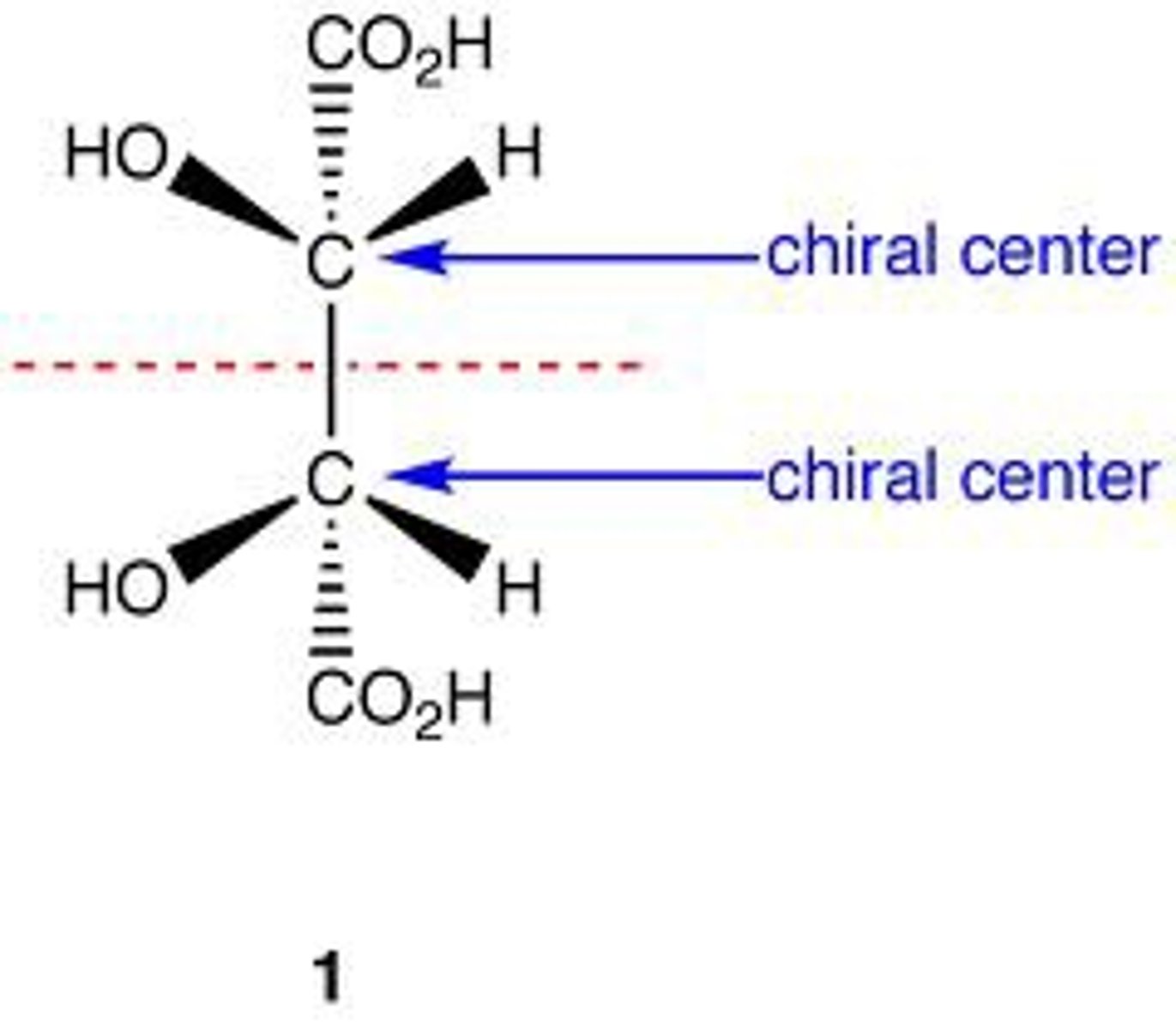
meso compound
An achiral molecule that contains 2 or more chiral centers, but has an internal mirror plane
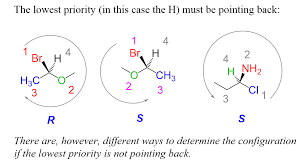
R & S Nomenclature for chiral centers
assign priority to each atom attached to the chiral center
higher atomic # → higher priority
highest priority #1, lowest #4
I > Br > Cl > S > F > O > N > C > H
if no difference at the first attached atom, move along the chain until there is a difference
multiple bonded atoms are equivalent to the same number of single bonds
orient the molecule so the lowest priority group (#4) is in the back
starting with the highest priority #1, move to #2, then #3
clockwise → R
counterclockwise → S
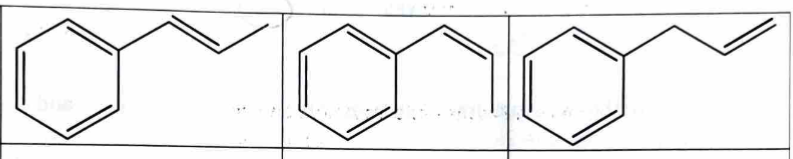
Possible Structures (of elimination products, lab 7)
looking at the double bonds:
trans
cis
terminal
MS (mass spectrometry)
ionize sample
separate and detect mass of ion [and its fragments]
detects “+” charged ions/fragments by mass (unit is M/Z)
Br detection:
1:1 M ratio → [M]+ contains Br79, and [M]2+ contains Br81
Cl detection:
3:1 ratio → [M]+ contains Cl35, and [M]2+ contains Cl37
![<ol><li><p>ionize sample</p></li><li><p>separate and detect mass of ion [and its fragments]</p></li></ol><ul><li><p>detects “+” charged ions/fragments by mass (unit is M/Z)</p></li><li><p>Br detection:</p><ul><li><p>1:1 M ratio → [M]+ contains Br<sup>79</sup>, and [M]2+ contains Br<sup>81</sup></p></li></ul></li><li><p>Cl detection:</p><ul><li><p>3:1 ratio → [M]+ contains Cl<sup>35</sup>, and [M]2+ contains Cl<sup>37</sup></p></li></ul></li></ul><p></p>](https://assets.knowt.com/user-attachments/edf612e4-a0c3-4079-86dd-d85d1558ccc7.png)
