Exam 3 Question Set to Study for Exams
1/152
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
153 Terms
Which organism from the case study was resistant to all treatments and has been found to cause disease in battled injured soldier from Iraq?
Acinetobacter baumannii
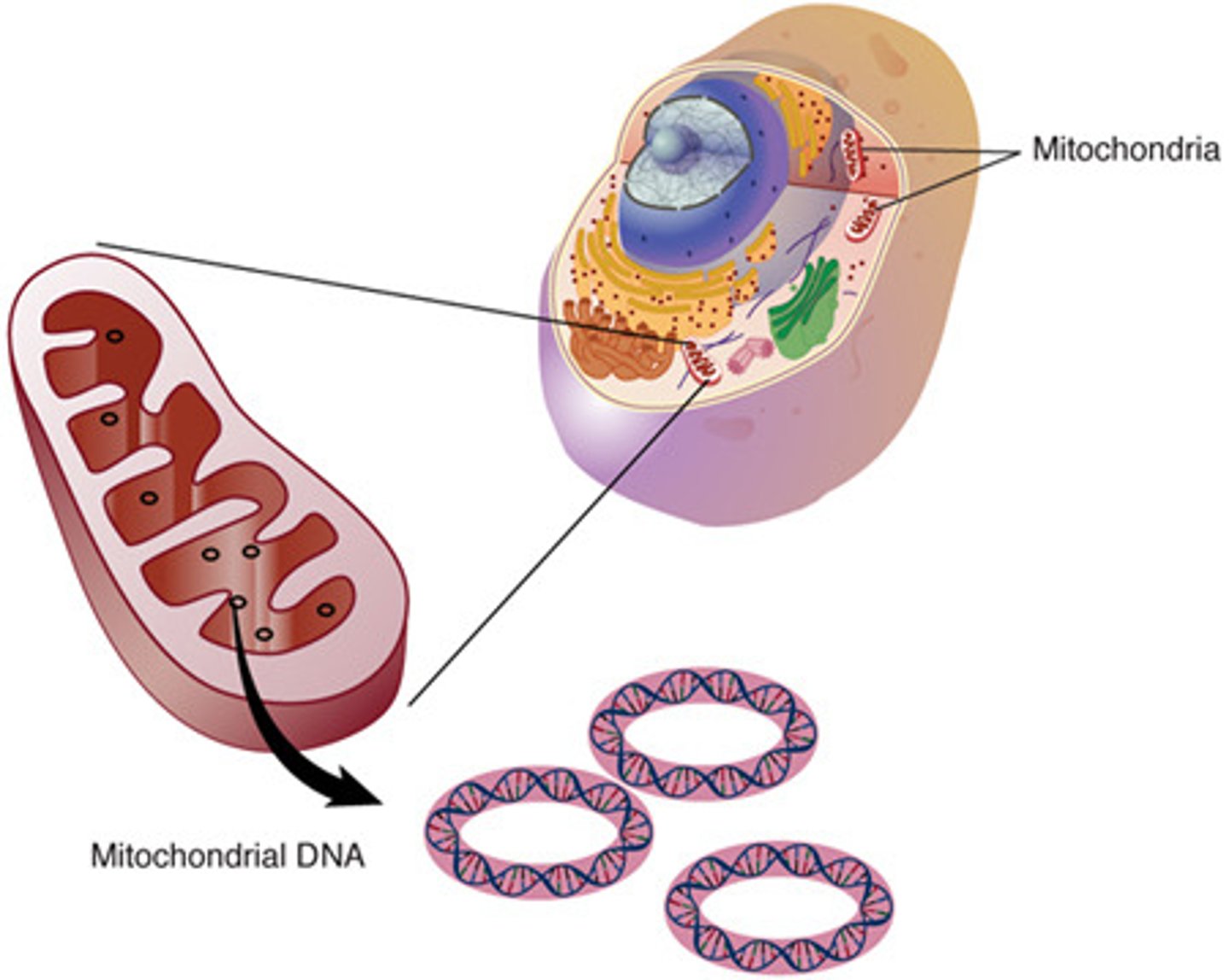
_______ and _______ are extracellular DNA molecules that can be found only in eukaryotic cells.
Mitochondrial and chloroplast DNA
What nucleic acid/s can make up the eukaryotic genome?
DNA
What nucleic acid/s can make up the prokaryotic genome?
DNA
What nucleic acid can make up the viral genome?
