T1 Fricker- Regulation of the cell cycle + Mitosis
1/14
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
15 Terms
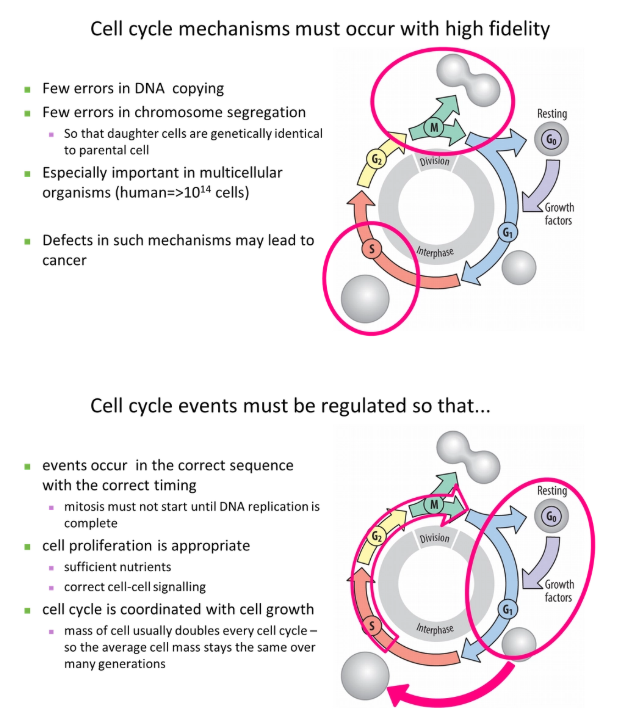
why must the cell cycle occur with high fidelity and be tightly regulated?
eg. parts of metabolism might be stopped during dividision to reduce mutations eg. to prevent the production of reactive oxygen species
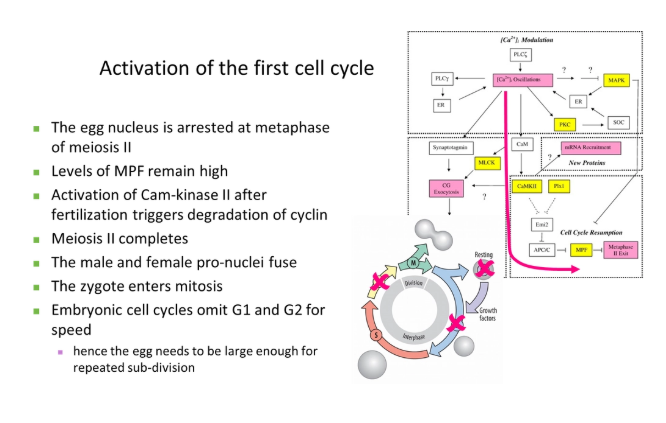
how is the first cell cycle activated?
MPF = maturation promoting factor (composed of the catalytic subunit cyclin dependent kinase and the regulatory subunit cyclin B)- conserved in all eukaryotes but have different names
this was chemically identified using temperature-sensitive mutations + recombination with different plasmids containing wild type genes
G1 and G2 can be omitted in the early embryo because it is faster, and the egg cell already contains loads of maternal resources for sub-division
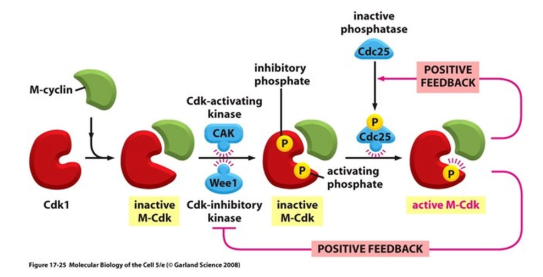
how is the cell cycle regulated?
the maturation promoting factor (MPF) is composed of the catalytic subunit cyclin dependent kinase (Cdk) and the regulatory subunit cyclin B
to progress into certain stages of the cell cycle, Cdk has to bind to the relevant cyclin to partially activate it
full activation of Cdk is also dependent on phosphorylation by Cdk-activating kinase (CAK), which is only carried out dependent on cellular conditions eg. sufficient size + lack of DNA damage
this allows the transition into the next stage by the phosphorylation of different target proteins
this cyclin is then degraded, deactivating Cdk- this is a ratchet that prevents the cycle moving backwards
Cdk activity is under additional regulation, allowing control at different checkpoints:
inhibitory phosphates can also be added to Cdk to block its active site and prevent progression into the next stage
additional Cdk inhibitor (CKI) proteins can block its activity eg. these get transcribed following detection of DNA damage
positive feedback loops in this process cause massive amplification to activate multiple Cdks, including those that degrade the cyclins
different cyclins (and in mammals, different Cdks) function at different stages of the cell cycle, dependent on the species
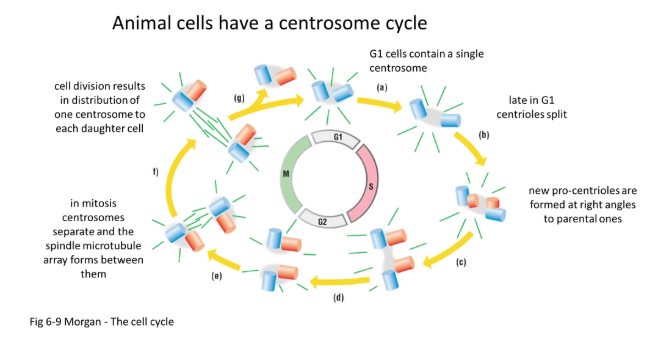
what is the centrosome cycle?