Infrared (IR) Spectroscopy
IR causes molecular vibration if the energy is at the appropriate level (energy is reported in wave numbers, in units of cm-1)
molecular vibrations depend on:
bond strength (of any given A—B bond); stronger bond, the more E it takes to move atoms ( higher E, higher wave numbers)
atom size; larger atom, lower wave number
things to look out:
C=O stretch (1780-1630cm-1)
O—H stretch (3400-2400) → carboxylic acid
N—H stretch (3500-3100) → amide
C—H stretch (2700-2800) → aldehyde
none → ketone or ester
C—H stretch
<3000 (sp3 C—H)
>3000 (sp2 [alkyne or arene] or sp C—H [alkyne])
Broad O—H stretch
3500-3200: alcohol
3400-2400 + carbonyl stretch
sharp N—H stretches (3500-3100)
2 bands= R—NH2 or R—C=O NH2
1 band= RNHR or RC=O NHR
triple bonds (2300-2100)
C triple bond C (2150) → alkyne
C triple bond N (2250
![<ul><li><p>IR causes molecular vibration if the energy is at the appropriate level (energy is reported in wave numbers, in units of cm-1)</p></li><li><p>molecular vibrations depend on:</p><ul><li><p>bond strength (of any given A—B bond); stronger bond, the more E it takes to move atoms ( higher E, higher wave numbers)</p></li><li><p>atom size; larger atom, lower wave number</p></li></ul></li><li><p>things to look out:</p></li></ul><ol><li><p>C=O stretch (1780-1630cm-1)</p><ol><li><p>O—H stretch (3400-2400) → carboxylic acid</p></li><li><p>N—H stretch (3500-3100) → amide</p></li><li><p>C—H stretch (2700-2800) → aldehyde</p></li><li><p>none → ketone or ester</p></li></ol></li><li><p>C—H stretch </p><ol><li><p><3000 (sp<sup>3</sup> C—H)</p></li><li><p>>3000 (sp<sup>2</sup> [alkyne or arene] or sp C—H [alkyne])</p></li></ol></li><li><p>Broad O—H stretch </p><ol><li><p>3500-3200: alcohol</p></li><li><p>3400-2400 + carbonyl stretch</p></li></ol></li><li><p>sharp N—H stretches (3500-3100)</p><ol><li><p>2 bands= R—NH<sub>2</sub> or R—C=O NH2</p></li><li><p>1 band= RNHR or RC=O NHR</p></li></ol></li><li><p>triple bonds (2300-2100)</p><ol><li><p>C triple bond C (2150) → alkyne</p></li><li><p>C triple bond N (2250</p></li></ol></li></ol><p></p>](https://assets.knowt.com/user-attachments/a11a19a0-6b79-44d4-a2fc-a663eb99427a.png)
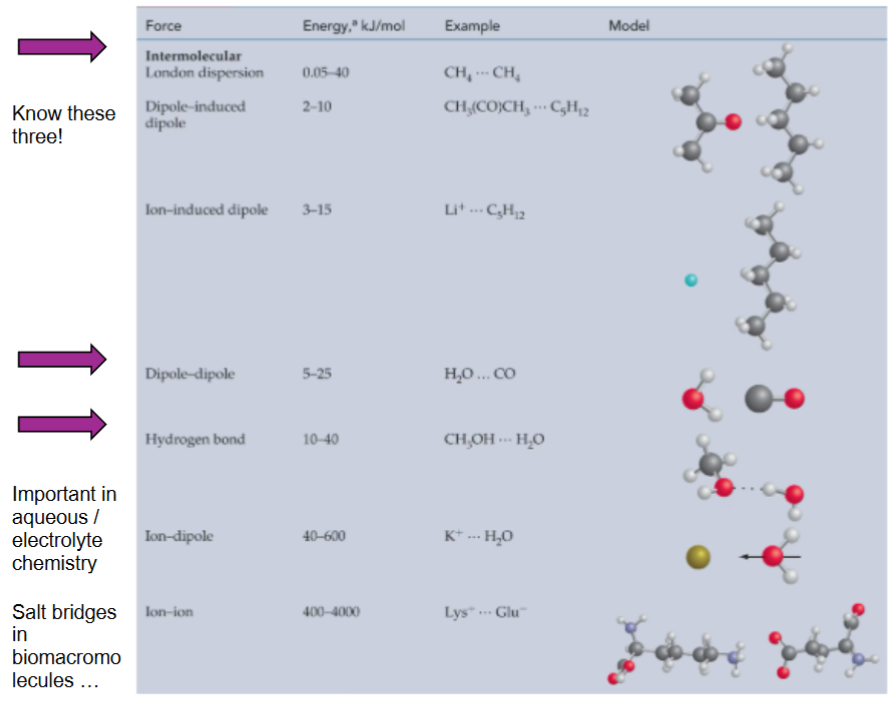
Intermolecular Forces
London dispersion forces (weak)
larger the mass, more the dispersion force, higher bp
more linear the structure, more the dispersion force, higher bp
dipole-dipole interaction (medium)
polar moelcules align according to the direction of their net dipole moments
hydrogen bonds (strong)
special type of dipole-dipole interaction involved H-atom and EN atoms
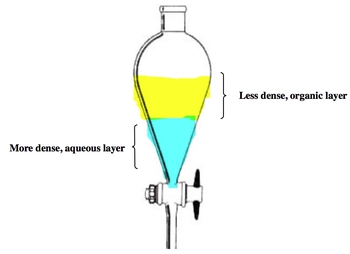
Lab 1- Aqueous Organic Extraction and recrystallization
2 solvents separate based on density (for this, it’s ethanol solution on top, water on bottom)
desired product is in aqueous layer (H2O), so it can be drained out
for this lab (based on exam 1 question), the following is correct:
top layer is aqueous HCI
after HCI addition, the species in the top layer is CH3(CH2)3NH2
bottom layer is dichloromethane (organic solvent)
after HCl addition, the species in the bottom layer is pentane
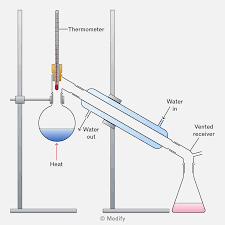
Lab 2- Distillation and Boiling Point
Compound that will be collected last is the one with a more stable structure. determine this with its ability to form hydrogen bonds with other molecules (lone pairs), maybe resonance, etc.
