Veterinary Genetics (AnSci 361) Exam 1
1/99
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
100 Terms
Closed chromatin structure
- more nucleosomes and densely packed DNA, no gene expression
- RNA polymerase cannot get access to the DNA
Open chromatin structure
- fewer nucleosomes, less densely packed DNA, genes expressed
- transcription by RNA polymerase can occur
Chromatin
- DNA structure wrapped around histones
- makes up chromosomes
DNA
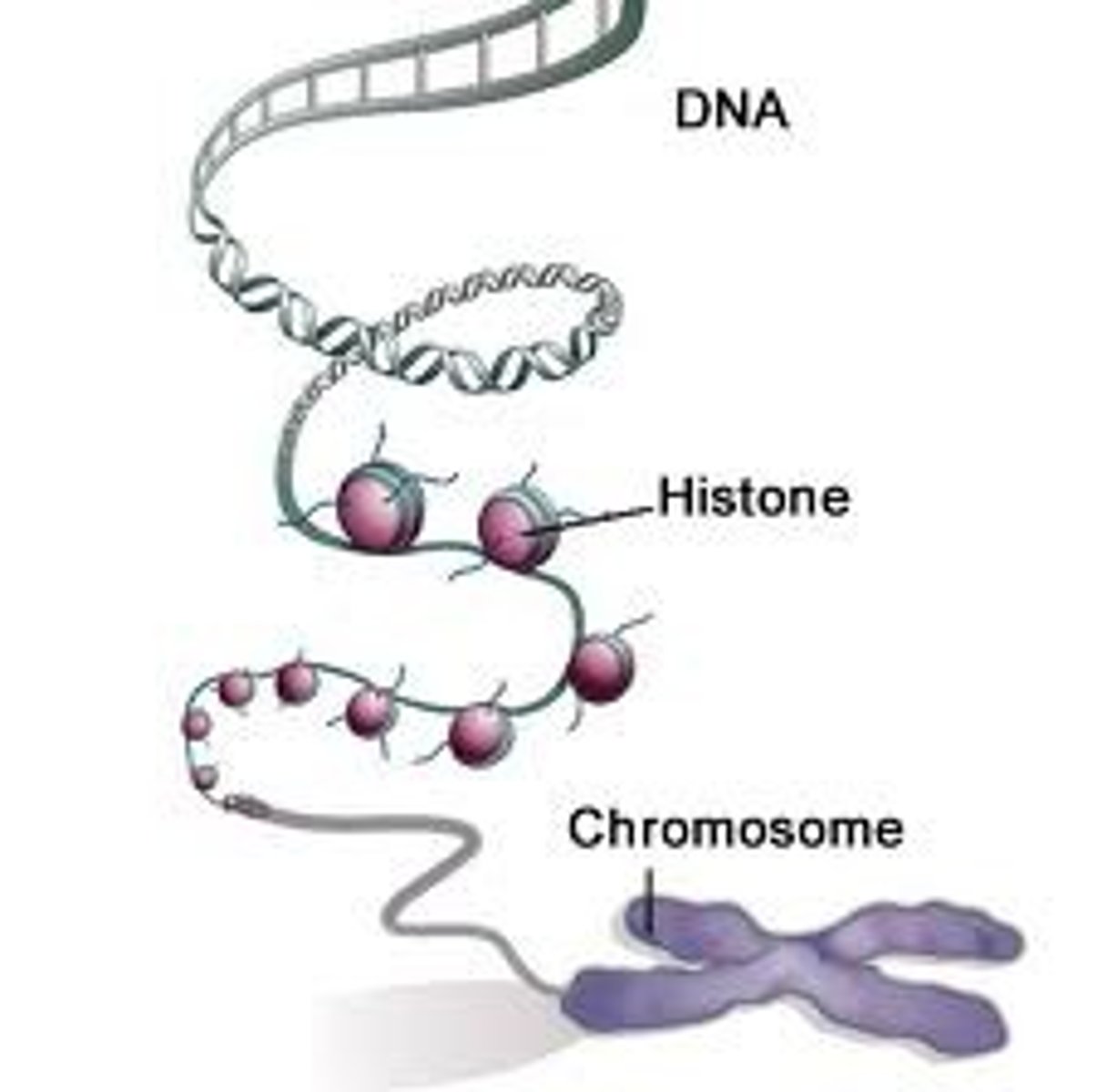
Nucleoside
nitrogenous base + sugar
Nucleotide
nitrogenous base + sugar + phosphate
Purines
- two-ring structures
- adenine (A)
- guanine (G)
Pyrimidines
- one-string structures
- thymine (T)
- cytosine (C)
Structure of DNA (polynucleotide) chain
- polynucleotide chains have polarity
- one end has 5'-phosphate and other end has 3'-OH
- pairings of C-G bonds have more chemical energy (3 hydrogen bonds)
- A-T is bonded via two hydrogen bonds
DNA strands are antiparallel
- sequences are written 5' to 3'
- left strand: 5' CTGGAC 3'
- right strand: 5' GTCCAG 3'
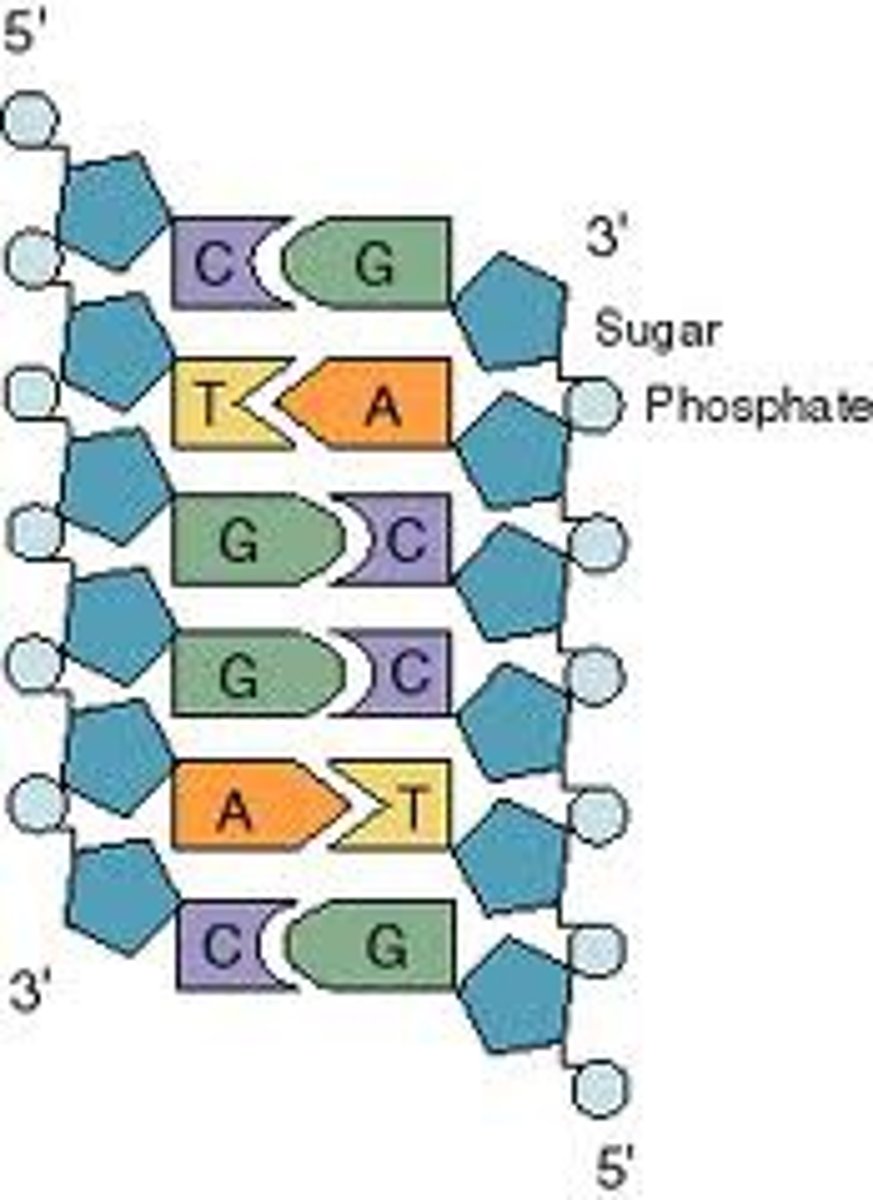
Restriction enzymes
- naturally occurring enzymes that serve to protect the bacterial cell by disabling the DNA of bacteriophages (viruses) that attack it
- the enzymes are named after the organism from which they were derived
- gave the ability to reproduce a certain segment of DNA and make millions of copies
Recognition sequence
- restriction enzymes cleave DNA at a specific sequence
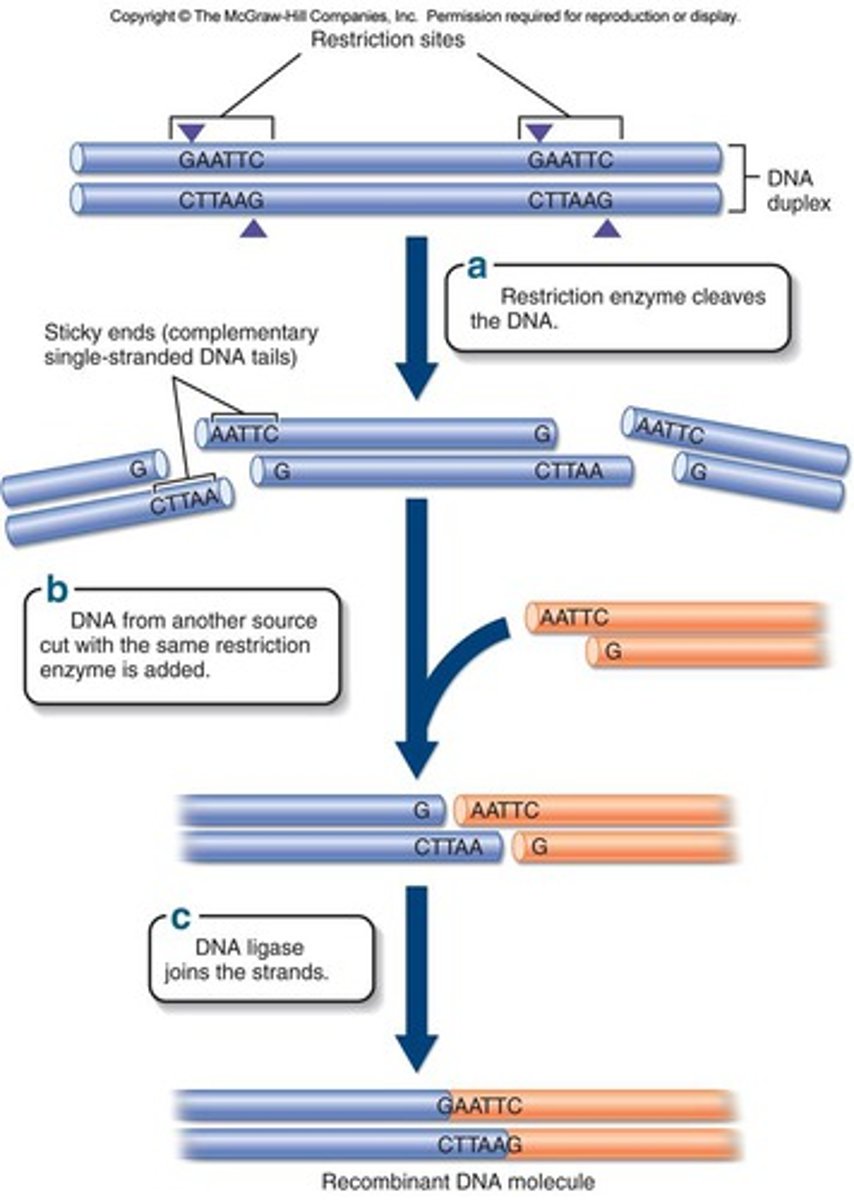
Properties of restriction enzymes
- DNA recognition sequence is usually 4-8 bp.