DNA or RNA
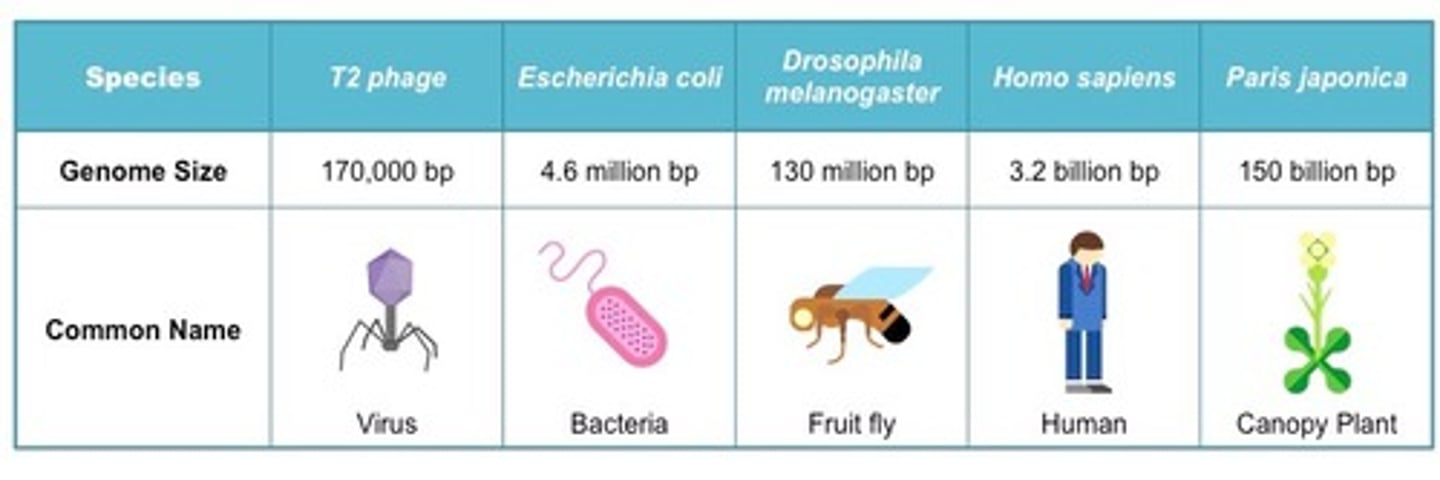
The size of the genome correlates to the size of the microbe. True or False
False
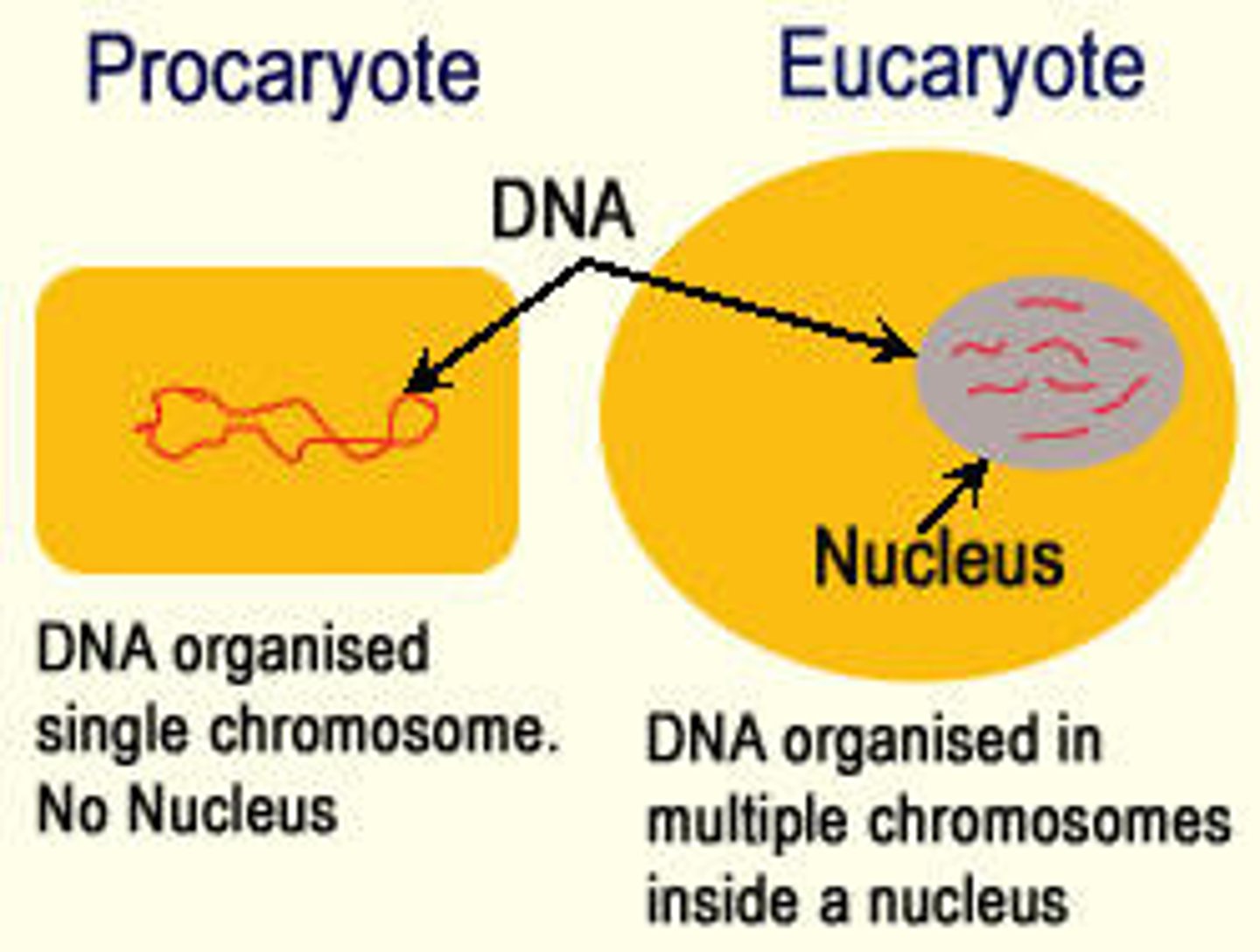
What is the main difference between eukaryotic and prokaryotic chromosomes?
•Prokaryote: Single double-stranded DNA circular loop in the nucleoid
•Eukaryote: Multiple, linear double-stranded DNA in the nucleus
Do eukaryotes have plasmids?