Blue section: where the compound is being condensed, going from a vapor to a liquid (using water), and then being collected in the tube
for boiling point:
the larger the mass, the more dispersion force, the higher the boiling point
london dispersion force- depends on size and shape
the more linear the structure, the more dispersion force, the higher the boiling point
Labs 5, 6, 10- Reaction Kinetics and Mechanism
image is for lab 5 specifically
intermediates: look at what bonds are formed/broken (stepwise rxn)
RDS: step with the highest TS
kinetics: how fast the reaction is
rate= k * [reaction A]a * [reactant B]b etc…
k= rate constant, different for each rxn
[reactant X]= amount of reactants, usually in mole or M
a, b, ..= order (number of molecules involved in RDS)
![<ul><li><p>image is for lab 5 specifically</p></li><li><p>intermediates: look at what bonds are formed/broken (stepwise rxn)</p></li><li><p>RDS: step with the highest TS</p></li><li><p>kinetics: how fast the reaction is</p><ul><li><p>rate= k * [reaction A]<sup>a</sup> * [reactant B]<sup>b</sup> etc…</p></li><li><p>k= rate constant, different for each rxn</p></li><li><p>[reactant X]= amount of reactants, usually in mole or M</p></li><li><p>a, b, ..= order (number of molecules involved in RDS)</p></li></ul></li></ul><p></p>](https://assets.knowt.com/user-attachments/faf45928-3060-4916-bc6d-109878f73317.png)
Labs 1 and 8- Product melting point analysis
purity of the isolated and dry solid can be estimated by recording its melting point on a melting point apparatus
the compound can be identified by its melting point
can be confirmed by recording the melting point of a mixture of the isolated compound and an authentic sample of the compound (a mixed melting point)
if the melting point of the mixture is the same as that recorded for the isolated compound, then the identity of the compound has been confirmed
Labs 7, 9, and 10- GC-MS
mass spec- measure the m/z of the ion ([M]+)
GC:
separate organic compounds based on volatile nature (~ boiling point)
the more volatile (~ lower BP) compounds reach the detector earlier, the less volatile the compounds later. time when the detector detects the compound is retention time (RT)
quantify of the compound = the “area-under-the-curve” of the peak
MS provides spectrum of abundance vs mass of ions generated during the analysis of the compound
each compound will fragment in a pettern unique to that compound
GC chromatogram: graph of abundance (total ion count) vs time (top of image)
RT of the peak identifies pure compounds
integration of each peak (area % of total) can be used to determine the relative percent composition of each compound in mixture
Mass spectra: abundance vs mass (mass/charge ratio) (bottom of image)
each GC chromatogram peak will have a separate mass spectra
the likely molecular weight and fragmentation pattern can be used for product identification