- the recognition sequence may be ambiguous: for example, (A/G)GCGC(C/T)
- most restriction enzymes recognize a single restriction site
- the recognition site is recognized without regard to the source of DNA
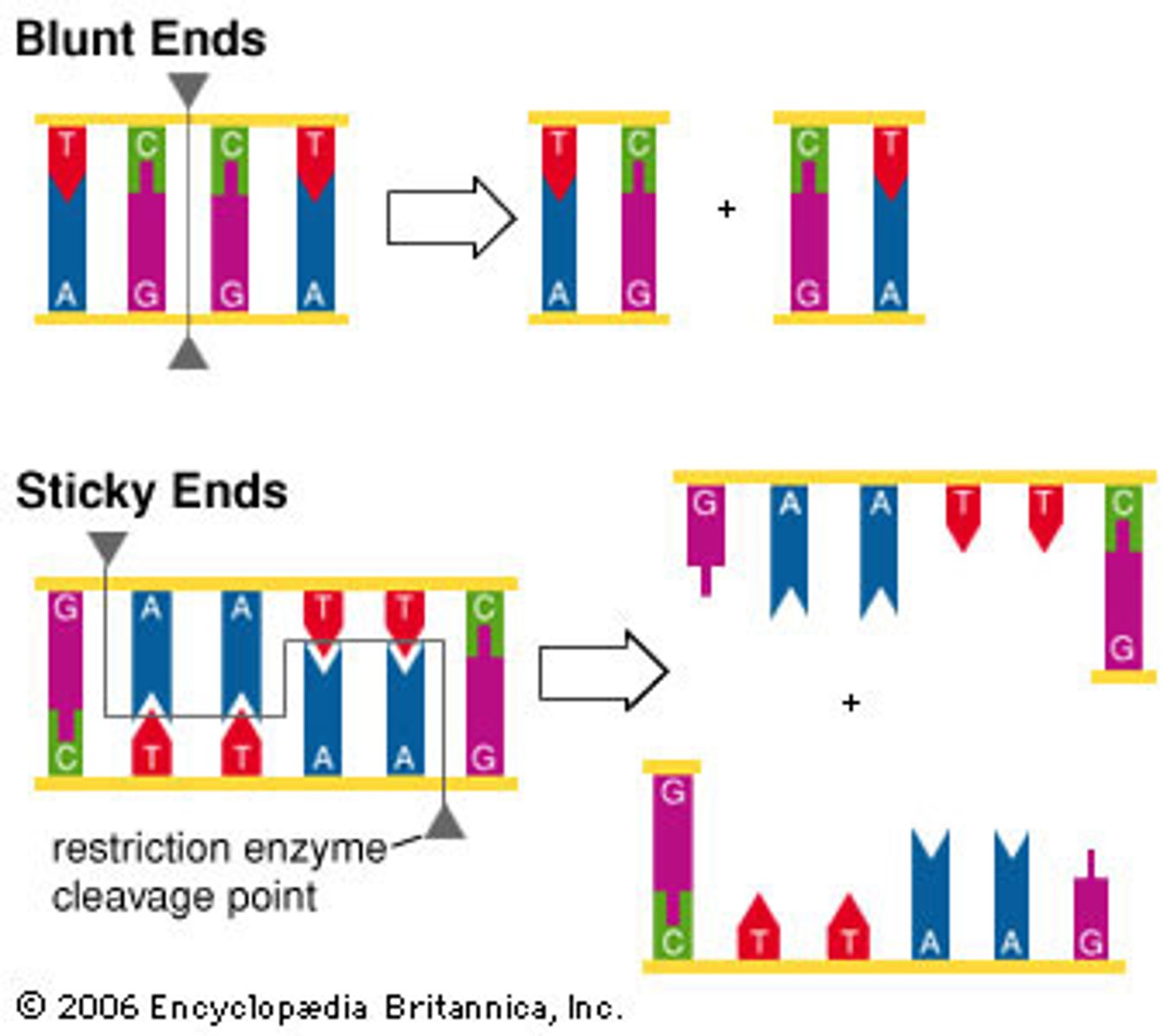
- the cuts may result in blunt or sticky ends
Blunt ends
- restriction fragments with no overlapping ends and that never combine with another type of DNA
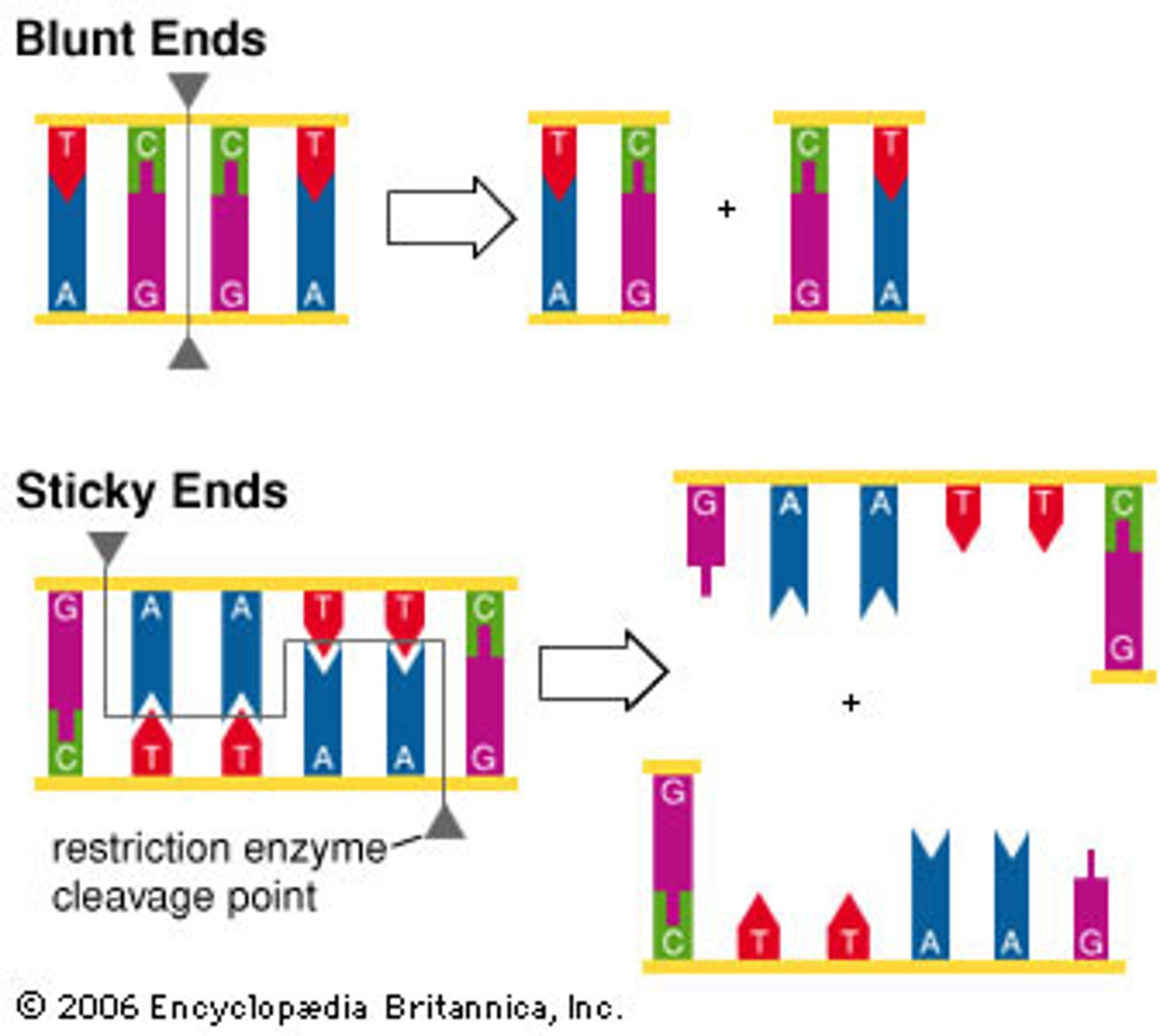
Sticky ends
- single stranded ends of DNA left after cutting with enzymes
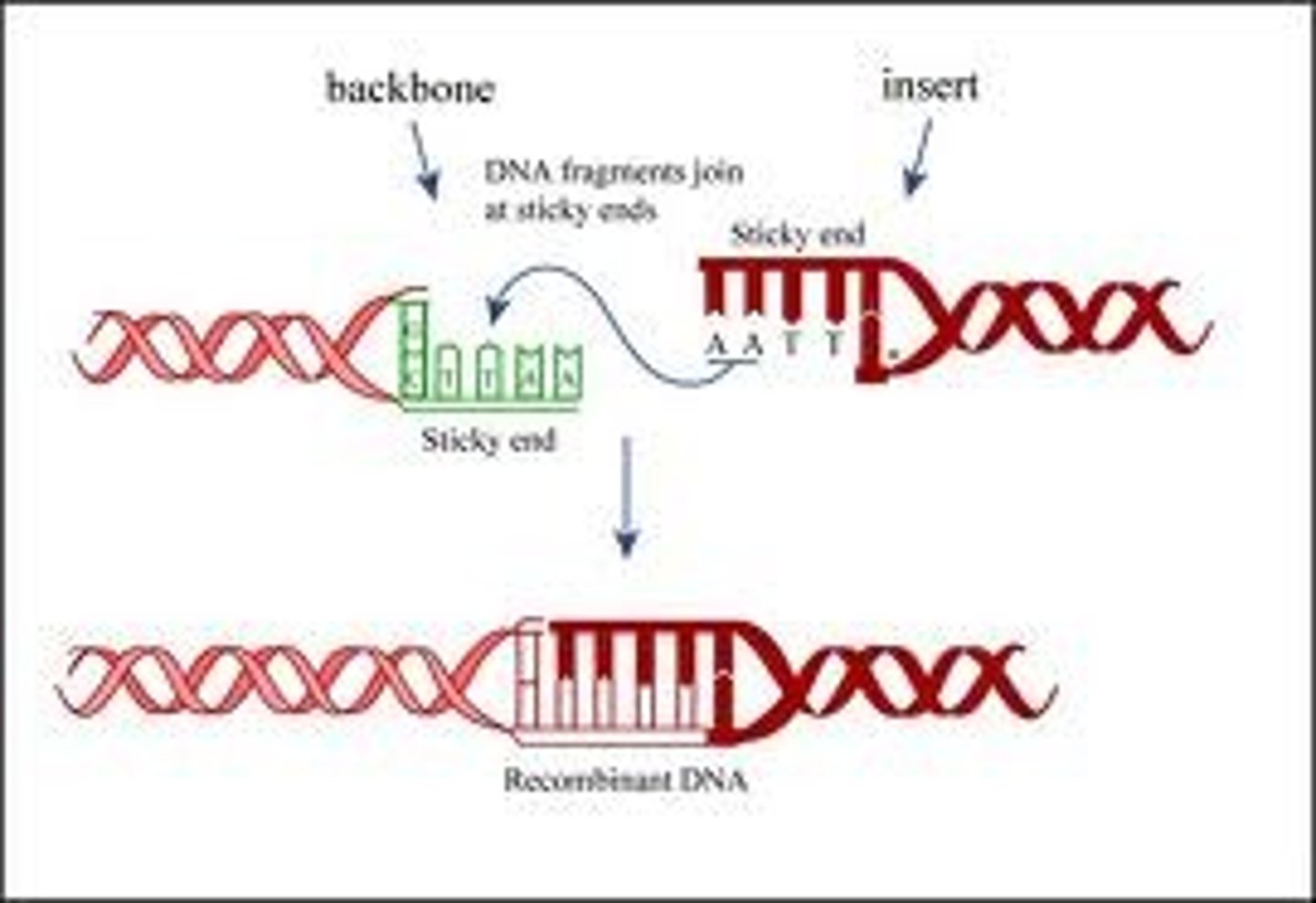
Ligase
- enzyme used to rejoin DNA fragments
Size separation of DNA fragments by electrophoresis is agarose gels
- DNA is negatively charged due to phosphates on its surface, so it moves towards the positive pole
- smaller fragments will move faster and farther through the gel
- agarose gel is stained with ethidium bromide which binds to DNA and fluoresces under ultraviolet light
Restriction site polymorphism
- a mutation in the recognition sequence of a restriction enzyme can lead to the gain or loss of a cutting size in a DNA sequence
- "normal": has a restriction site which allows the DNA to be cut and replicated
- "abnormal": has a mutation and no restriction site and therefore the DNA cannot be cut and replicated
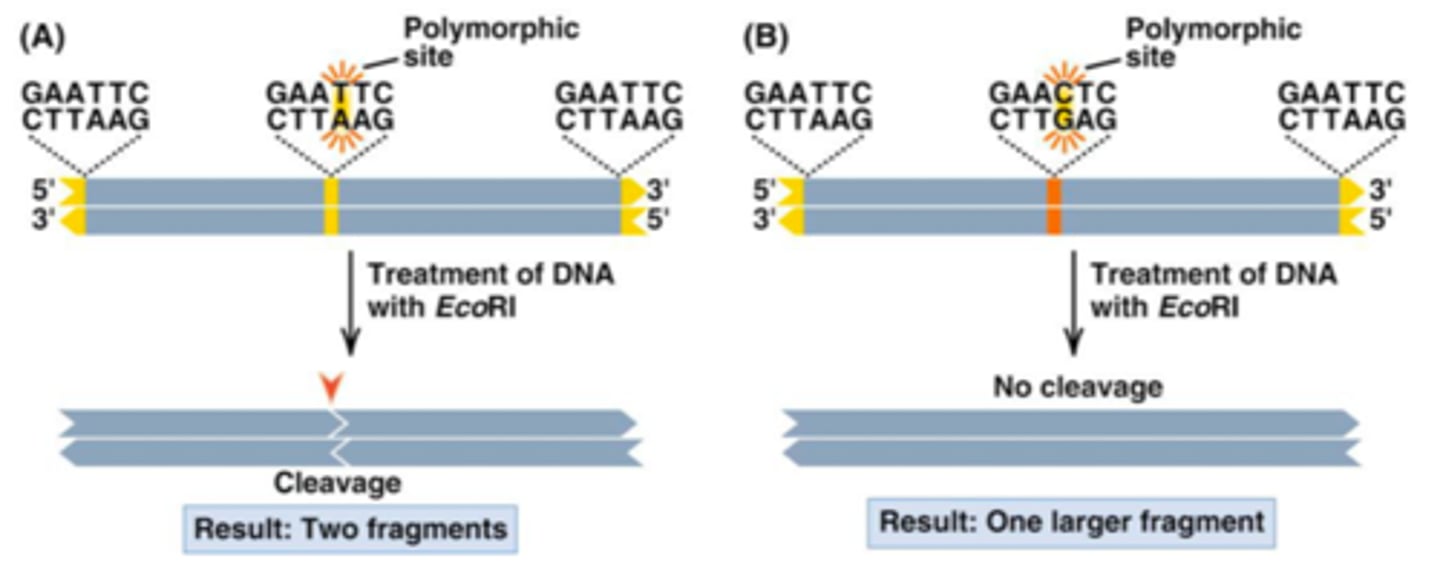
Polymorphic site (polymorphism)
- a restriction sequence with more than one variant
Polymorphism, variant, mutation
- all used at various times to refer to the same thing, a change in genomic sequence
- when applied in genotyping animals: genetic marker
Alleles
- alternate forms of genes or markers
Restriction fragment length polymorphism (RFLP)
- variation in restriction nuclease cutting sequence at a particular site creates DNA fragments of different sizes
- homozygous organism: the fragments are the same size
- heterozygous organism: the fragments are different sizes
DNA polymerase
- synthesizes a strand of DNA complementary to a template sequence (duplication of DNA)
Polymerase chain reaction (PCR)
- in vitro amplification of DNA fragments
- length limit: sequences longer than ~5000 base pairs are difficult to amplify by conventional PCR procedures
Components of PCR
- DNA polymerase: an enzyme that forms the sugar-phosphate bonds between adjacent nucleotides
- primers: short, ~20-24 base single-stranded DNA corresponding to ends of segment targeted for duplication
- nucleotides ("dNTPs")
- buffer (controlled pH, salt, MgCl2 concentration)
Steps of PCR
1. Denaturation: breaking hydrogen bonds between complementary DNA strands at 95*C
2. Hybridization of primer (annealing): attaching primers to single strands of DNA for new strand synthesis
3. DNA synthesis (extension): Taq DNA polymerase is used to survive the high temperatures - extension of new DNA strands from the primers
Genetic markers
- sources of genetic variation
- microsatellites
- single nucleotide polymorphisms (SNPs)
- insertions and deletions (indels)
Molecular techniques
- gel electrophoresis
- polymerase chain reaction (PCR)
- restriction fragment length polymorphisms (RFLP)
- sequencing
What is the basis of our phenotypic and genetic variation?
