BCH4024 Exam 3 (Dr. Aleman)
1/215
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
216 Terms
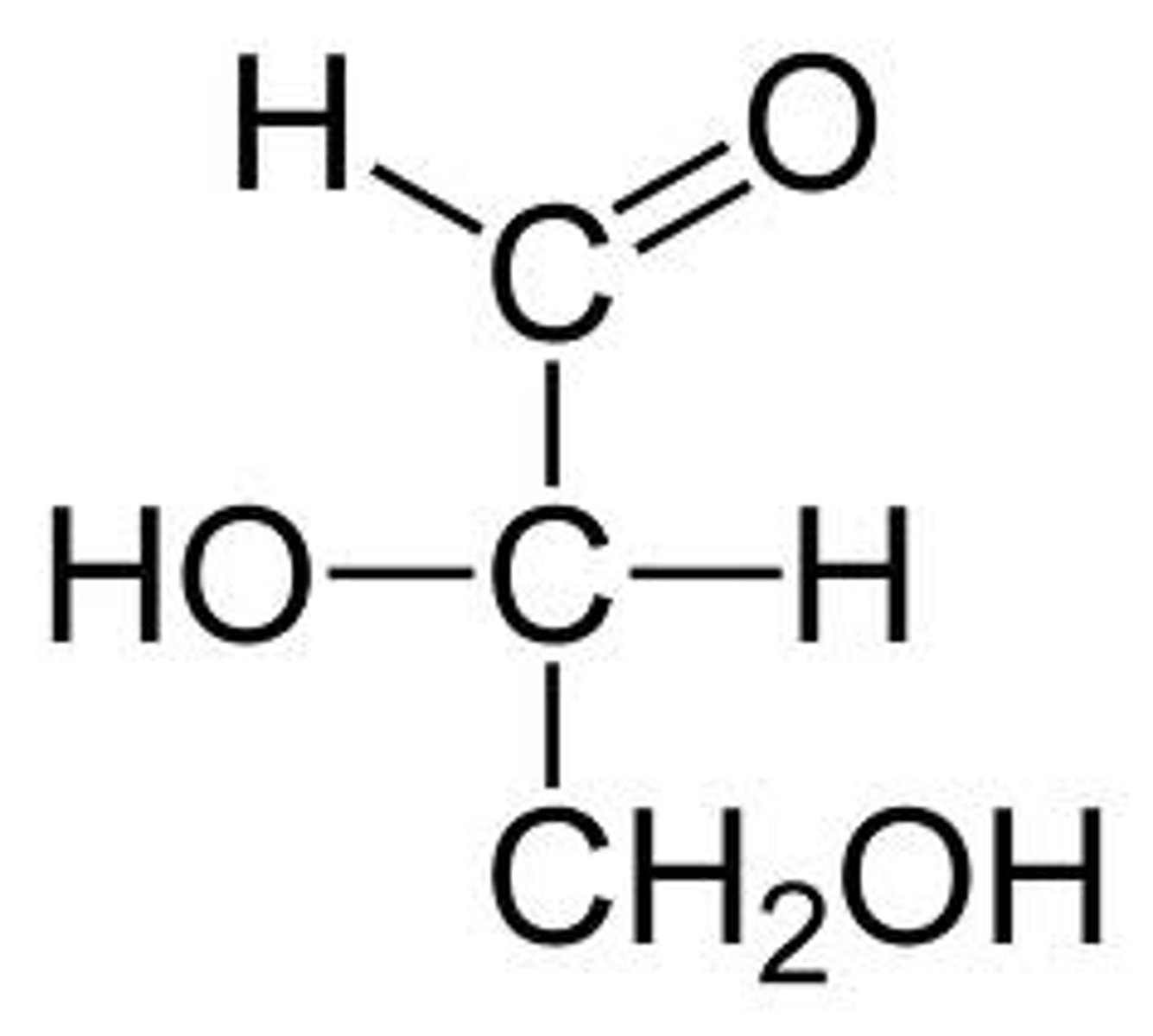
D-glyceraldehyde
aldotriose (aldose, three carbons)
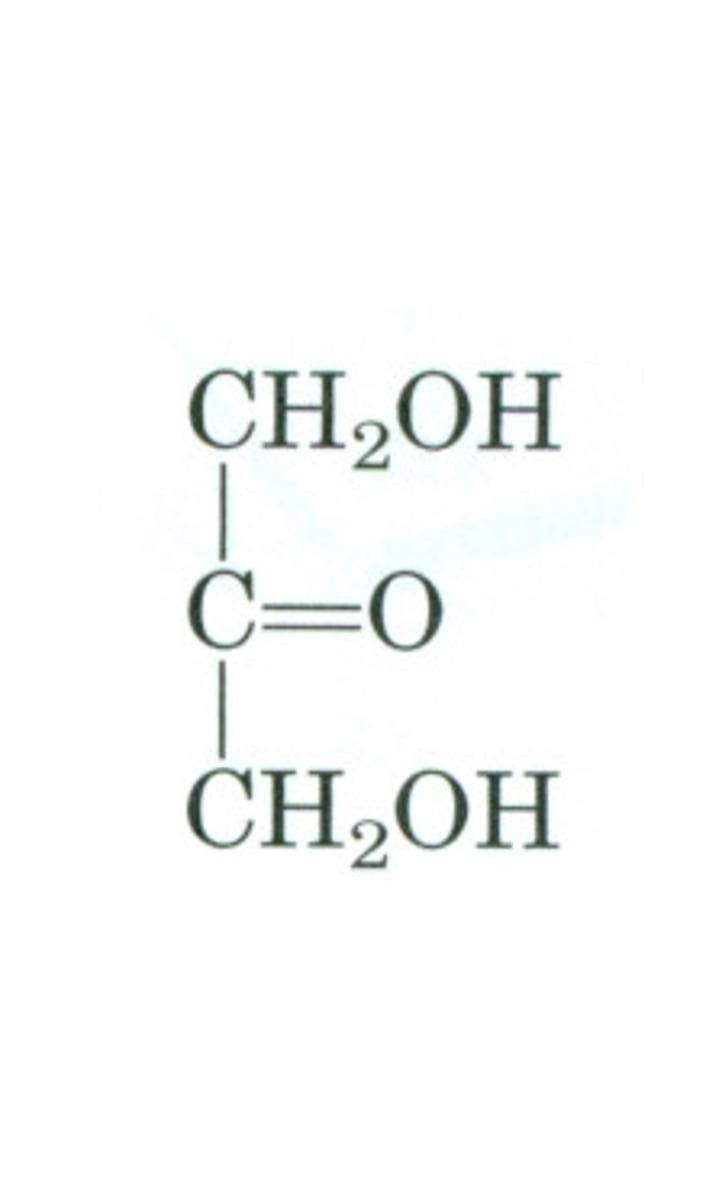
dihydroxyacetone
ketotriose (ketose, three carbons)
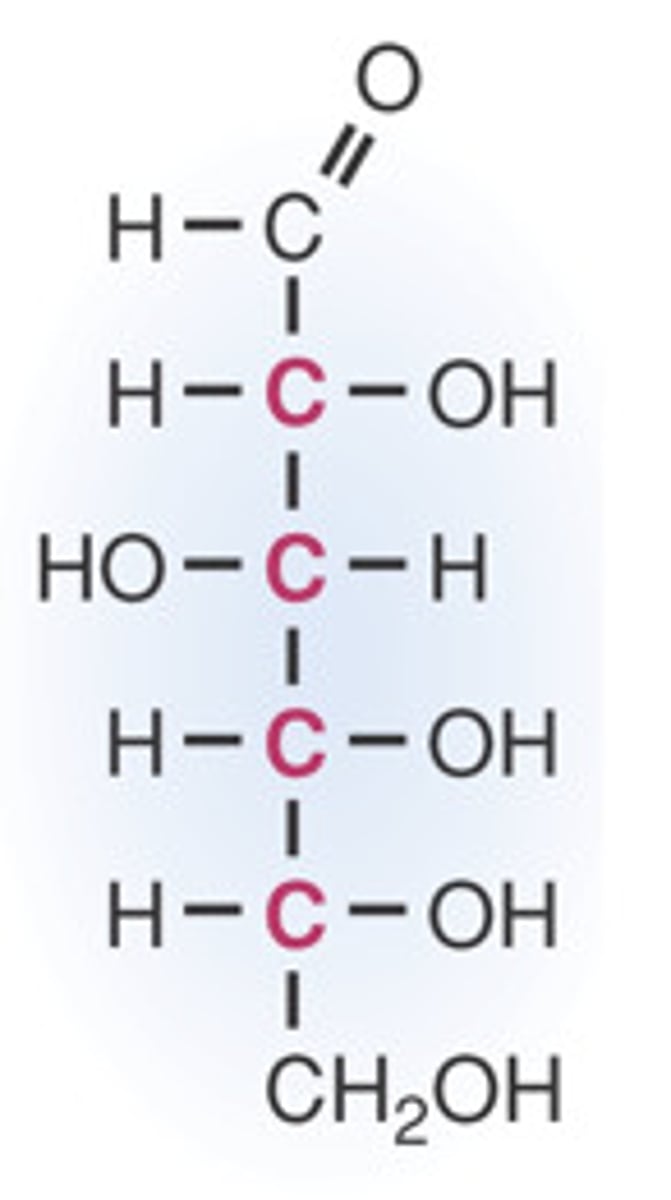
D-glucose
aldohexose
D-fructose
ketohexose
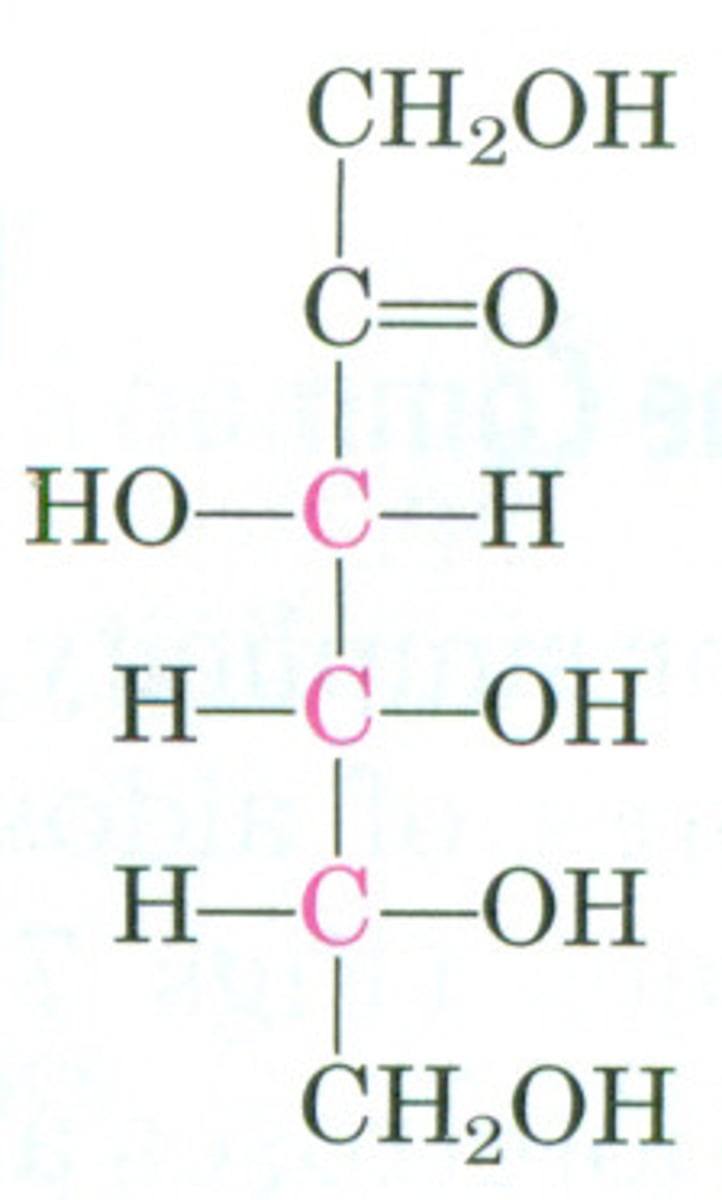
D-ribose
aldopentose
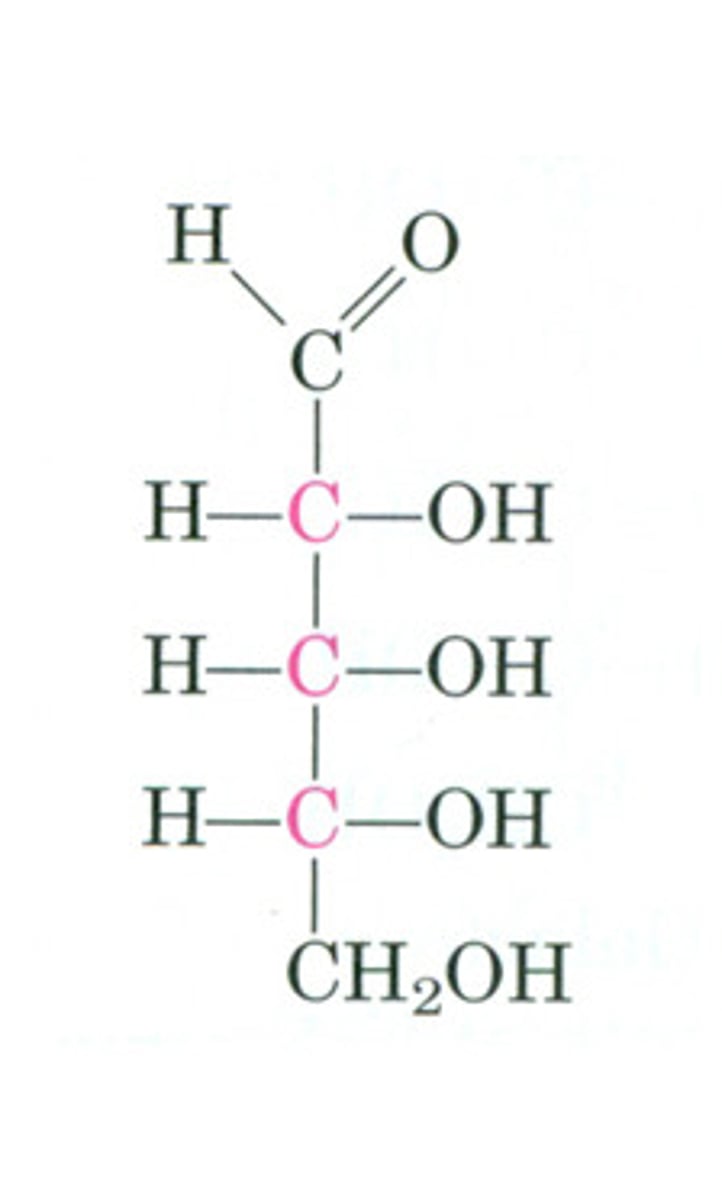
2-deoxy-D-ribose
aldopentose
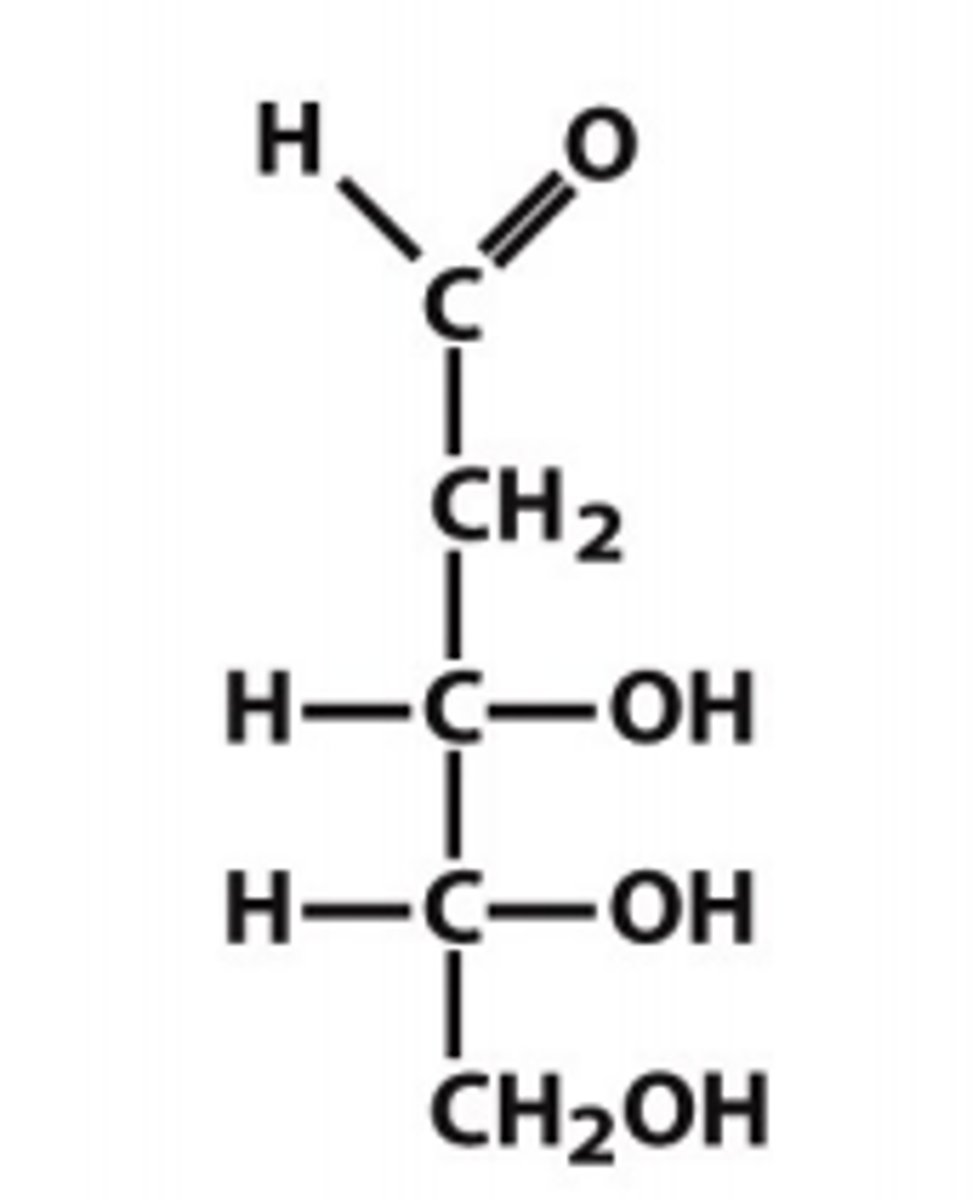
reference carbon
the chiral carbon most distant from the carbonyl carbon
aldose
aldehyde group at the end
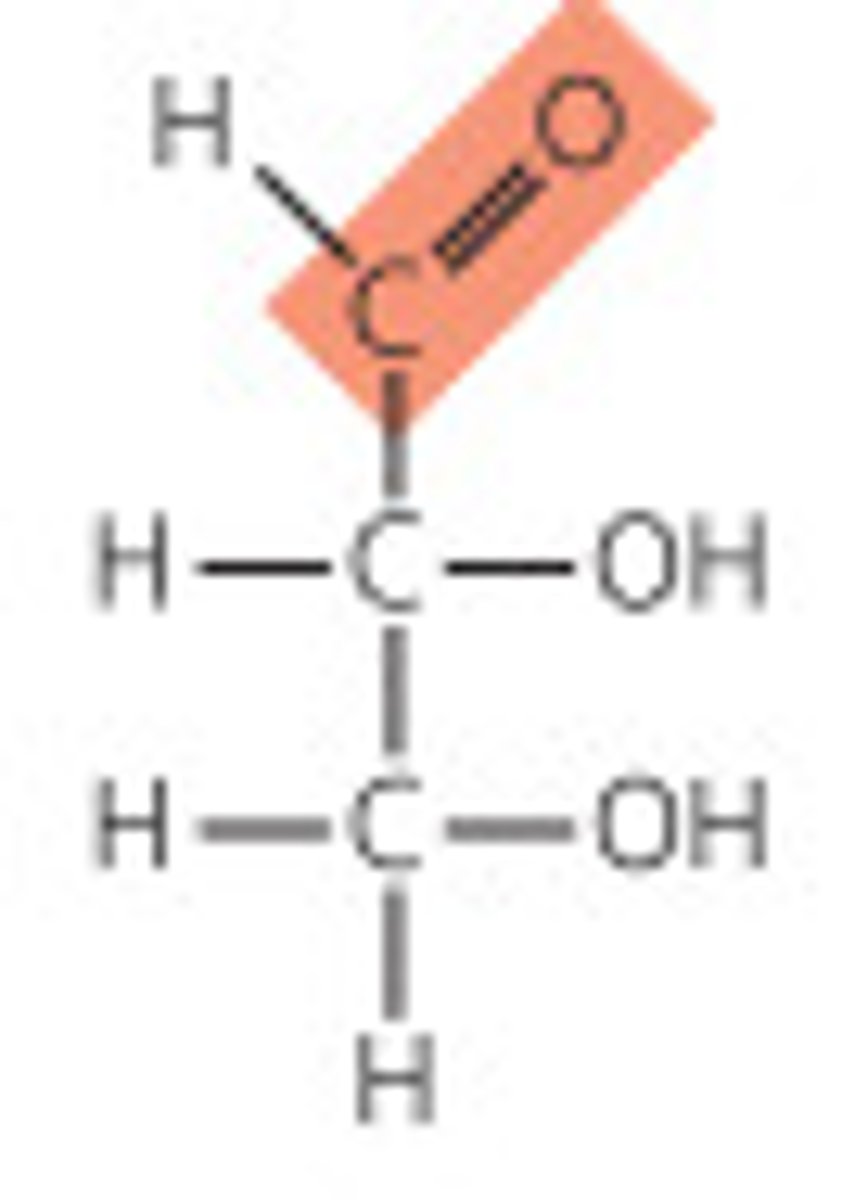
ketose
ketone group at the end
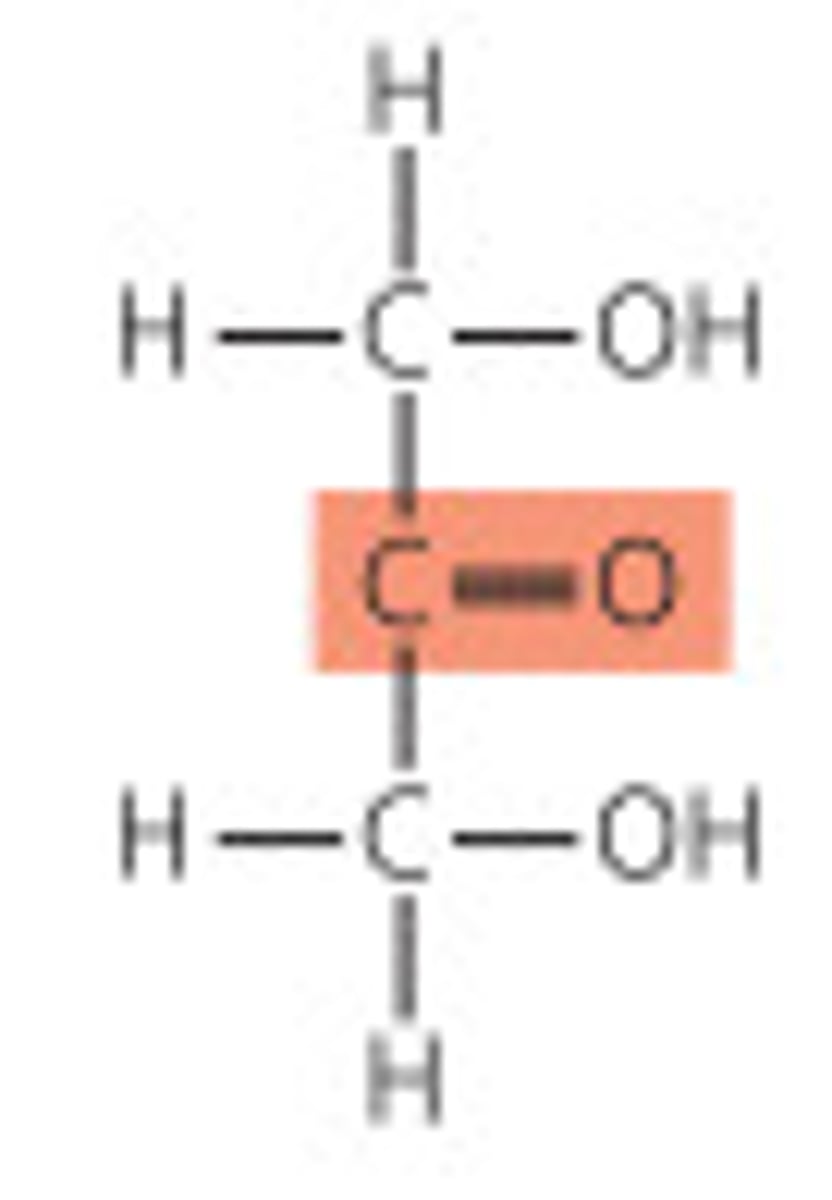
epimers
two sugars with ONLY one chiral center that has opposite configuration
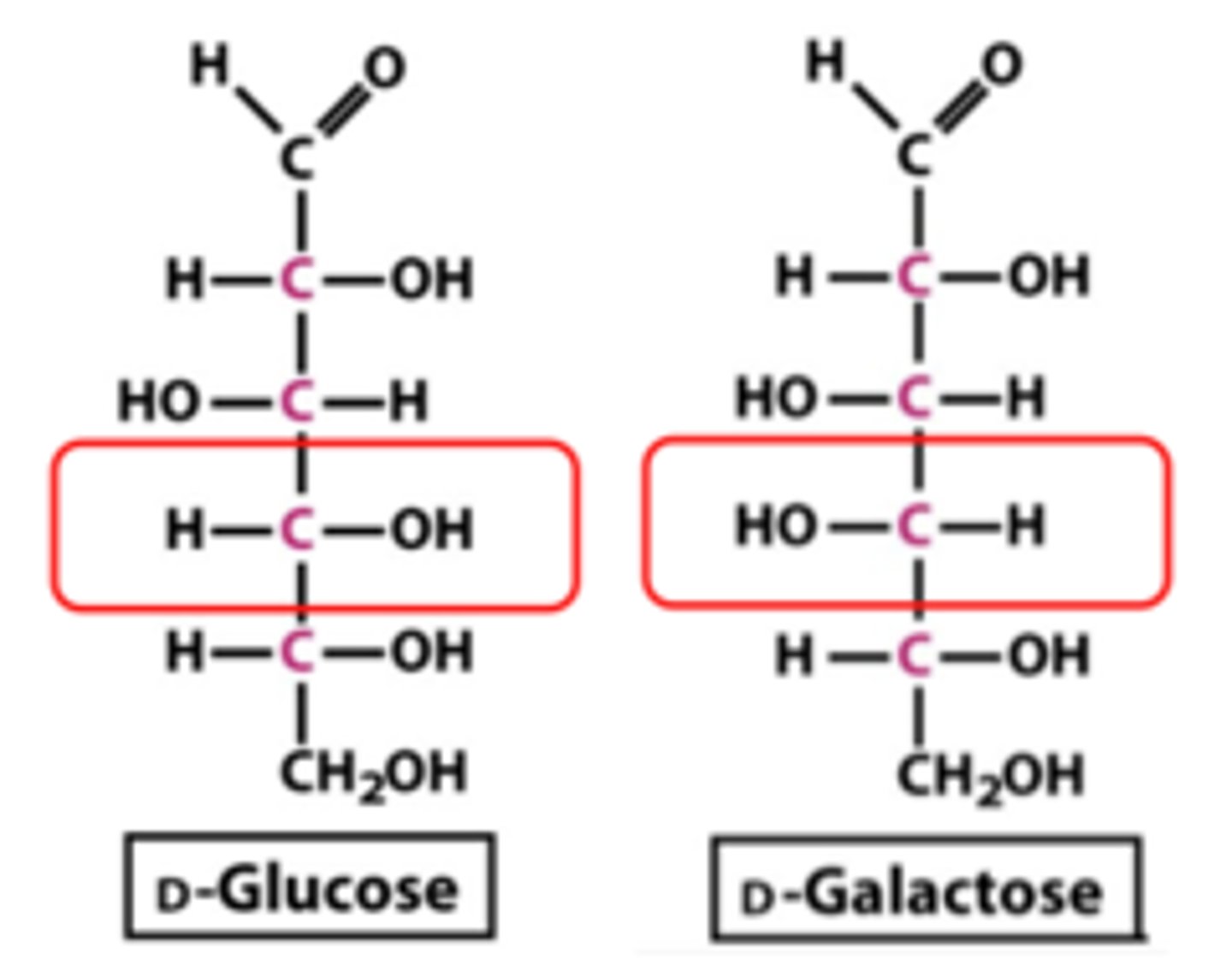
anomeric carbon
only carbon attached to two oxygens, -OH may point up or down. used to be a carbonyl
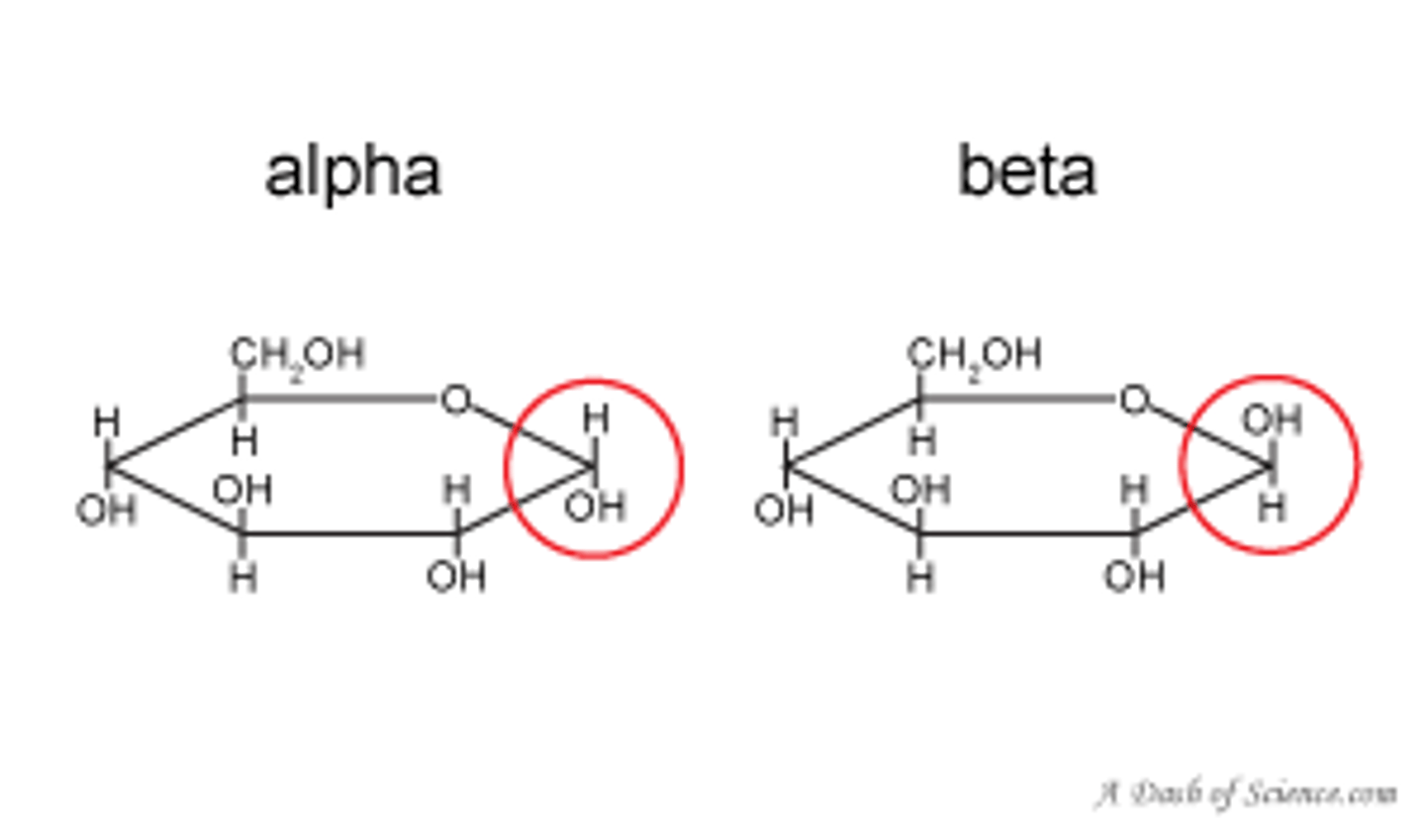
reducing sugars
has a hemiacetal (OH next to C-O) on either anomeric carbons that can form glycosidic bonds
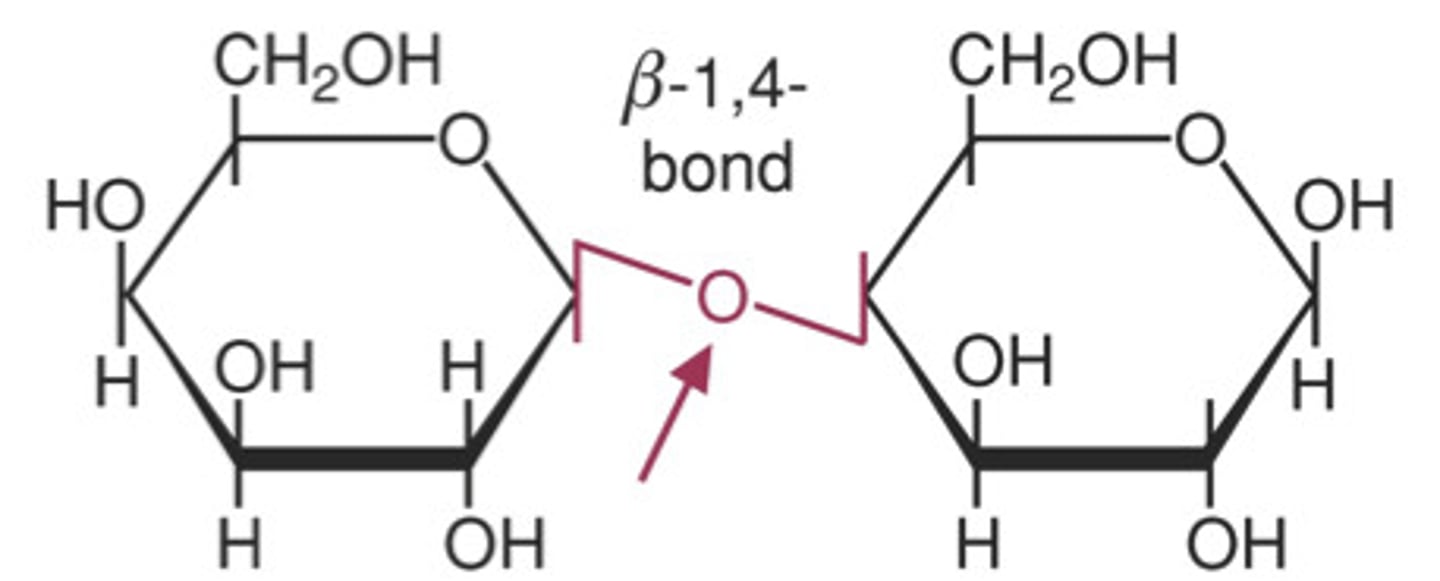
non-reducing sugars
does not have a hemiacetal on either anomeric carbons
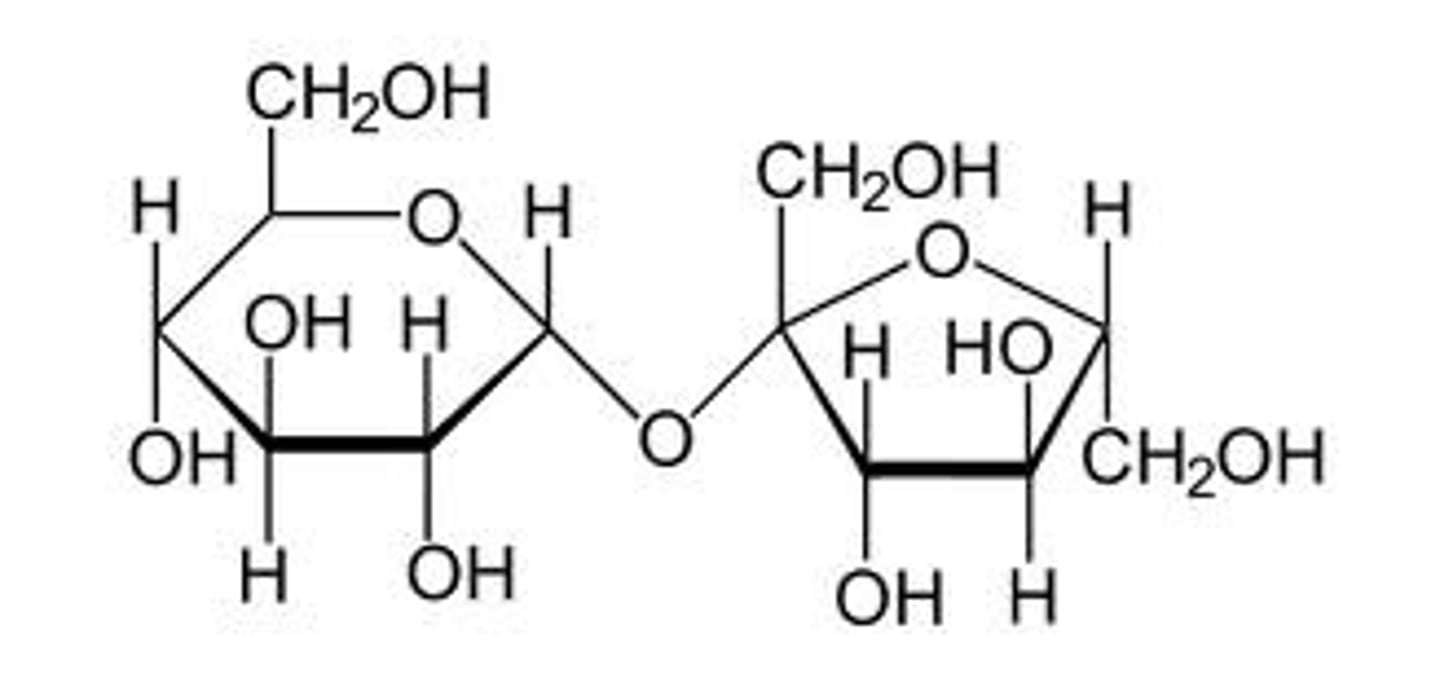
pyranose
6-membered heterocyclic ring
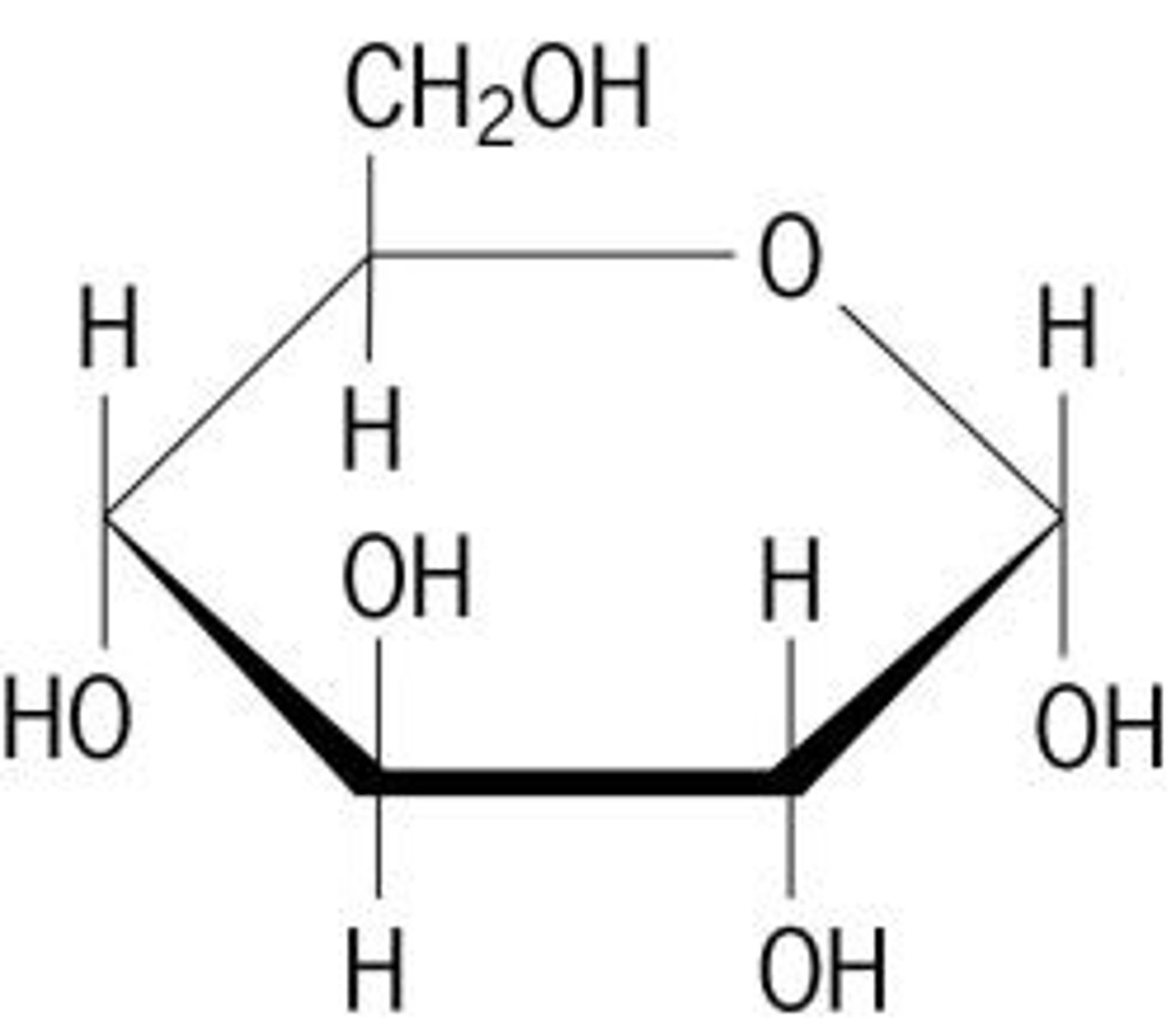
furanose
5-membered heterocyclic ring
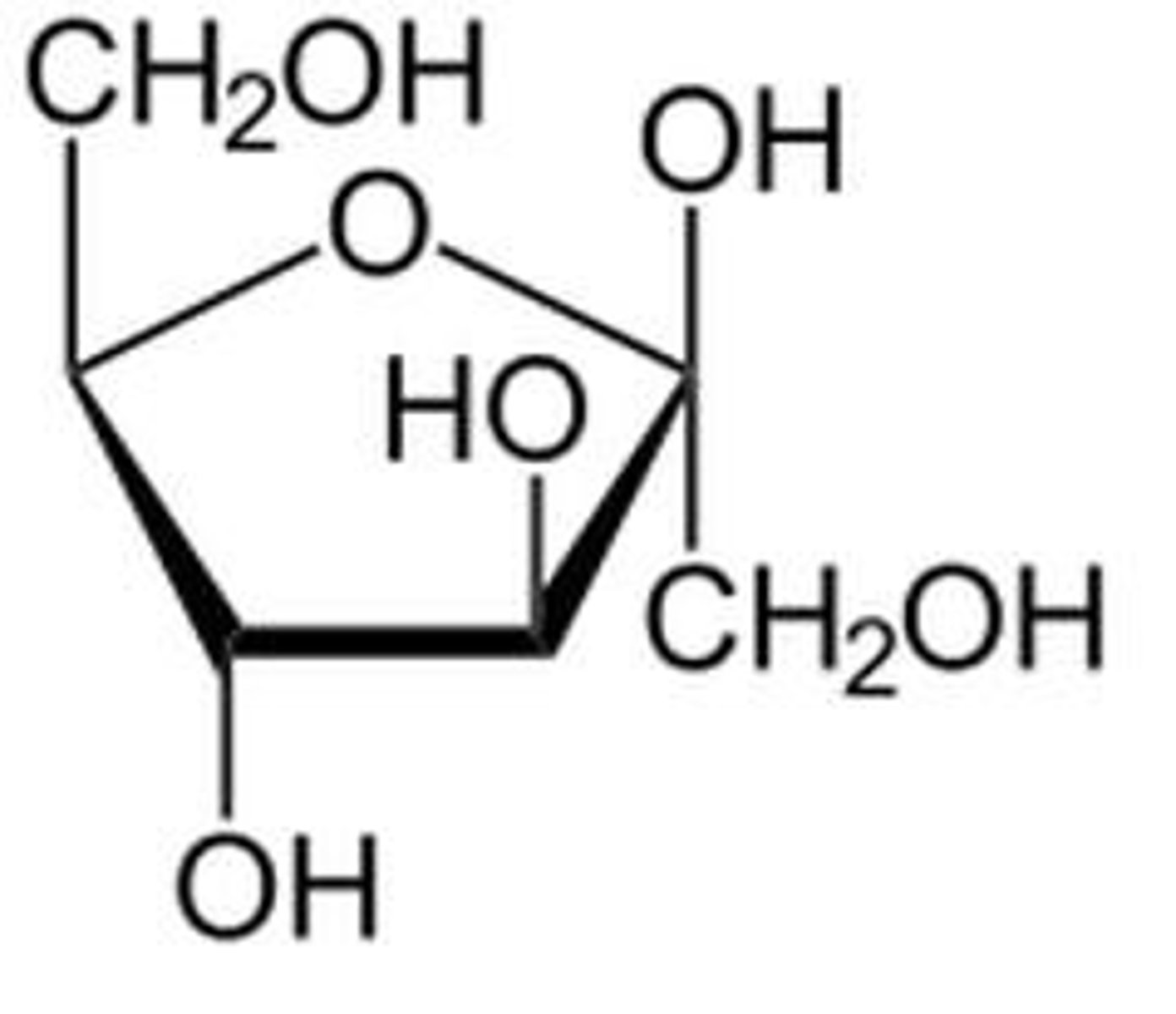
hemiacetal
single carbon bonded to two oxygens (RO-C-OH)
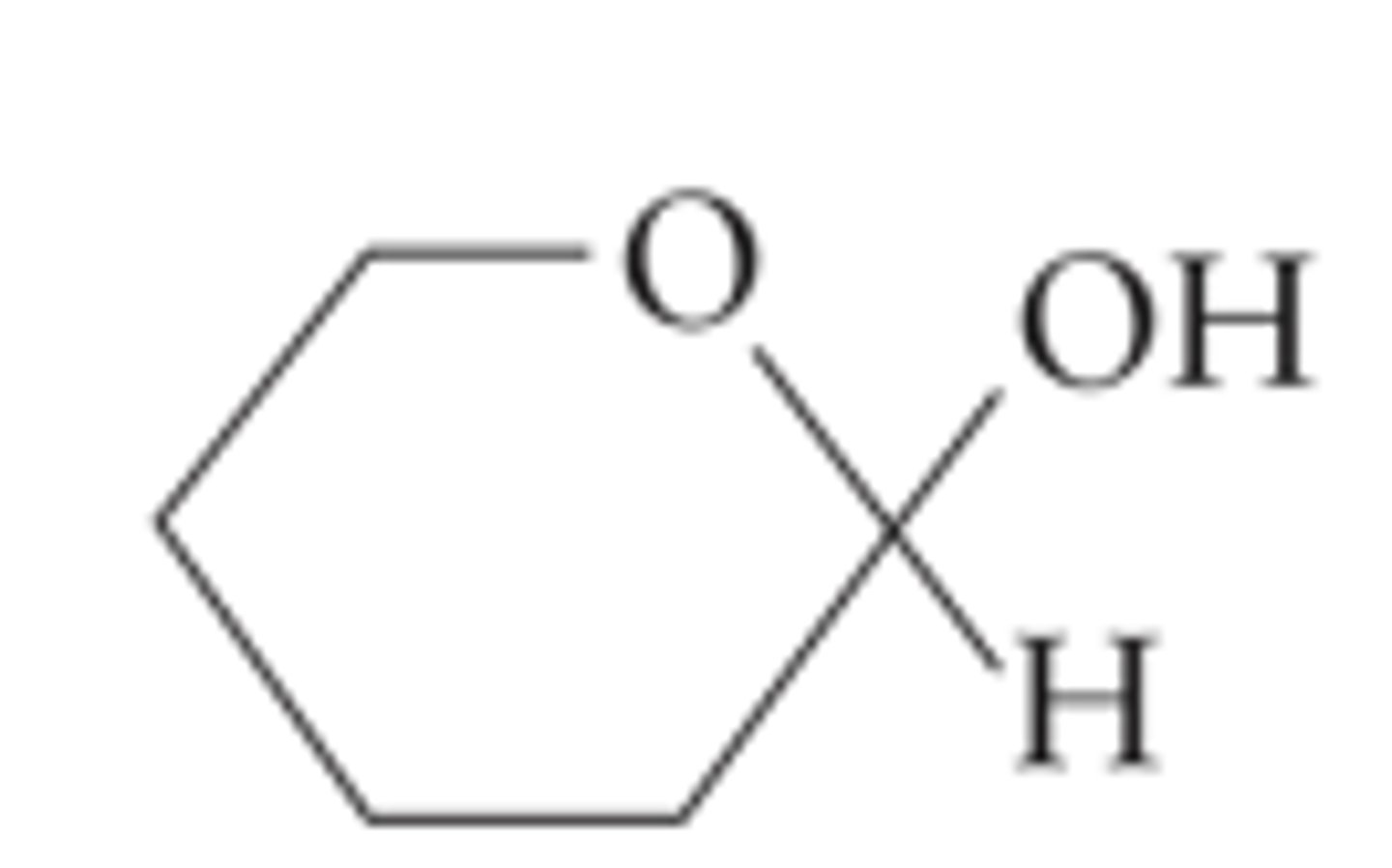
glycosidic bond (α)
covalent bond between two cyclic sugars, O's are on the same plane
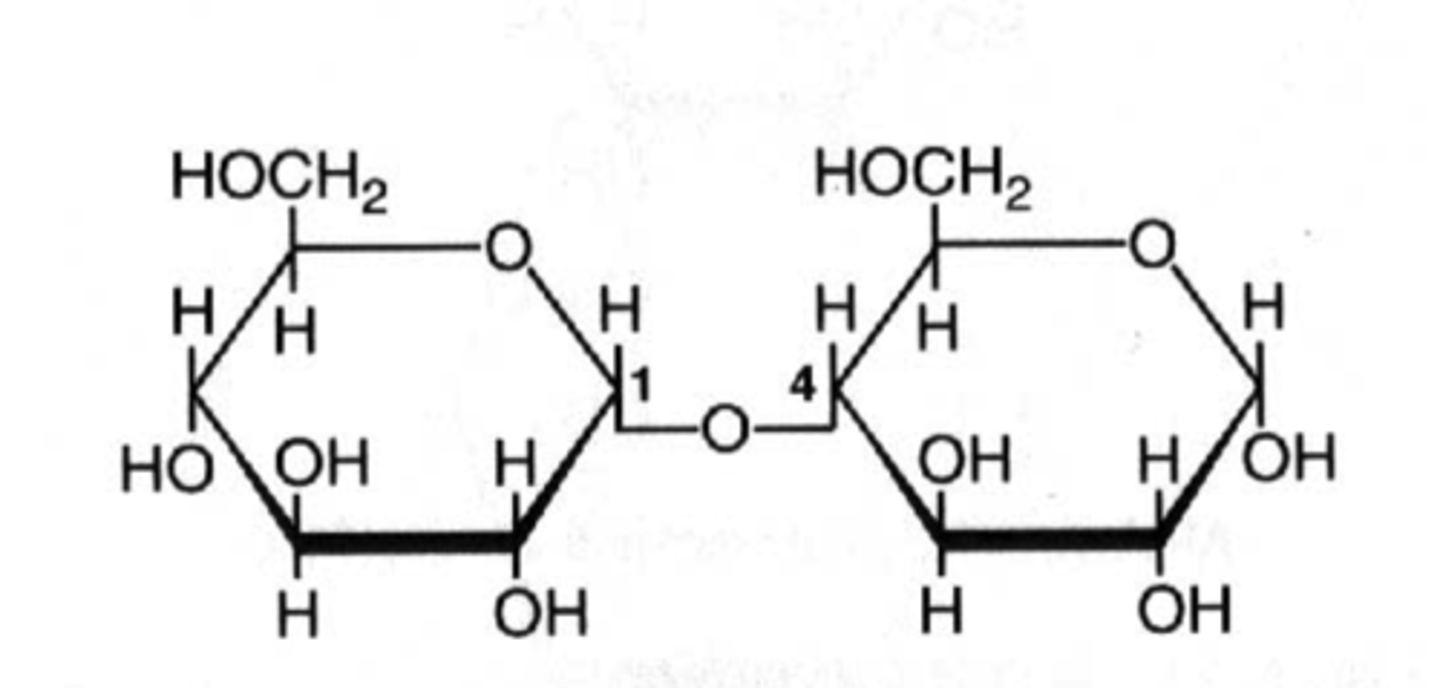
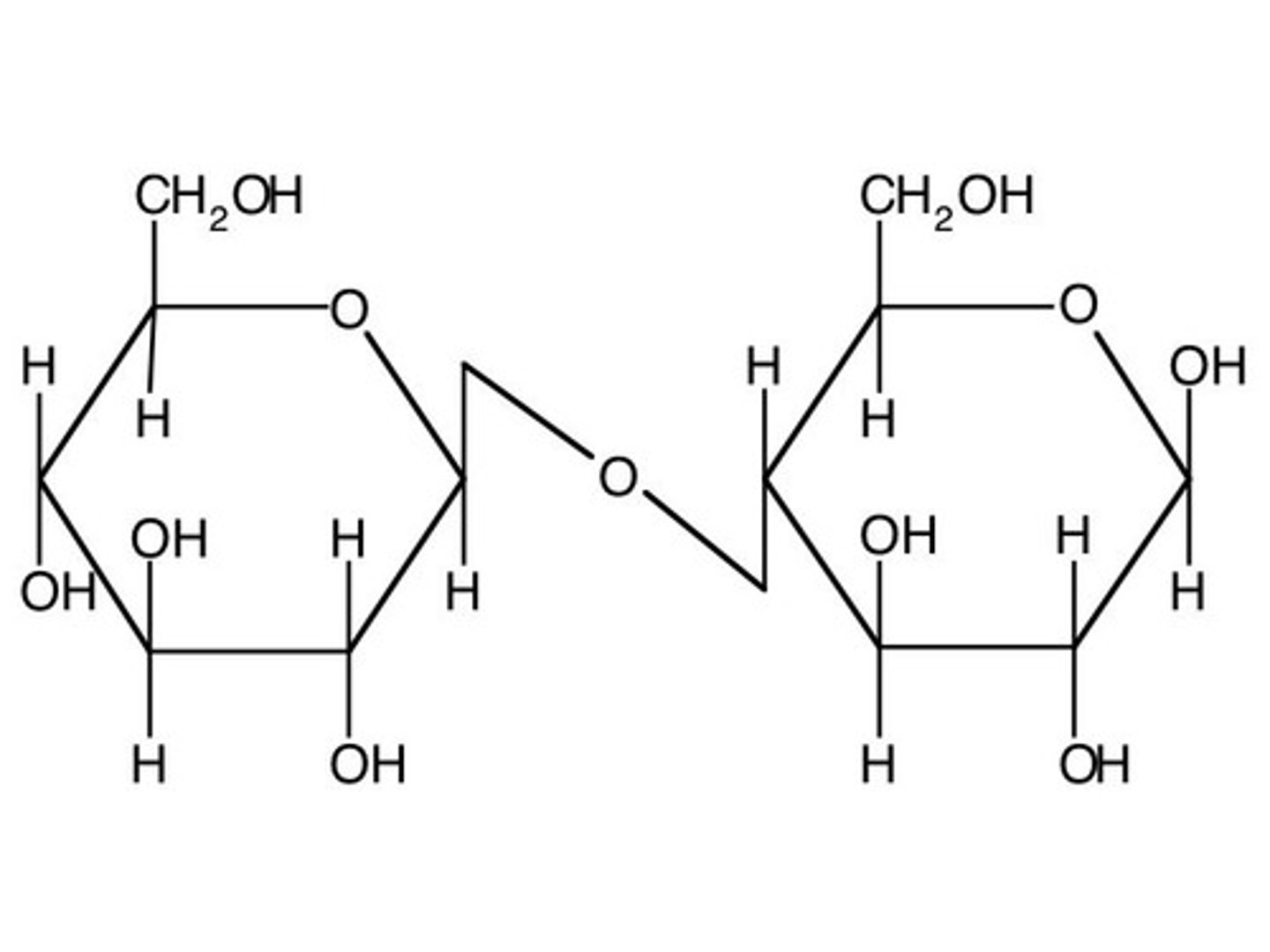