Yes some protozoa and fungi have plasmids
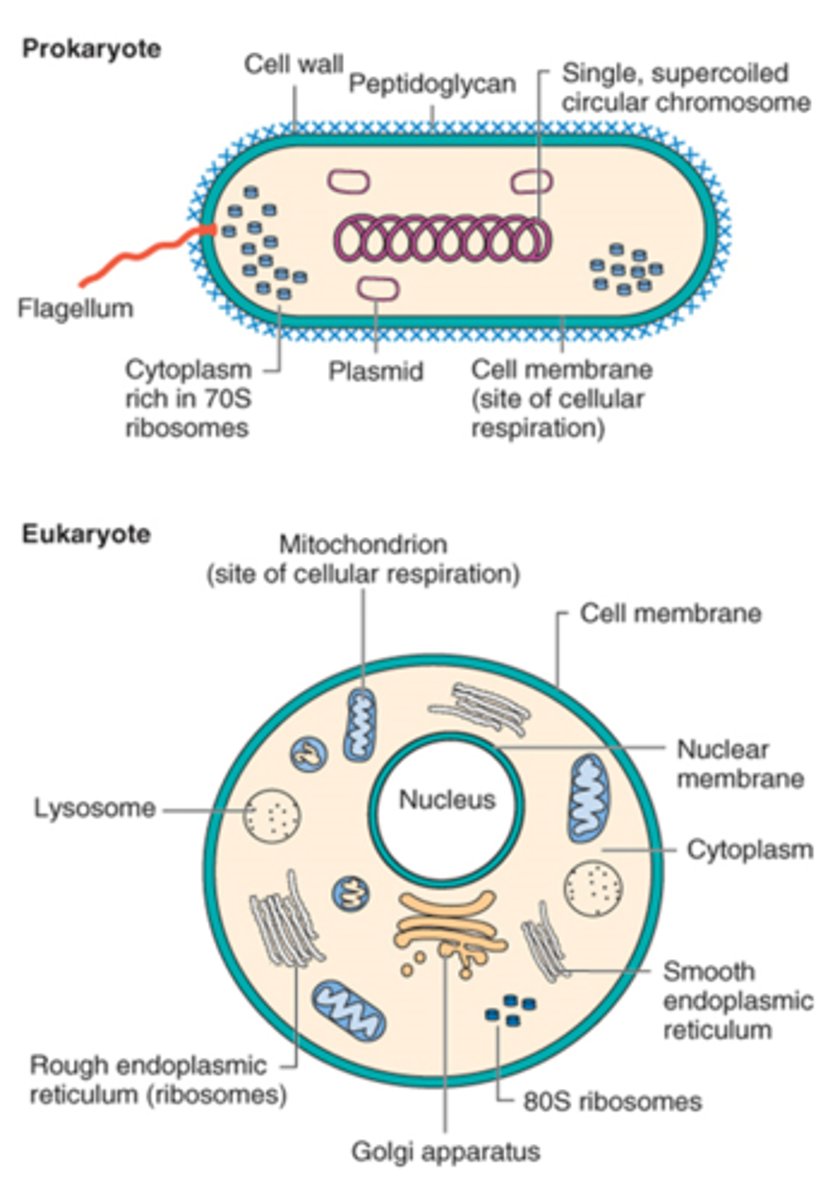
Bacteria use this enzyme to package their DNA.
Topoisomerase or gyrase
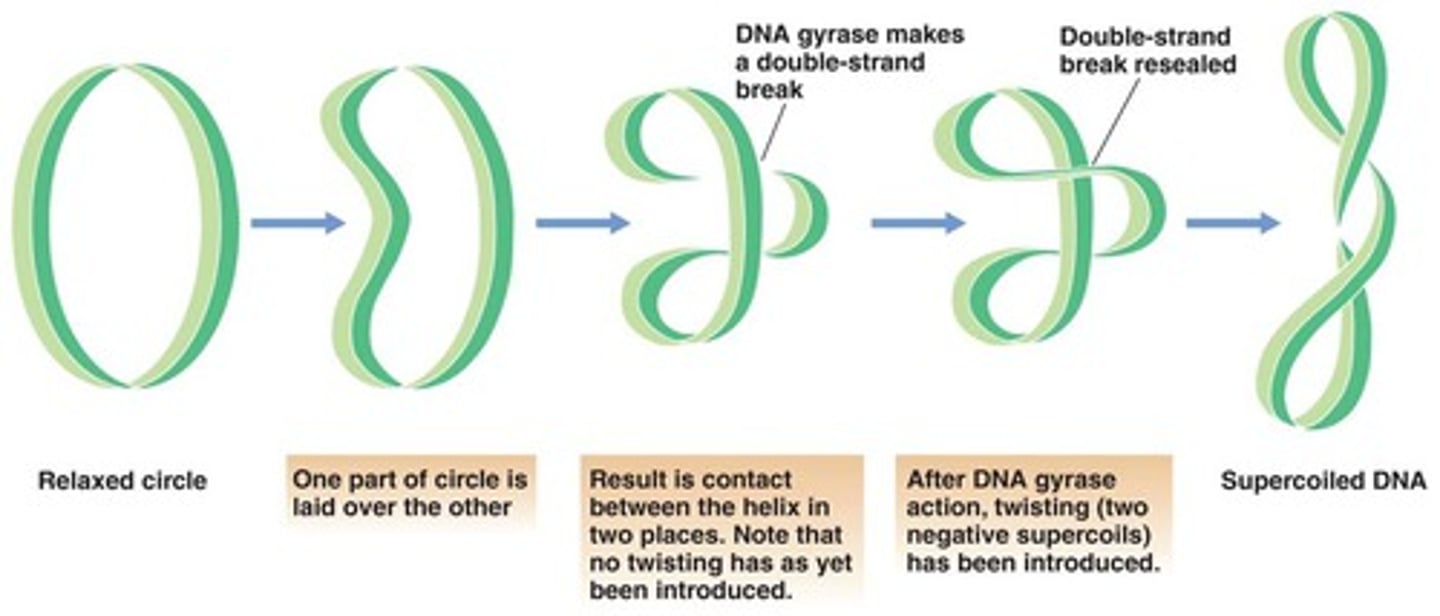
What are the three basic categories of genes?