- independent inheritance of chromosomes
- recombination
- mutation
- modification affecting gene expressions (epigenetics)
- gene by environment interactions
- DNA of any two individuals within a mammalian species are 99.9% identical
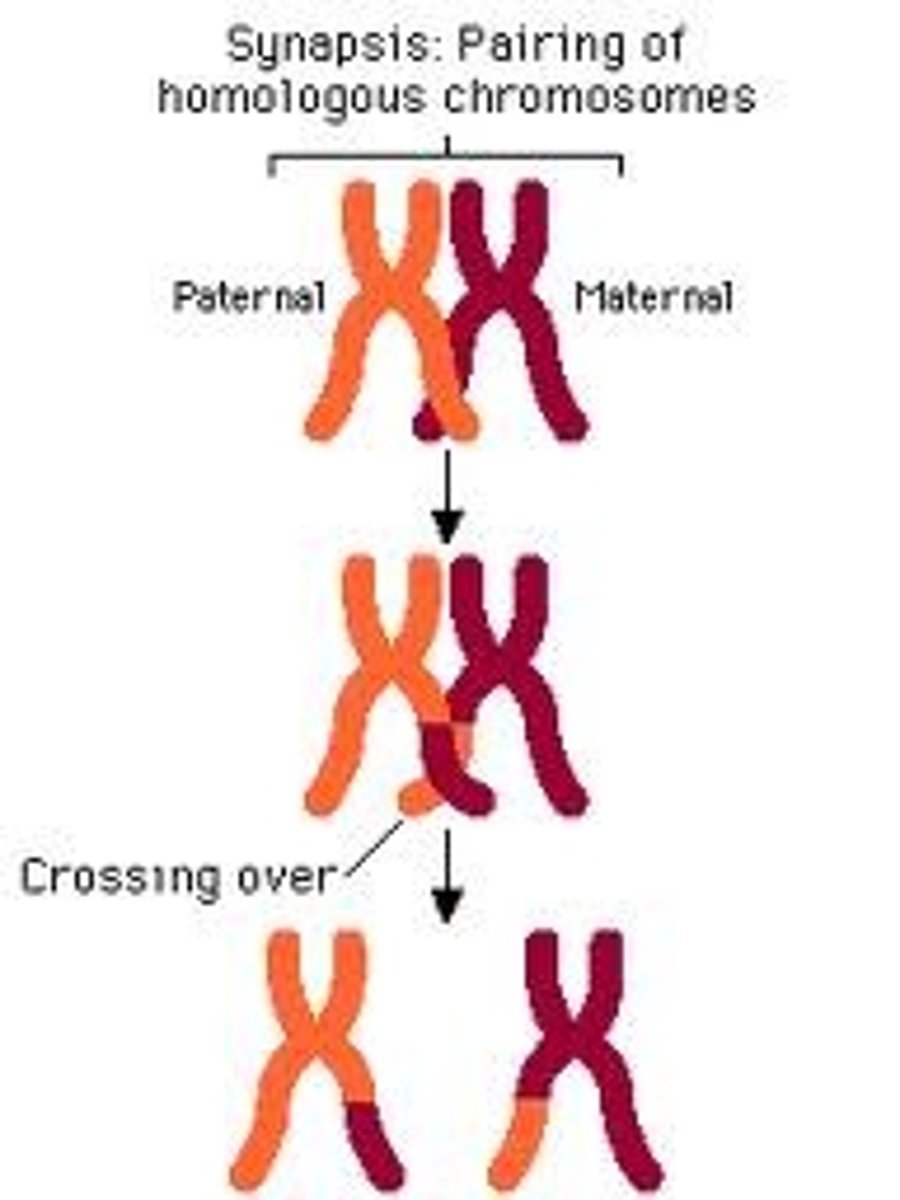
Recombination
- the genetic process by which one chromosome breaks off and attaches to another chromosome during reproductive cell division
- linkage maps are based on determining recombination rate between loci
microsatellites (short tandem repeats)
- markers with repetitive sequences and multiple alleles
- naturally occurring polymorphism
- short sequences of DNA repeating in tandem arrays
- different alleles have different lengths of microsatellites (different number of repeat units)
- not typically found in exons (if found in exons always a 3 base repeat)
Why are microsatellites useful as genetic markers?
- the number of repeated sequenes varies between individuals within populations
- have multiple alleles (highly informative)
- widely dispersed in genome
- readily genotyped using PCR and capillary electrophoresis
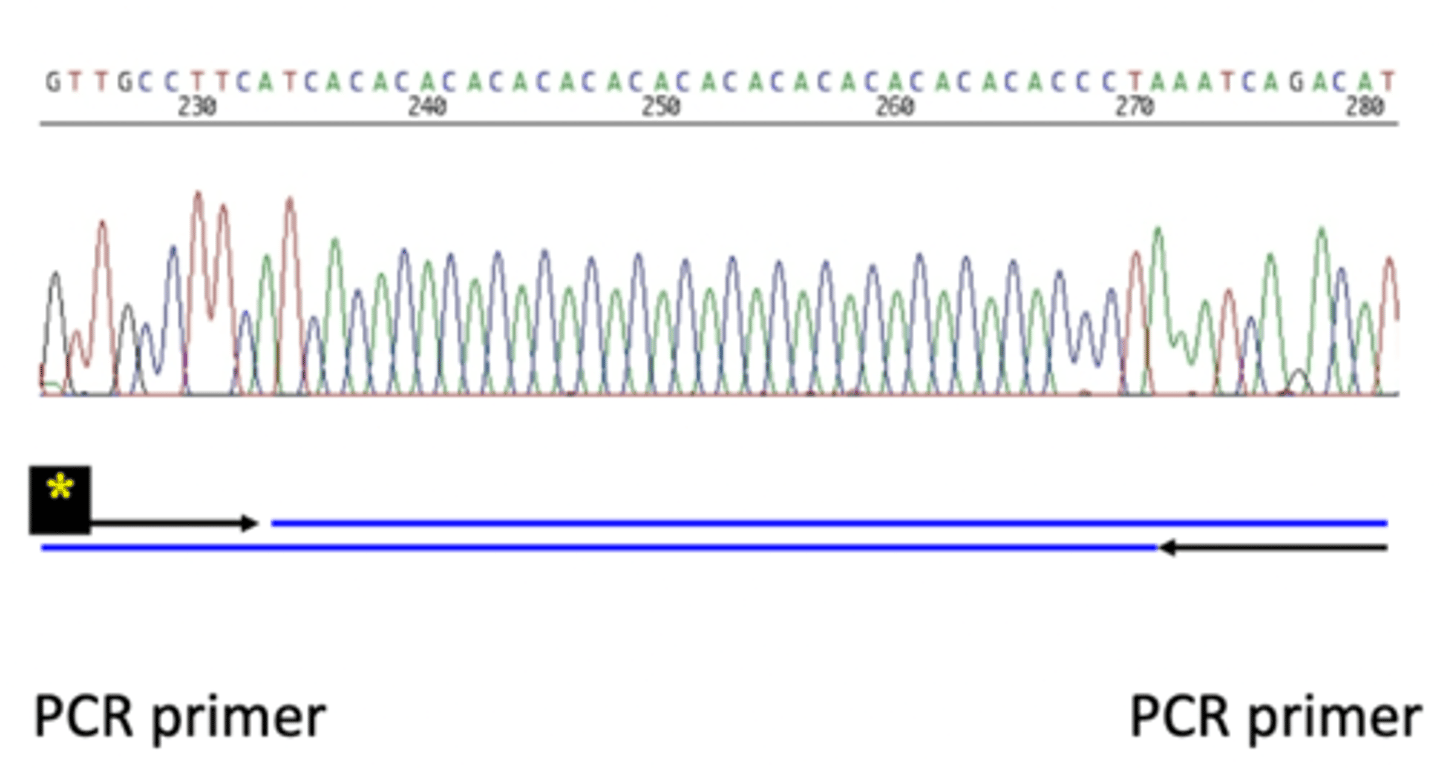
Microsatellite sequence data
- AC repeat in the microsatellite
- PCR primers flank the sequence from the outside
- primer needs to begin at a unique sequence
- if primer is located in the microsatellite sequence you would get multiple products
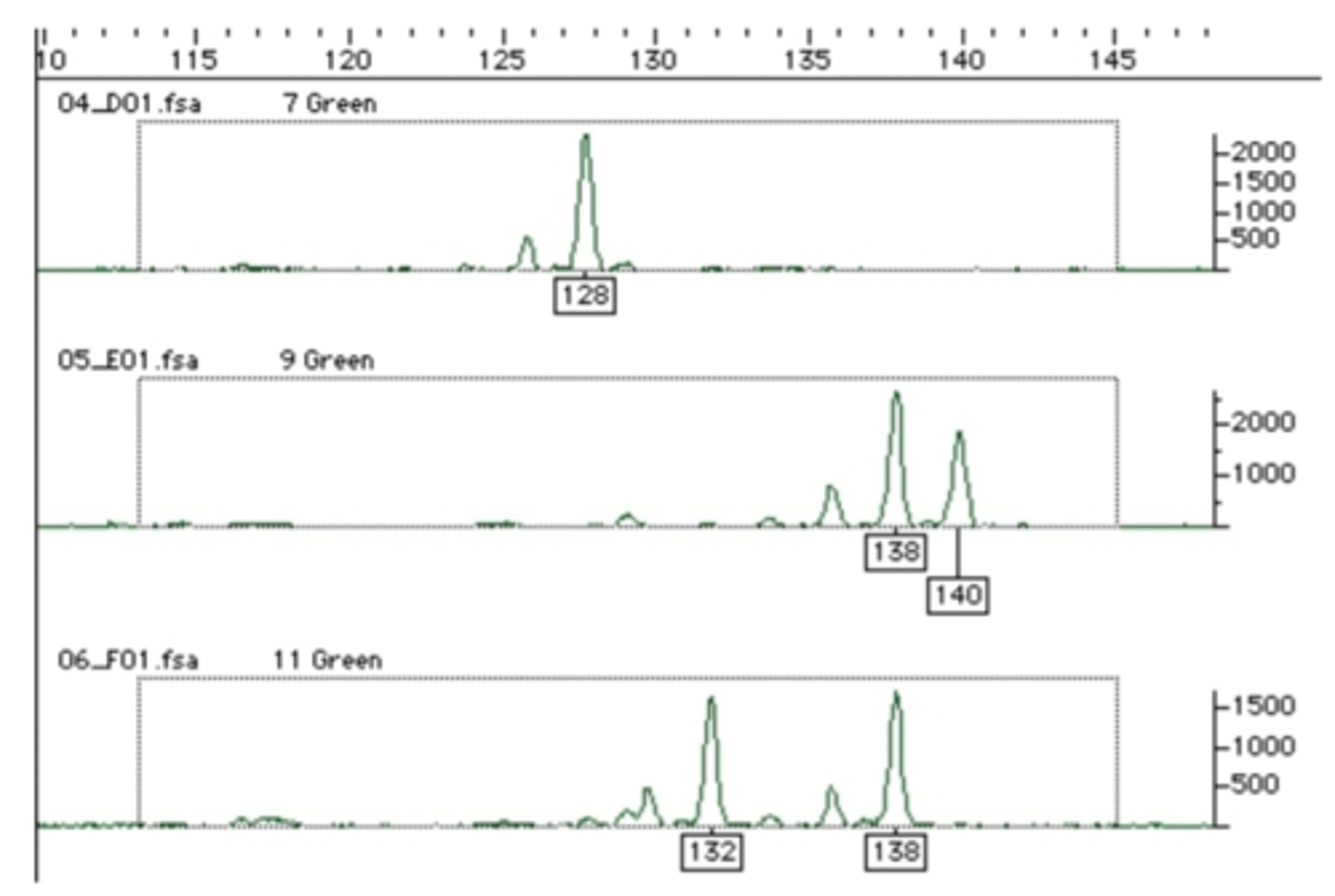
Microsattelite genotyping
- one peak: homozygous for 128 bp (first graph)