glycosidic bond (β)
covalent bond between two cyclic sugars, O's are on different planes
glycogen
an extensively branched glucose storage polysaccharide, broken down at low glucose levels
monosaccharides
simple sugars, one polyhydroxyl aldehyde or ketone unit. freely soluble in water
disaccharides
oligosaccharides with two monosaccharide units
polysaccharides
sugar polymers with 10+ monosaccharide units
sucrose
glucose + fructose
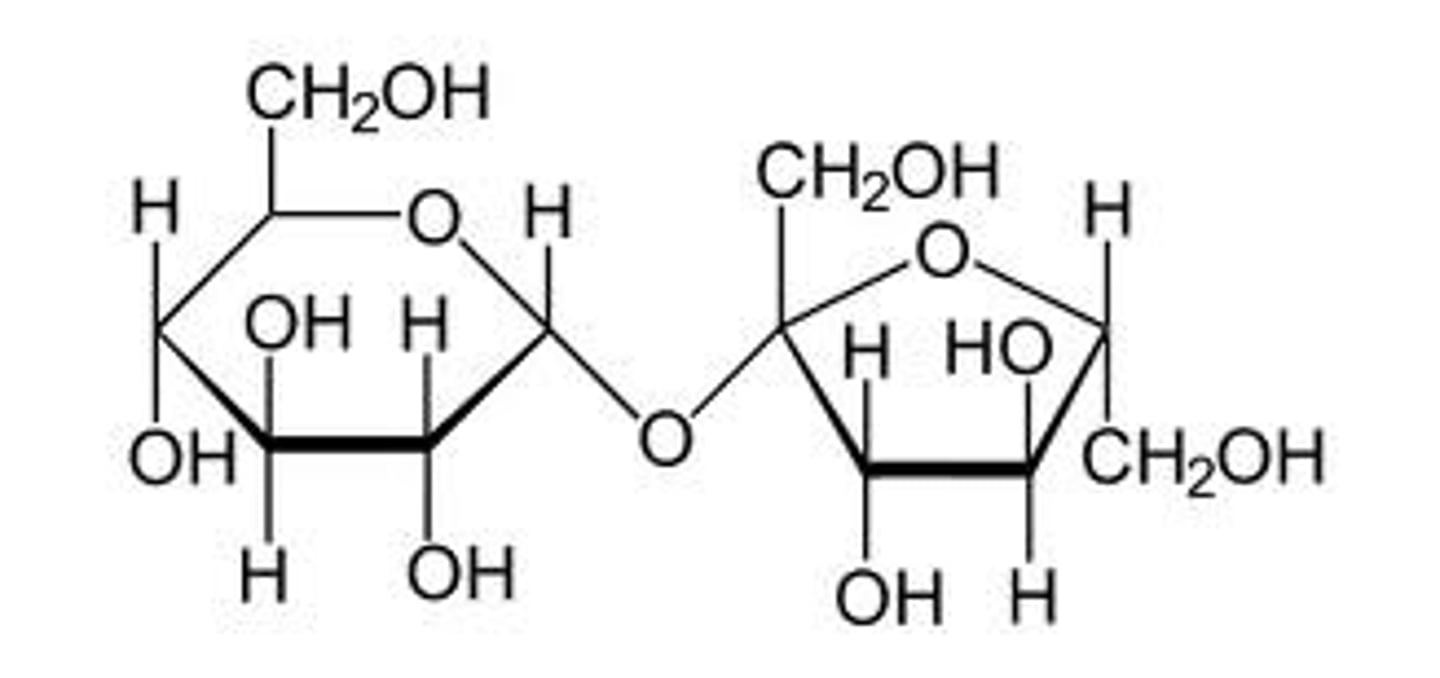
lactose
glucose + galactose
maltose
glucose + glucose
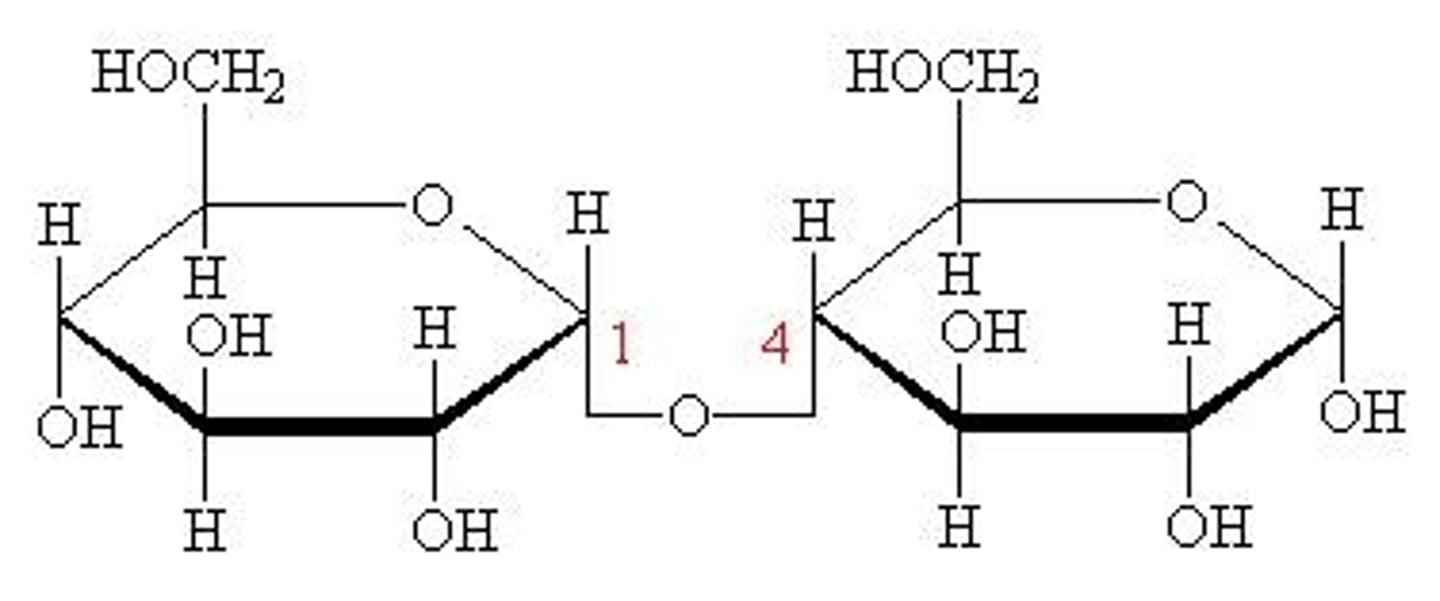
carbohydrate functions
- oxidized to release energy, twice as less kJ than lipids
- provide structure (cellulose, chitin)
- important for recognition and regulation of functions (glycoproteins)
stereiosomer number
2^n (n = # of chiral centers)
L vs D sugar configuration
hydroxyl group on reference carbon is on the left (L) or right (D)
cyclization
- hydroxyl group from reference carbon attacks carbonyl carbon (aldose & ketoses)
- makes a cyclic compound with a hydroxyl that points down (alpha) or up (beta)
naming glycosidic bonds
- anomeric carbon configuration-(# -> #)
- numbering from anomeric, clockwise
e.g. α (1->4) bond
storage polysaccharide characteristics
- branched vs. unbranched
- degree of branching (higher degree = higher storage)
glycoproteins
used in messaging, sugar moieties are always on outside of the cell
catabolism
degradative phase of metabolism, releases energy, usually oxidation
anabolism
building phase of metabolism, requires energy, usually reduction
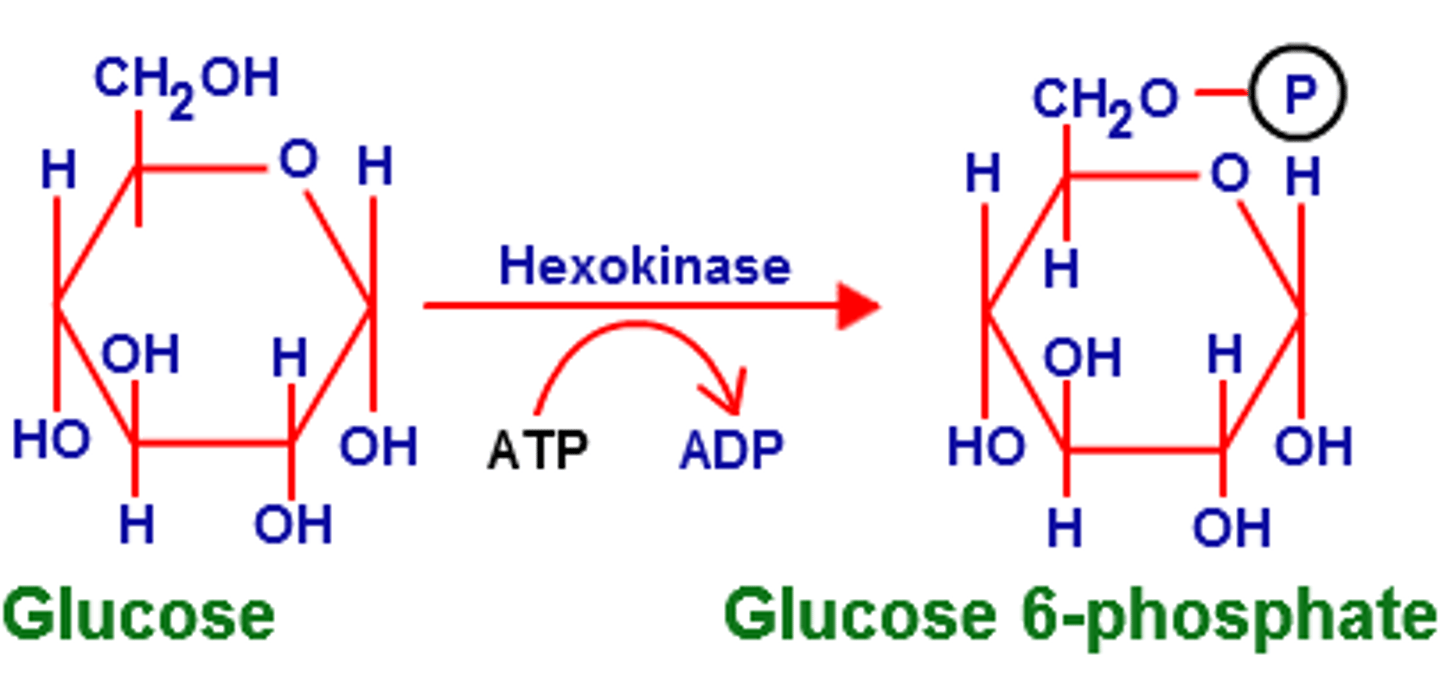
1st step of glycolysis
reactant: glucose
enzyme: hexokinase
product: glucose 6-phosphate