in animals:
during the S phase, the pair of centrioles (perpendicular to each other) self-replicate to form two pairs of daughter centrioles
they begin to produce spindle fibres
during prophase they begin to move to the poles by pushing away from each other using antiparallel spindles (plus ends together)
following mitosis, one centrosome (pair of centrioles) is distributed in either daughter cell
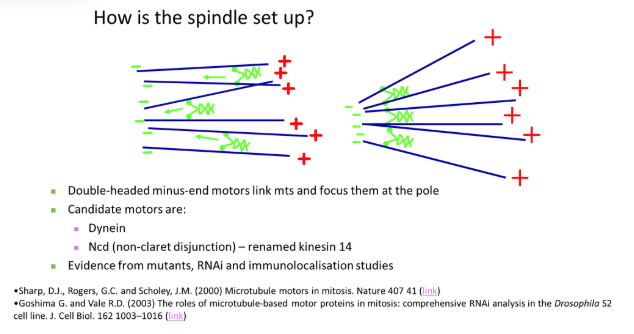
how are spindle fibres set up?
double-headed minus-end motors link adjacent microtubules together, drawing them in to focus at the minus pole
plus-end motors are anchored at the centrosome to tether the focus pole
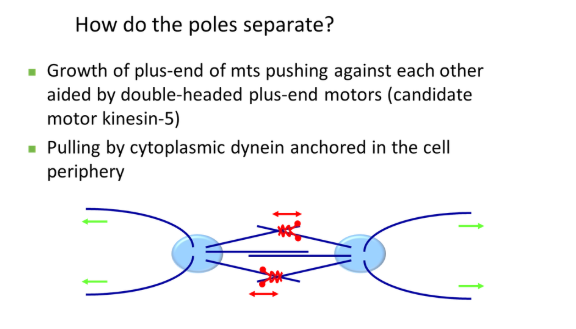
how do centrosomes separate to the poles of the cell in prophase?
the pair of centrosomes produce spindle fibres that overlap antiparallel (plus ends together) and push each other away
this is carried out by double-headed plus-end motors that walk towards the overlapping region
also, cytoplasmic dynein (minus-end motor), anchored at the plasma membrane, pulls the centrosome away from the centre via astral microtubules
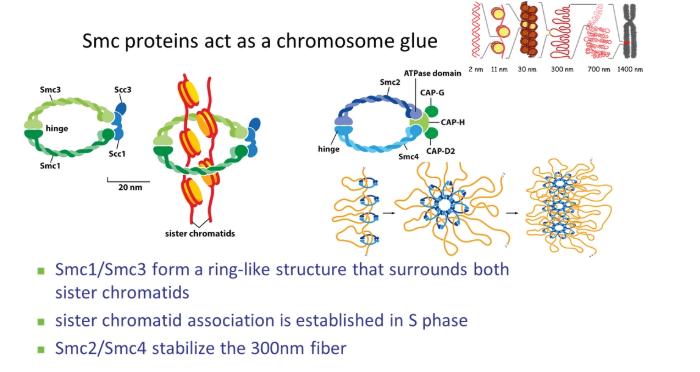
how does chromosome condensation happen in prophase?
Cdk phosphorylates histones
SMC (stable maintenance of chromosomes) proteins condense the chromosomes using their hinged ring structures
condensin stabilises the 300nm fibre by organising nucleosome loops
cohesin clamps the two sister chromatids together
how is nuclear envelope breakdown initiated in pro-metaphase?
the mesh of nuclear lamins (intermediate filaments) on the inner side of the nuclear membrane is phosphorylated by Cdk
this causes the filaments to dissociate from each other and break down the nuclear membrane by vesiculation
how do microtubule dynamics change in metaphase?