•Genes that code for proteins: Structural genes
•Genes that code for RNA
•Genes that control gene expression: Regulatory genes
Name the three components of a DNA nucleotide.
5 carbon sugar: Deoxyribose
Nitrogenous base: Adenine, thymine, guanine, cytosine
Phosphate group
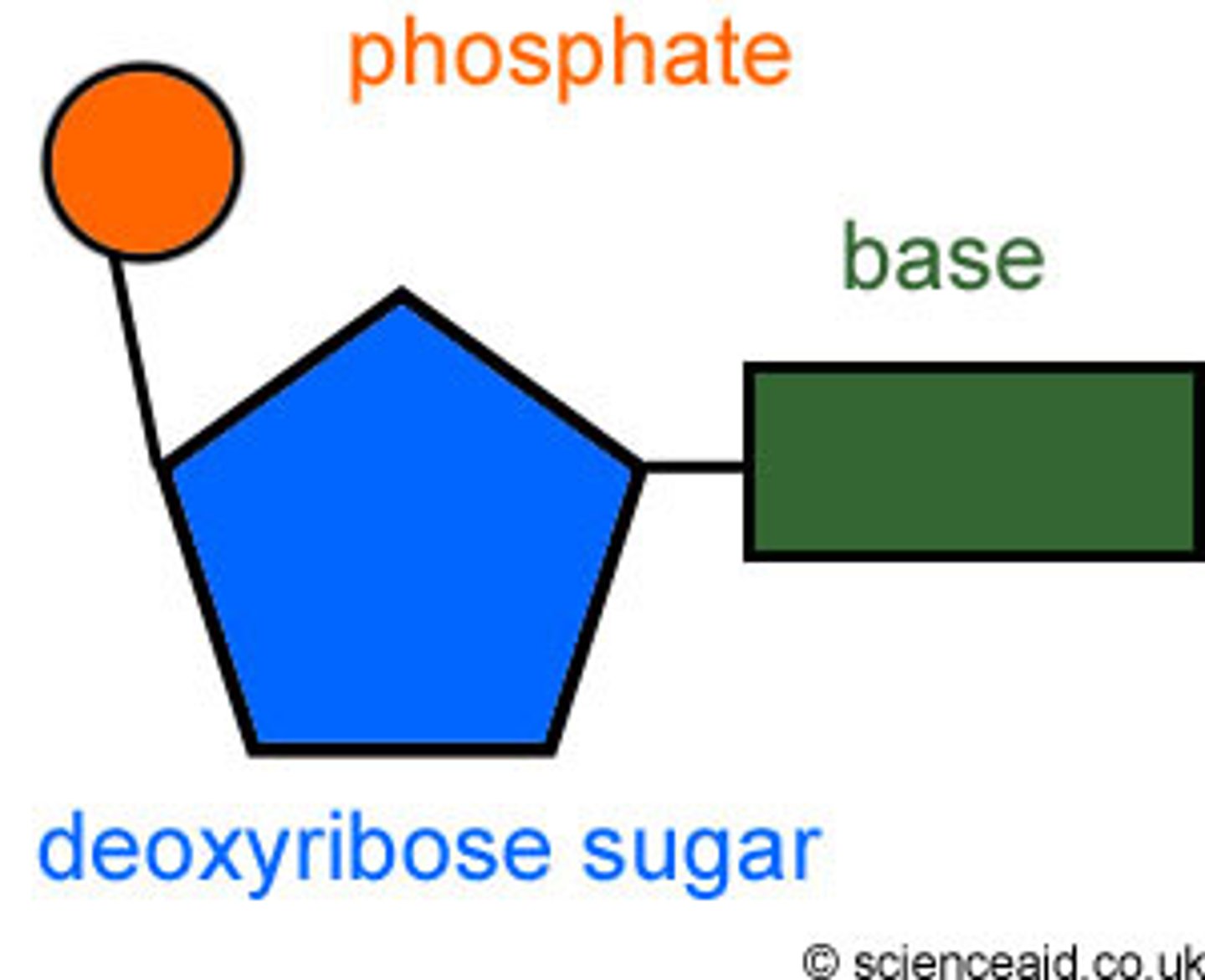
Type of bond used to hold nucleotides together in one strand
Covalent bonds/Phosphodiester bond
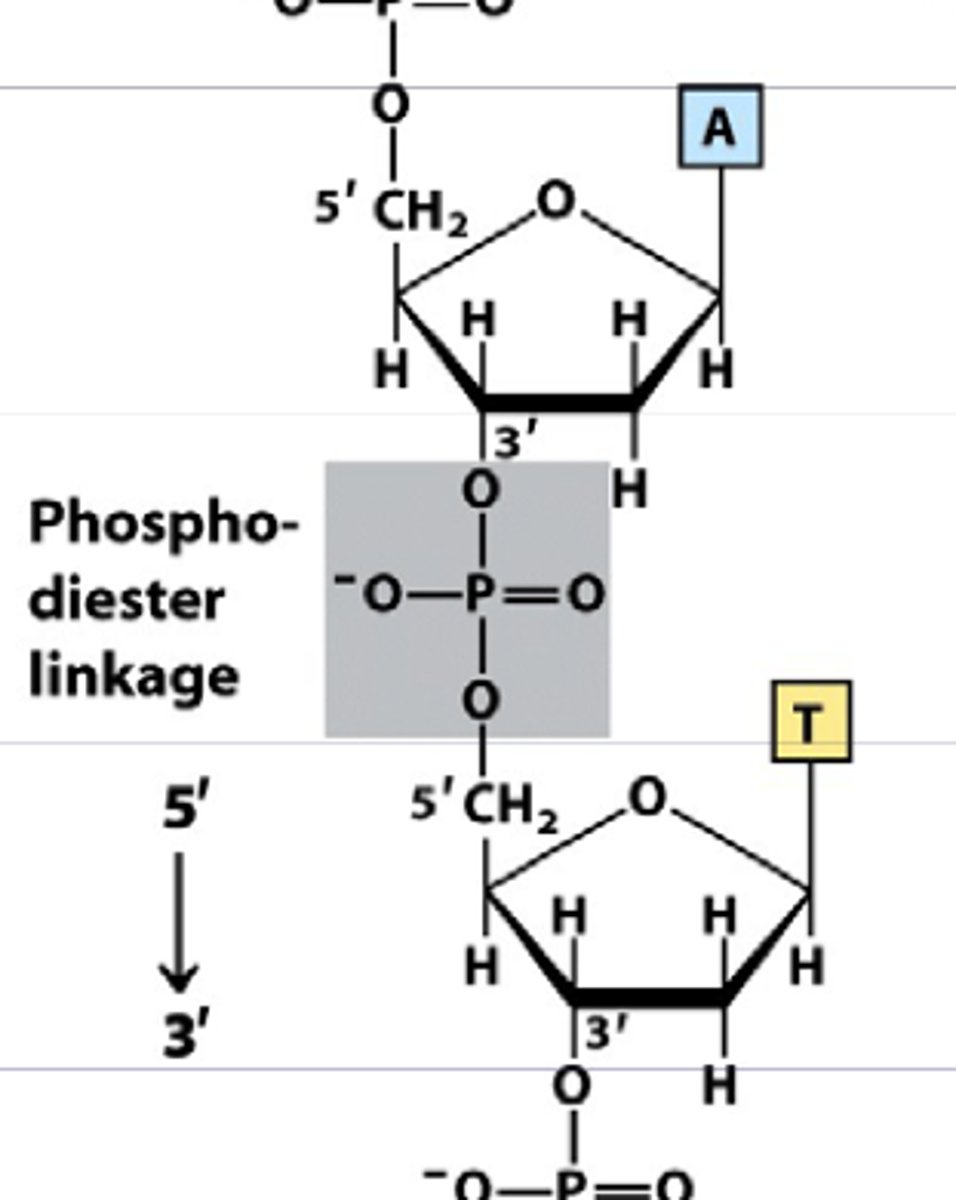