![<ul><li><p>mass spec- measure the m/z of the ion ([M]+)</p></li></ul><p>GC:</p><ol><li><p>separate organic compounds based on volatile nature (~ boiling point)</p></li><li><p>the more volatile (~ lower BP) compounds reach the detector earlier, the less volatile the compounds later. time when the detector detects the compound is retention time (RT)</p></li><li><p>quantify of the compound = the “area-under-the-curve” of the peak</p></li></ol><ul><li><p>MS provides spectrum of abundance vs mass of ions generated during the analysis of the compound</p><ul><li><p>each compound will fragment in a pettern unique to that compound</p></li></ul></li><li><p>GC chromatogram: graph of abundance (total ion count) vs time (top of image)</p><ul><li><p>RT of the peak identifies pure compounds</p></li><li><p>integration of each peak (area % of total) can be used to determine the relative percent composition of each compound in mixture</p></li></ul></li><li><p>Mass spectra: abundance vs mass (mass/charge ratio) (bottom of image)</p><ul><li><p>each GC chromatogram peak will have a separate mass spectra</p></li><li><p>the likely molecular weight and fragmentation pattern can be used for product identification</p></li></ul></li></ul><p></p>](https://assets.knowt.com/user-attachments/d61d0a9f-b373-420b-95e5-f44452aa890d.png)
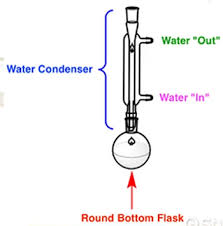
Lab 11- Reflux setup, MS fragmentation
molecular fragments: fragmentation gives the most stable carbocation/carbon radical
for this lab, either 1 degree alcohol or 2 degree alcohol as possible products
reflux setup (image): condenser traps solvent vapors by cooling them and forcing the condensed solvent back into our rxn flask
observe rxn mixture to determine when the process of reflex starts (condensation around the side of the round bottom flask)
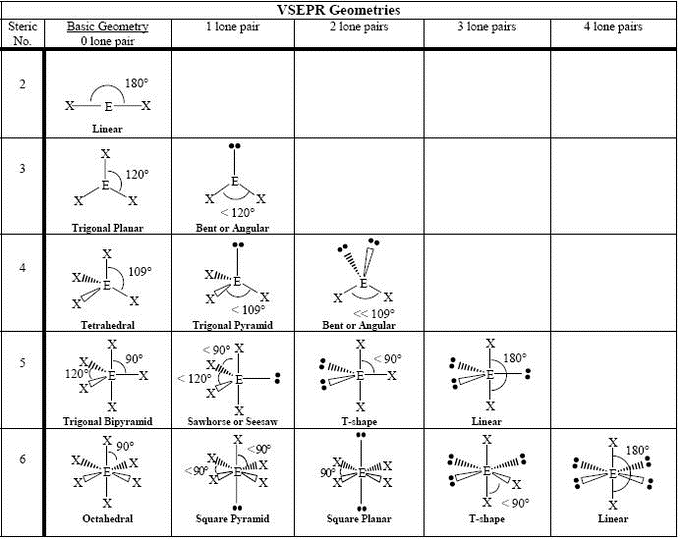
VSPER
4e- domain: tetrahedral-like
angle (tetrahedral)- 109 degrees
3e- domain: trigonal planar-like
angle (trigonal planar)- 120 degrees
2e- domain: linear angle 180 degrees
double/triple bonds count as a 1e- group
Polarity of chemical bonds
EN: ability of an atom to compete for e- with other atoms
quantified between 0.7 and 4.0 (figure will be on exam)
less than 0.5 difference= nonpolar
0.5-2.0= polar
2.0+= ionic
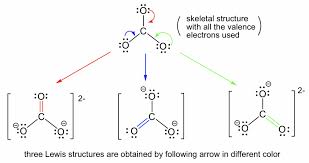
resonance structure
if a compound contains less than or equal to 2 RS, then none of the individual RS lewis structures represent the TRUE structure of the compound
true structure- hybrid of all of them
best RS → closest to the true structure
2nd best RS → hints at reactivity of compoudn
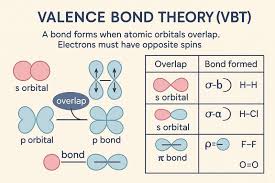
valence bond theory
atomic orbitals can undergo hybridization to generate hybrid orbitals
# of hybrid oribtal = # of atomic orbitals combines for hybridization
2p orbitals are 90 degrees with respect to each other
single bonds between atoms- sigma bond
pz orbitals= double/triple bonds, which are pi bonds
double bond=1 pi bond (1 pz)
triple bond= 2 pi bonds (1 py and 1 pz)
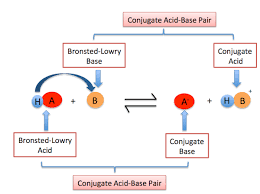
Bronsted acid-base
acid= any species that donates a proton (H)
