Nucleic Acids and Enzymes
1/64
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
65 Terms
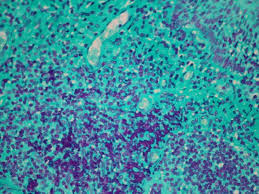
Feulgen reaction purpose (2)
demonstrate DNA, DNA quantification for tumor pathology
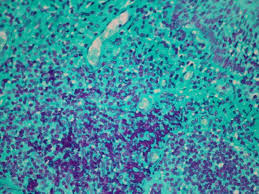
Feulgen reaction Mode of Action
HCl causes hydrolysis of DNA, removes purine bases (A and G), creates aldehyde groups that can be stained with Schiff
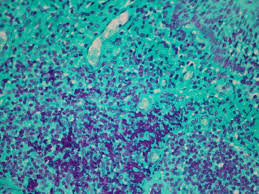
Feulgen Reaction reagents
Hydrozyler:
Primary stain:
Differentiator:
Counter stain:
Hydrozyler: HCl
Primary stain: Schiff reagent
Differentiator: sulfurous acid
Counter stain: light green
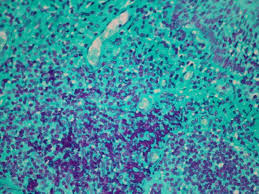
Feulgen reaction results
reddish purple:
light green:
reddish purple: DNA
light green: cytoplasm
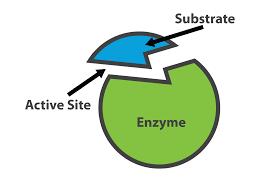
Fixative rules for Enzymes (2 exceptions)
No fixatives used except:
Naphthol ASD chloroacetate esterase uses methanol with formaldehyde
phosphorylase uses cold acetone
Fixative rules for nucleic acids (1 exception)
10% NBF used except:
Feulgen uses any fixative EXCEPT bouins
Feulgen reaction QC
thickness:
Feulgen reaction QC
thickness: 4 um
Methyl green pyronin y purpose (2)
differentiate between RNA and DNA
identify plasma cells or immunoblasts in tissue
methyl green pyronin y mode of action
DNA is more polymerized so methyl green binds to it, and RNA is less polymerized so pyronin y binds to it
methyl green pyronin y QC:
thickness:
section must contain:
thickness: paraffin, 4 um
section must contain: plasma cells
methyl green pyronin y reagents:
pH:
nucleic acid stain:
methyl green purifier:
pH: acetate buffer solution
nucleic acid stain: methyl green pyronin y staining solution
methyl green purifier: chloroform
methyl green pyronin y results:
green to blue green:
red to rose:
mint green:
pale pink to colorless:
intense red:
green to blue green: DNA and nuclei
red to rose: RNA
mint green: goblet cells
pale pink to colorless: backgrounds
intense red: immunoblast and cell cytoplasm
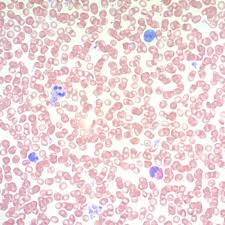
May grunwald giemsa purpose (2)
diffferentiate cells in bloody tissue
demonstrate microorganisms
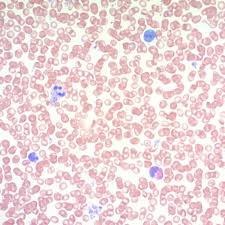
may grunwald giemsa mode of action
combining basic methylene blue and acidic eosin gives a wide color range due to impurities
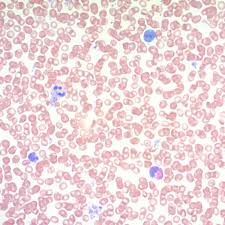
may grunwald giemsa QC:
thickness:
tissue:
thickness: paraffin, 3-4 um
tissue: spleen
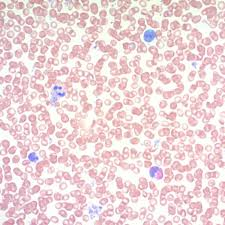