Type of bond used to hold two DNA strands together in a helix
Hydrogen bonds
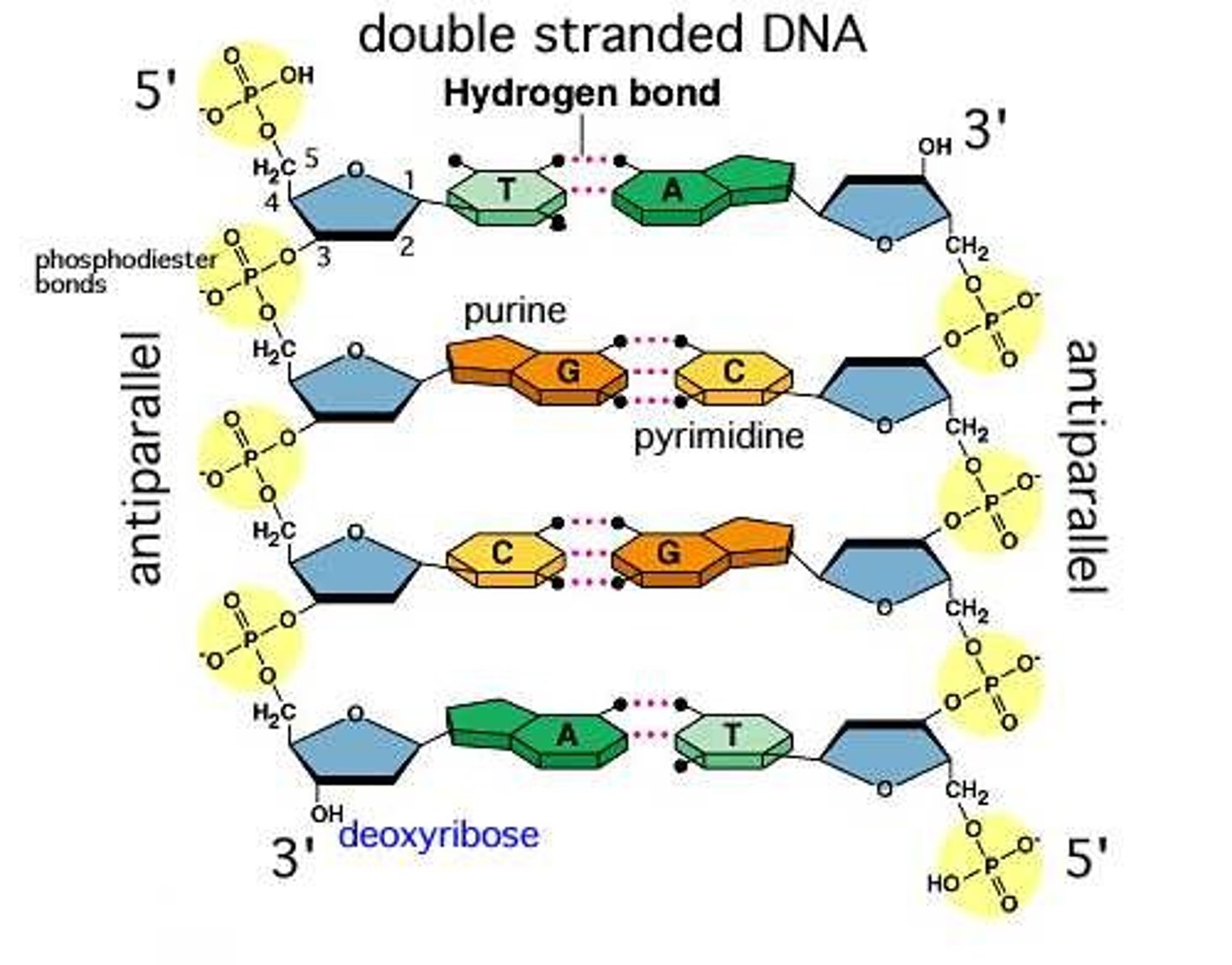
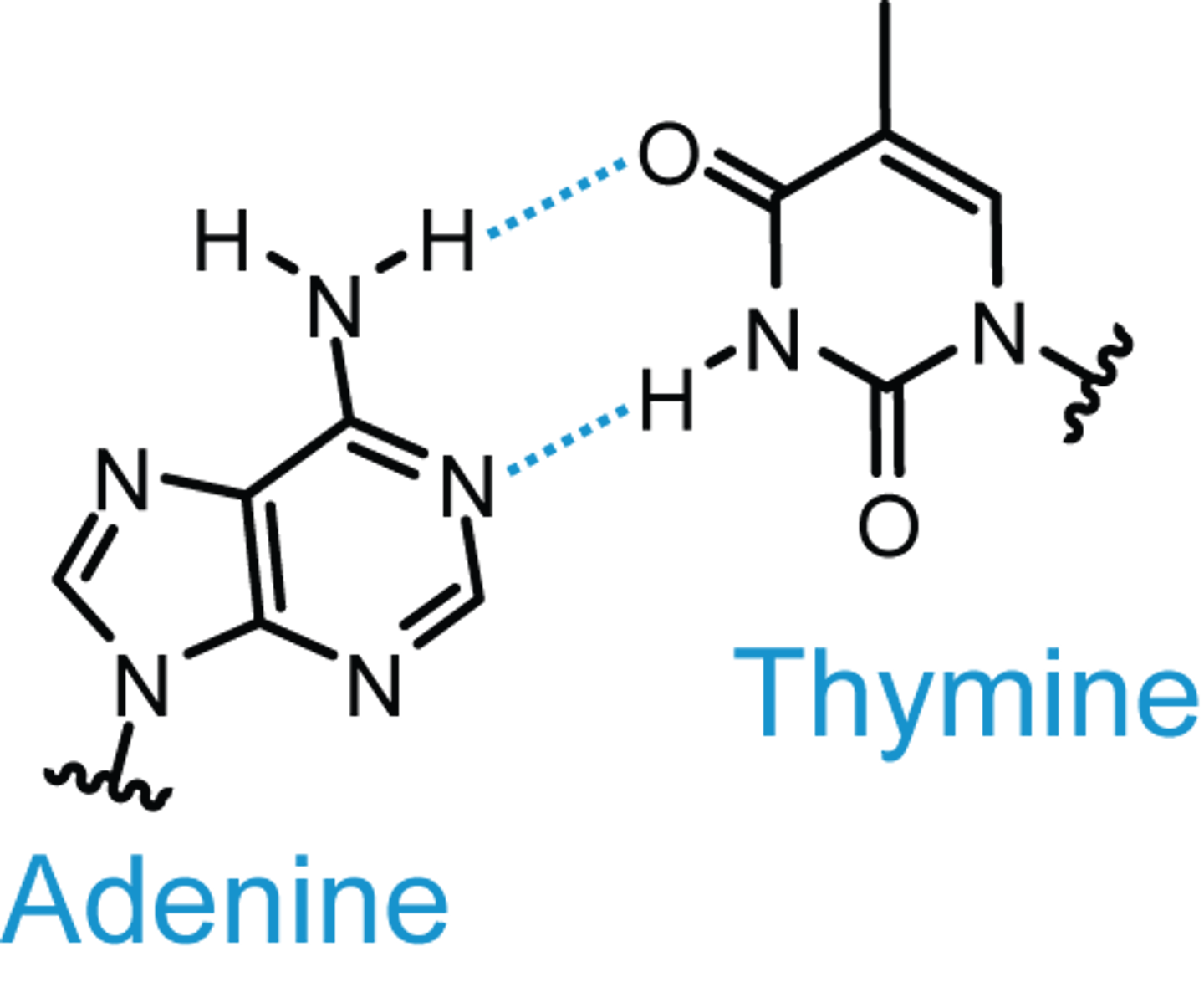
What nitrogen base pairs with adenine in DNA?