- ATP hydrolyzed
- highly regulated
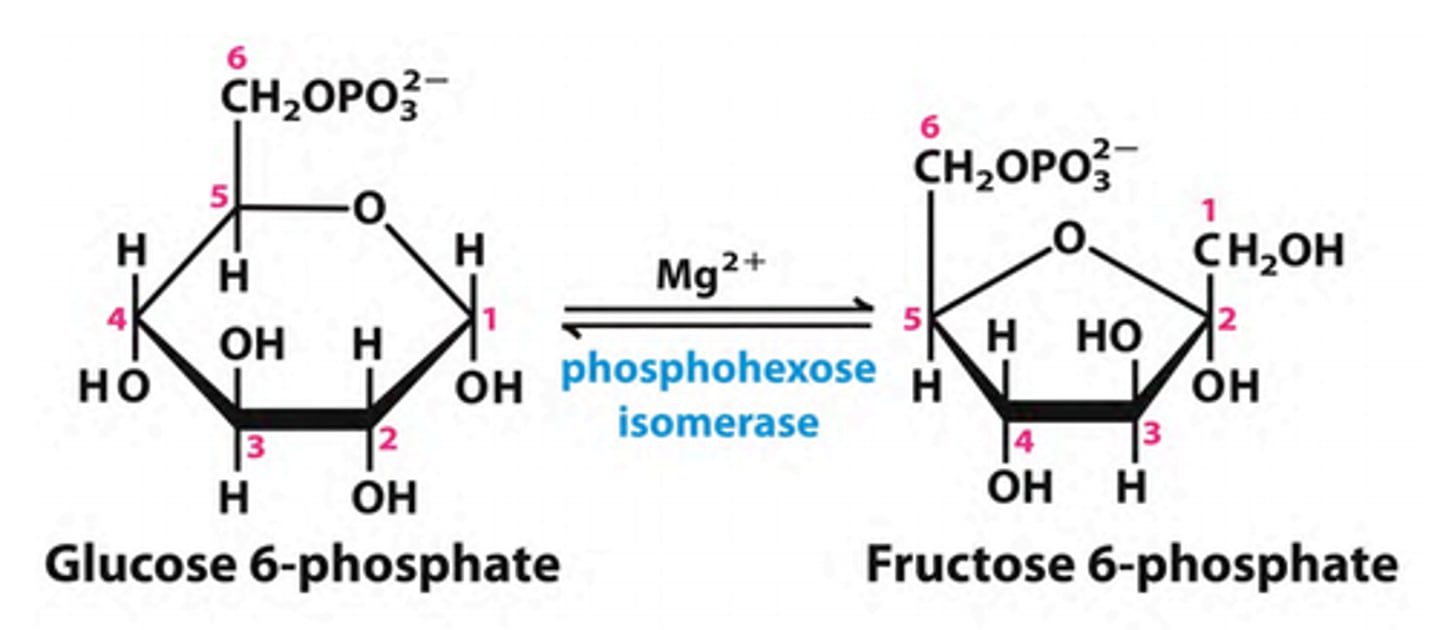
2nd step of glycolysis
reactant: glucose 6-phosphate
enzyme: phosphohexose isomerase
product: fructose 6-phosphate
- requires Mg2+
- makes molecule more symmetrical
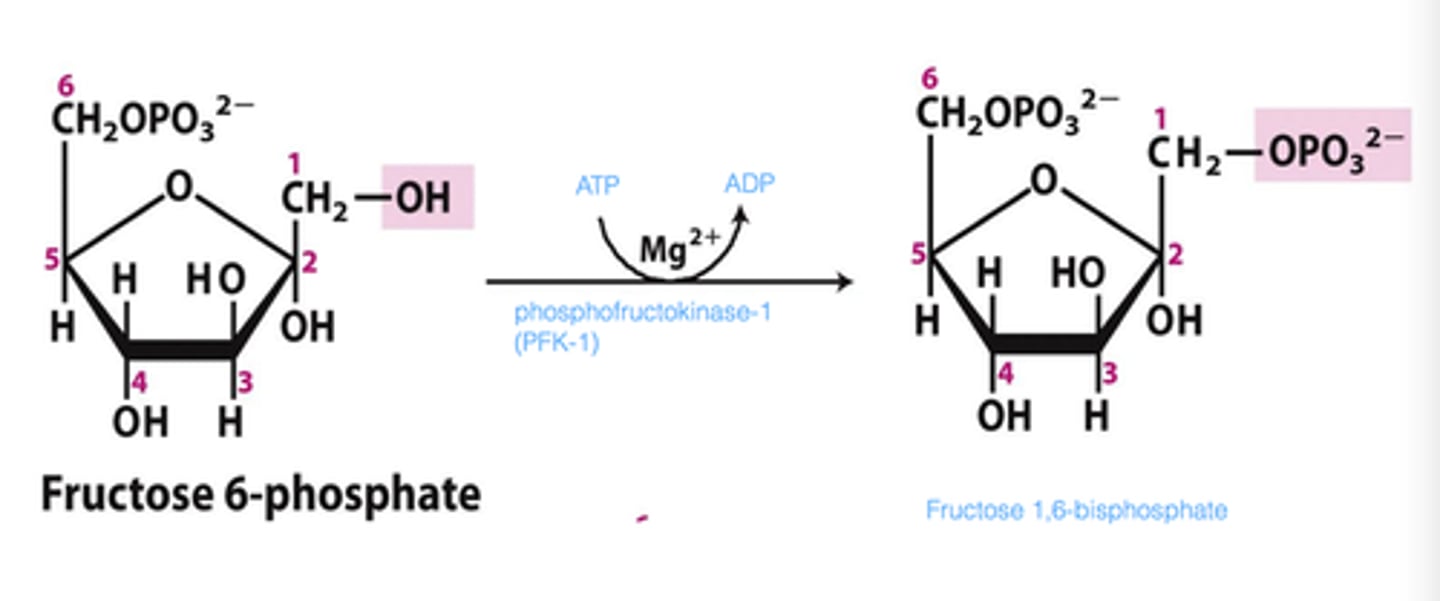
3rd step of glycolysis
reactant: fructose 6-phosphate
enzyme: phosphofructokinase-1 (PFK-1)
product: fructose 1,6-bisphosphate
- ATP hydrolyzed
- rate limiting step, reaction will commit to glycolysis
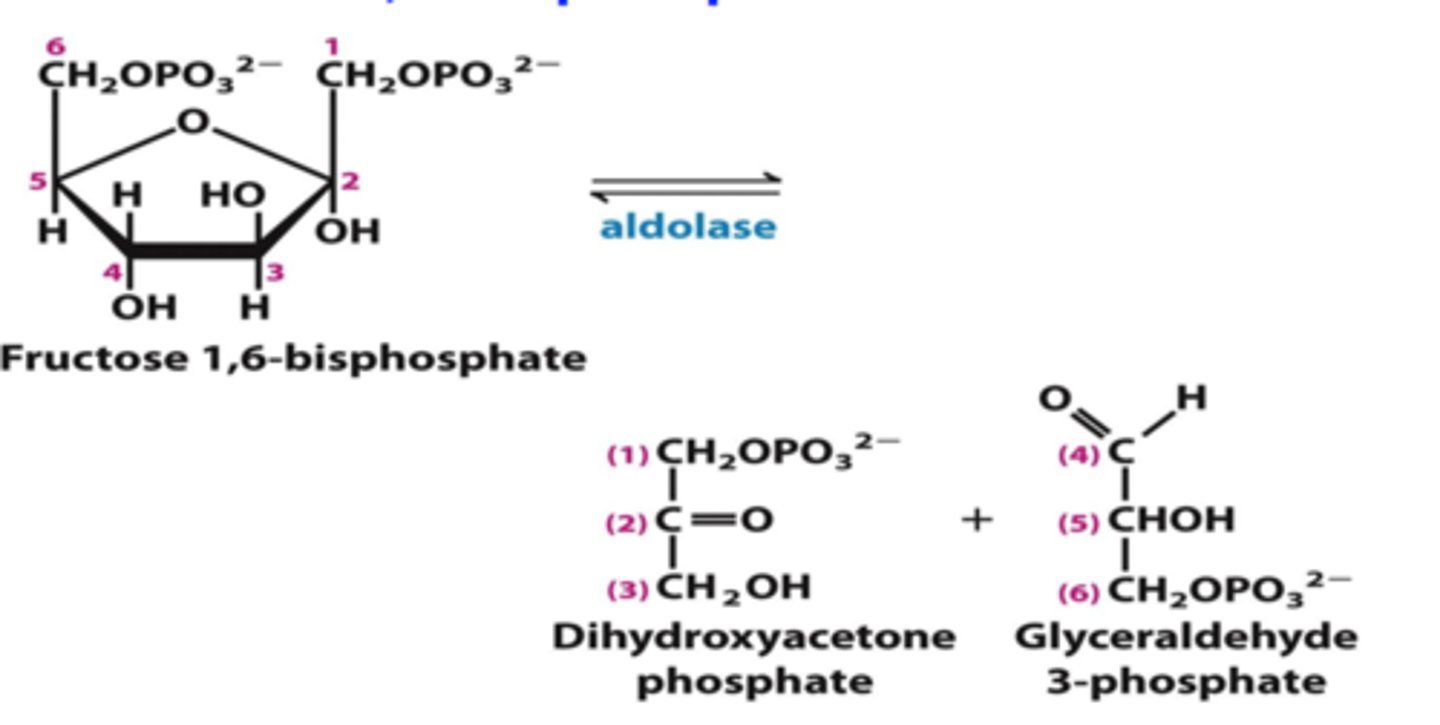
4th step of glycolysis
reactant: fructose 1,6-bisiphoshate
enzyme: aldolase
product: dihydroxyacetone phosphate AND glyceraldehyde 3-phosphate
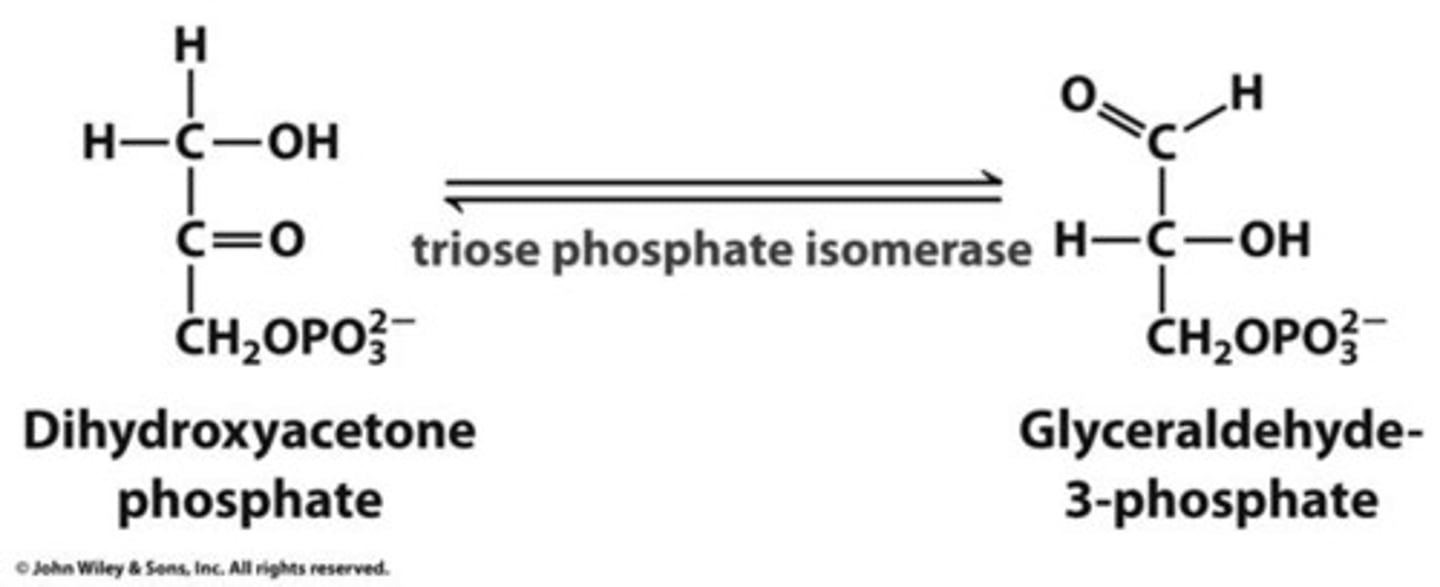
5th step of glycolysis
reactant: dihydroxyacetone phosphate
enzyme: triose phosphate isomerase
product: glyceraldehyde 3-phosphate
- 2 glyceraldehyde 3-phosphate per 1 glucose
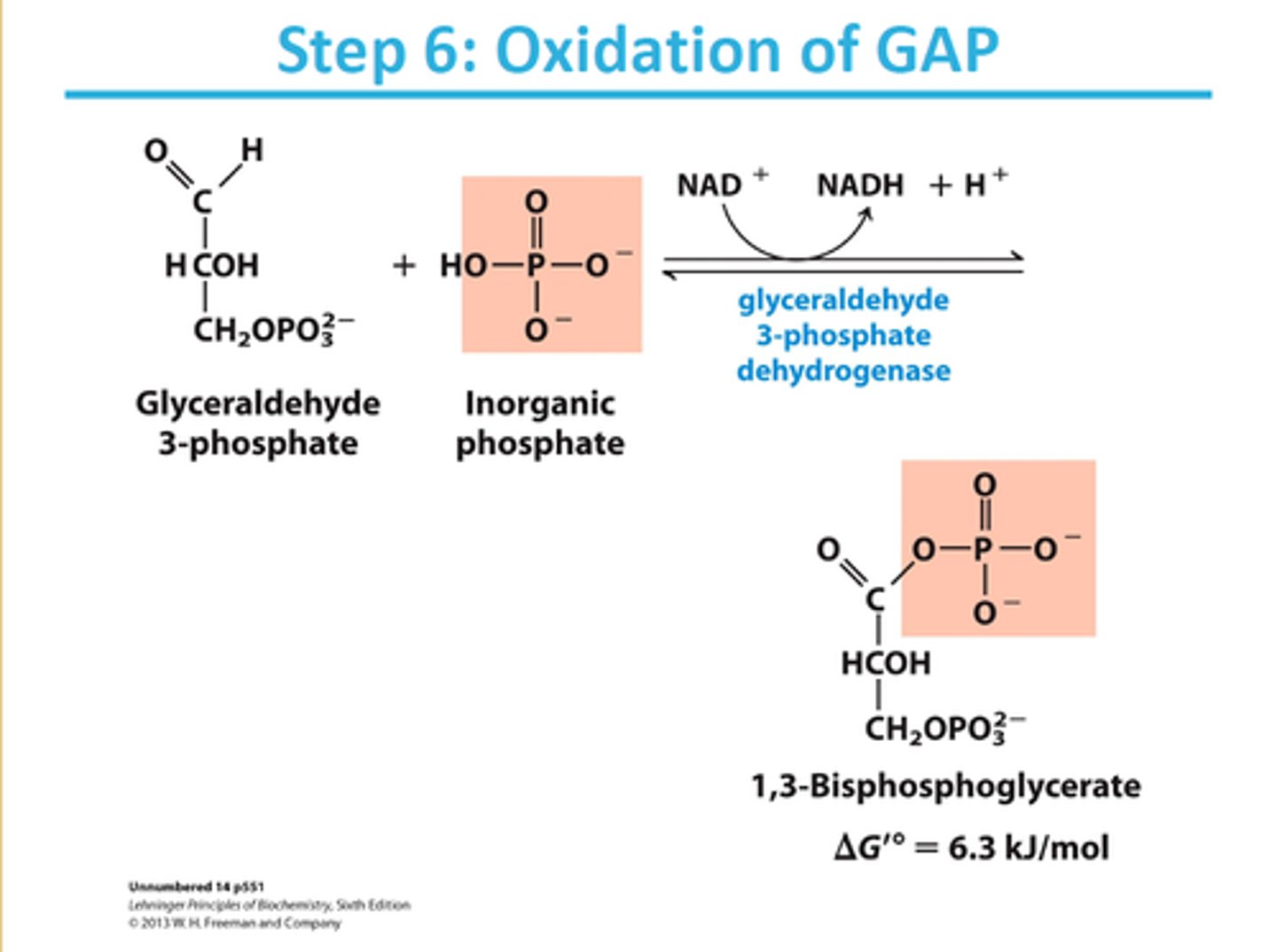
6th step of glycolysis
reactant: glyceraldehyde 3-phosphate
enzyme: glyceraldehyde 3-phosphate dehydrogenase
product: 1,3-bisphosphoglycerate
- NAD+ reduced to NADH + H+
- phosphate group added
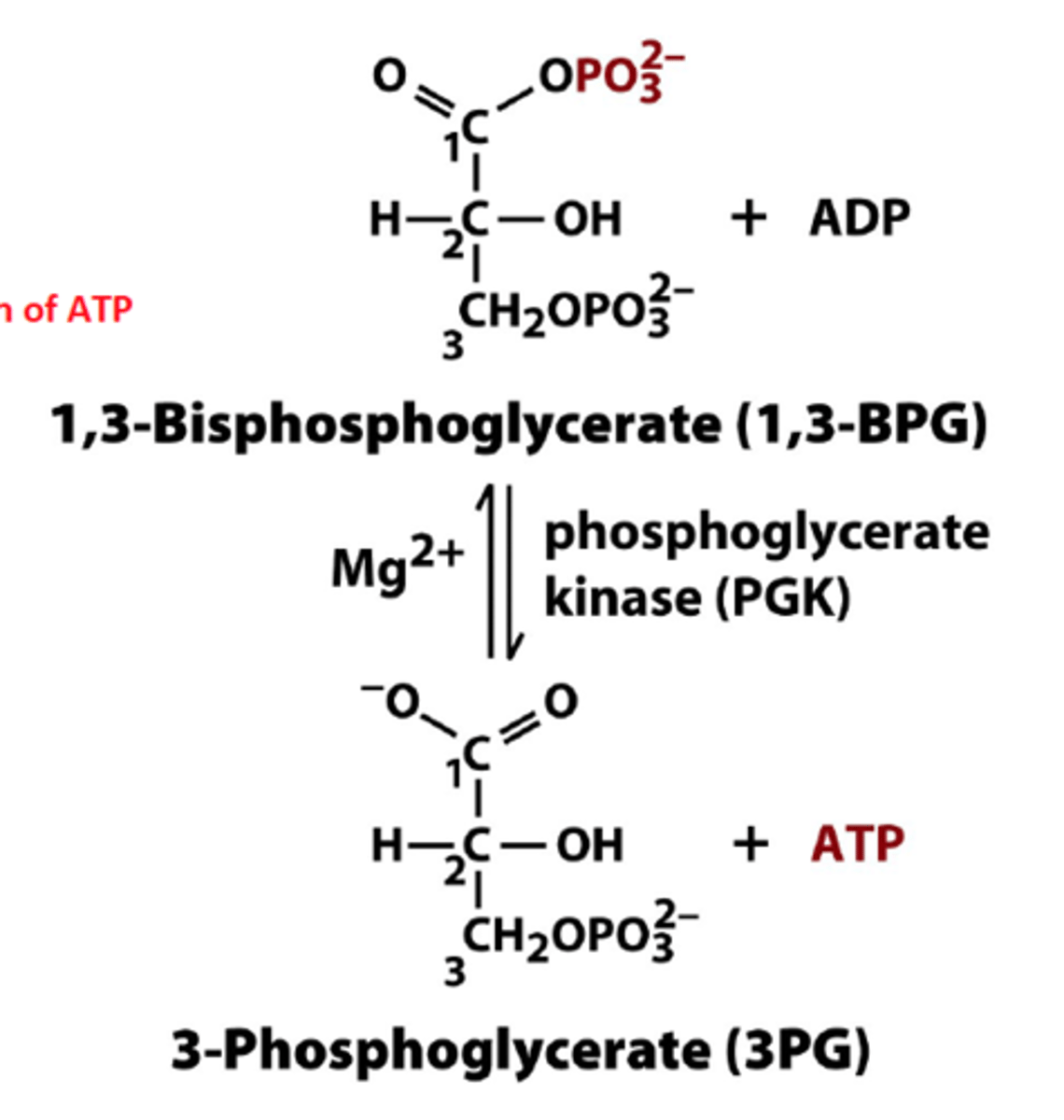
7th step of glycolysis
reactant: 1,3-bisphospoglycerate + ADP
enzyme: phosphoglycerate kinase
3-phosphoglycerate + ATP
- first substrate-level phosphorylation, 2 ATP made
- Mg2+ required
- reaction coupled so it is reversible
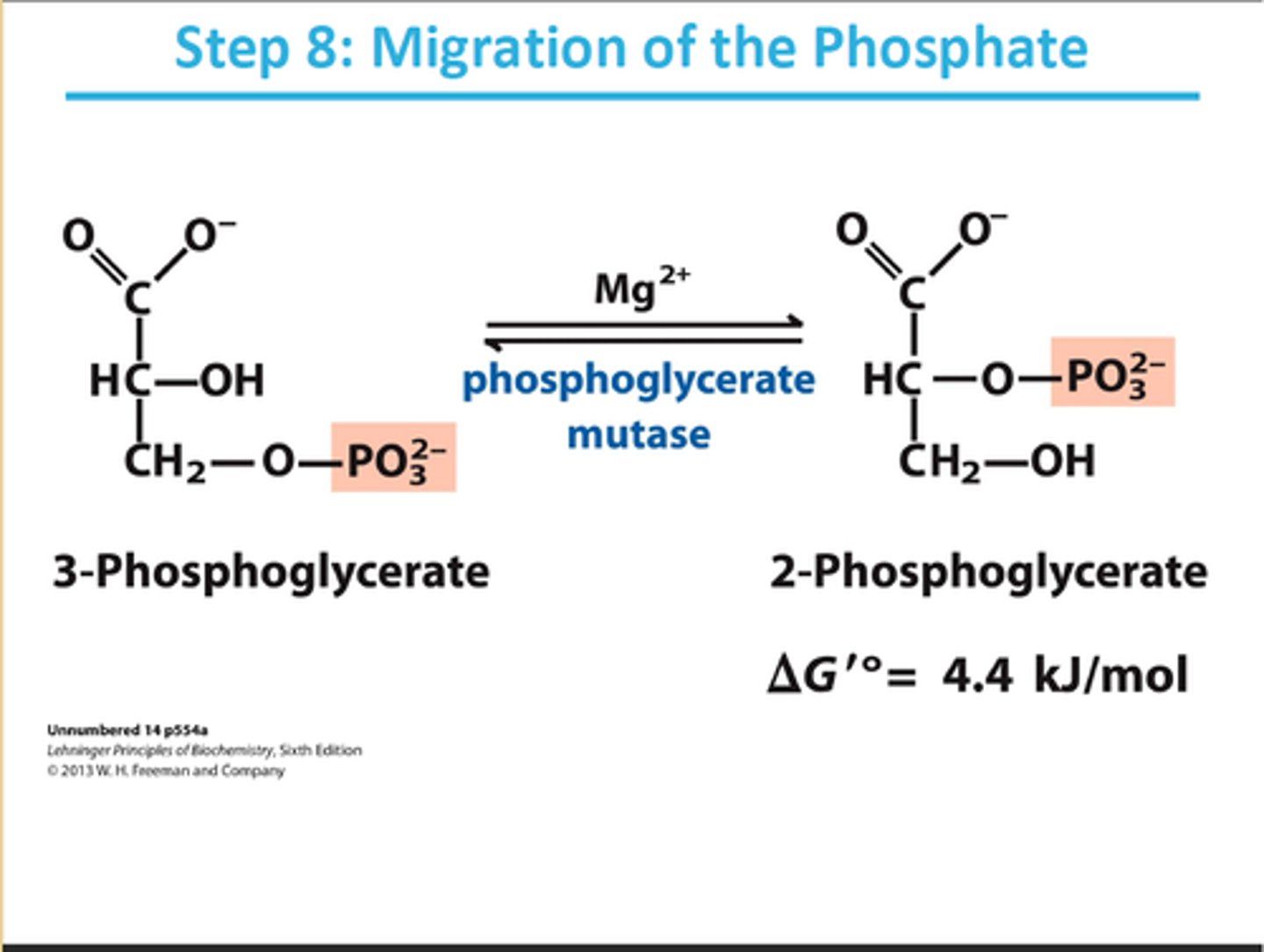
8th step of glycolysis
reactant: 3-phosphoglycerate
enzyme: phosphoglycerate mutase
product: 2-phosphoglycerate
- requires Mg2+
- moves PO32- to 2nd carbon
9th step of glycolysis
reactant: 2-phosphoglycerate
enzyme: enolase
product: phosphoenolpyruvate (PEP)