may grunwald giemsa reagents:
primary stain:
secondary stain:
differentiator:
secondary fixative:
primary stain: giemsa solution
secondary stain: jenner solution
differentiator: acetic acid
secondary fixative:methanol
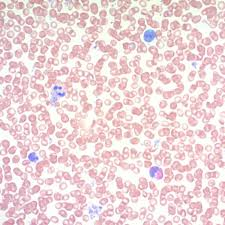
may grunwald giemsa results:
blue:
pink/gray/blue
blue: nuclei and microorganisms
pink/gray/blue: white blood cells
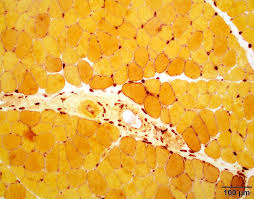
alpha naphthyl acetate esterase purpose (2)
differentiate between type 2 muscle and neurogenic atrophy
demonstrate motor end plates and lysosomes
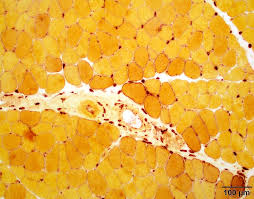
alpha naphthyl acetate esterase mode of action (3)
esterases hydrolyze aliphatic and aromatic ester bonds
classified as specific or nonspecific
stains histiocytes and acetylcholinesterase in denervated fibers
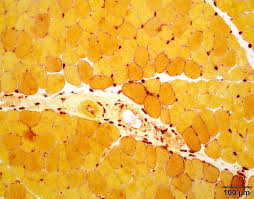
alpha naphthyl acetate esterase equation (3)
basic fuchsin + sodium nitrate → hexazotized pararosaniline
alpha naphthyl acetate + water —-esterases→ alpha naphthol + acetic acid
alpha naphthol + hexazotized pararosaniline → azo dye
alpha naphthyl acetate esterase reagents:
incubation solution:
adjust pH:
incubation solution:
pH 7.2 phosphate buffer
pararosaniline
adjust pH: HCl or NaOH
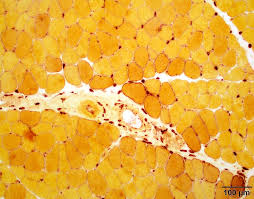
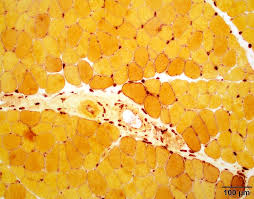
alpha naphthyl acetate esterase results:
brick red:
pale yellow:
dark red-brown:
brick red:motor endplates
pale yellow: normal muscle fibers
dark red-brown: denervated muscle fibers, macrophages, lysosomes
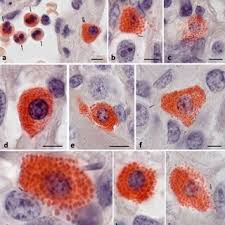
naphthol ASD chloroacetate esterase (Leder) purpose
identify granulocytes in the classification of leukemias or chloromas
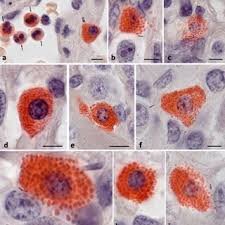
naphthol ASD chloroacetate esterase (Leder) mode of action (2)
esterases hydrolyze aliphatic and aromatic ester bonds
specific esterase stain
naphthol ASD chloroacetate esterase (Leder) reagents:
incubation solution:
adjust pH:
incubation solution:
pararosaniline
sodium nitrate
adjust pH: HCl or Na barbitol
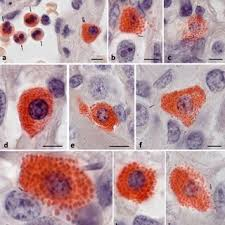
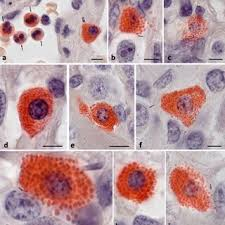
naphthol ASD chloroacetate esterase results:
bright red:
blue:
unstained:
bright red: positive cells, granulocytes, mast cells
blue: nuclei
unstained: backgrounds
atpase purpose (2)
differentiate between type 1, 2A, 2B muscle fibers
separate myopathic from nuropathic processes
atpase mode of action (2)
ATPase is pH dependent, on the type of muscle fiber