Thymine
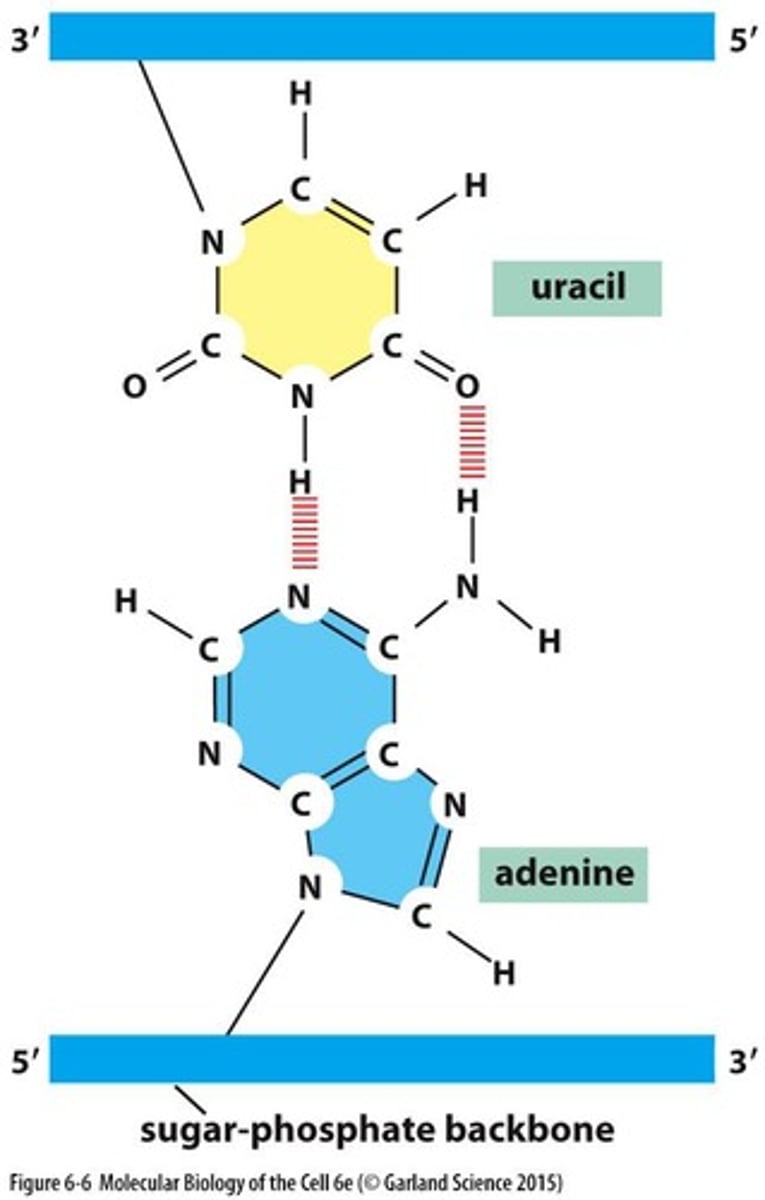
What nitrogen base pairs with adenine in RNA?
Uracil
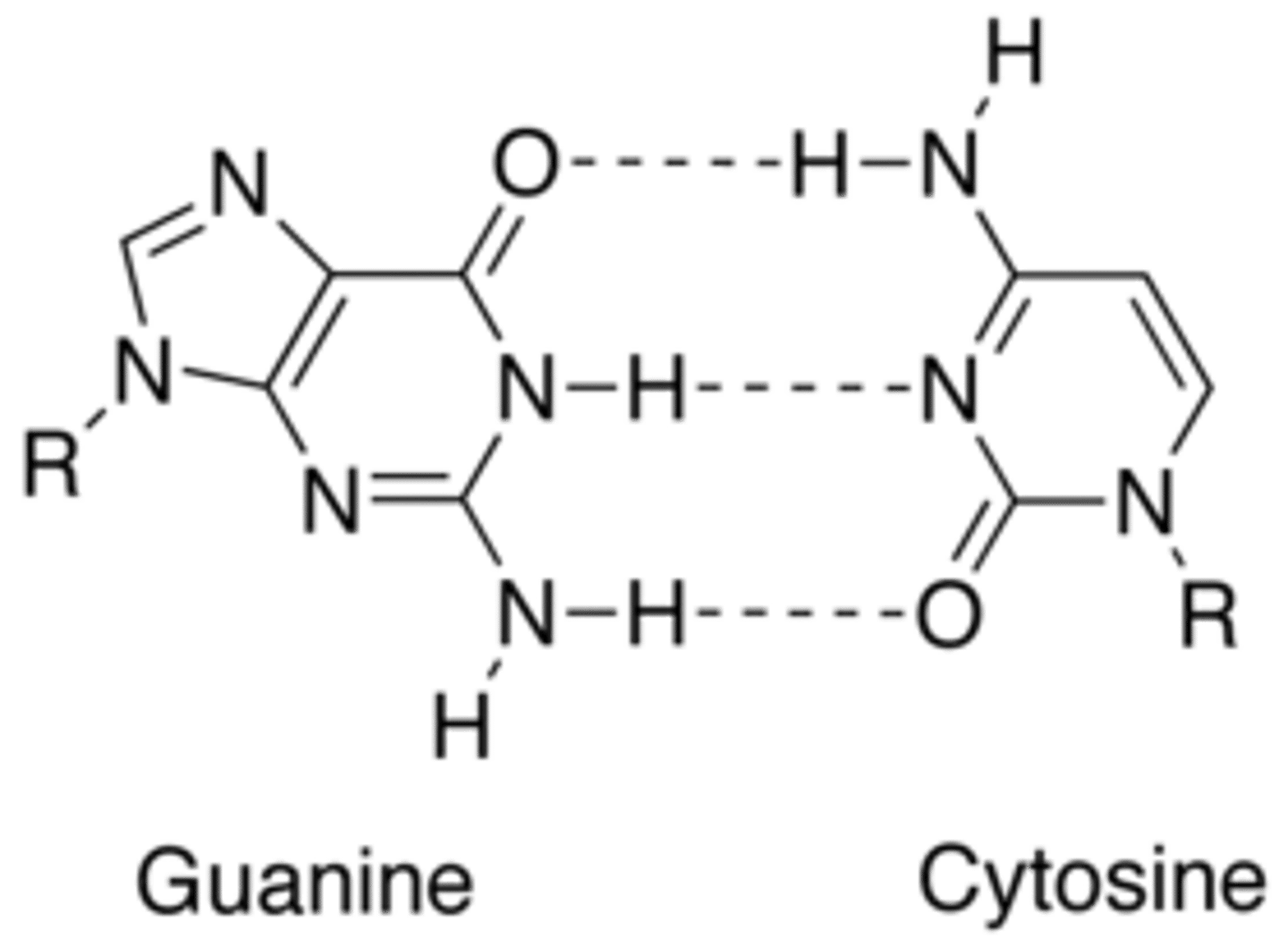
What nitrogen base pairs with cytosine in DNA?