- dehydration, H2O eliminated
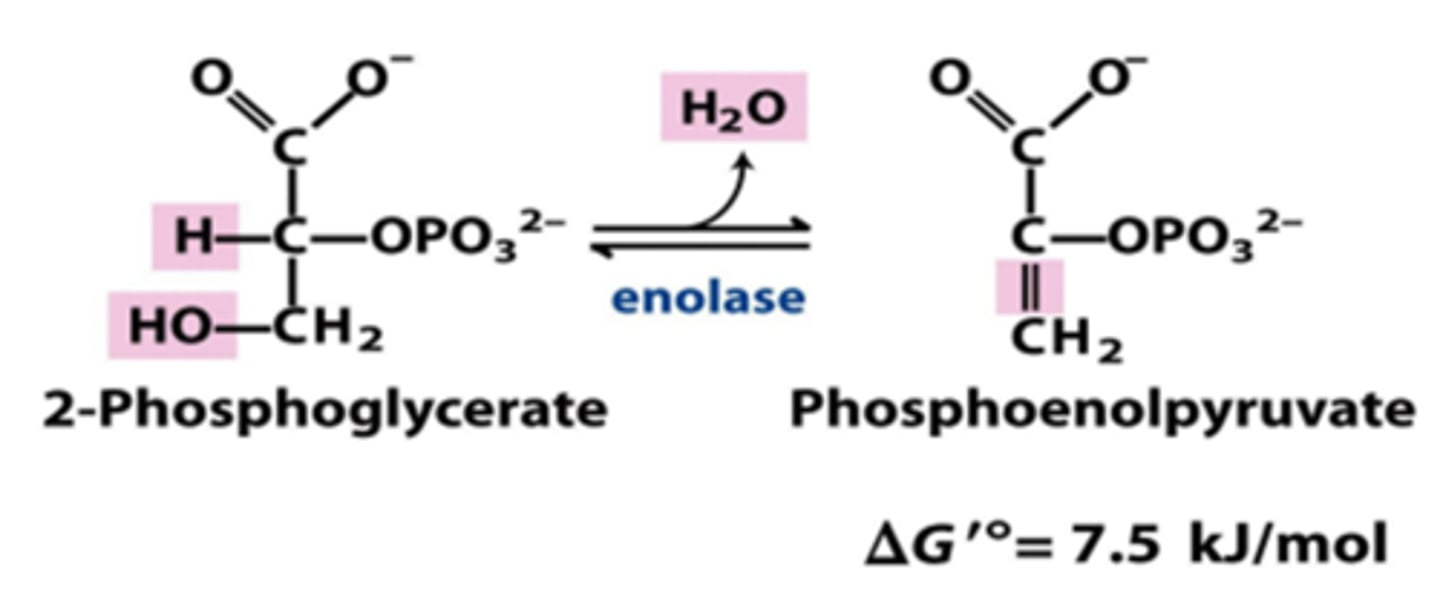
10th step of glycolysis
reactant: phosphoenolpyruvate + ADP
enzyme: pyruvate kinase
product: pyruvate + ATP
- 2nd substrate-level, 2 ATP made phosphorylation
- requires Mg 2+ & K+
- very large negative delta G
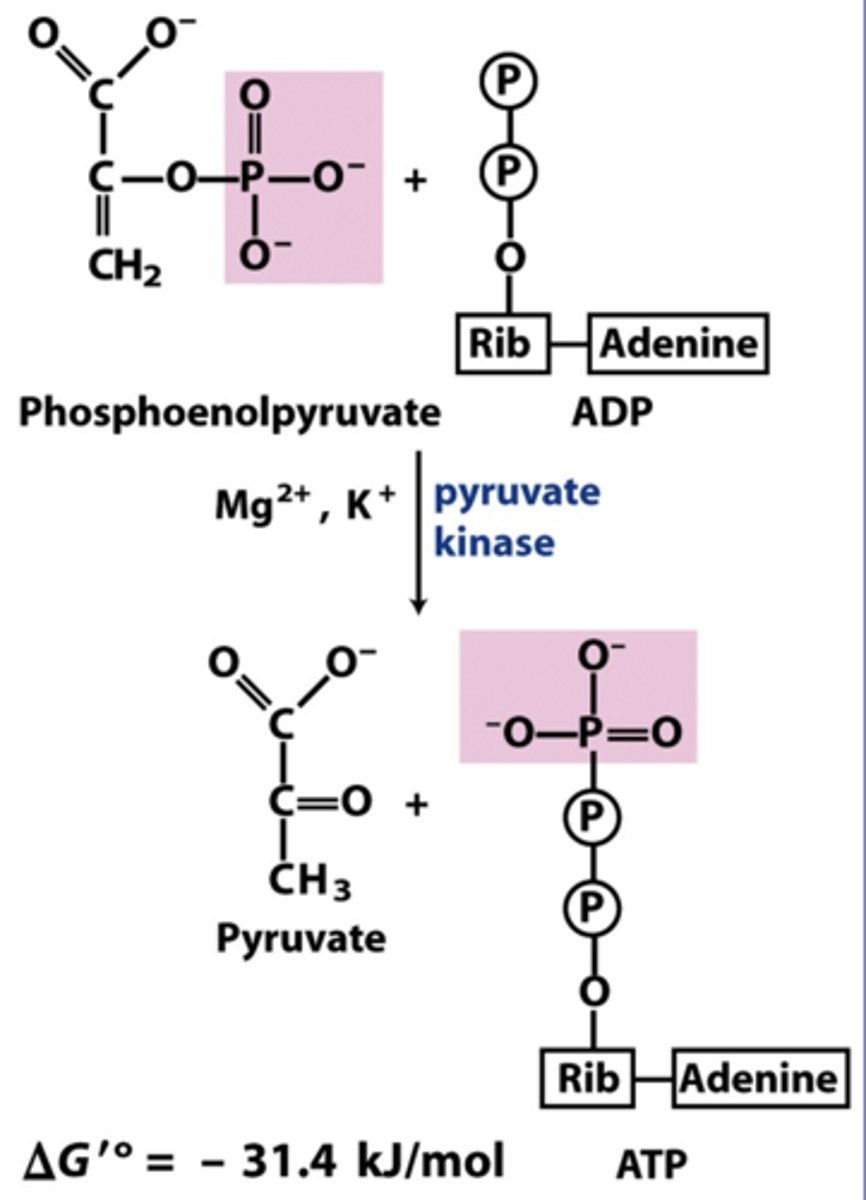
substrate level phosphorylation
enzyme-catalyzed formation of ATP by direct transfer of a phosphate group to ADP from an intermediate substrate
glycolysis overall reaction
2 ATP + 2 NAD+ + 2 ADP + 2 Pi --> 2 pyruvate + 2 ADP + 2 NADH + 2 H+ + 4 ATP + 2H2O
fate of pyruvate
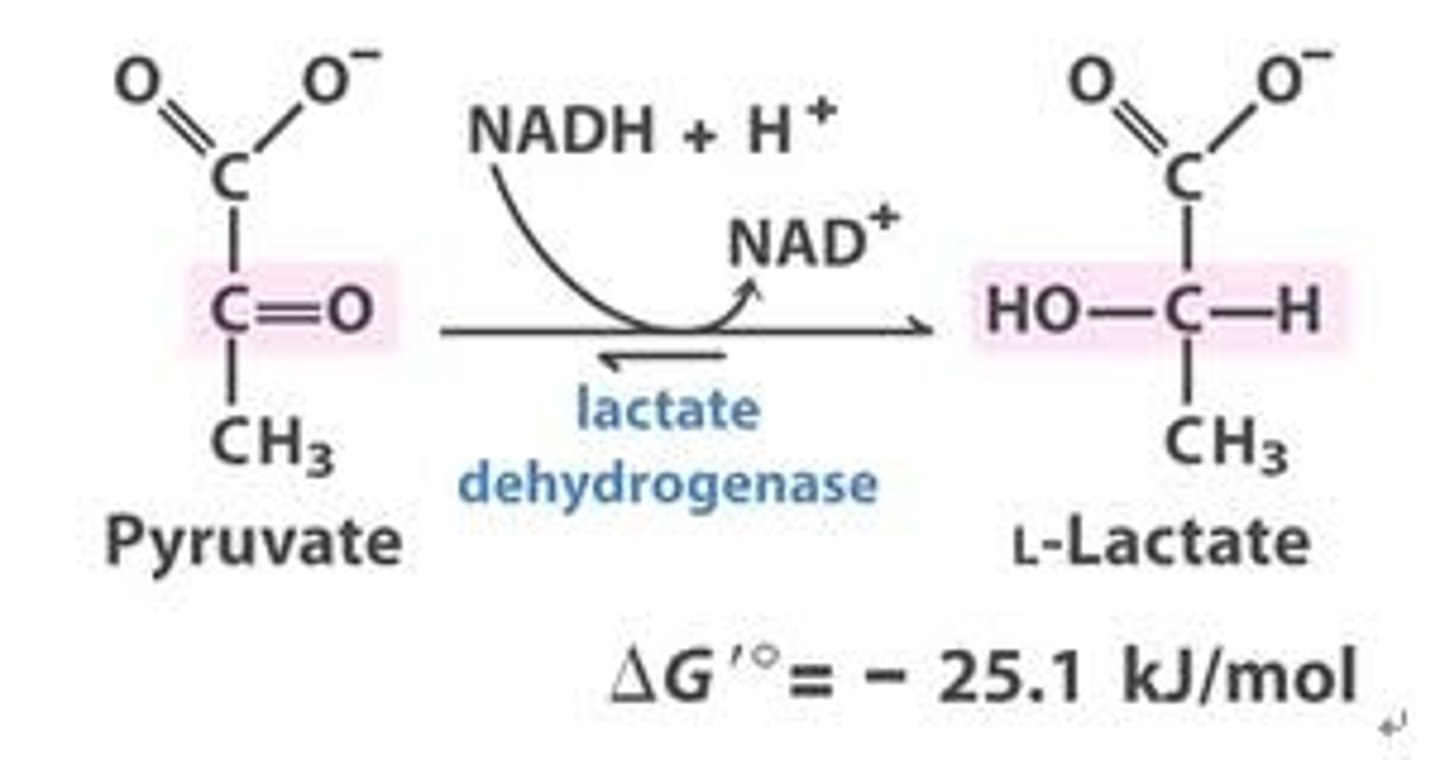
reactant: pyruvate
enzyme: lactate dehydrogenase
product: L-lactate
- regenerates NAD+ which is necessary for redox rxns (glycolysis)
Warburg effect
- cancer cells do much more glycolysis than other cells even when oxygen is present
- important for treatment
glycerol entry point of glucose
- FFA breakdown -> TAGs broken down -> glycerol groups released
- glycerol kinase uses ATP to make glycerol 3-phosphate
- glycerol 3-phosphate dehydrogenase uses NAD+ to make dihydroxyacetone phosphate
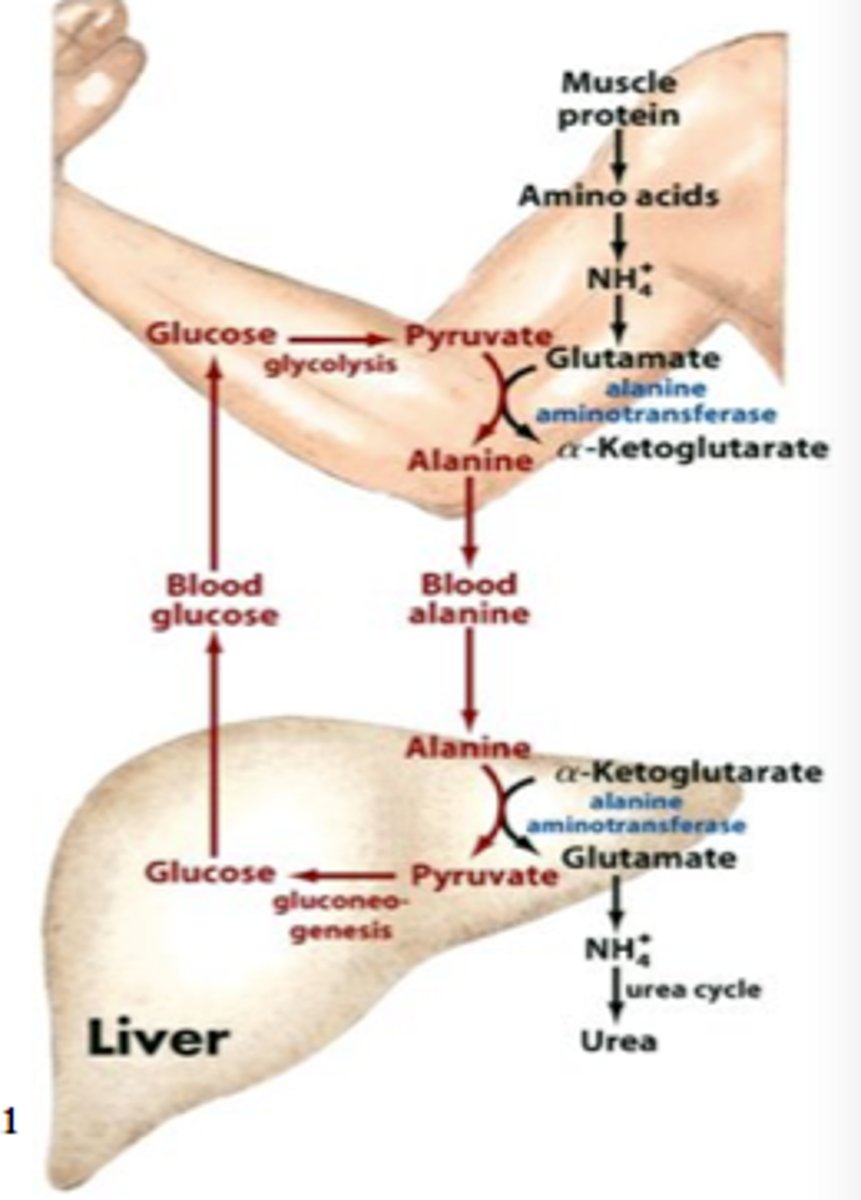
glucose-alanine cycle
- entry point
- (M) pyruvate to alanine through alanine aminotransferase
- (L) alanine to pyruvate to glucose
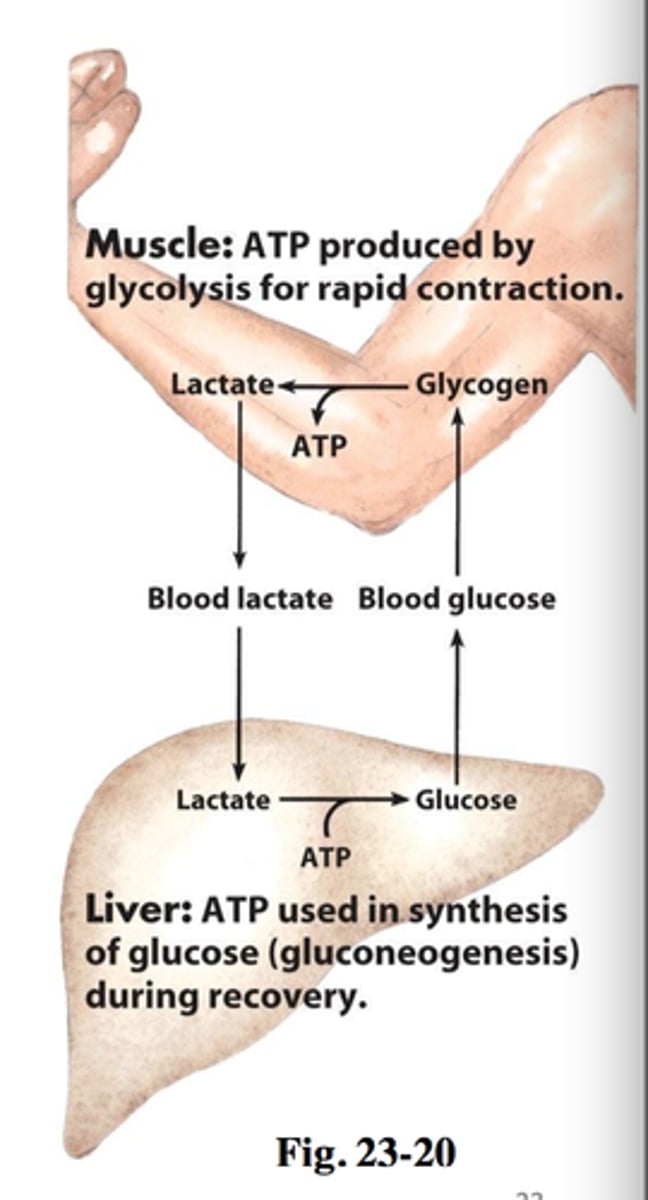
cori cycle
- (M) low glucose = glycogen in muscles are broken down
- (M) glucose from glycogen -> pyruvate -> lactate (ATP released)
- (L) lactate -> pyruvate -> glucose (ATP used)
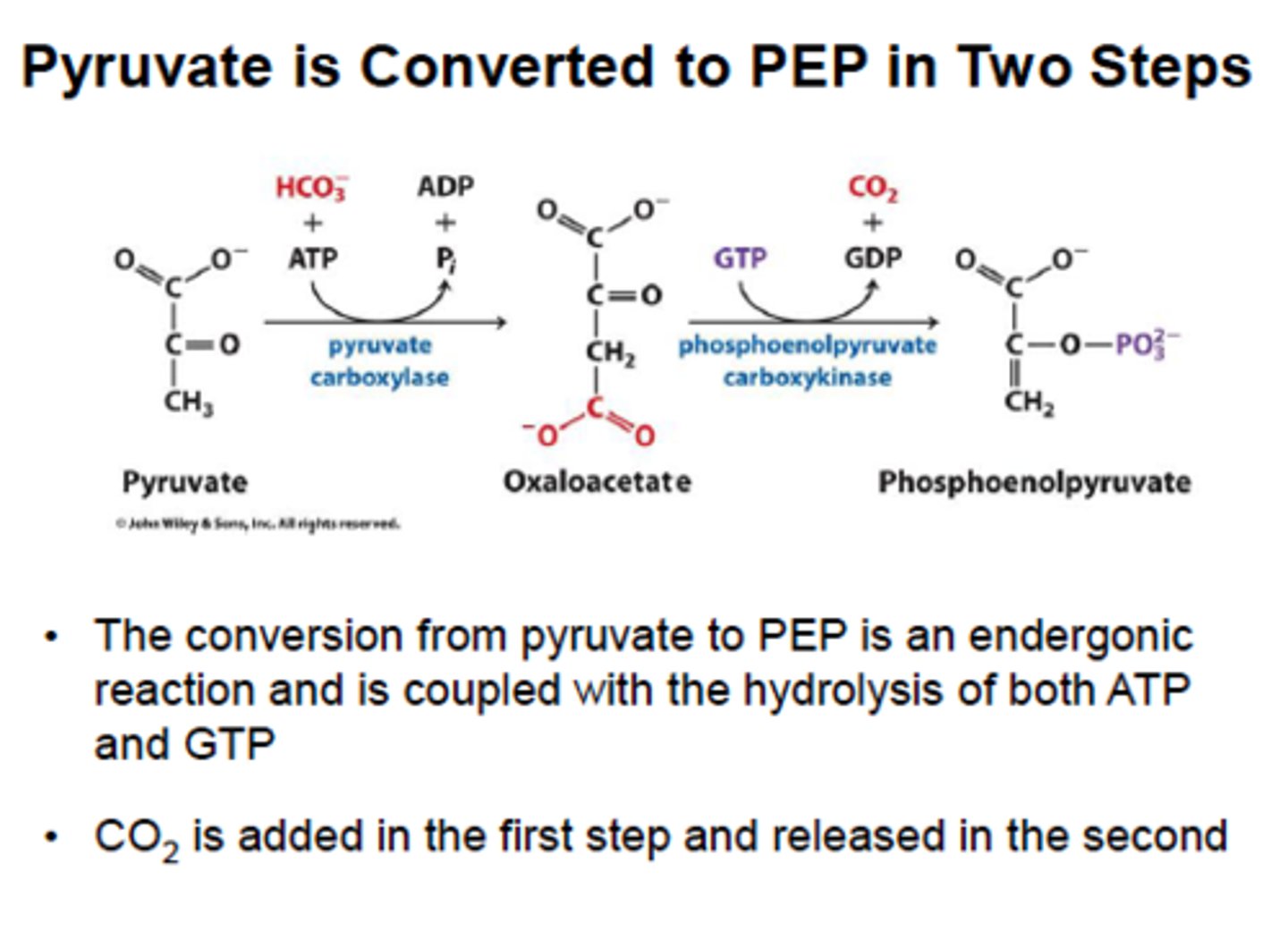
1st (step 1) bypass (gluconeogenesis)
reactant: pyruvate
enzyme: pyruvate carboxylase
product: oxaloacetate