- 128 would be the true allele
- smaller peak would have skipped over a base pair resulting in it being smaller
- multiple peaks: heterozygous for 138 and 140 base pairs (second and third graph)
- each true allele would have a 2 base pair band (smaller peak behind it)
- small peaks are from DNA polymerase making mistakes - adding or taking away repeat units
Single nucleotide polymorphism (SNP)
- SNPs are bi-allelic (two alternative alleles)
- a SNP genotype is the combination of two alleles at one locus for one individual: GG (homozygous), TT (homozygous), or GT (heterozygous)
- three possible genotypes for bi-allelic SNP

SNP markers
- millions of SNPs in livestock genomes
- widely dispersed in genome
- found in exons, introns, and intergenic regions (everywhere)
- can by typed by PCR - based methods or other methods
- amendable to high-throughput systems
Small insertions/deletions (indels)
- insertion or deletion of one or multiple nucleotides
- indels can be considered like SNPs, considering the first inserted or deleted nucleotide
- usually bi-allelic

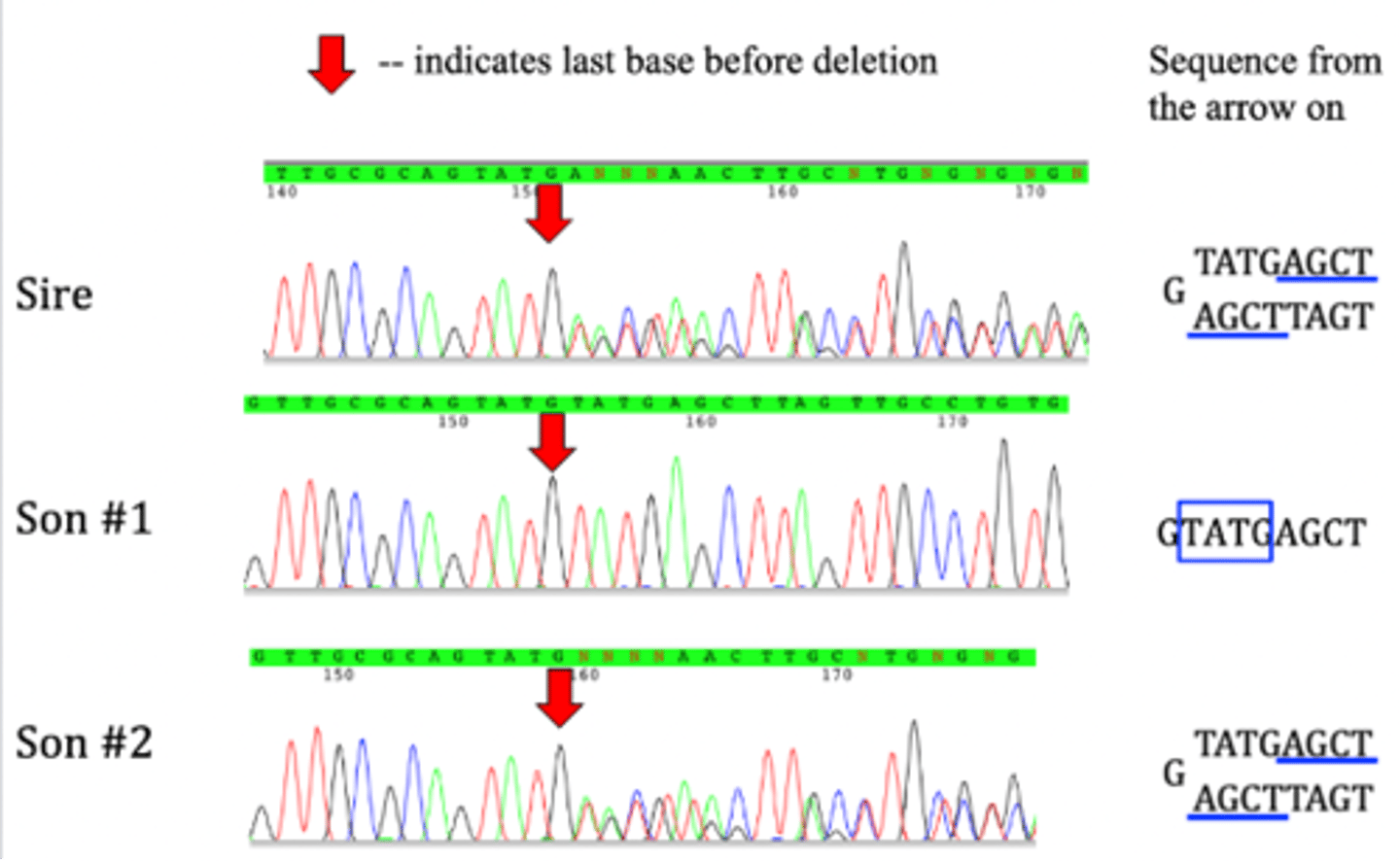
Indel example
- sire and son 2 have sequence overlap
- 4-base deletion
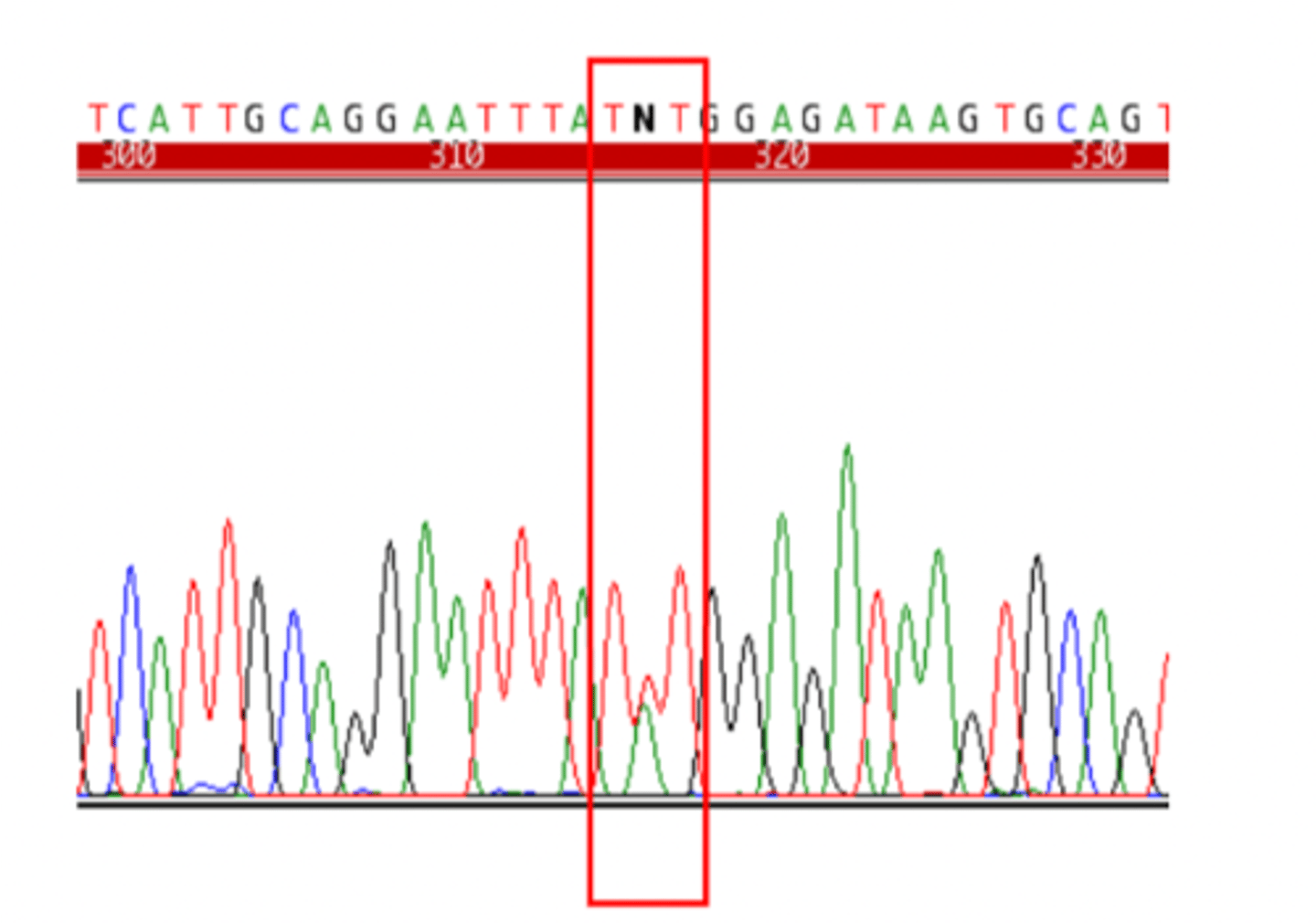
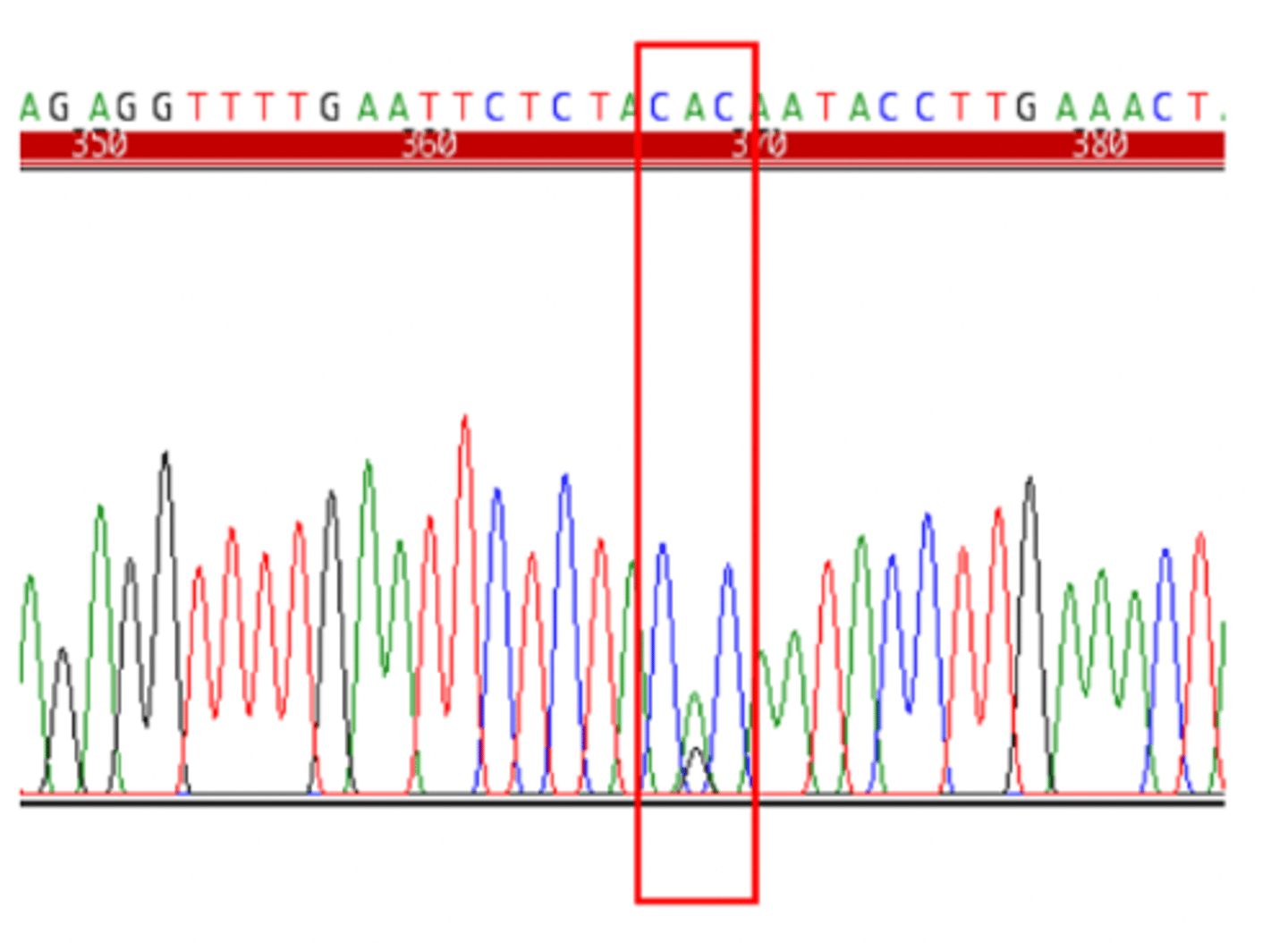
SNP discovery - sequence an individual (Heterozygous)
- identify sites were an individual is heterozygous
- overlapping peaks
SNP discovery - sequence an individual (Homozygous)
- identify sites where individuals differ
How has the cost of sequencing a genome changed?