base= any species that accepts a proton
equilibrium arrow- species on both sides of the equation are present
if equilibrium favors product/right side- strong acid, large Ka, and small/lower pKa
if equilibrium favors reactant/left side- weak acid, small Ka, and large/higher pKa
general trends:
conjugate acid with higher pka gets the proton
stronger acid HA makes a weak base A-, and a weak acid HA makes a strong base A-
examples:
acid- water, HCl
base- water, methoxide anion, azide anion
Strengths of different acids
The more stabilized A- is, the more willing HA is to give its H+
EN and induction effect
looking at the row
ex: F > O > N > C
F- > OH- > NH2- > CH3-
F > H
polarizability and size: large atoms can diffuse e- density over a larger atomic radius
looking at column (comes first in determining stronger acid, ie I vs F)
ex: I > Br > Cl > F (stability of I- > Br- > Cl- > F-)
S > O / H3CS- > H3CO-
resonance: delocalization of ‘-’ charge and/or e- to more evenly share the “burden”
hybridization: higher % of s-character in hybrid orbital, then more stabilized e- density (sp3, sp2, sp)
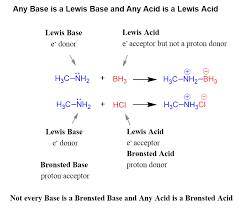
Lewis Acid-Base reactions
acid (LA): electron acceptor
e- deficient/poor → electrophile (E+)
base (LB): electron donor
e- rich → Nucleophile (Nu)
in general, this rxn involves formation of a new covalent bond
all Bronsted acids/bases are Lewis acids/bases, BUT, not all lewis acids/bases are bronsted acids/bases
examples:
LA- water, BF3, Na+, Trityl cation, HCl
LB- water, methoxide anion, azide anion
strengths of acids and bases:
nucleophilicity- tendency of Nu to form a (covalent) bond with E+ (often trends qualitatively with the pKa of their conjugate bronsted acid)
electrophilicity- tendency of E+ to form a (covalent) bond with Nu
E1 reactions
weak-moderate base
good LG
electrophile: 3 > 2
polar solvent
1st order kinetics
stereochemistry: E and Z isomers
rearrangement common
2 steps
E2 reactions
strong base (KOtBu, LDA, also heat)
good LG
3 > 2 > 1
polar solvent
2nd order kinetics
stereochemistry: anti-coplanar geometry
no rearrangement
1 step
SN1 reaction
just needs any nucleophile
good LG
3 > > 2 electrophile
polar protic solvent good (water, ethanol)
1st order kinetics
stereochemistry: racemic mixture
common rearrangement
2 steps
SN2 reaction
strong nucleophile needed
good LG
electrophile: methyl > 1 > 2
polar aprotic solvent good (DMF, DMSO)
2nd order kinetics
stereochemistry: inversion of configuration
no rearrangement
1 step
Nucleophile properties
anionic Nu stronger than neutral Nu (OH- stronger than H2O)
periodic trends → nucleophilicity
increases right to left
more EN, worse Nu
less EN, good Nu
increases top to bottom
size and polarizability
LG properties
e- withdrawing group, ie polarizable C—LG bond
LG- needs to be stable on its own after it leaves, via resonance or polarizability
common LGS: leave as anion (Cl-, Br-, etc.) OR neutral molecule (H2O, etc.)
poor LG: OH, C—OH, OR-, NR2-, CH3-, H-, F-, etc.