- uses 1 ATP, in mitochondria to prevent futile cycle
- requires biotin to carry CO2 from bicarbonate
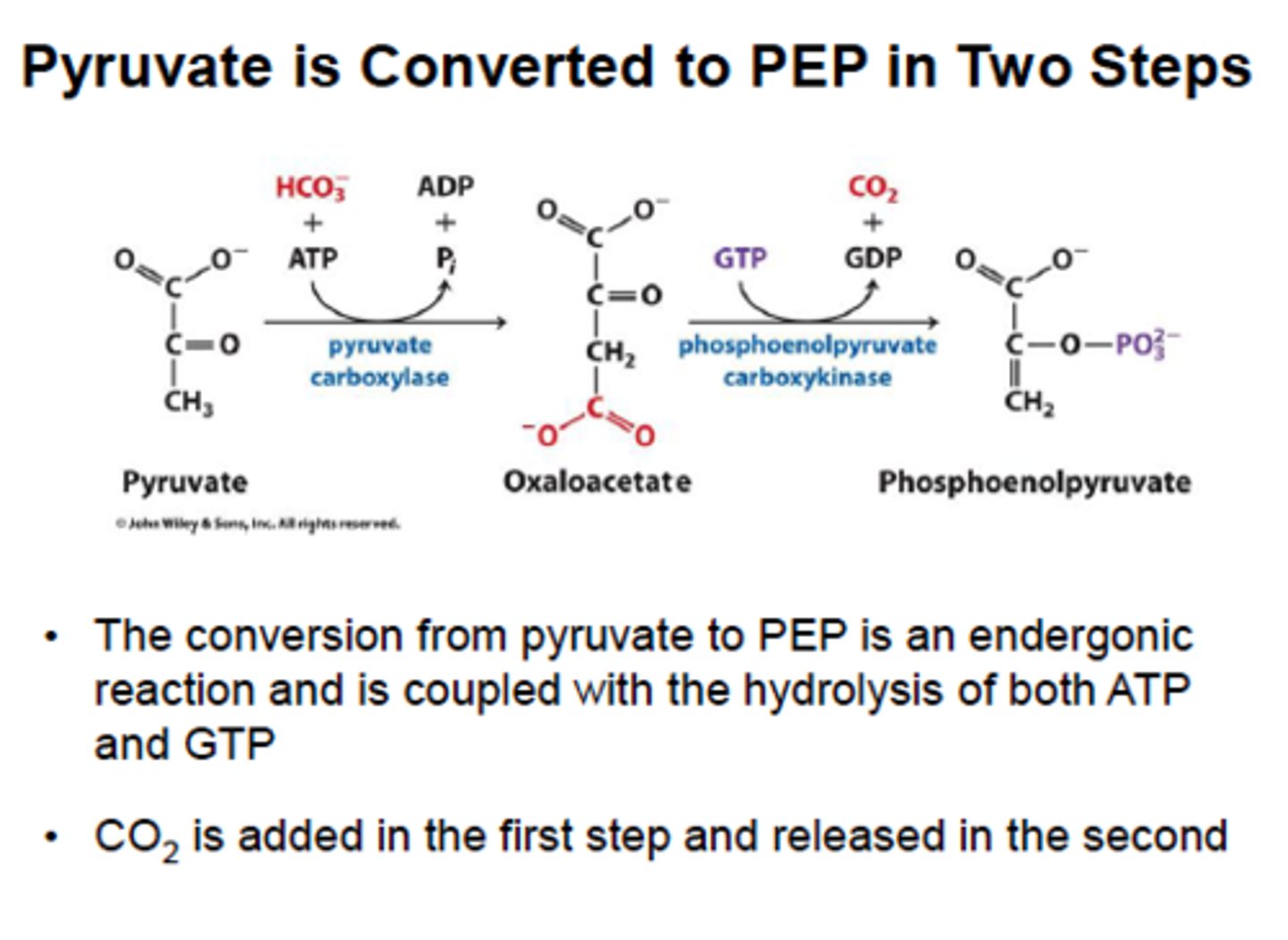
1st (step 2) bypass (gluconeogenesis)
reactant: OAA + GTP
enzyme: PEP carboxykinase
product: PEP + GDP + CO2
- OAA first moved to cytosol as malate then becomes OAA again (malate dehydrogenase, requires NADH/NAD+)
- GTP hydrolyzed, decarboxylation
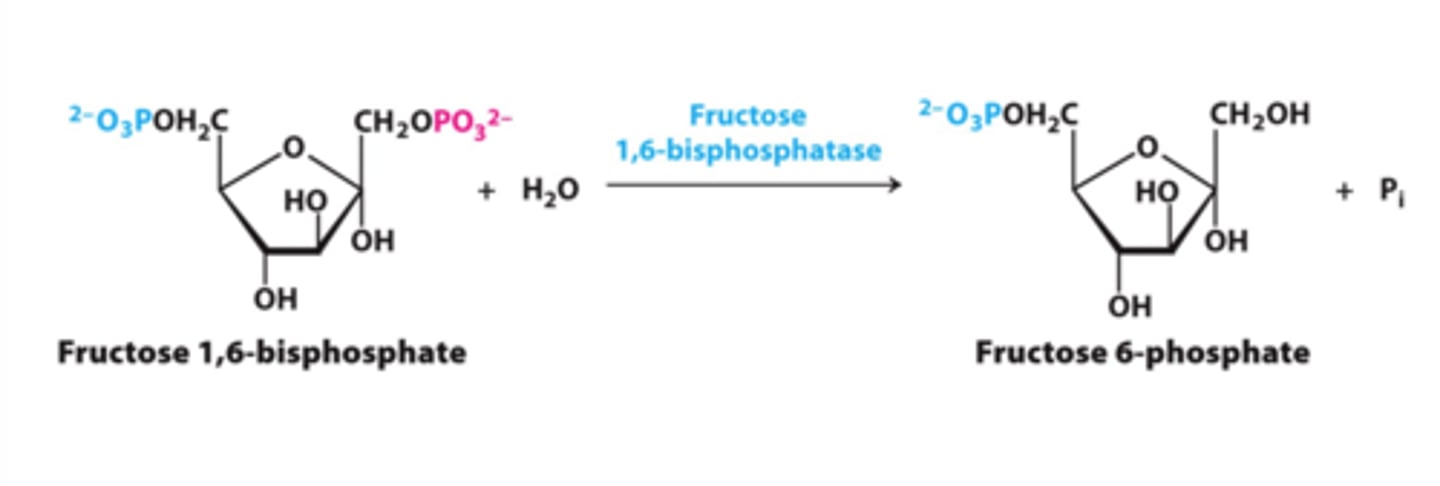
2nd bypass (gluconeogenesis)
reactant: fructose 1,6-bisphosphate + H2O
enzyme: fructose bisphosphatase-1 (FBPase-1)
product: fructose-6-phosphate + Pi
- large negative delta G (irreversible)
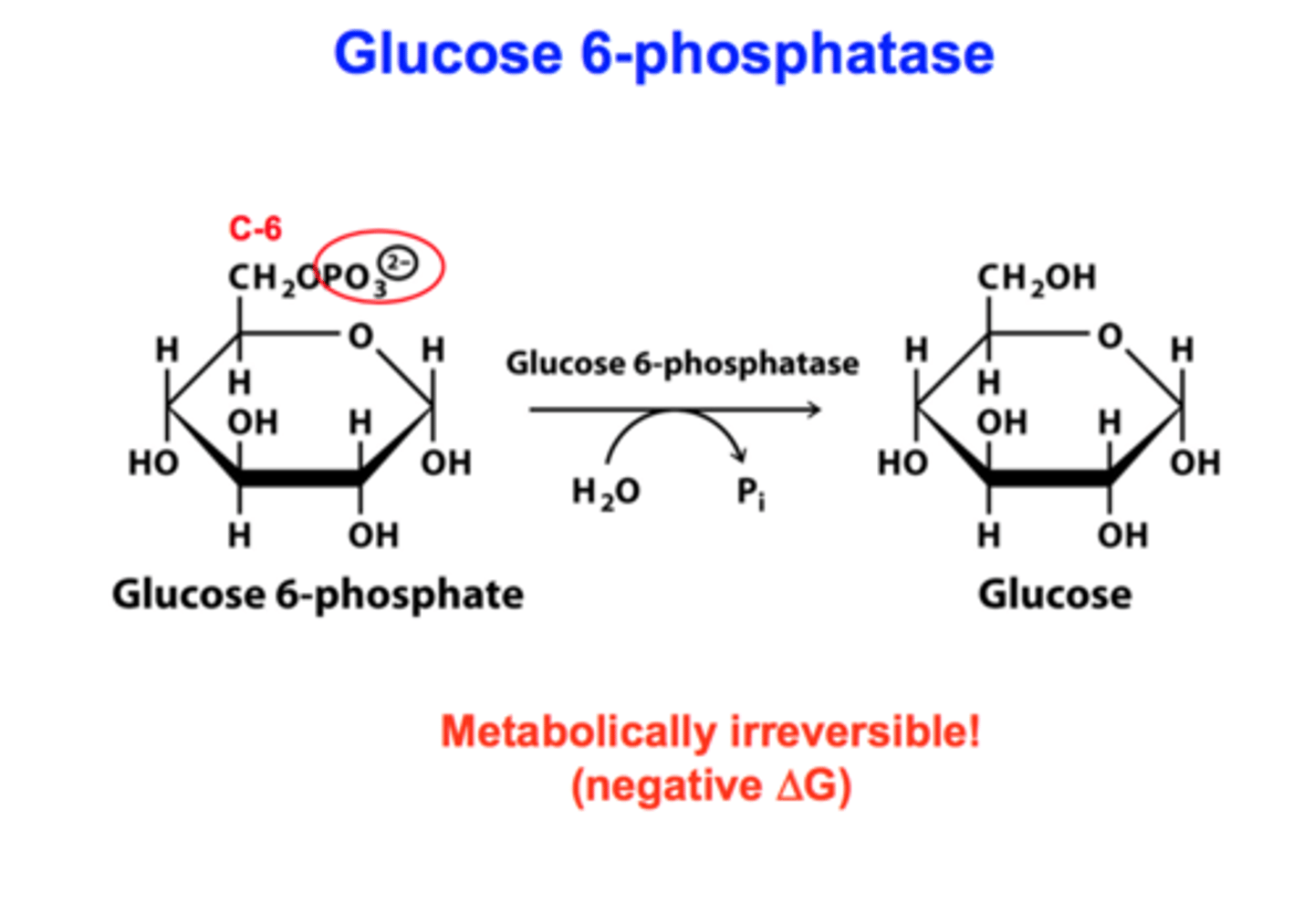
3rd bypass (gluconeogenesis)
reactant: glucose 6-phosphate + H2O
enzyme: glucose 6-phosphatase
product: glucose + Pi
- large negative delta G (irreversible)
- only found in lumen of ER of liver & kidney cells
gluconeogenesis overall reaction
2 pyruvate + 4 ATP + 2 GTP + 2 NADH + 2 H+ + 4 H2O -> glucose + 4 ADP + 2 GDP + 2 NAD+ + 6 Pi
hexokinase regulation
- inihibits: G6P
- enhances: Pi
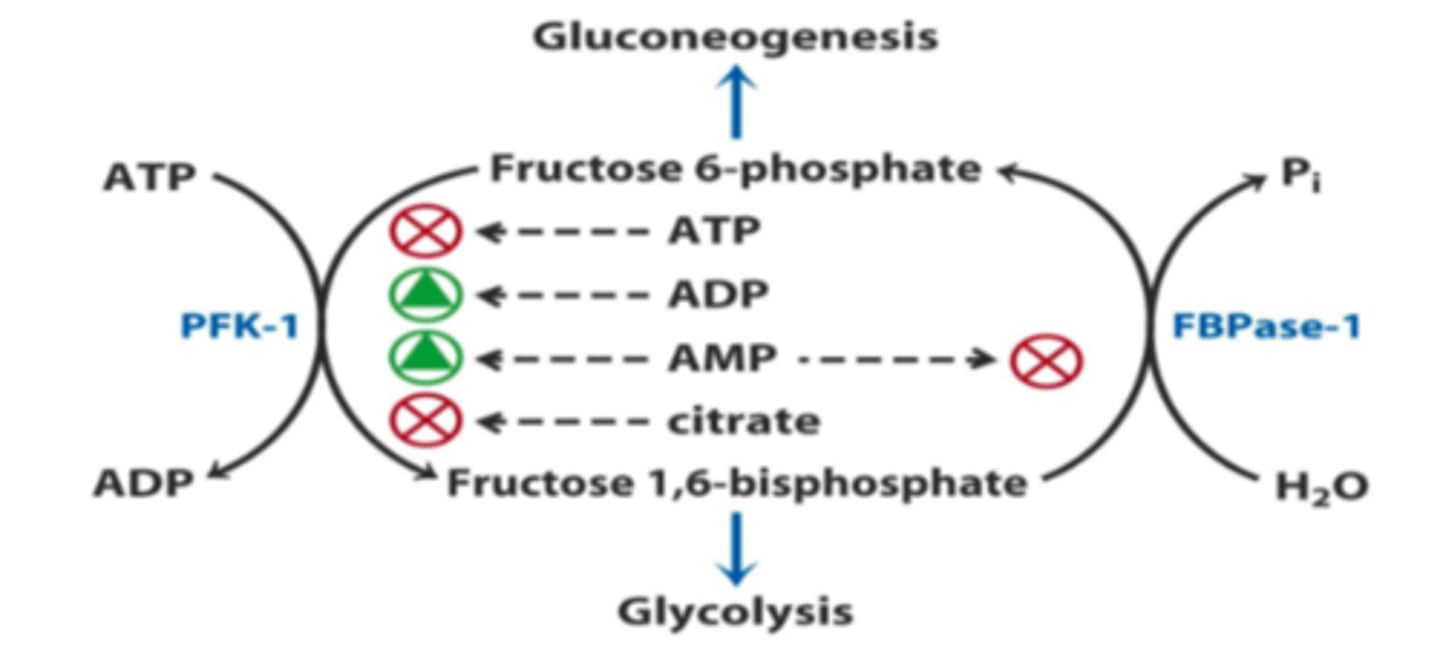
PFK-1 regulation
- inhibits: ATP, citrate
- enhances: AMP, ADP, fructose 2,6-bisphosphate
- regulation of rate-limiting step
- high ATP shuts down all of glycolysis
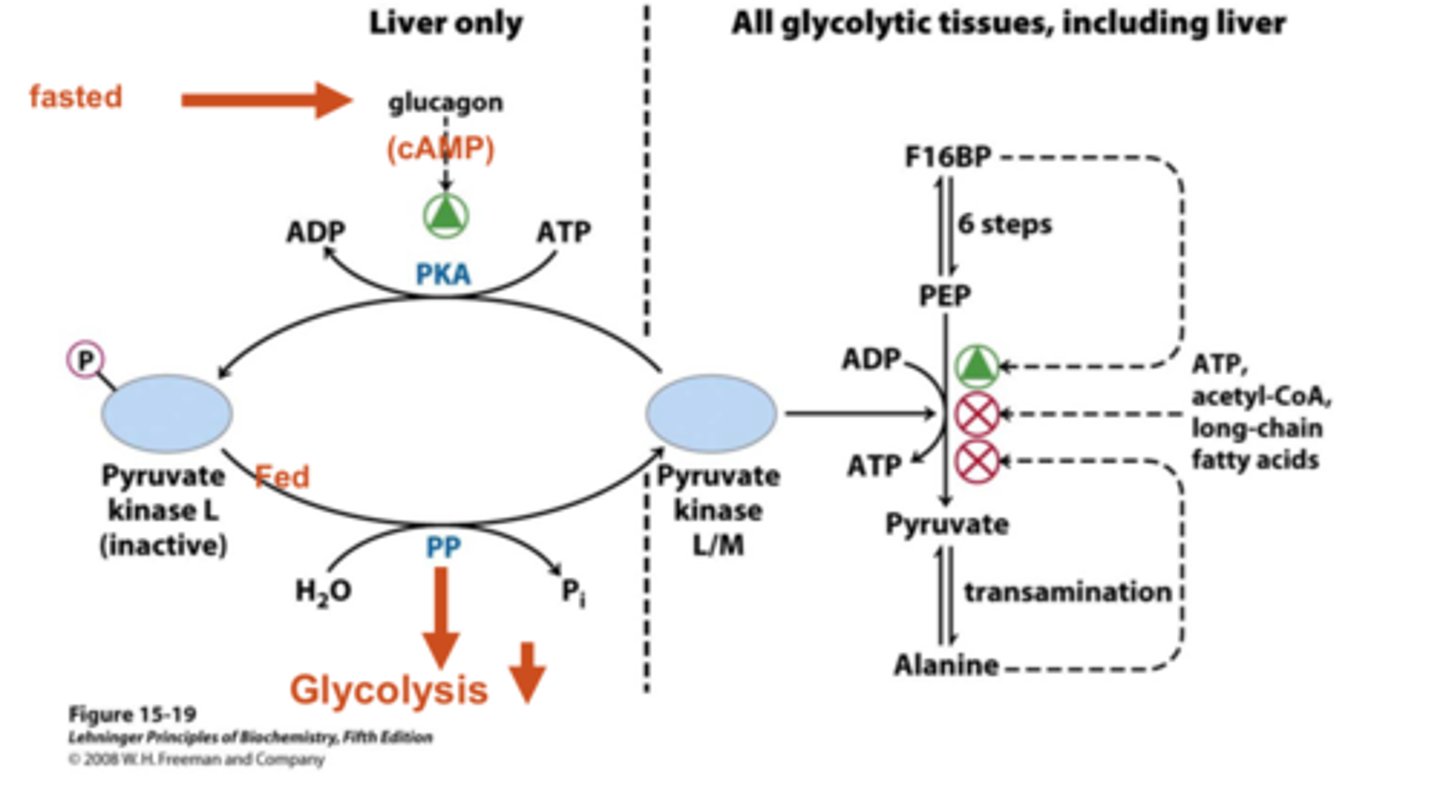
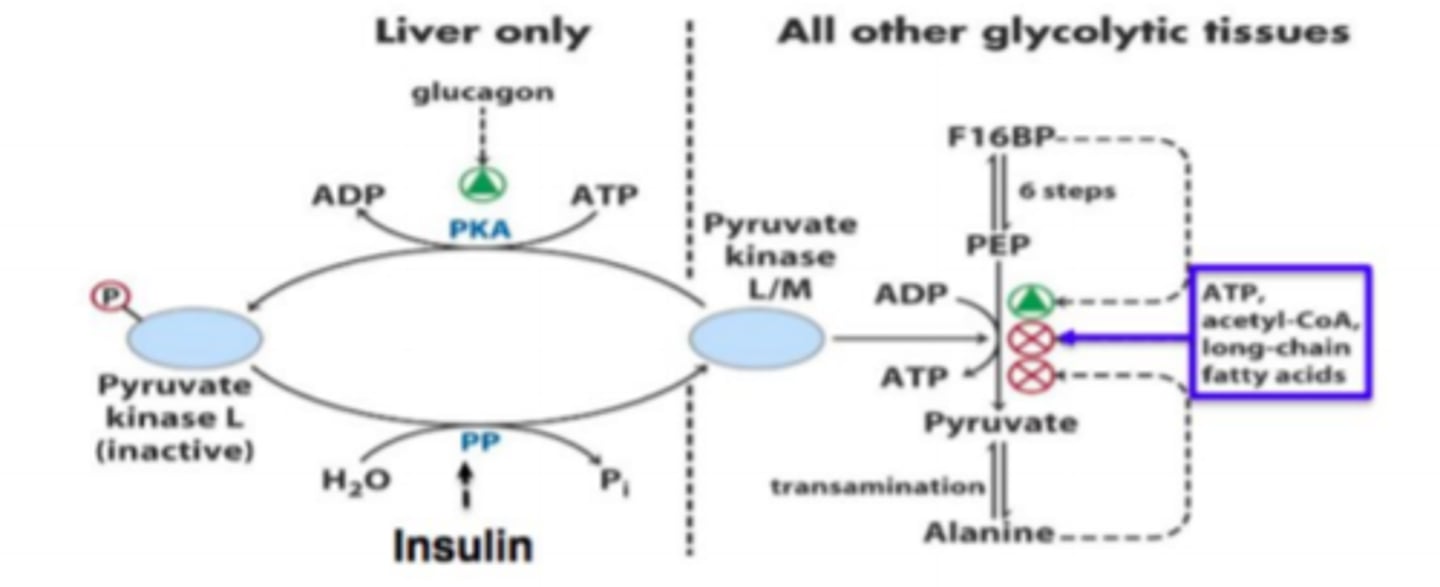
pyruvate kinase regulation
liver only (low blood glucose):
- glucagon enhances PKA = phosphorylates = inactivates pyruvate kinase
all tissues (high blood glucose):
- insulin dephosphorylates pyruvate kinase = activates
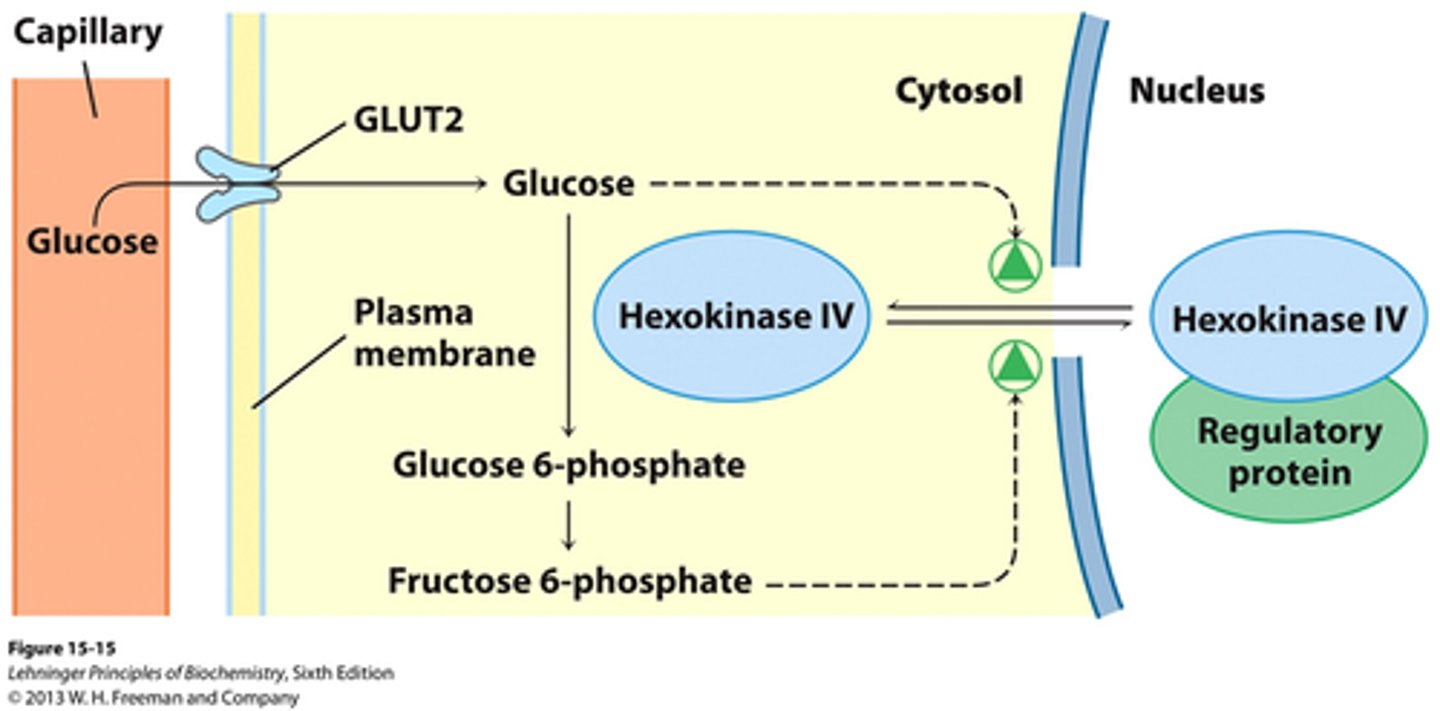
glucokinase (aka hexokinase IV)
- only in liver, has a low affinity for glucose but high capacity to decrease blood glucose concentration.