- gone down extremely over time
SNPs v. Microsatellites
SNPs
- two alleles
- widely dispersed
- more than 90 million
- high throughput genotyping
- low mutation rate (<1 x 10^-8) per generation
Microsatellites
- multiple alleles
- widely dispersed, but not in coding sequence unless 3-base pair repeat
- hundreds of thousands
- semi-automated genotyping, not high throughput
- high mutation rate (1 x 10^-3) per generation
Genotype
- combination of alleles at a locus
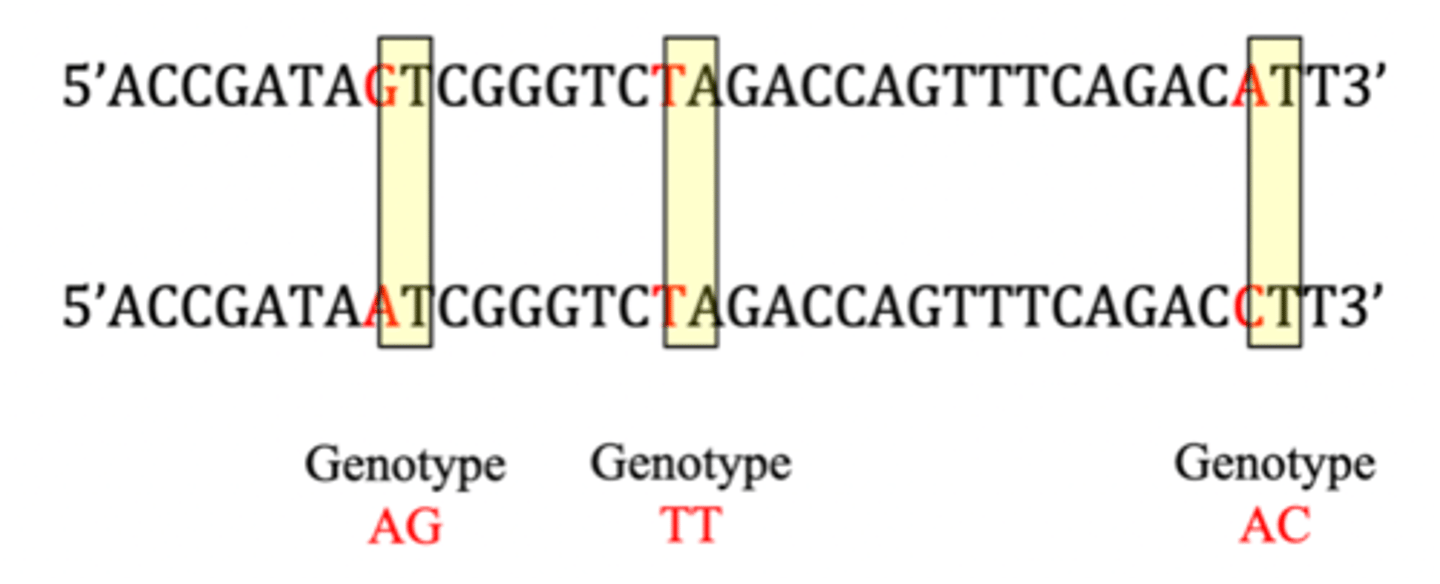
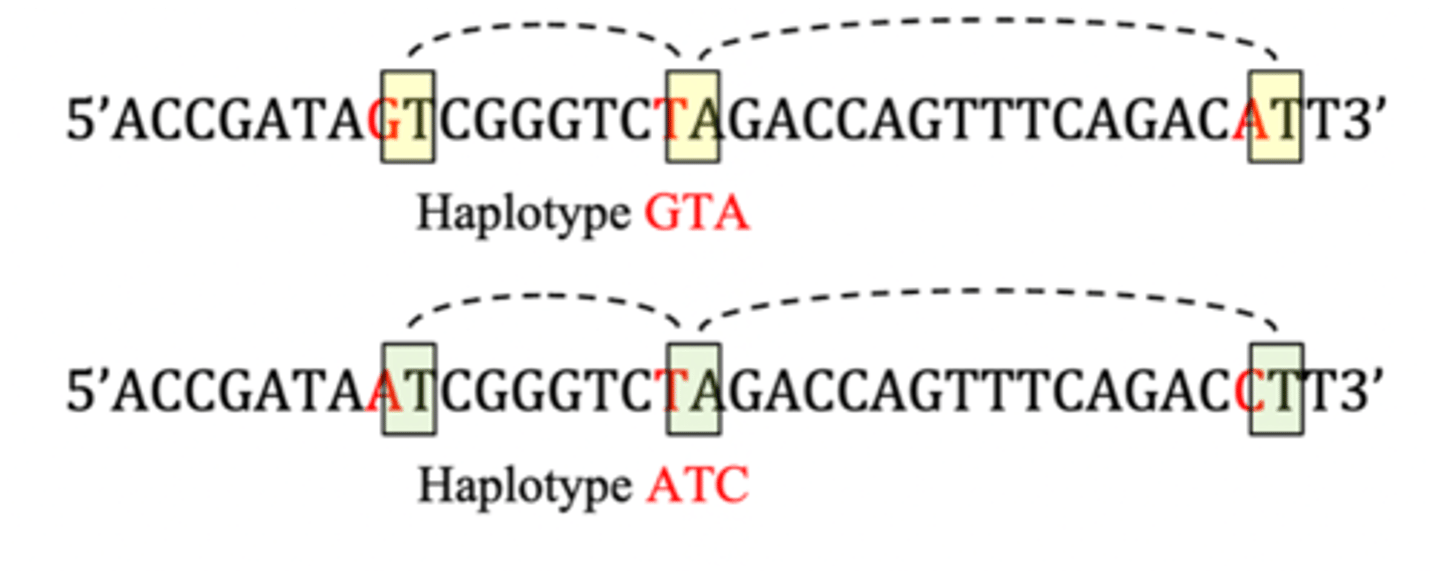
Haplotype
- combination of alleles at two or more consecutive loci on one chromosome strand
Why learn about forensics and parentage testing in animals?
- verify the parentage and determine the authenticity of pedigrees
- verify the identity of an animal
- identify species from a tissue sample
- determine parentage for pedigree recording
- epidemiology (trace diseased animal back to farm of origin)
What are the applications of genomics in human forensics?
- crime solving
- identify accident victims
- identify members of armed forces in war
- paternity testing
- immigration testing (are two people related)
- identify missing persons from remains
- convicted felons databases
Sources of DNA for forensic testing
- blood
- bones
- urine
- hair with roots
- hair without roots
- semen
- salive
- envelopes/stamps
- skin/other tissues
- old blood stains
- ancient bones
- cheek swabs
- toothbrush
- dead animals
- beverage cans
- bite marks
- gum
- cigarette butts
- fingernail scrapings
What is NOT a source of DNA forensic testing?
- red blood cells
- not true in mammals but true in birds!