metal precipitation of hydrolyzed orthophosphoric acid with calcium
atpase equations (4)
ATP + water —-ATPase→ ADP + H3PO4 + energy
H3PO4 + CaCl2 → CaHPO4 + 2H + 2Cl
CaHPO4 + CoCl2 → CoHPO4 + CaCl2
CoHPO4 + (NH4)2S → CoS + (NH4)2HPO4
atpase reagents:
adjust pH:
incubation solution:
activate muscle and inhibit mitochondrial atpase:
adjust pH: HCl or NaOH
incubation solution:
sodium barbitol
activate muscle and inhibit mitochondrial atpase: calcium chloride
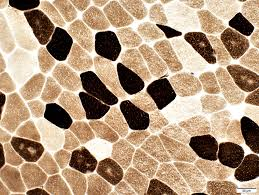
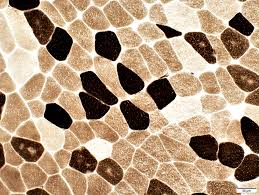
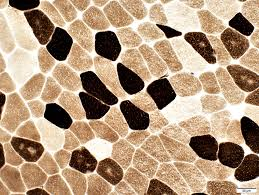
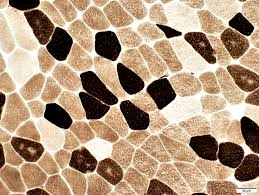
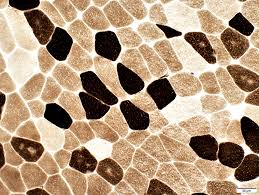
atpase results:
pH 9.4 slide:
dark:
light:
pH 4.3 slide:
dark:
light:
pH 4.6 slide:
dark:
light:
medium:
pH 9.4 slide:
dark: type 1
light: type 2
pH 4.3 slide:
dark: type 1
light: type 2
pH 4.6 slide:
dark: type 1
light: type 2A
medium: type 2B
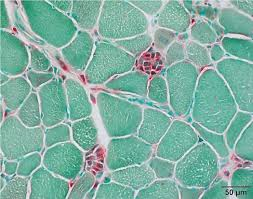
acid phosphatase purpose (2)
demonstrate inflammatory cells/lysosomes in muscle biopsies
demonstrate acid maltase deficiency: pompe disease, lysosomal storage disease
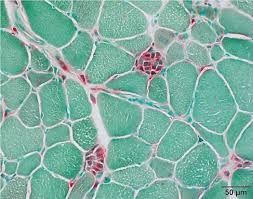
acid phosphatase mode of action
acid phosphatase hydrolyzes esters of orthophosphoric acid
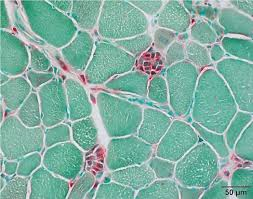
acid phosphatase equations (2)
naphthol ASBI phosphate + water —acid phosphatase→ naphthol ASBI + orthophosphatate
naphthol ASBI + hexazotized pararosaniline → red azo dye
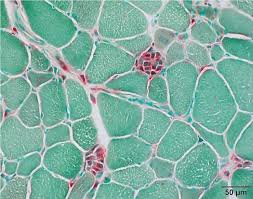
acid phsophatase reagents:
incubation medium:
pH:
counterstain:
incubation medium:
pararosaniline hydrochloride
sodium nitrate
pH: veronal acetate buffer 5.0
counterstain: methyl green
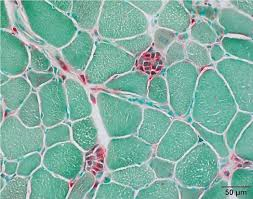
acid phosphatase results:
red:
green:
red: sites of acid phosphatase activity
green: background
alkaline phosphatase purpose (2)
detection of regenerating muscle fibers
diagnose inflammatory myopathy
alkaline phosphatase mode of action
alkaline phosphatase hydrolyzes esters of orthophosphoric acid
alkaline phosphatase equations (2)
naphthol ASBI phosphate + water —alkaline phosphatase→ naphthol ASBI + orthophosphatase
naphthol ASBI + fast red violet → red azo dye
alkaline phosphatase reagents:
incubation mediun:
pH:
primary stain:
counterstain:
incubation mediun:
pH: tris buffer 8.74
primary stain: fast red violet
counterstain: harris/mayer hematoxylin
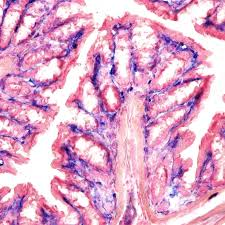
alkaline phosphatase results