- not inhibited by pdt
glucokinase/hexokinase IV regulation
- inhibit: fructose 6-phosphate binds HK4 to regulatory protein in nucleus
- enhance: glucose stimulates release into cytosol
F6P accumulation = no glycolysis = inhibits HK4
PFK-1 regulation graph
- largely enhances: F2,6BP
- enhances: AMP
- inhibits: ATP & F2,6BP
- largely inhibits: ATP or citrate
enhances: shifts left, less concentration needed to activate
inhibits: shifts right, more concentration needed to activate
FBPase-1 regulation
- inhibit: AMP
- active during low energy to make glucose (2nd bypass)
fructose 2,6-bisphosphate
mediates glycolysis (activates PFK-1) and gluconeogenesis (FBPase). - more F2,6BP = during higher blood glucose.
- less F2,6BP = during lower blood glucose
bifunctional enzyme
- contains PFK-2 and FBPase-2
- controls flux through glycolysis or gluconeogenesis
PFK-2/FBPase-2 function
- PFK-2 phosphorylates and increases F2,6BP
- FBPase-2 dephosphorylates and decreases F2,6BP
regulation of bifunctional enzyme
- high blood glucose = insulin = PP1 = dephosphorylated = PFK-2 active = more F2,6BP = glycolysis
- low blood glucose = glucagon = PKA = phosphorylated = FBPase-2 active = less F2,6BP = gluconeogenesis
pyruvate kinase
- PEP -> pyruvate
- dephosphorylated form active at high blood glucose
- inhibit: ATP, acetyl-CoA, FAs
- enhance: F1,6BP (precursor to pyruvate)
β-granules
- glycogenin dimer: prrimer
- tiers of glucose residues are α(1->4) linkage, with α(1->6)-linked branches
- tiers: glycogenin -> primer -> second tier -> etc.
- branches provide non-reducing ends
glycogen structure
linear: α(1->4) glycosidic bonds
branch points: α(1->6 glycosidic bonds).