Standard tool for forensic testing
- microsatellite marker panels
- lengths of microsatellites may quite variable (i.e. multiple alleles)
- use of microsatellites is attractive: relative ease of use and accuracy of typing
- ability to employ PCR; in criminal casework only minute samples of DNA may be available
PCR-based genotyping
- one primers from each primer pair is end-labeled with fluorescent dye
Microsatellite array for horse parentage testing and identification
- each microsatellite has a unique range
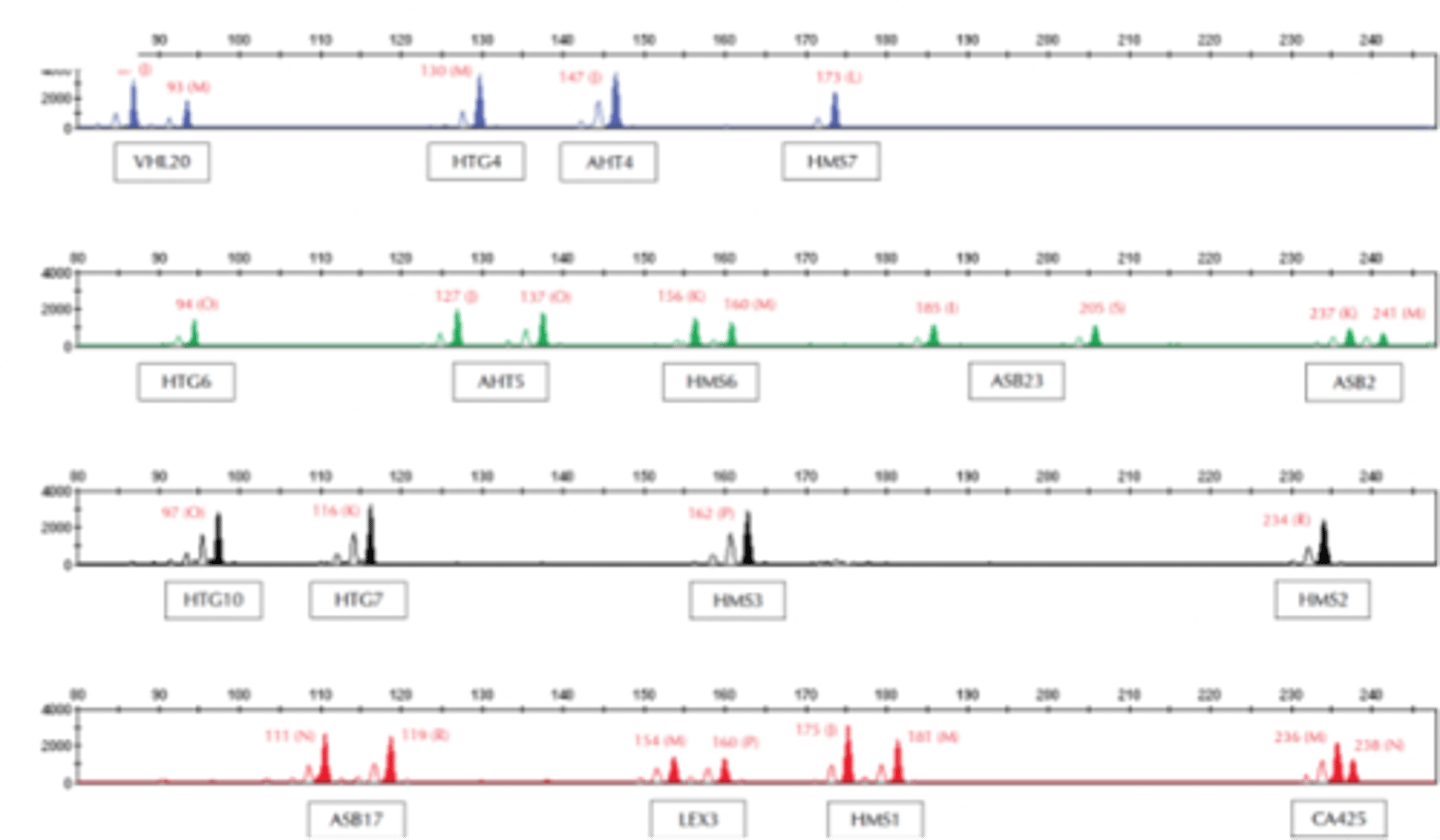
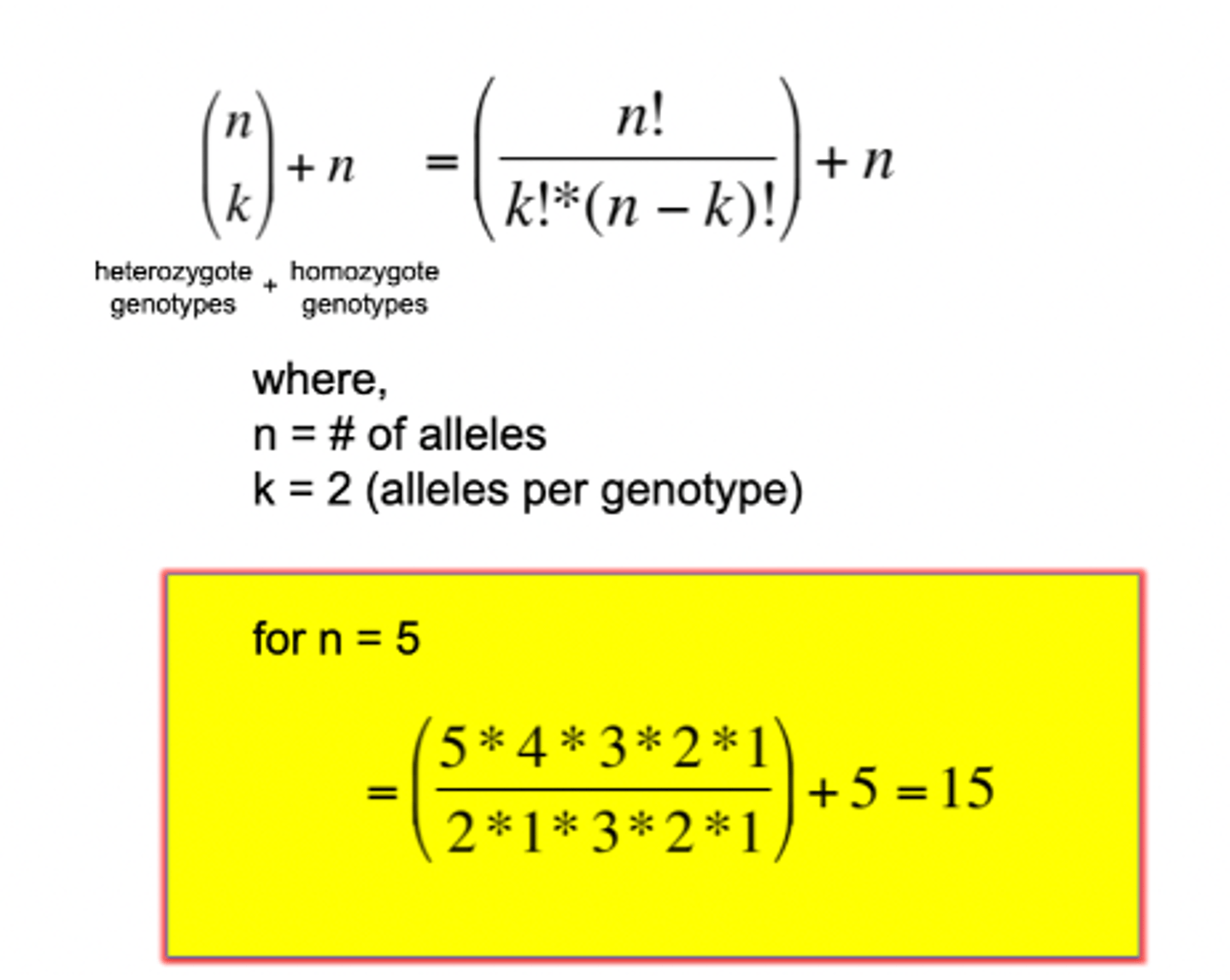
Identity testing - how many genotypes from one marker?
- everything after the first two numbers (5x4) will cancel out

Three types of animal DNA evidence
1. the animal as a victim - animal abuse, identifying the remains of a lost pet, theft of an animal, endangered species
2. the animal as the "suspect" - attack on a person or other animal, animal causing an accident, responsible for property damage
3. the animal as a witness - using animal DNA to link a suspect with a crime
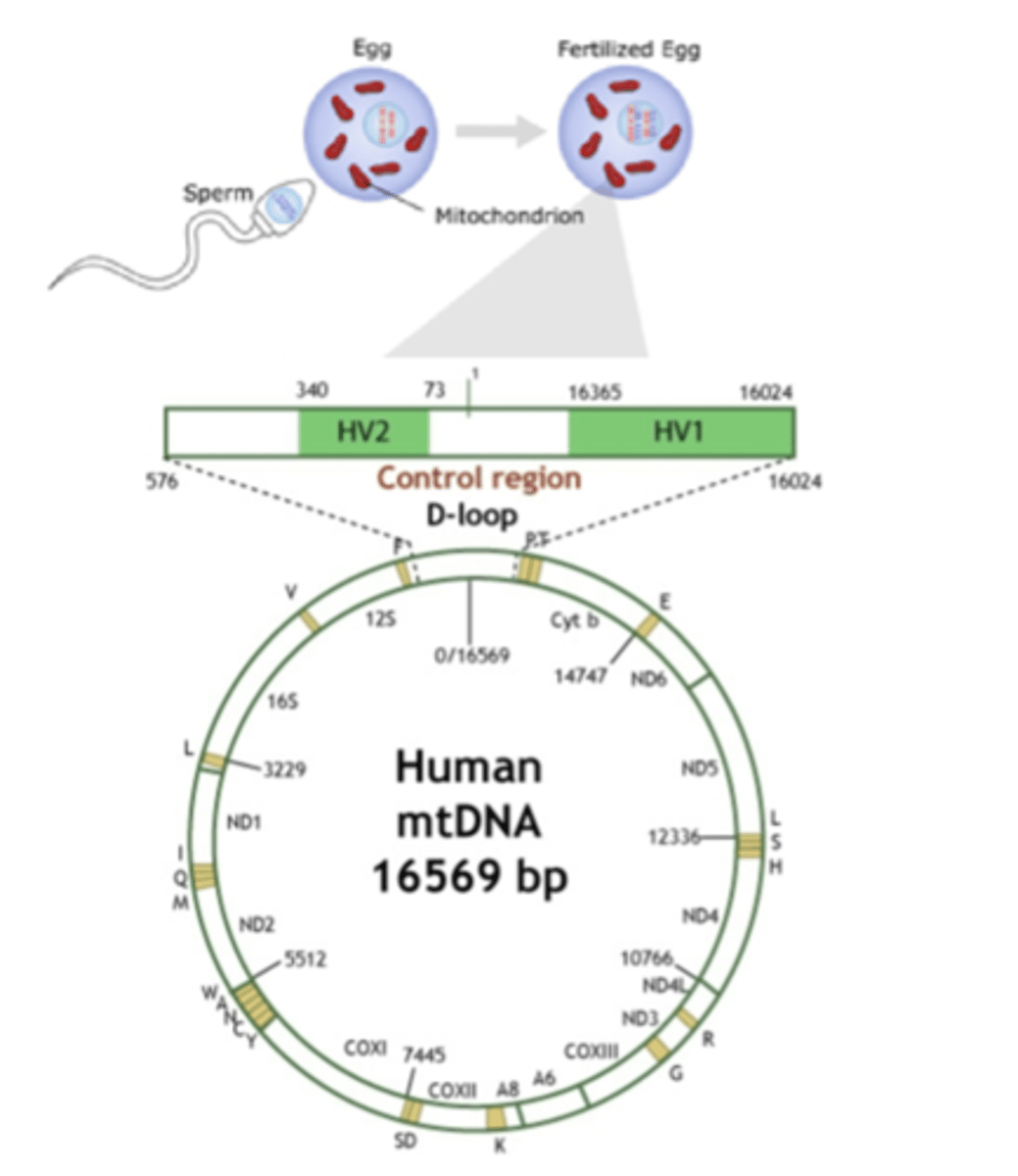
Mitochondrial DNA (mtDNA)
- hundreds to thousands of mitochondria per cell
- 5-10 mtDNA copies per mitochondrion
- many more copies per cell compared to nuclear DNA
- maternally inherited
mtDNA testing
- cytochrome B gene: variation is limited within species, but good for between-species comparisons
- D-loop region: contains two hyper-variable regions, good for within-species comparisons
- HV1: hypervariable region - target here for in-species identification because there is more differentiation
Nuclear DNA (STR) markers preferred for forensic work, unless:
- sample degradation or sample type make recovery of nuclear DNA problematic
- comparisons of matrilineal relatives is desired
- limitation of mtDNA: some sequences (haplotypes) are too common
What are reasons for parentage testing?
- determine parentage for seedstock when sire unknown
- verify parentage of potentially valuable breeding stock
- verify parentage in progeny testing herds (misidentification rate in dairy herds can be high)
- improve accuracy of breeding value estimation (EPDs)
What is an effect of misidentified parentage?