pink red to red:
blue:
pink red to red: sites of alkaline phosphatase activity
blue: nuclei
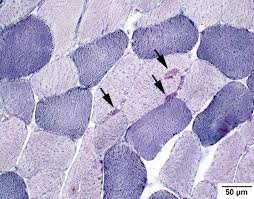
NADH diaphorase purpose (2)
demonstrate abnormalities in mitochondria, z band material, sarcoplasmic reticulum
demonstrate architectural changes in central cores, target fibers, nemaline rods, targetoid fibers
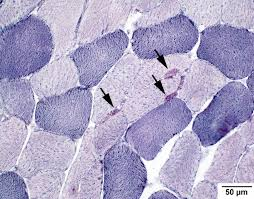
NADH disphorase mode of action (2)
diahoresis reaction
tetrazolium salts accept hydrogen, but is reduced by flavoprotein enzymes
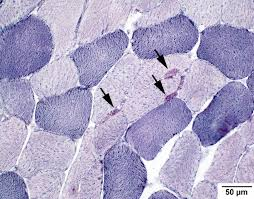
NADH disphorase equations (2)
NADH —-disphorase→ NAD+ + H
NBT (nitro blue tatrazolium) + H → formazan
NADH diaphorase reagents
incubation medium:
pH:
primary stain:
incubation medium:
pH: phosphate buffer 7.4
primary stain: NBT (nitro blue tetrazolium)
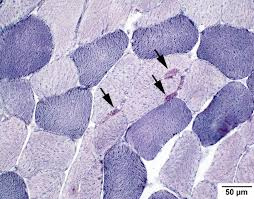
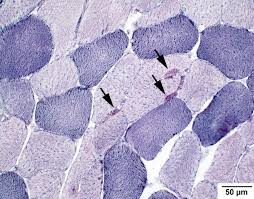
NADH diaphorase results:
dark purple deposits:
dark purple:
light purple:
dark purple deposits: sites of NADH diaphorase enzyme activity
dark purple: type 1 muscle fibers
light purple: type 2 muscle fibers
QC Rules for enzymes: cut at ____ um and use _____ tissue (1 exception)
___ uses spleen as a control
QC Rules for enzymes: cut at 10 um and use skeletal muscle tissue
naphthol chloroacetate esterase (Leder stain) uses spleen as a control
QC Rules for nucleic acids: cut at ____ um AND:
___ must contain plasma cells
___ uses spleen tissue
QC Rules for nucleic acids: cut at 4 um AND:
methyl green pyronin y must contain plasma cells
may grunwald giemsa uses spleen tissue
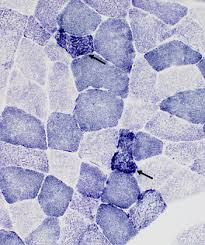
succinic dehydrogenase (SDH) purpose
identify the source of NADH diaphorase activity, because only mitochondria show positive SDH activity
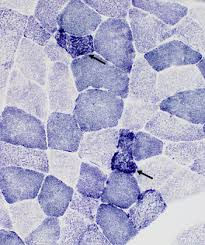
succinic dehydrogenase (SDH) mode of action
SDH substrate is incubated with nitro blue tetrazolium and formazan pigment is created
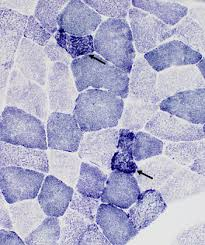
succinic dehydrogenase (SDH) equations (2)
succinate —succinic dehydrogenase (SDH)→ fumarate + hydrogen
NBT (nitro blue tetrazolium) + hydrogen -→ formazan
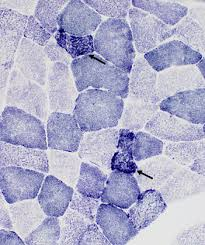
succinic dehydrogenase (SDH) reagents:
incubation medium:
pH:
rinse:
secondary fixative:
incubation medium:
NBT solution
pH: phosphate buffer 7.6
rinse: physiological saline
secondary fixative: formaline saline
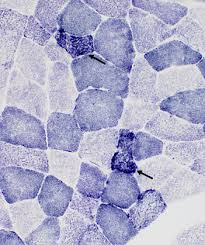
succinic dehydrogenase (SDH) results:
blue:
blue: sites of succinic dehydrogenase (SDH) activity
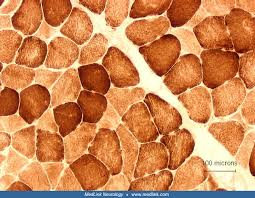
cytochrom c oxidase (COX) purpose
identify cytochrome c oxidase activity
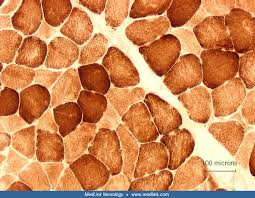
cytochrome c oxidase (COX) mode of action
diaminobenzidine (DAB) is an electron donor to show cytochrome c oxidase (COX) activity which will form a soluble polymer
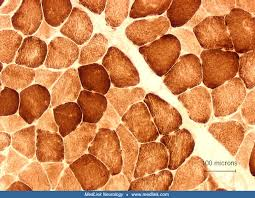
cytochrome c oxidase (COX) reagents:
incubation medium:
blocking:
pH:
mounting media:
incubation medium:
blocking: catalase
pH: sorenson buffer 7.4
mounting media: toluene based
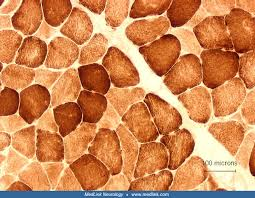
cytochrome c oxidase (COX) results
brown:
brown: sites of cytochrome c oxidase (COX) activity
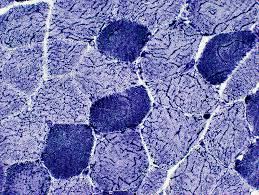
phosphorylase purpose
diagnose mcardle disease
phosphoylase mode of action
phosphorylase creates unbranched amylose, whose length is proportional to phosphorylase activity
phosphorylase reagents:
incubation medium:
prevent excessive branching:
primer:
pH:
activator:
visualizer:
mounting medium:
incubation medium:
prevent excessive branching: alcohol
primer: glycogen
pH: acetate buffer 5.9
activator: insulin
visualizer: gram iodine
mounting medium: iodine-glycerin medium
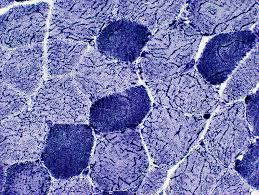
phosphorylase results:
blue/puple:
colorless:
blue/puple: phosphorylase activity
colorless: mcardle disease
modified gomori trochrome reagents:
gomori trichrome solution:
myofibril stain:
muscle feature stain:
differentiator for muscle:
pH:
differentiator at end:
nuclei stain:
gomori trichrome solution:
myofibril stain: fast green fcf
muscle feature stain: chromotrope 2R
differentiator for muscle: phosphotungstic acid
pH: glacial acetic acid
differentiator at end: acetic acid
nuclei stain: harris hematoxylin
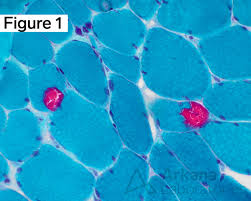
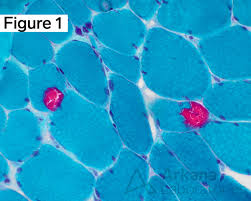
modified gomori trichrome results:
red/purple:
blue/green:
red:
red/purple: nuclei
blue/green: myofibrils
red: intermyofibrillar sarcoplasm, myelinated nerve twigs, nemaline rods, abnormal mitochondria
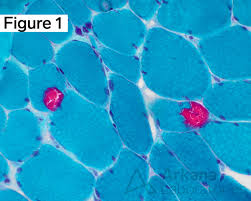
modified Gomori trichrome purpose
identify structural abnormalities in muscle to check for mitochondrial dysfunction
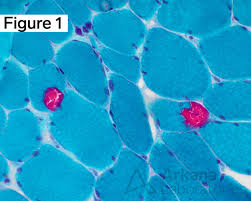
modified gomori trichrome mode of action
large tungstate ion is taken up by thick collagen fibers and it stains this complex
small chromotrope 2R molecules taken up by thin muscle fibers, and it stains the complex