- highly branched to reduce osmotic pressure by having 1 large molecule
glycogenin
core protein found at the center of glycogen, branches build from this protein
glycogenolysis
the breakdown of glycogen into glucose through glycogen phosphorylase and debranching enzyme
glycogen phosphorylase
- uses Pi to cleave glucose from chain on nonreducing end
- phosphorylates glucose (final pdt glucose 1-phosphate)
- cuts until 4 residues away from branch point
debranching enzyme
- transfers 3 of the 4 remaining glucose on branch to the main chain
- glucosidase activity cuts branching glucose
phosphoglucomutase
(M) reversibly catalyzes glucose 1-phosphate conversion to glucose 6-phosphate
- saves 1 ATP from first step of glycolysis
fates of glucose 6-phosphate
- (M) G6P enters glycolysis
- (L) glucose 6-phosophatase (from gluconeogenesis) converts G6P to glucose in the ER to increase blood glucose
- prevents futile cycle
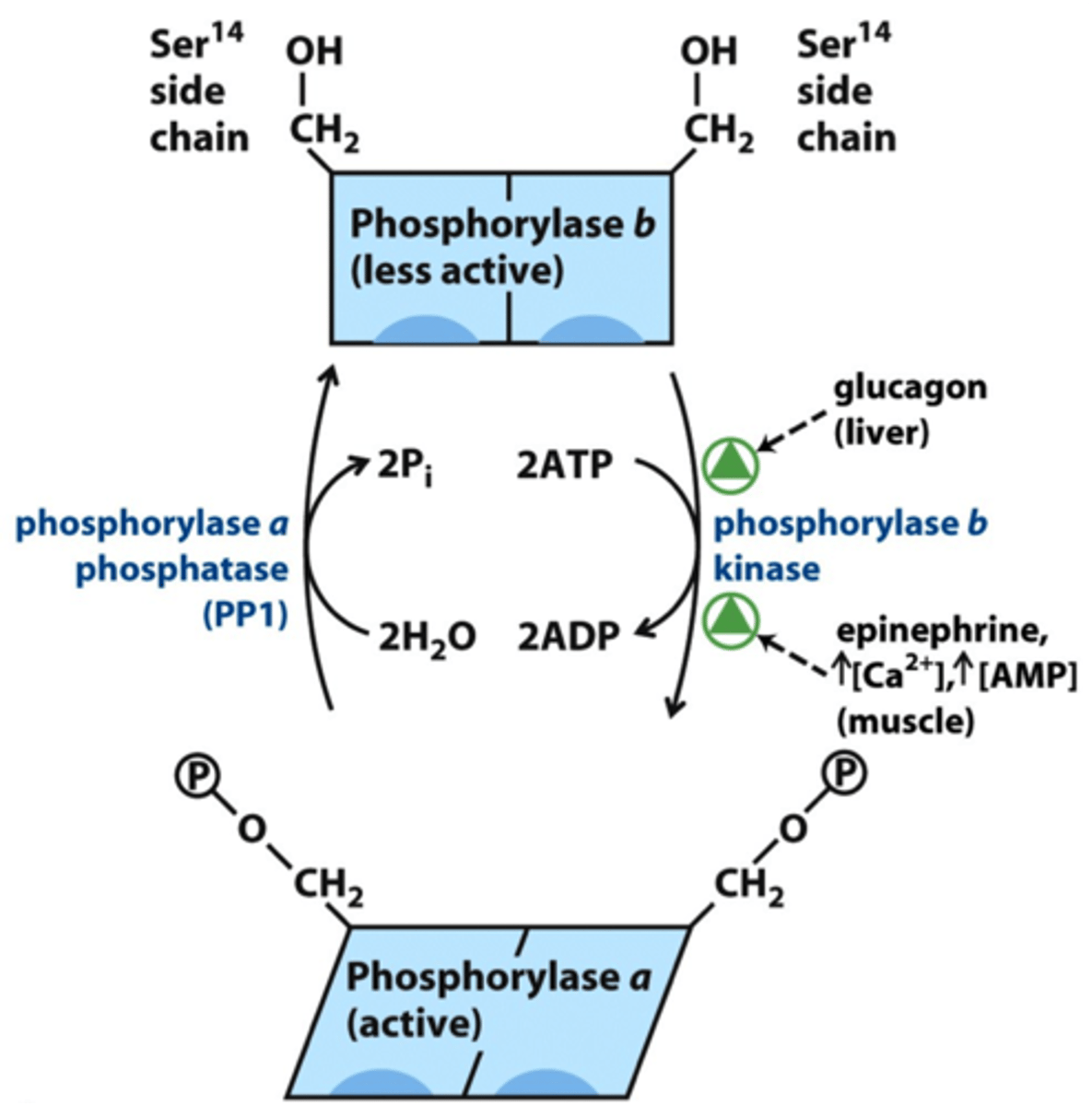
glycogen phosphorylase a
- active, phosphorylated
- breaks down glycogen into glucose 1-phosphate
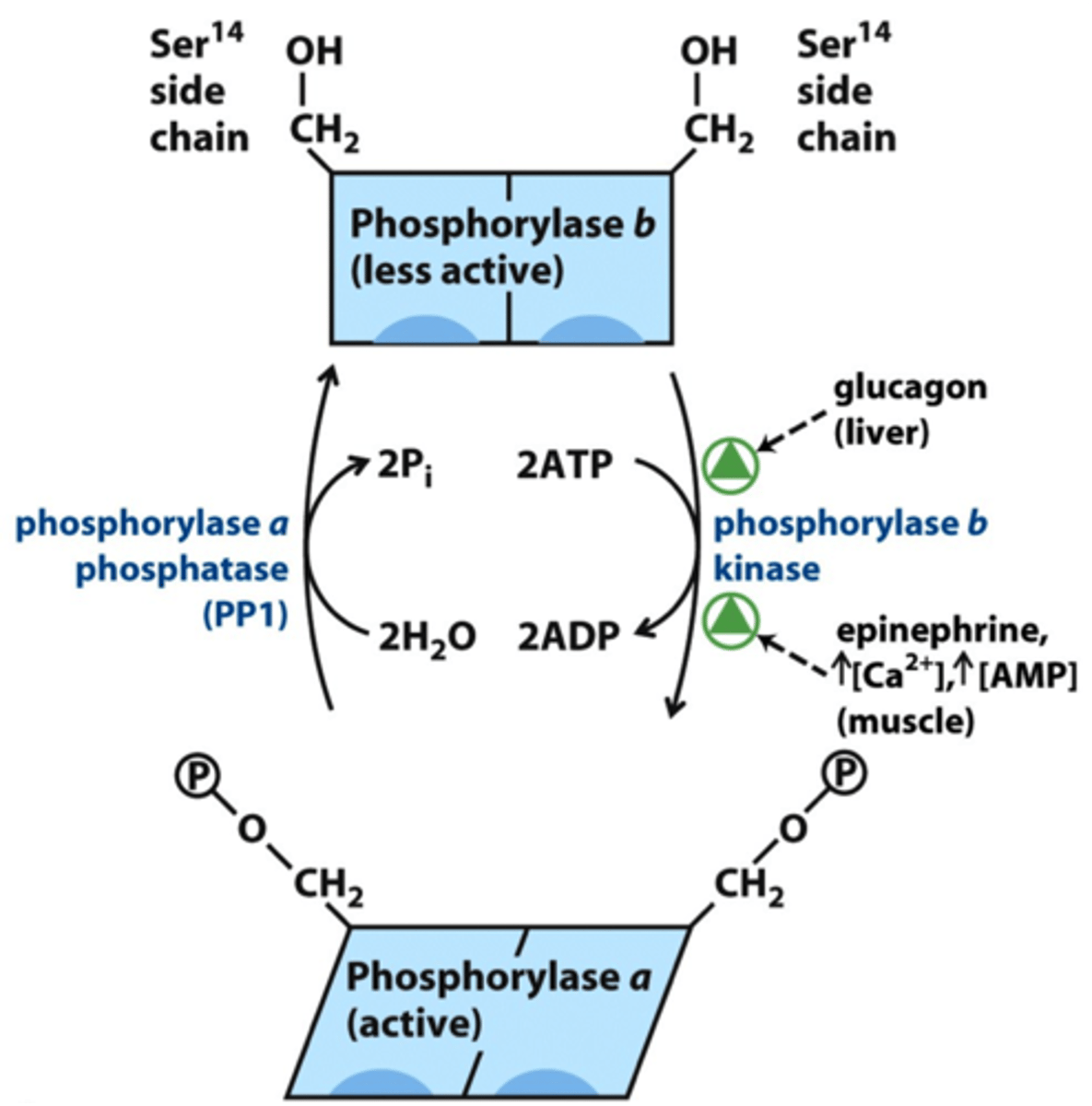
glycogen phosphorylase b
- inactive, dephosphorylated
- phosphorylase b kinase changes phosphorylase b to phosphorylase a
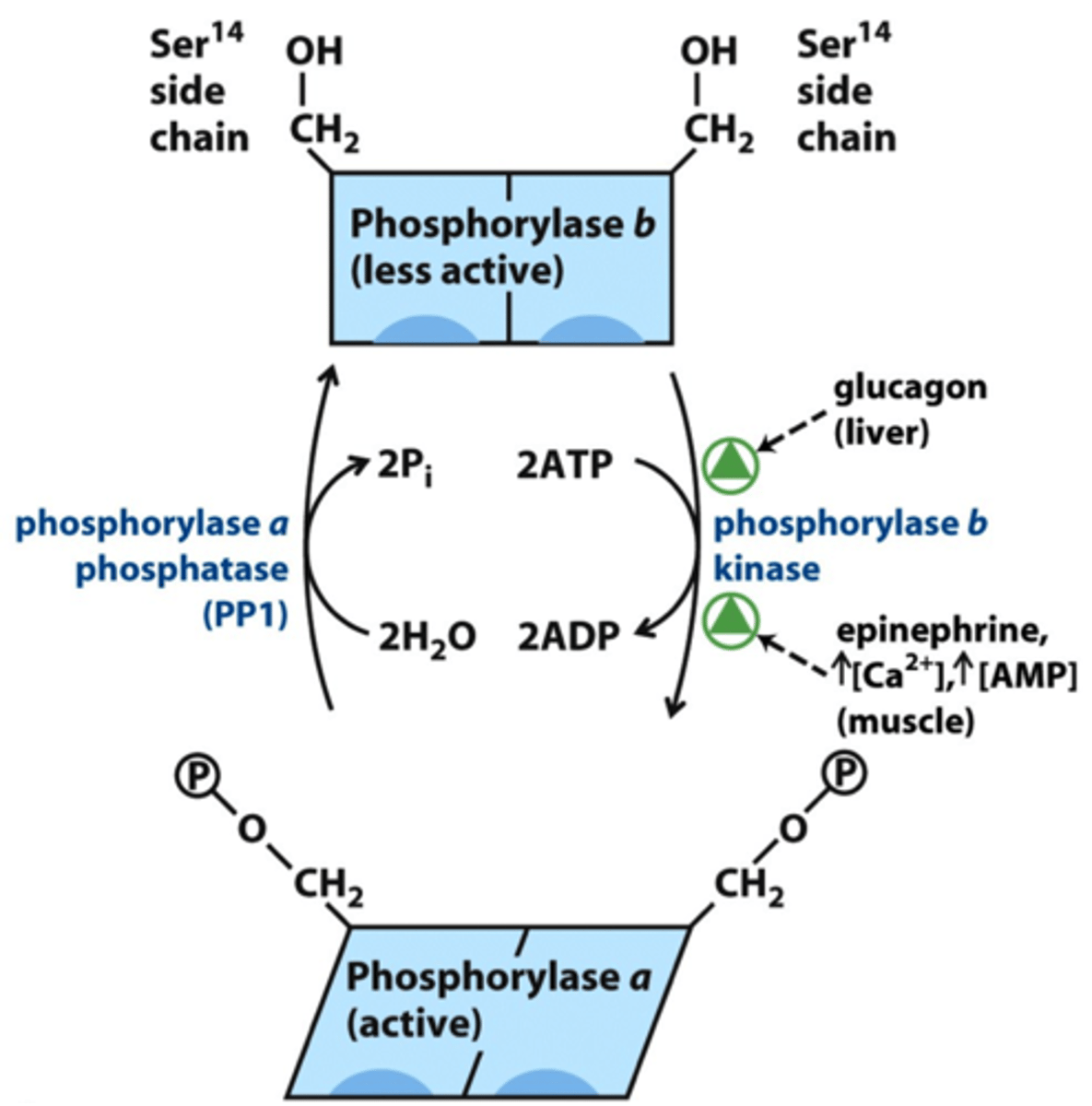
glycogen phosphorylase regulation
allosteric:
- Ca2+: (M) phosphorylase b kinase activated, phosphorylase b -> phosphorylase a
- AMP: (M) phosphorylase b -> phosphorylase a (low energy)
- ATP: (M) phosphorylase a -> phosphorylase b
hormonal:
- epinephrine (M) & glucagon (L) phosphorylates phosphorylase b -> phosphorylase a
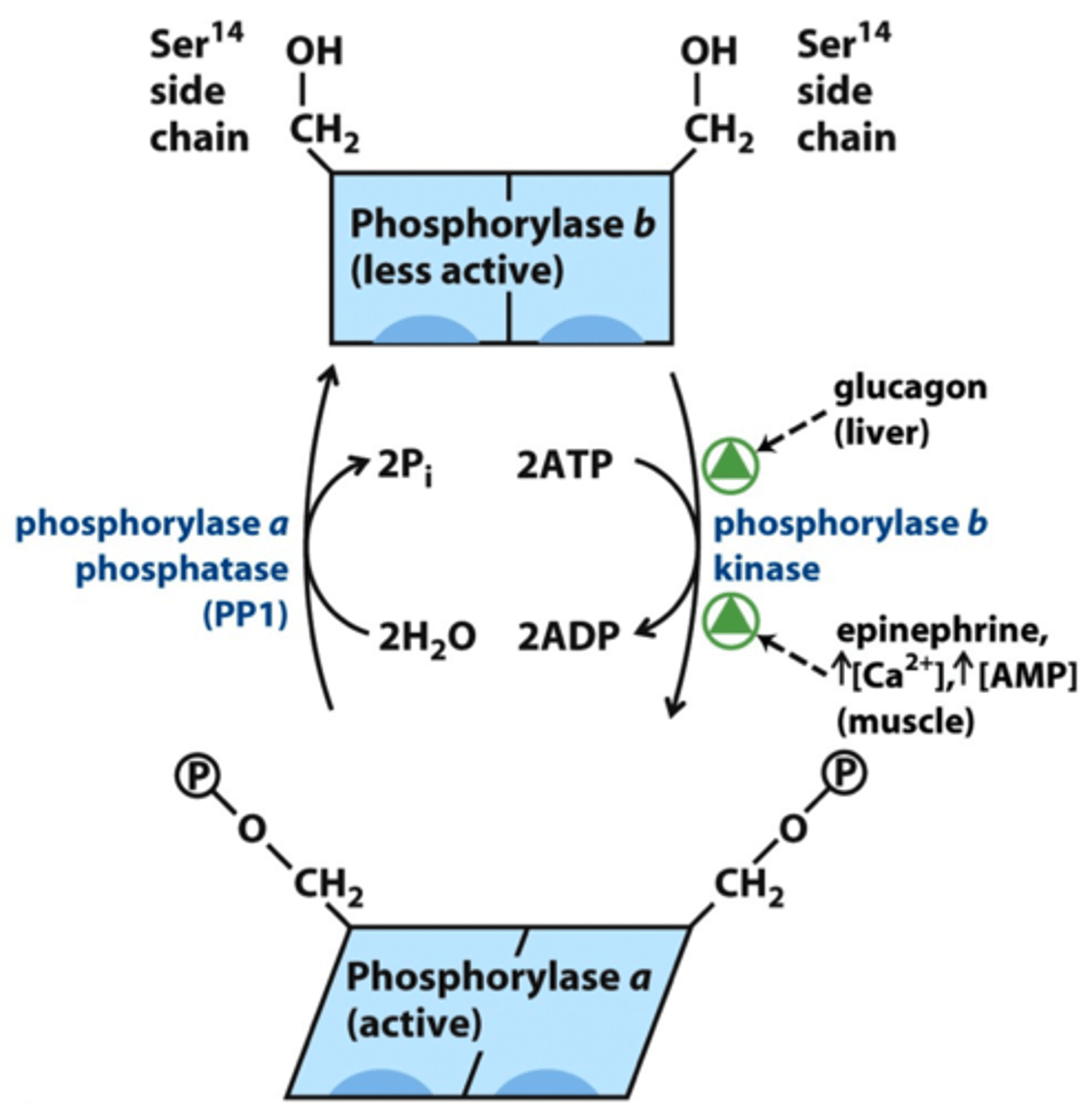
phosphorylase b kinase
- phosphorylated by PKA
- activated allosterically by Ca2+ (muscle contraction)
- phosphorylates phosphorylase b (requires 2 ATP) in low blood glucose
glucose sensor of glycogen phosphorylase
- glucose allosterically binds to phosphorylase a at high glucose levels
- insulin dephosphorylates phosphorylase a and inactivates it (glycogenolysis stops)
glycogenesis
building glycogen with glucose molecules during high blood glucose levels
UDP-glucose
an activated sugar nucleotide that donates glucose for glycogenesis
UDP-glucose phosphorylase
UTP + glucose 1-phosphate -> UDP-glucose + PPi
glycogen synthase
- catalyzes transfer of glucose from UDP-glucose to nonreducing end of glycogen chain with >4 residues
- forms α(1->4) glycosidic bonds
branching enzyme
- takes 7-10 residues and transfers them 4 residues in on a linear chain
- forms α(1->6) glycosidic bond
glycogen synthase regulation
- enhances a: insulin, G6P
- inhibits a: glulcagon, epinephrine
- inhibits b: insulin prevents phosphorylation (inhibits glycogen synthase kinase 3)
- high glucose (b -> a) dephosphorylates enzyme to make glycogen through synthase a