- reduce genetic gain from selection
Parentage testing
- compare marker genotypes of offspring with: dam and/or potential sires
- genotyping will exclude certain sires
- microsattelites are more informative individually
- SNPs less informative individually, but current chips genotype tens of thousands
- SNP genotyping now commonly used in dairy industry to predict genetic merit
Single-gene disorders
- disorders caused by a mutation/alteration within one gene
Polygenic diseases (complex, multifactorial)
- disorder caused by collective action of alleles in many genes
- gene x environment interactions
- more common scenario than single gene disorders
Recessive inheritance
- recessive phenotypes require two copies of recessive allele
- "aa"
Dominant inheritance
- one copy of dominant allele is sufficient to cause dominant phenotype
Additive inheritance
- additional allele has incremental effect
- as you go from A1 --> A2 there is incremental increase in phenotypic expression
- A1A1, A1A2, A2A2
- A1A2 half-way between homozygotes (exactly in the mdidle)
Incomplete dominance
- heterozygote intermediate between homozygotes, but effect is not incremental
- not necessarily exactly in the middle
Overdominance
- "heterozygote superiority"
- heterozygote is superior to either homozygote
Inheritance of single-gene disorders
- autosomal dominant
- autosomal recessive
- X-linked dominant
- X-linked recessive
- Y-linked
- mitochondrial
Characteristics of recessive phenotypes
- disorder may skip generations
- all offspring of two affected parents are affected
- equal number of males and females affected
Autosomal recessive pedigree
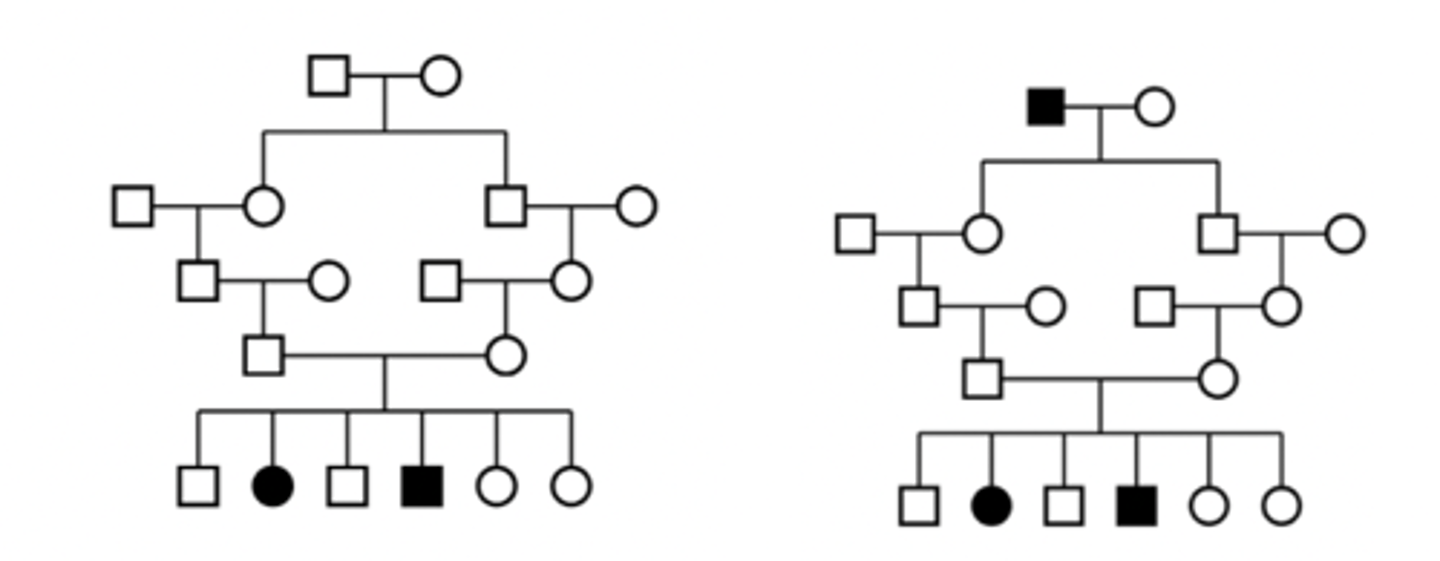
Characteristics of dominant phenotypes
- disorder does not skip generations
- every affected offspring has at least one affected parent
Autosomal dominant pedigree
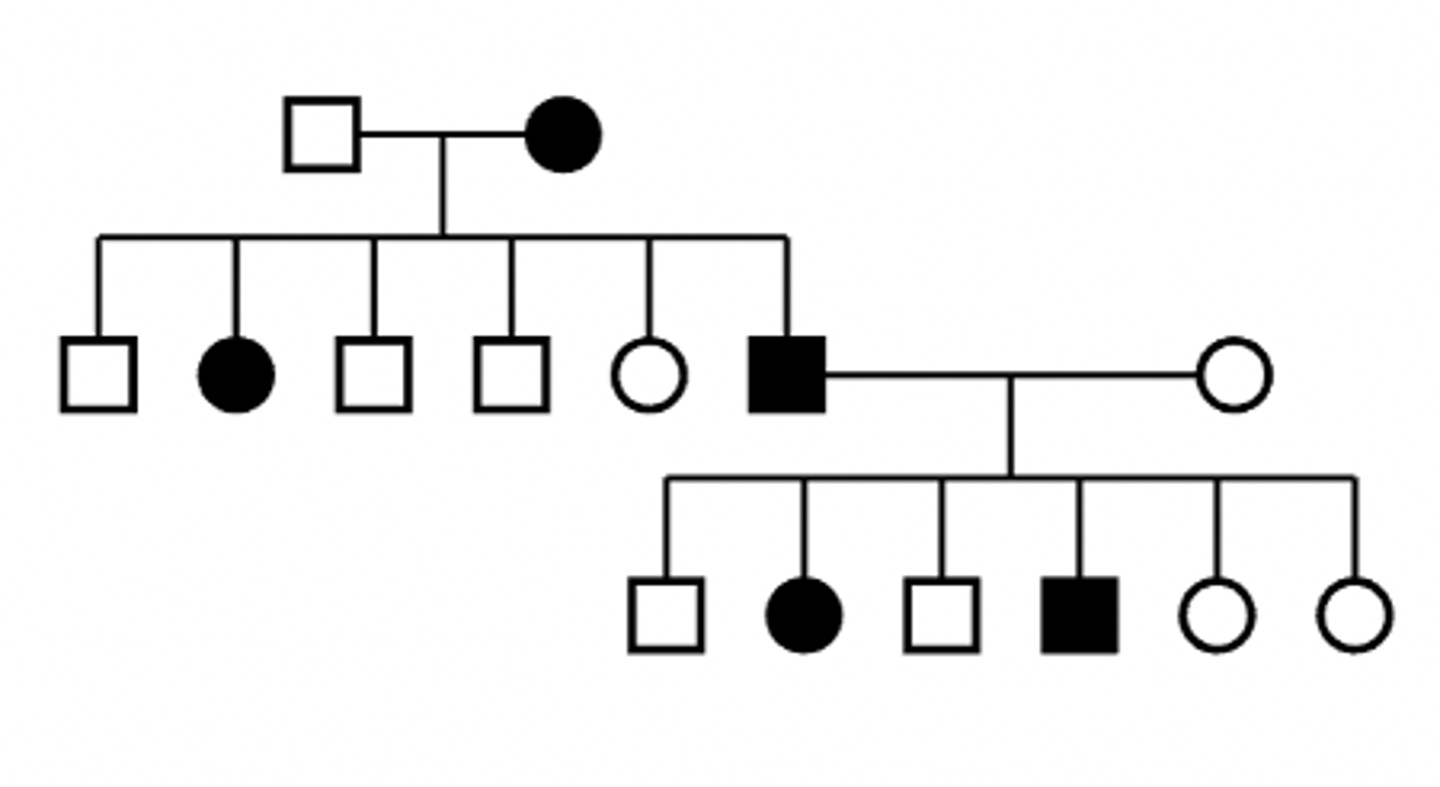
Characteristics of X-linked dominant
- affected males transmit disorder to all daughters and no sons
- male will always give the X chromosome to all of his daughters, where the Y chromosome will go to all of the sons
Hemizygous
- x-linked gene is affected in heterozygote
- X^RY and X^rY
X-linked dominant pedigree
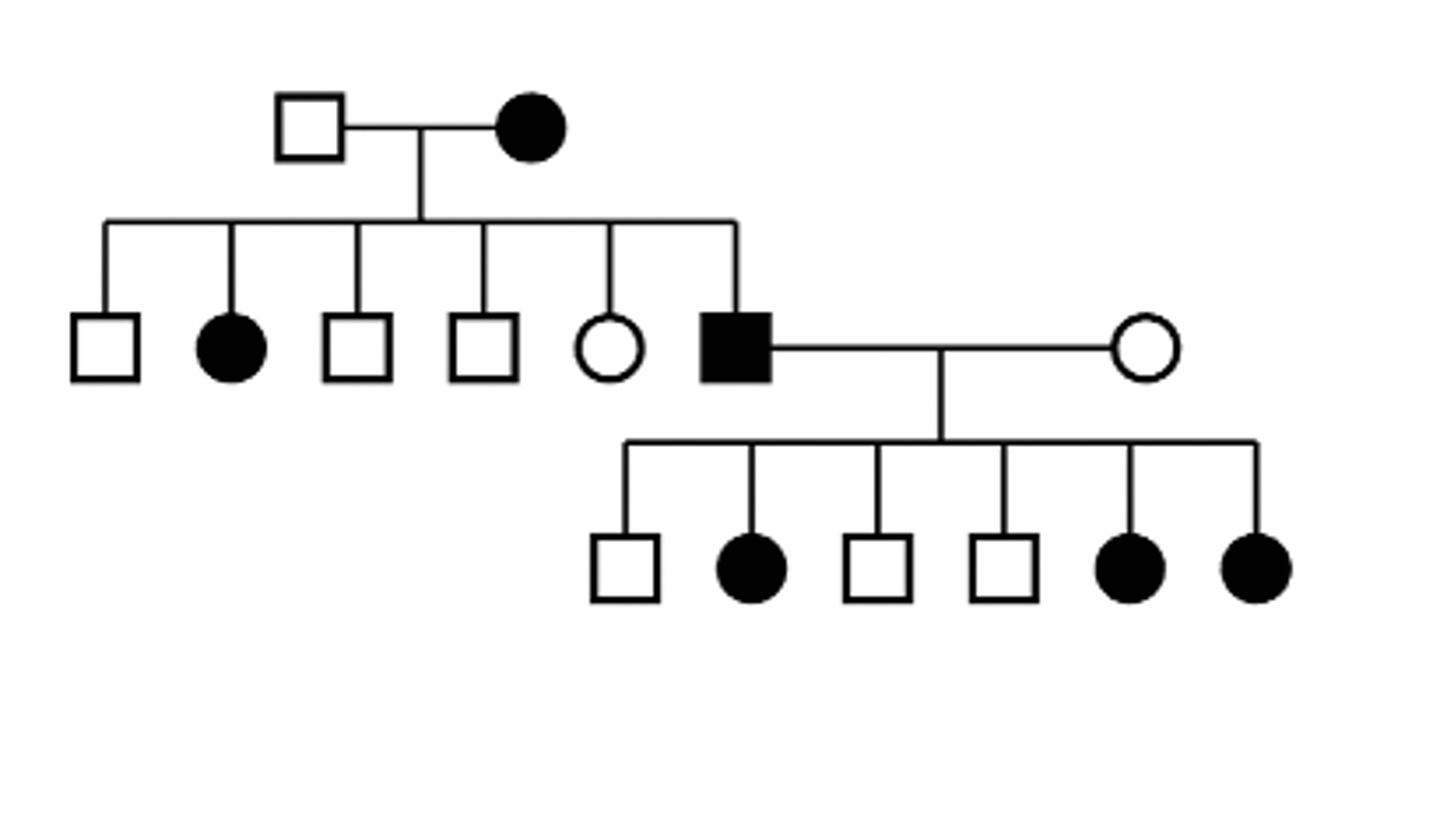
Characteristics of X-linked recessive
- disorder may skip generations
- all offspring of affected parents are affected
- incidence in females is lower than in males
X-linked recessive pedigree
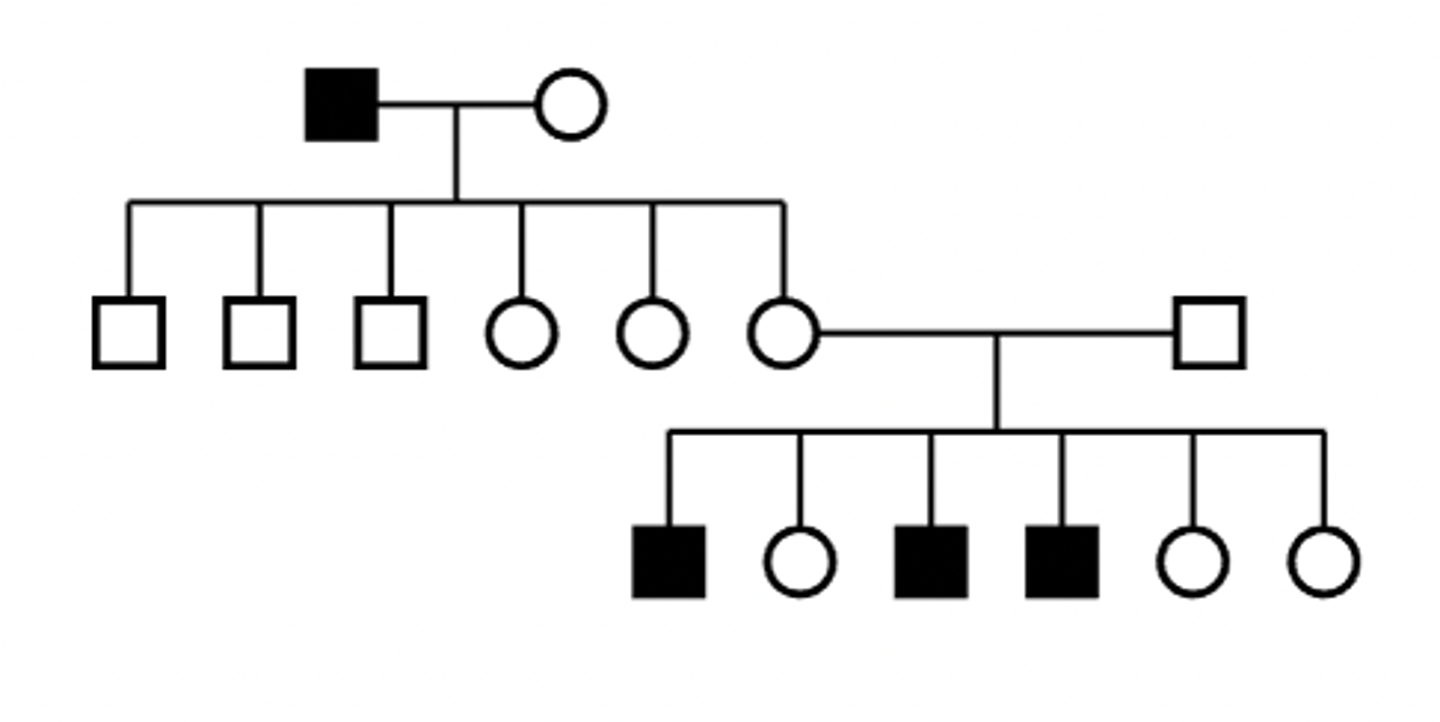
Characteristics of mtDNA disorder
- males and females affected
- sporadic, with different manifestations in affected individuals
- mothers may or may not show symptoms of disease
- pattern of transmission is non-Mendelian
mtDNA disorder pedigree
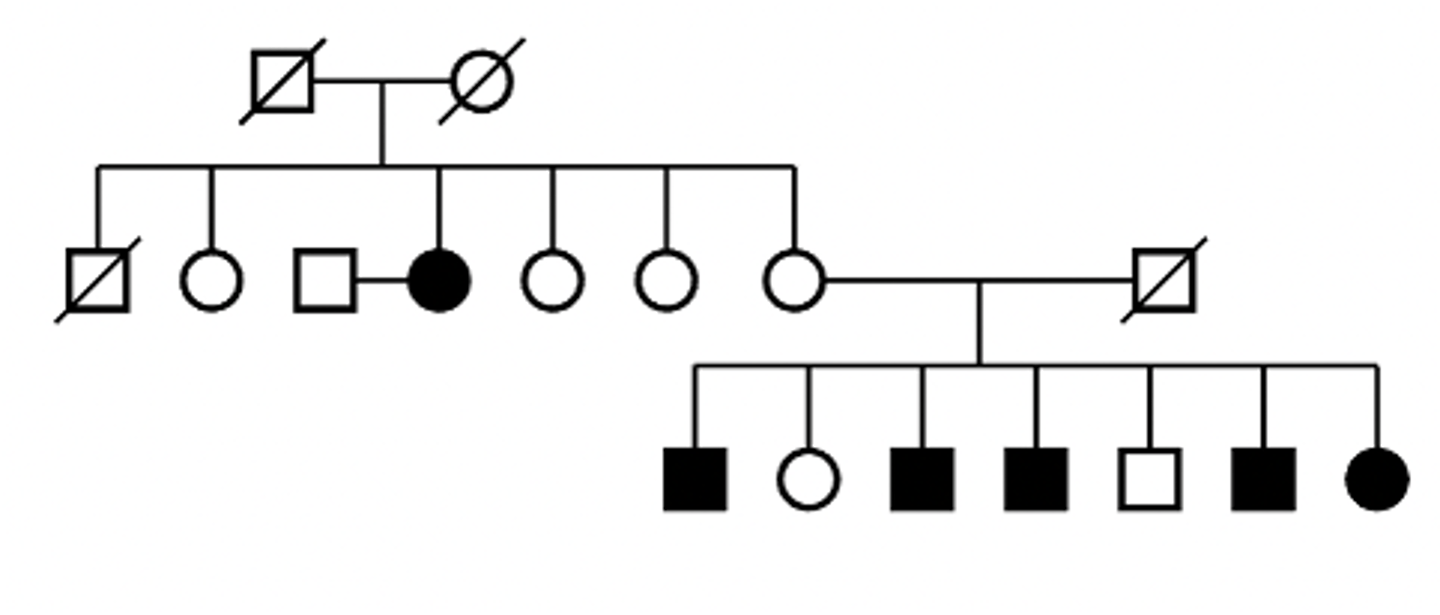
mtDNA disorders
- three differences distinguish disease caused by mtDNA mutations from Mendelian diseases:
1. maternal inheritance
2. heteroplasmy
3. a combination of wild-type and mutant mitochondria at the level of a single cell
- threshold effect: it takes a certain number of mitochondria that are affected for the effect to be seen
Maternal inheritance
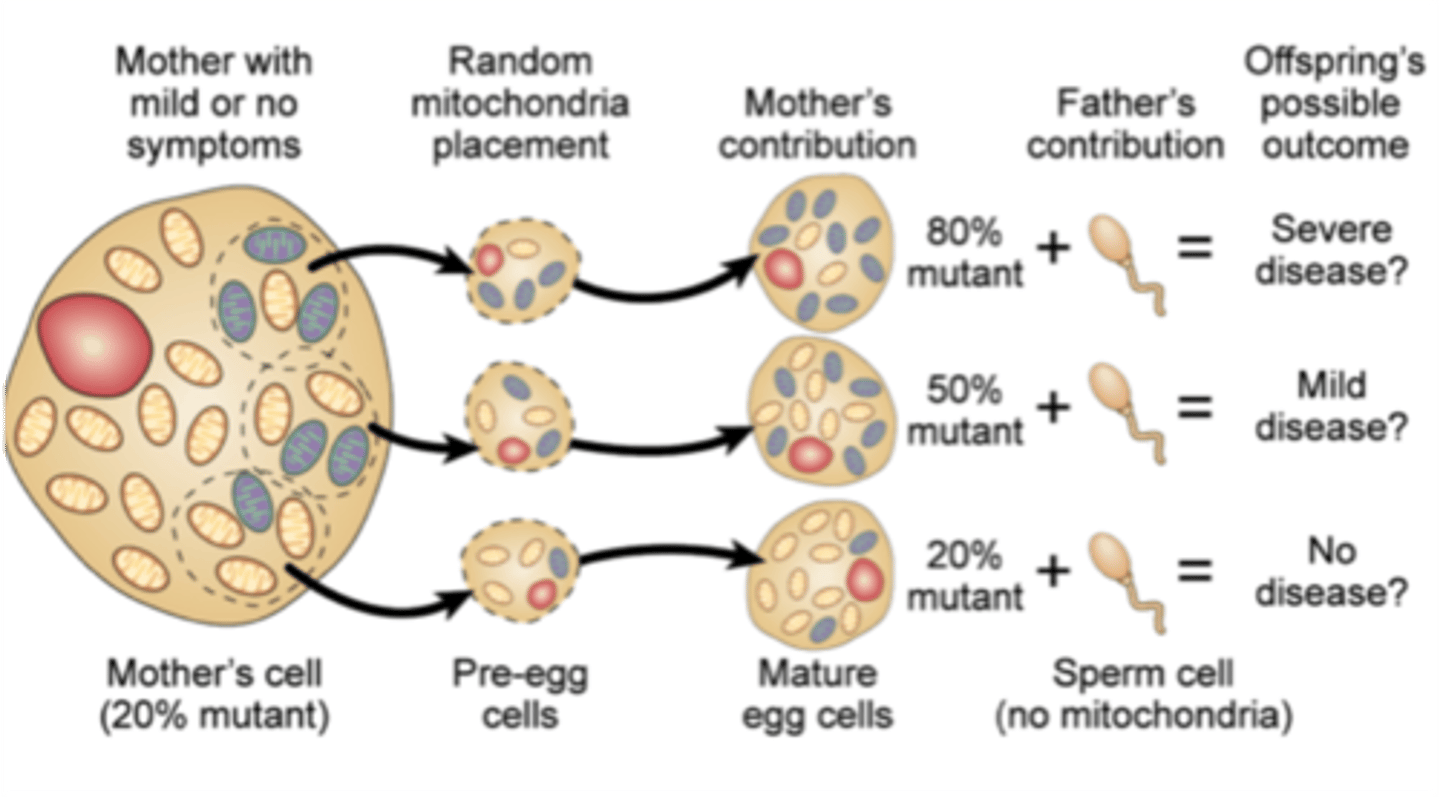
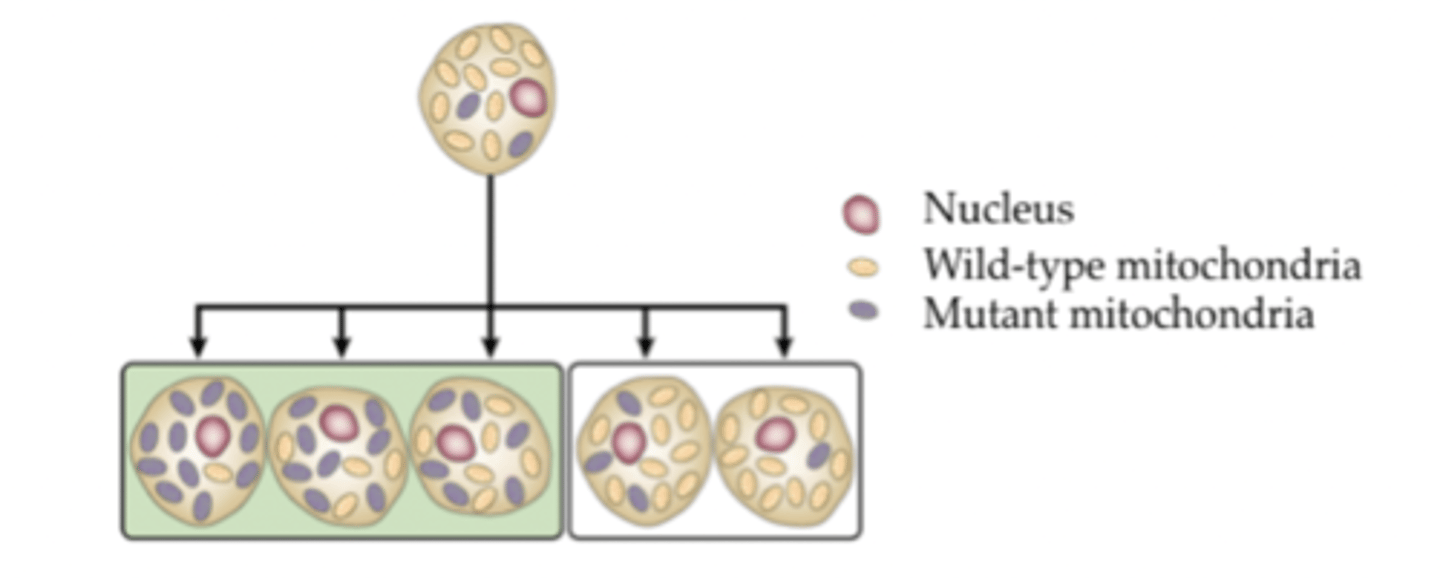
Heteroplasmy
