Gene-L5-Genome III: chromosomes, telomeres and gene silencing
1/11
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
12 Terms
how is the human eukaryotic genome organised on chromosome level?
22 autosomes, 2 sex chromosomes
p arm- short
q arm- long
what happens to chromosomes during the cell cycle- interphase and M phase
interphase
DNA is replicated and each chromosome is duplicated- has 2 copies of each homologous chromosome
M phase- mitosis
chromosome condenses into visible chromosomes
nuclear envelope breaks down
chromosomes are captured by mitotic spindles and pulled apart to opposite poles
nuclear envelope reforms
what are centromeres, what is their structure? how do they function during cell division?
centromeres-DNA sequences that directs movement of chromosomes into daughter cells during cell division
serves as the assembly site for specific proteins: H3 histone variant CENP-A forming a specialised nucleosomes
additional proteins assemble on the centromere- kinetochores which attaches chromosomes to the mitotic spindle
centromere is located within the heterochromatin- silenced through deacetylation and methylation
what features of chromatin are found at the centromere? how do they affect gene activity?
centromeric chromatin- heterochromatin
has special H3 variant
histone deacetylation
histone methylation
makes compact and stable chromatin structue
what are the origins of replication- how do they differ in prokaryotes and eukaryotes?
origin of replication: DNA sequence where replication begins
prokaryotes: one single origin as they have less DNA to replicate
humans: multiple origins per chromosome. at AT rich regions- easier to unwind due to 2 hydrogen bonds
what are telomeres? where are they located and what is their function?
telomeres: end structures of eukaryotic chromosomes- not proteins, just DNA sequnces
composed of repeated sequences
G-rich sequences- makes a 3’ single stranded overhang of 15 nucleotides
can be extended by telomerase enzyme
chromosome end stability: DNA folds back into itself to make a t loop. prevents the exposure of single stranded DNA and stops activation of DNA damage repair pathways
prevention of information loss during replication: compensates for loss of DNA due to primer removal on lagging strand
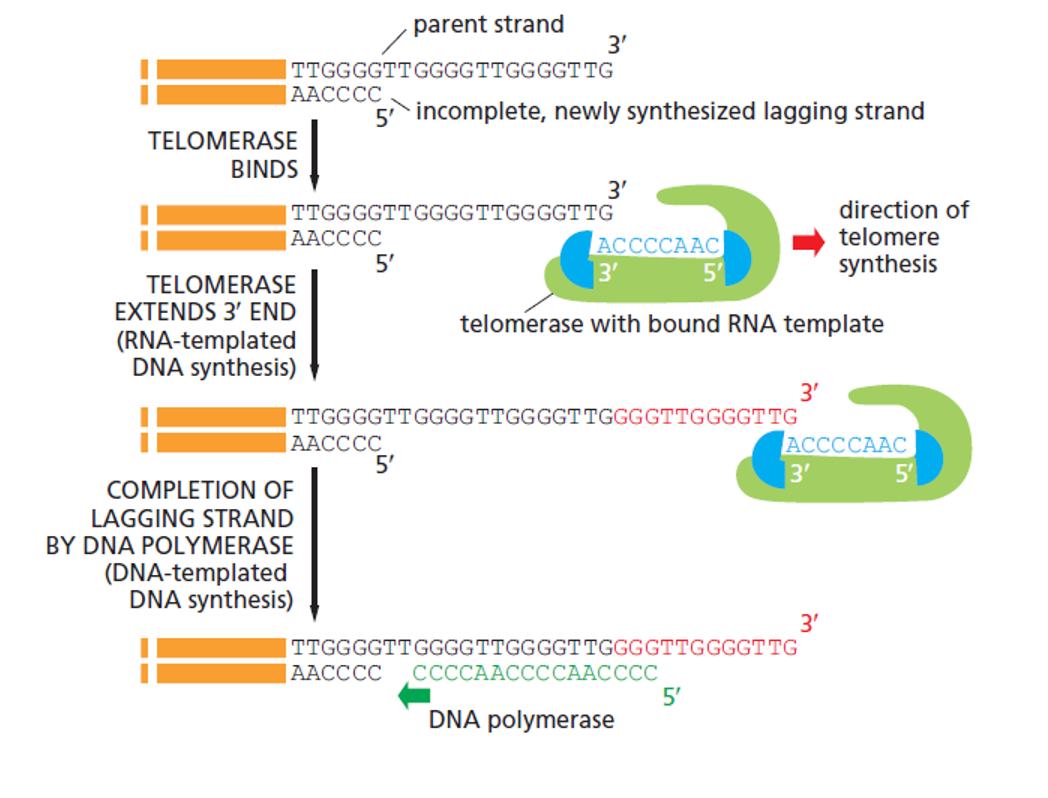
how does telomerase solve the end replication problem?
telomerase binds to 3’ end and has an RNA template where it attaches an extra telomere
now DNA polymerase has somewhere to bind and replicate the information that would have been lost
how does telomere shortening relate to ageing, disease and chromatin silencing?
somatic cells have little to no telomerase activity and therefore shorten over time
leads to cellular senescenc
short telomeres- accelerate ageing dyndrome
Down syndrome- accelerated telomere loss
how does telomere gene silencing occur in yeast? what do SIR proteins do? what does Rap1 do?
SIR- silence information regulator
yeast telomere-bound SIR protein complex recognises underacetlyated (H4) or methylated (H3) N-terminal
tails of selected histones and keeps
them deacetylatedRap1 recognises telomeric DNA that needs to be silenced and acts as a signal for SIR to be recruited
SIr assembles and spreads along the chromosome from this site- modifies the N terminal this
how does methylation lead to gene silencing? how is it recognised by the immune system?
DNA methylation is a gene silencing mechanism
occurs at cysteine bases- forms 5-methylcytosine
common at CpG C-G dinucleotides
in humans: DNA is usually hypermethylated (silenced)
in bacteria and viruses, DNA is undermethylated and immune system recognises this via TLR9 recognising undermethylated CpG DNA
how are DNA methylation patterns maintained after DNA replication?
after replication: parental strand is methylated and the new strand is ynmethylated
DNA is temporarily hemimethylated
DNA methyltransferase enzymes helps methylate the full DNA by
recognition of methylated cytosine on parental strand
methylation of the corresponding cytosine on the new strand
characteristics features of eukaryotic genes? 4
promotor region: DNA sequence where RNA polymerase and TFs bind and defines the start of transcription
regulatory proteins: have DNA sequences recognised by regulatory proteins to control gene expression levels
Exons- coding sequences which are transcribed into RNA and translated into proteins
introns: non coding sequences and in between cons and removed by splicing