- low glucose (a -> b) phosphorylates enzyme to stop glycogenesis
PDH complex
pyruvate from glycolysis moved to mitochondria -> acetyl-CoA product
- E1 : thiamine pyrophosphate (TPP)
- E2: coenzyme A (CoA-SH) & lipoic acid (lipoate)
- E3: FAD & NAD
- regulatory kinase & phosphate, large negative delta G
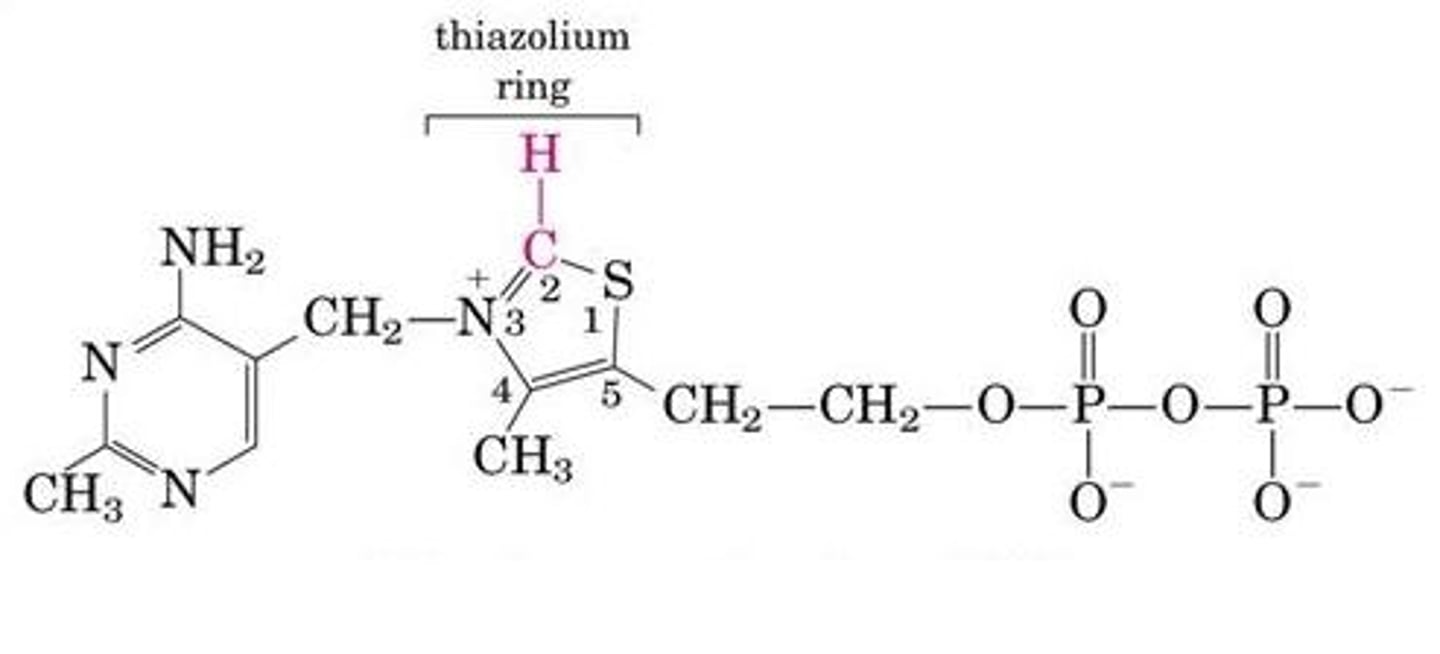
thiamine & thiamine pyrophosphate (TPP)
- thiamine (vitamin B1) in TPP, an enzyme cofactor in the E1 complex
- carbon on thiazolium ring is used during aldehyde transfer
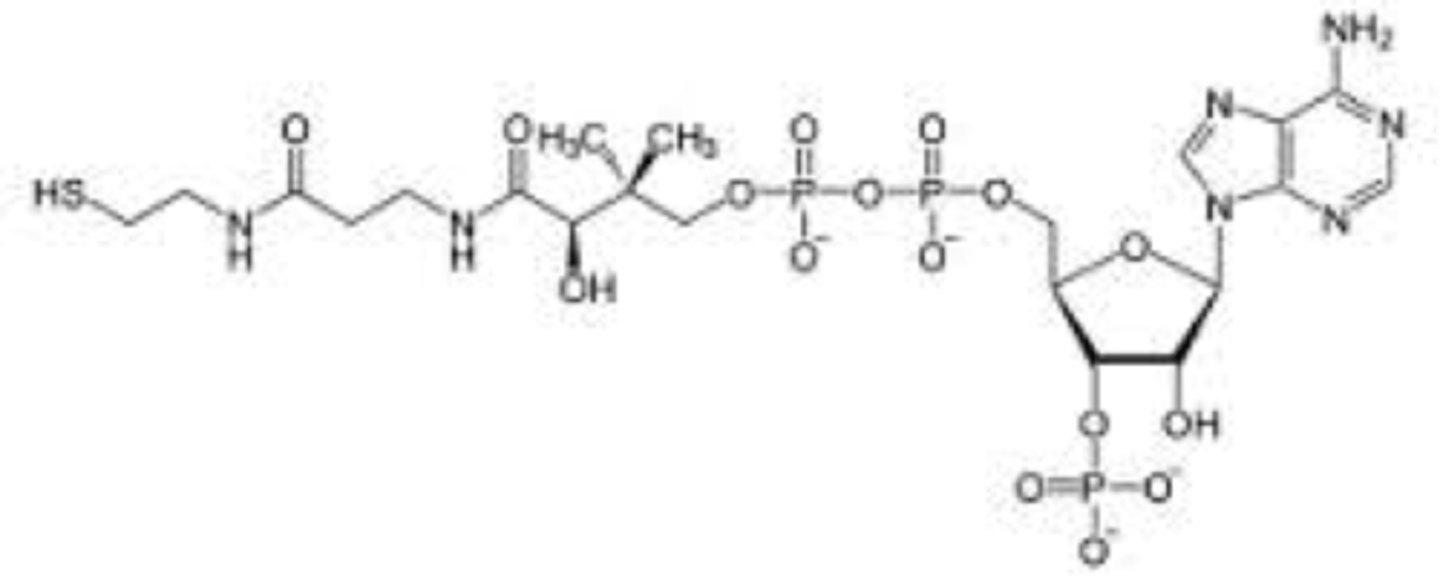
CoA-SH & acetyl-CoA
- cofactor of E2
- has reactive thiol group, forms thioester w/acetate to make acetyl-CoA
- has pantothenic acid (vitamin B5)
- functions as an acyl carrier
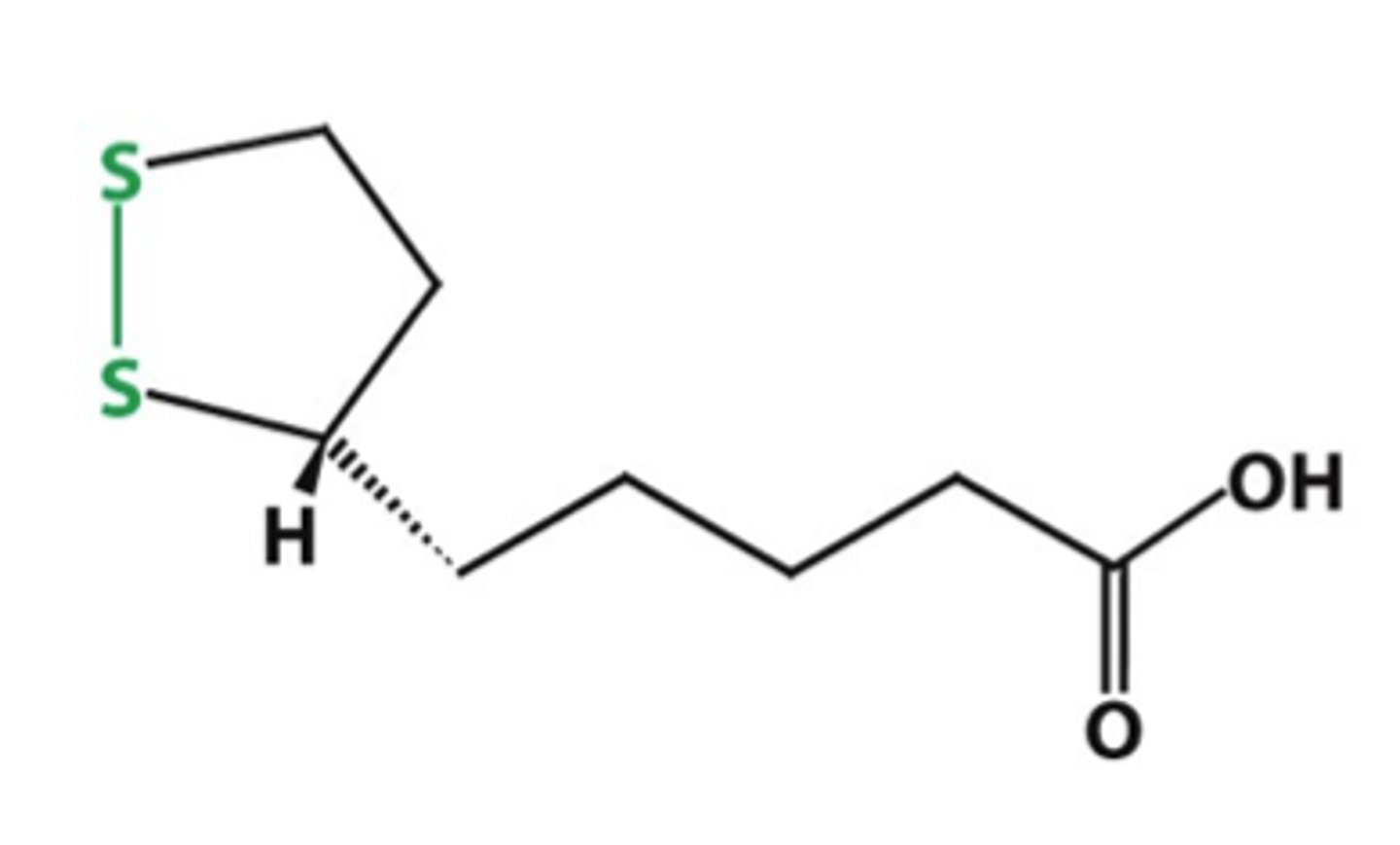
lipoic acid
- cofactor of E2
- has disulfide (reactive thiol), can accept acetyl group
- can be reduced (disulfide bond broken) to get acetylated
FAD and NAD+
- flavin: FAD, three rings
- niacin: NAD, one ring
- FADH2 is a cofactor in complex III and is oxidized to pass 2 e- to NAD+
E1 of PDH
- E1: pyruvate dehydrogenase
- TPP decarboxylates pyruvate -> hydroxyethyl TPP
- CO2 released
- rate limiting step
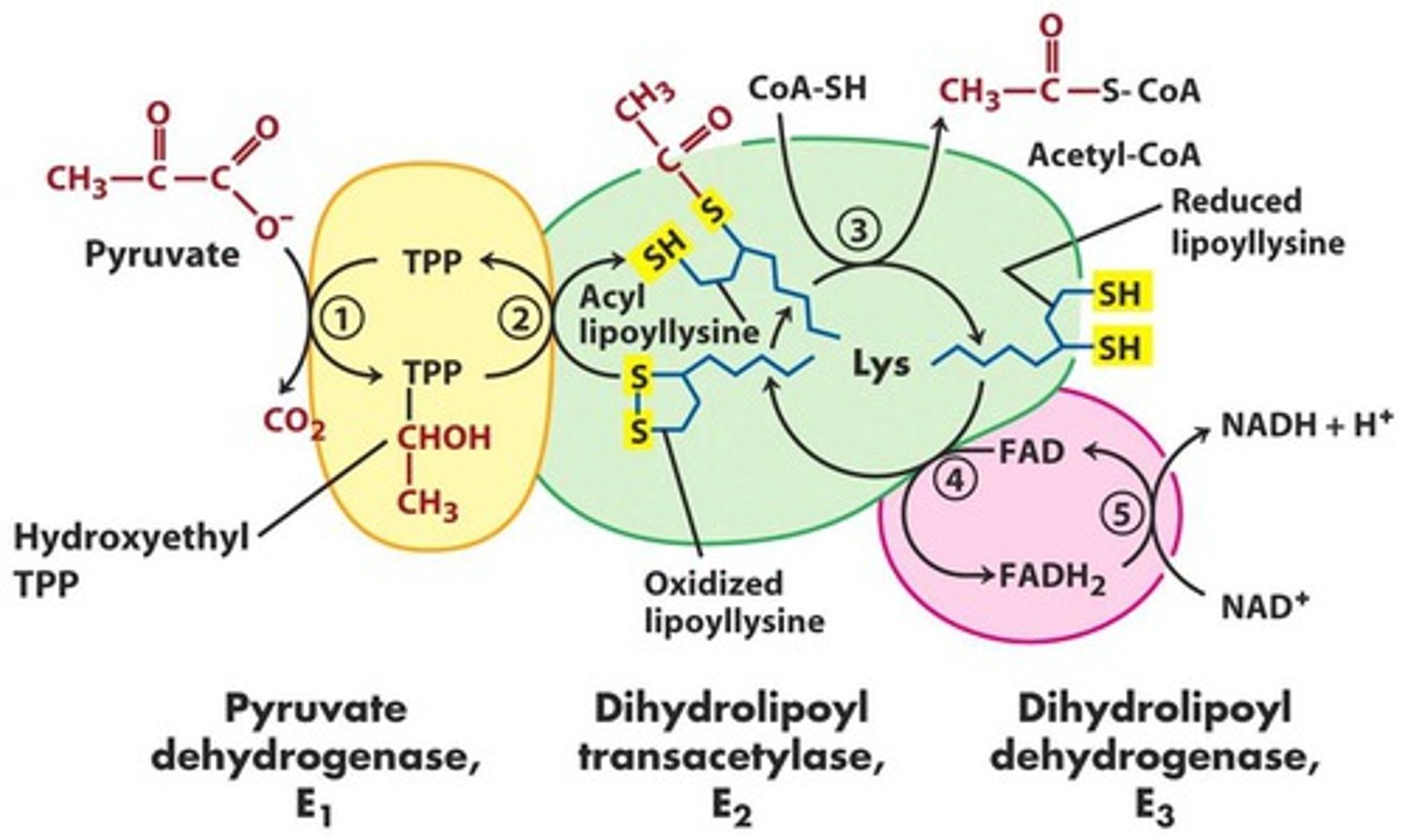
E2 of PDH
- E2: dihydrolipoyl transacetylase
- disulfide on lipoic acid is reduced (bond broken)
- transfers hydroxyethyl group to thiol group as acetyl
- acyl transfer to CoA -> acetyl-CoA
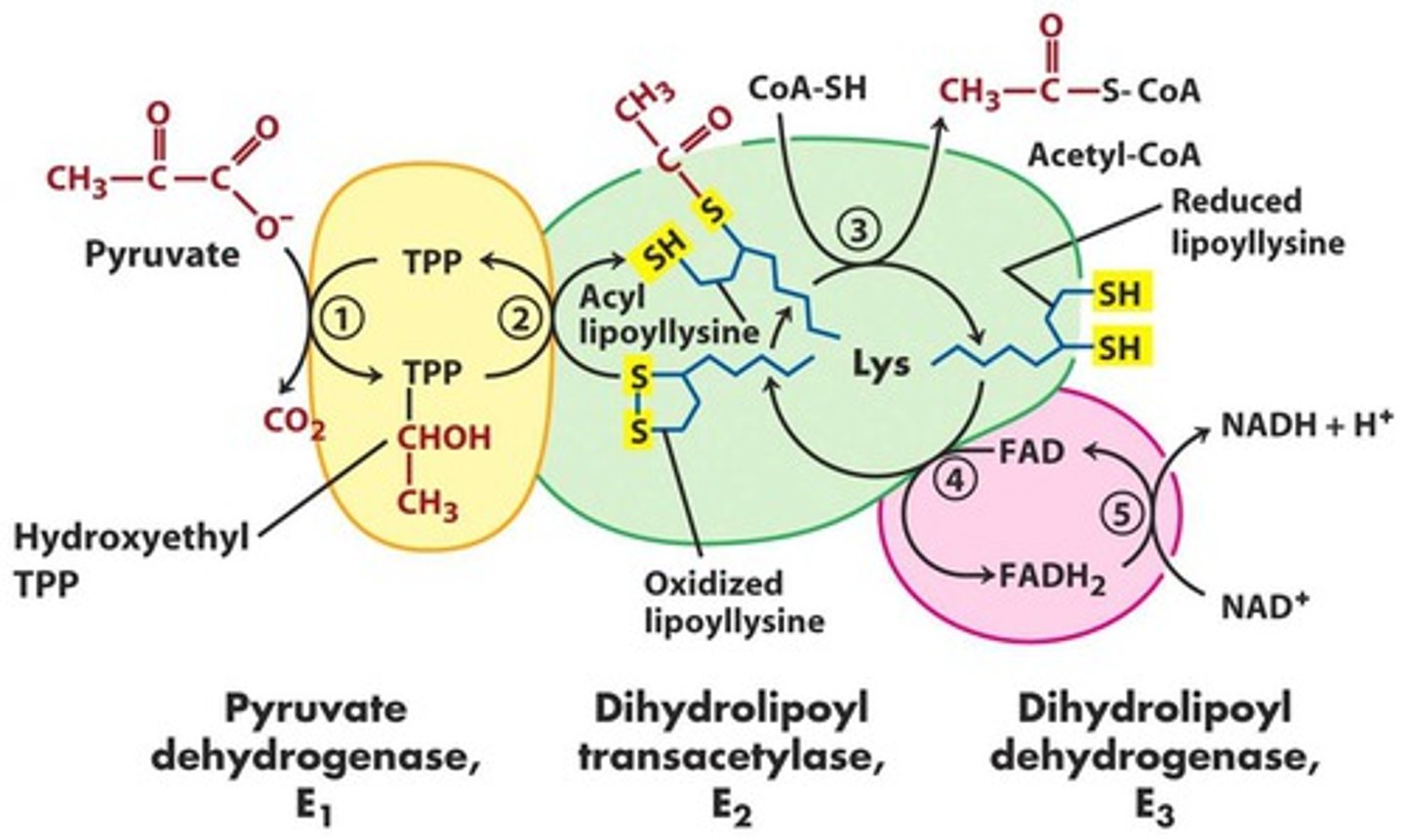
E3 of PDH
- E3: dihydrolipoyl dehydrogenase
- reduced lipoic acid are passed to FAD -> FADH2 (stuck in E3)
- FADH2 -> FAD and passes e- to NAD+ to enter ETC
PDH allosteric regulation
- inhibit: ATP, products (acetyl-CoA and NADH)
- enhance: AMP, reactants (CoA-SH and NAD+)
- lots of glycolysis = lots of ATP = stop PDH, don't need to make more acetyl-CoA for TCA
- little glycolysis = low energy = more PDH to make more acetyl-CoA for TCA = more ATP
PDH covalent regulation
- active: dephosphorylated by PDP from insulin
- inactive: phosphorylated by PDK (NOT BY GLUCAGON)
1st step of TCA
reactant: acetyl-CoA + OAA
enzyme: citrate synthase (H2O -> CoA-SH)
product: citrate
- Claisen condensation
- large negative delta G, highly regulated
- lots of energy from hydration of thioester in acetyl-CoA
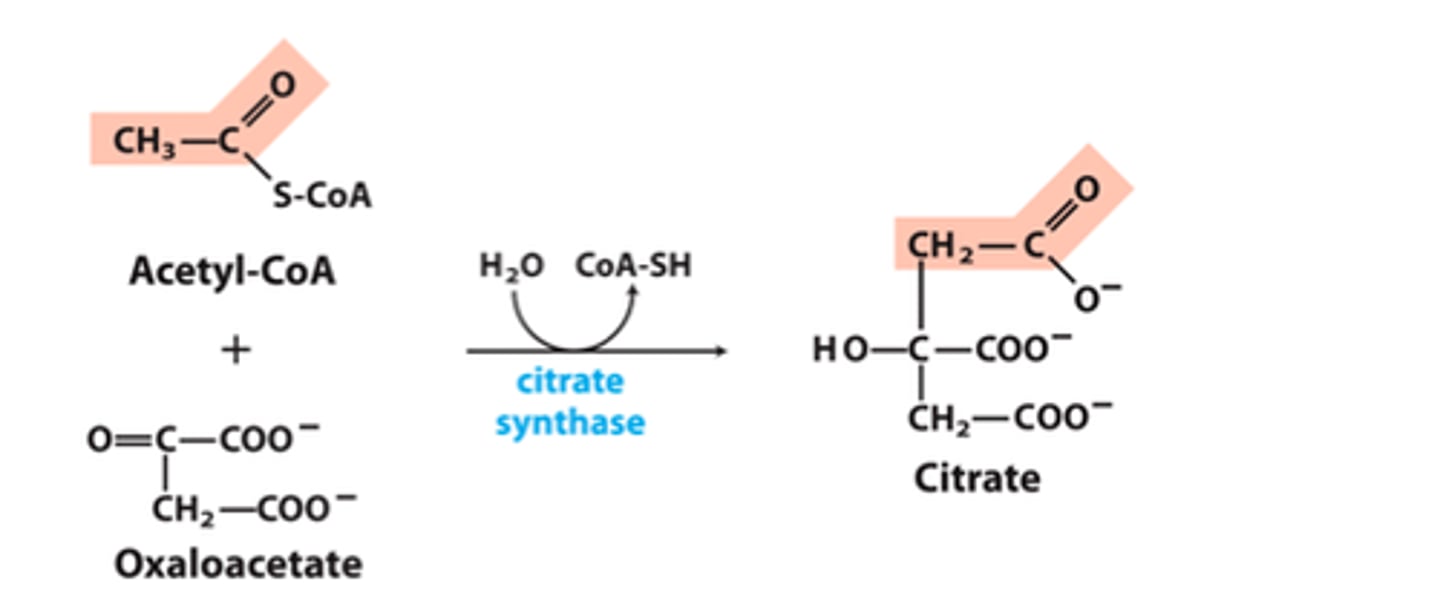
2nd step of TCA
reactant: citrate
enzyme: aconitase
product: isocitrate
- dehydration, rehydration
- moves OH group from C3 to C2 (tertiary -> secondary alcohol) for oxidation
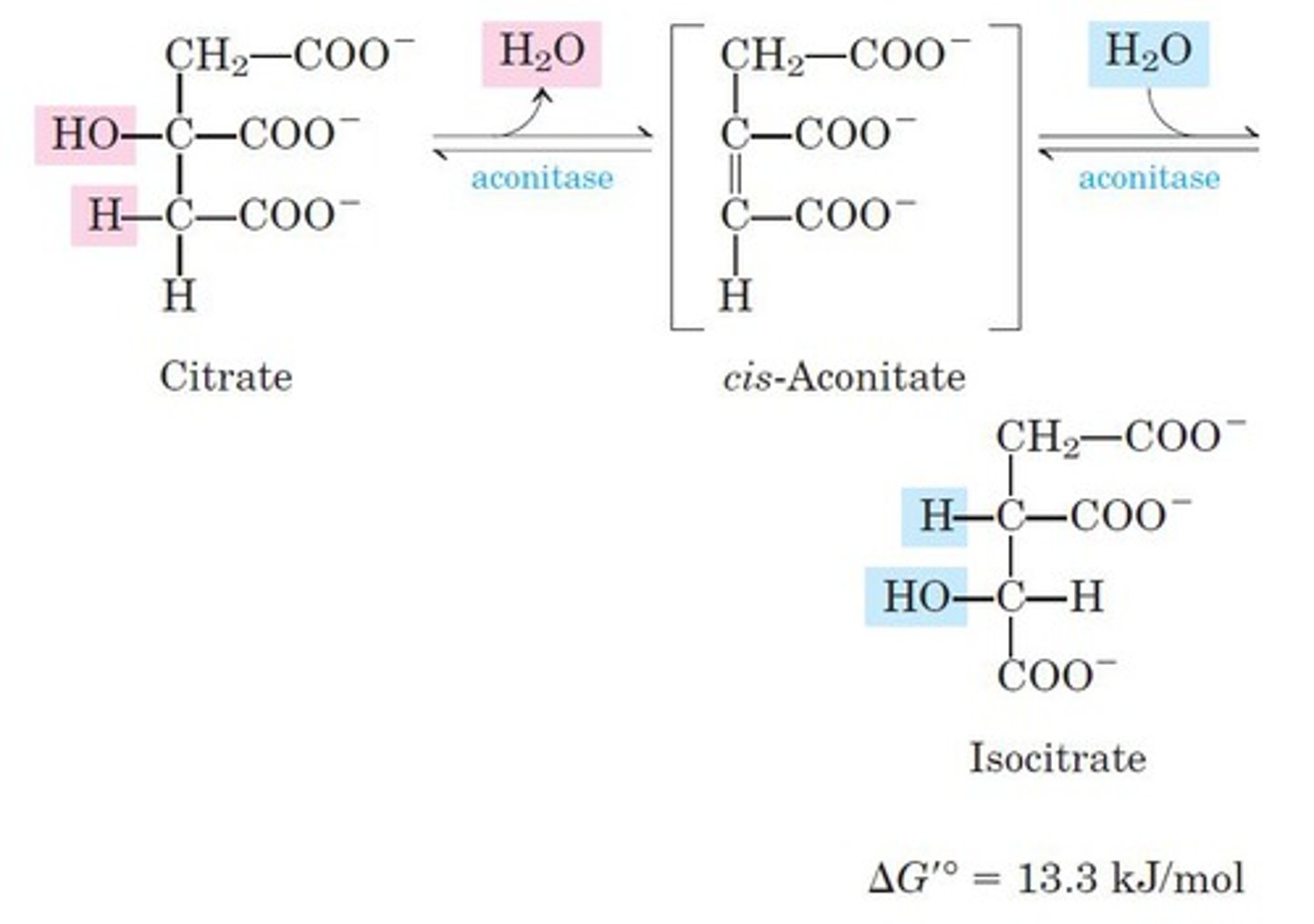
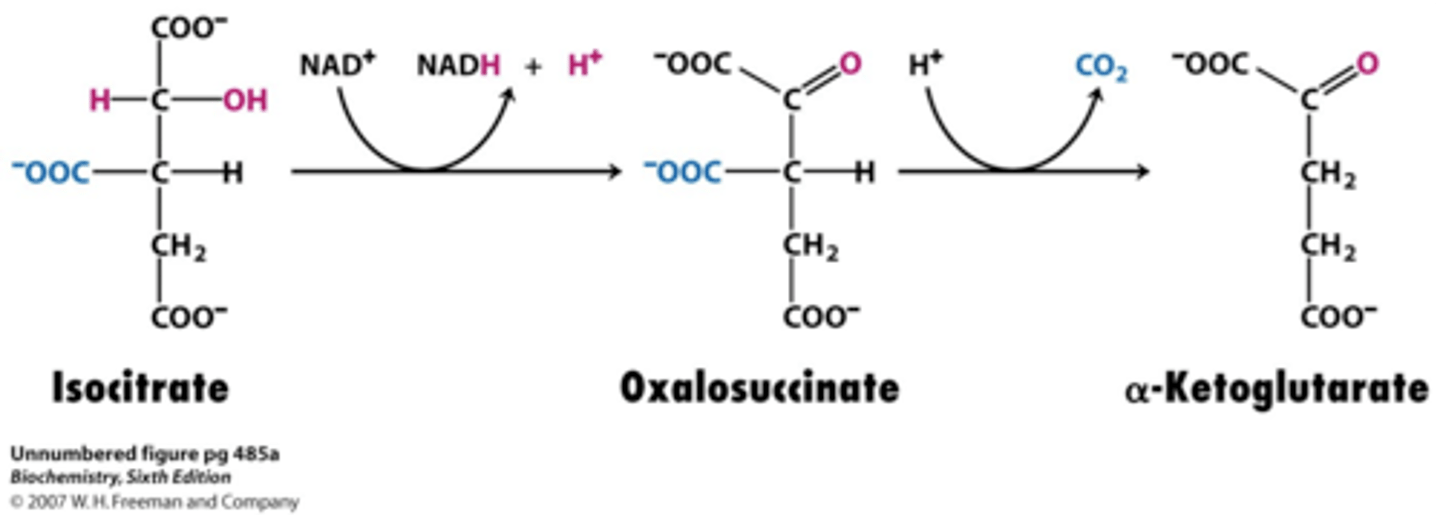
3rd step of TCA
reactant: isocitrate
enzyme: isocitrate dehydrogenase
product: α-Ketoglutarate
- oxidative decarboxylation of isocitrate
- NAD+ -> NADH + H+ (reduced e-carriers donates energy)
- NADP -> NADPH + H+ in cytosol
- CO2 released (6 -> 5 C's)
- highly regulated