- single cell has a combination of mutant and wild type mitochondria
- dividing cells unevenly distribute mutant mitochondria
- a critical number of mutant mitochondria in a cell can result in a disease phenotype
Threshold effect
- the number of mutant mitochondria that it takes for the effect to be seen
- different tissues have different energy requirements
- cell types consuming greater amounts of energy will be more sensitive to mutated mitochondria
Causes of single gene disorders
- missense mutation
- nonsense mutation
- structural variant
- non-synonymous point mutation
- synonymous point mutation
- insertions/deletions (indels) of single or multiple bases
Missense mutation
- a single base substitution in a codon that results in an amino acid substitution
- typically results in improper protein function
Nonsense mutation
- a single base substitution that changes a function codon into a stop codon
- typically results in a shortened protein
Structural variant
- deletion, duplication, translocation
Non-synonymous mutation
- alters amino acid encoded by codon
- both nonsense and missense mutations are non-synonymous
Synonymous point mutation
- does not alter amino acid encoded by codon
- not a cause of a genetic disorder
TATAA box
- promoter
- where RNA polymerase will bind to and begin transcribing
- transcription factors bind close to TATAA box to draw RNA polymerase in (can enhance or repress transcription)
Location of mutations in the gene
- mutations may occur in introns as well as exons
- mutations near either end of the intron may cause a change in splicing of exons and produce an abnormal protein
- DNA alteration flanking or intergenic locations may cause change in transcription or translation
Enhancer mutation
- DNA sequence to which activators of transcription bind
Silencer mutation
- DNA sequence to which repressors of transcription bind
Activators
- a transcription factor that binds to DNA and increases the rate of transcription
- protein
Repressors
- transcription factor that reduces the rate of transcription
- protein
Genetic heterogeneity
- mutations in different genes causing the same phenotype
- mutations in the same gene causing the same phenotype
F-value
- the more likely that a gene is related to the trait you're looking for, the higher the F-value
How can genes with recessive alleles that are lethal before birth be identified?
- reduced reproductive performance (lower apparent conception, pregnancy, or non-return rate)
- absence of homozygote genotypes among individuals in the population
Process to identify recessive alleles (Van Raden et al.)
1. large numbers of animals in breed genotyped with 50k SNP chip
2. haplotypes deduced from genotype and pedigree data
3. determine haplotype frequency
4. identify haplotypes that are relatively frequent yet absent in a homozygous state
5. test method for genomic locations of known lethals
6. examine haplotype effect on sire conception rate and stillbirth rate
What do you do with information of recessive lethal alleles?
- selection against carriers based on haplotypes
- plan matings to avoid carrier x carrier
- move on to identify gene and causative motive
Linkage equilibrium
- independence between genotype frequencies at two or more loci