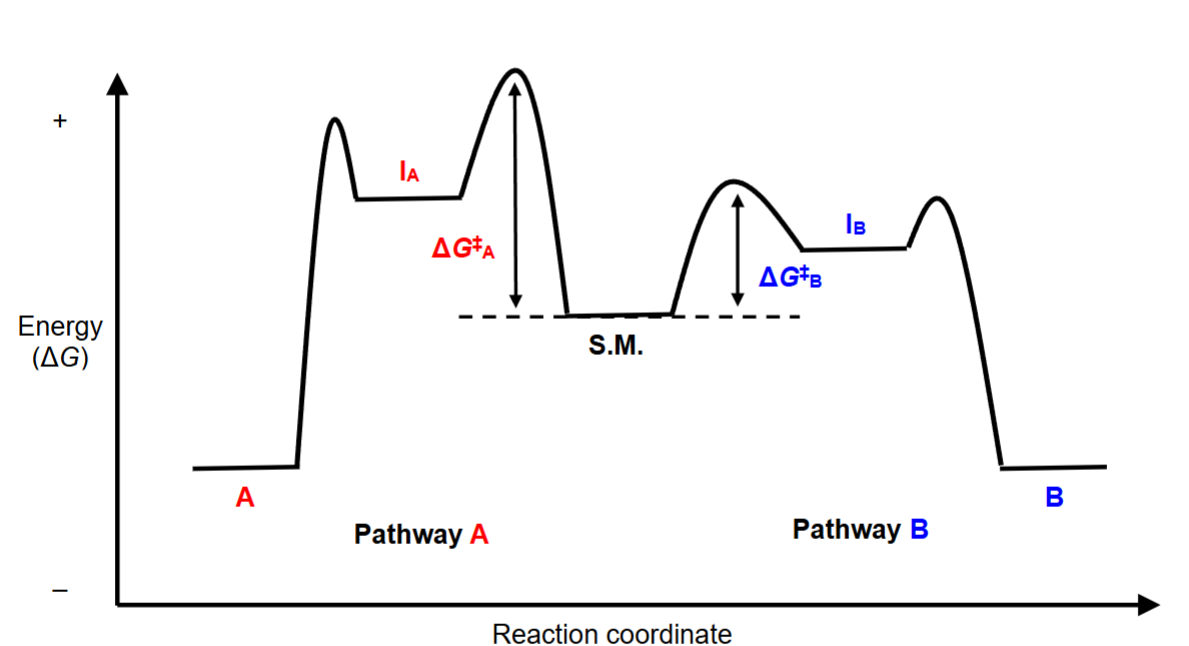
Reaction Coordinate Diagram
analyzing the image:
Pathway A vs Pathway B, 2 different products and 2 intermediates
IB is lower in energy (more stable) than IA
delta GB is lower energy than delta GA
therefore, Pathway B is the faster and preferred rxn pathway, which forms the most stable and lowest energy intermediate
transition state: high energy point where bonds are being formed/broken (NOT INTERMEDIATE)
thermochemistry: delta G ~ delta H
if delta H < 0, rxn is exothermic/favorable
if delta H > 0, rxn is endothermic/not favorable
Stabilizing Carbon reactive intermediates
resonance: donation of e- density from adjacent lp or pi bond
hyperconjugation: donation of e- from adjacent sigma bonds (sigma bonds adjacent to empty p-orbitals can share e- density)
3 > 2 > 1 > methyl (most hyperconjugation → least hyperconjugation AND most stable intermediate → least stable intermediate
free radical substitution reactions
substitution reactions
one-step sub rxn (concerted)
RDS: only one step - A—B C- → A- B—C
rate law: rate= k[A—B]1 * [C-]1
total order: 2, 2nd order
multi-step sub rxn (stepwise)
A—B C- → A- B+ C- → A- B—C
scenario 1: Step 1 is RDS
rate= k[A—B]1
total order: 1
scenario 2: step 2 is RDS
rate= k[B+]1 [C-]1
total order: 2
Addition reactions
syn addition (X—Y added at same time)
anti addition (X—Y added diff time through intermediate bridge)
random addition (X—Y added at dff time through planar intermediate [C cation or radical])
![<ol><li><p>syn addition (X—Y added at same time)</p></li><li><p>anti addition (X—Y added diff time through intermediate bridge)</p></li><li><p>random addition (X—Y added at dff time through planar intermediate [C cation or radical])</p></li></ol><p></p>](https://assets.knowt.com/user-attachments/4ccfcefb-014d-42bf-987b-c494714fe625.png)
Markovnikov’s Rule
when H—A is added to C==C, the H+ adds to the least substituted carbon (carbon with the most H)
C with most H gets 1 more H