in metaphase, spindle fibres have to try find the chromosomes
instead of growing out long distances and catastrophising when they reach the plasma membrane, which is very inefficient, their dynamics change
they grow out short distances, then break down, in an exploratory effort to find the chromosomes
this is because plus-end motors increase catastrophe frequency
this means microtubule turnover and dynamic instability increases greatly
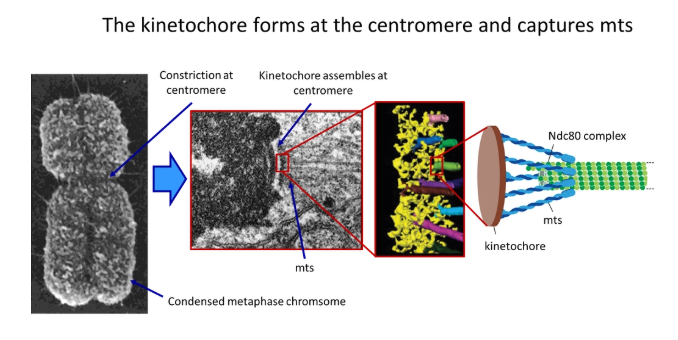
how do microtubules attach to the centromere? what process does this allow?
two kinetochores (protein complexes) assemble at the centromere of each chromosome, which grab onto the microtubules upon contact
this is carried out by a cluster of anchored minus-end protein motors called the Ndc80 complex
this holds around the sides of each microtubule so the end is still free to depolymerise and polymerise
this allows the poleward flux (while tubulin subunits are lost from the minus end, they are added at the plus end- constant length but still dynamic)
how are the chromosomes maintained along the metaphase plate?
tension from spindle fibres would pull all chromosomes into the centre of the cell- something has to keep them arranged along the metaphase plate
while the centromeres are pulled towards the centrosomes by minus-end motors at the kinetochore, the chromosome arms are pushed towards the metaphase plate by plus-end motors on the attached spindles
this is called the polar ejection force
how is correct centromere attachment by spindle fibres detected?
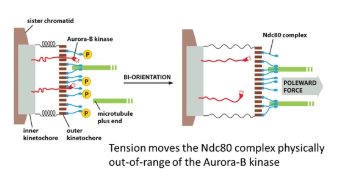
tension from spindle fibres on both sides of the chromosome is the only stable arrangement- this has to be sensed for every kinetochore
in the inner kinetochore, the Aurora-B-kinase enzyme is anchored, which phosphorylates the Ndc80 complex of the outer kinetochore (which tethers the microtubule)
however, when the kinetochore is under tension, the inner and outer layers are pulled apart and the Aurora-B-kinase is pulled out of range to phosphorylate Ncd80
this works because phosphorylation of Ndc80 reduces its affinity for microtubules, meaning it promotes the release of incorrectly attached microtubules
unattached kinetophores are bound by the mitosis arrest deficient protein (Mad), which prevents chromosome separation (by blocking securin ubiquitination by the APC) if any one kinetochore is not attached/aligned
how is chromosome separation triggered and prevented?
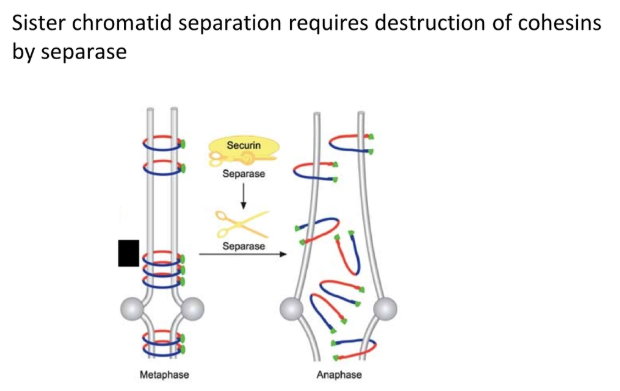
loss of the cohesin SMC proteins that bind sister chromatids together allows for chromosome separation
when the anaphase-promoting complex (APC) is activated by Cdk, it ubiquitinates the securin protein
this tags securin for degradation by protease, releasing the separase enzyme, which destroys cohesin
the APC also degrades cyclin B (by ubiquitination → protease break down) to deactivate Cdk so that mitosis can’t be reinitiated
unattached kinetophores are bound by the mitosis arrest deficient protein (Mad), which blocks the APC if any one kinetochore is not attached/aligned
this prevents chromosome separation if there is a single error (positive signal)
what drives chromosome separation at anaphase?
there are multiple redundant mechanisms to ensure correct, complete chromosome separation:
plus-end motors anchored at the centrosomes pull the microtubules in
minus-end motors at the kinetochore pull the microtubules towards the centrosome
the microtubules depolymerise at both ends
the late stage of anaphase is driven by plus-end motors pushing the overlapping spindle fibres away from each other at the centre (these are only activated at the M/A transition)
why are microtubules used for spindle fibres? why are intermediate filaments used for nuclear lamins?
spindle fibres- even though microtubules are more energetically expensive to produce than actin, they are used for spindle fibres because actin isn’t stable under compression (it buckles)- whereas microtubules can be pushed away from each other to separate the poles
nuclear lamins- intermediate filaments are persistent for a longer period of time than microtubules and actin filaments, because they are less dynamic