Guanine
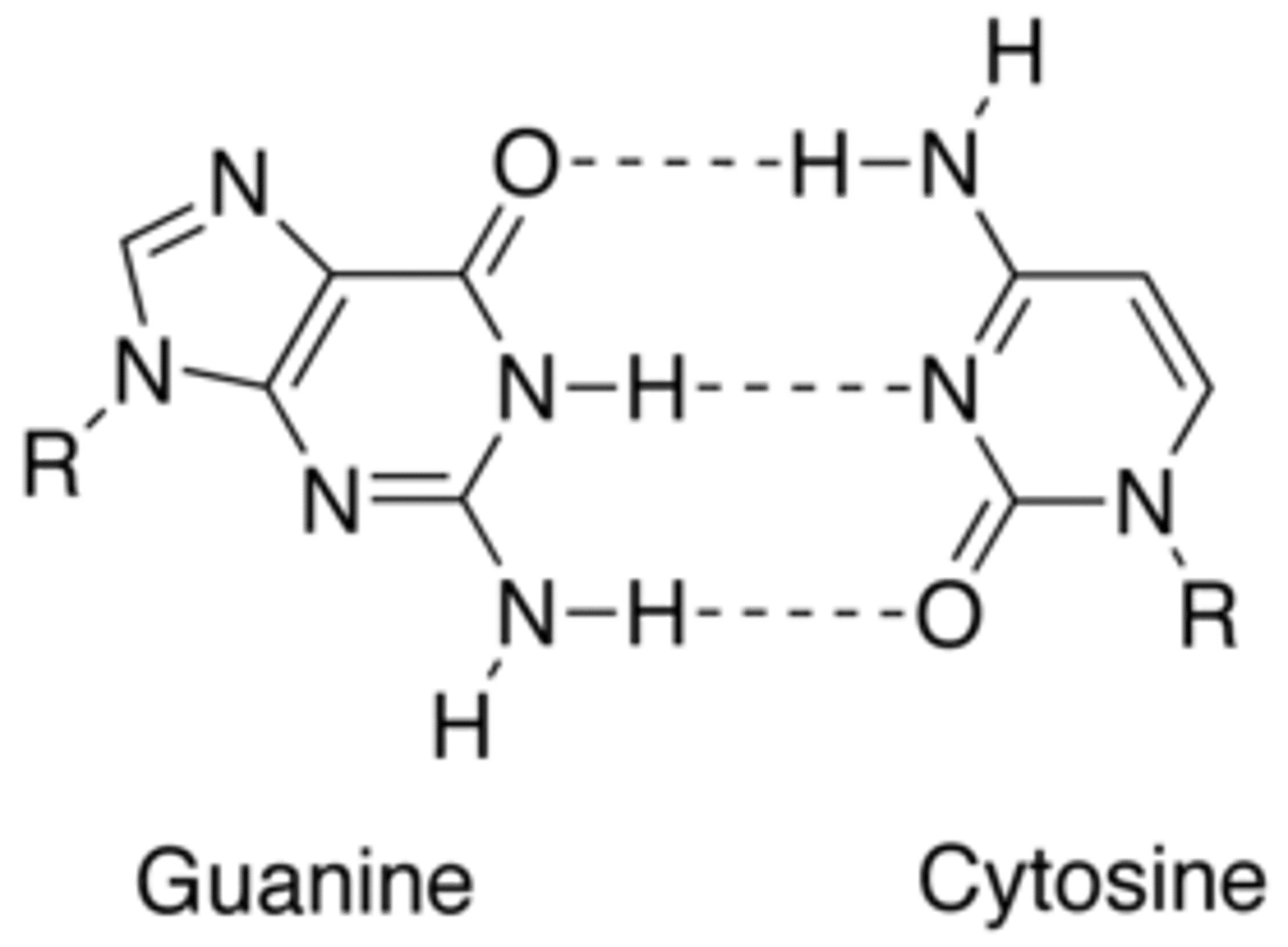
What nitrogen base pairs with cytosine in RNA?
Guanine
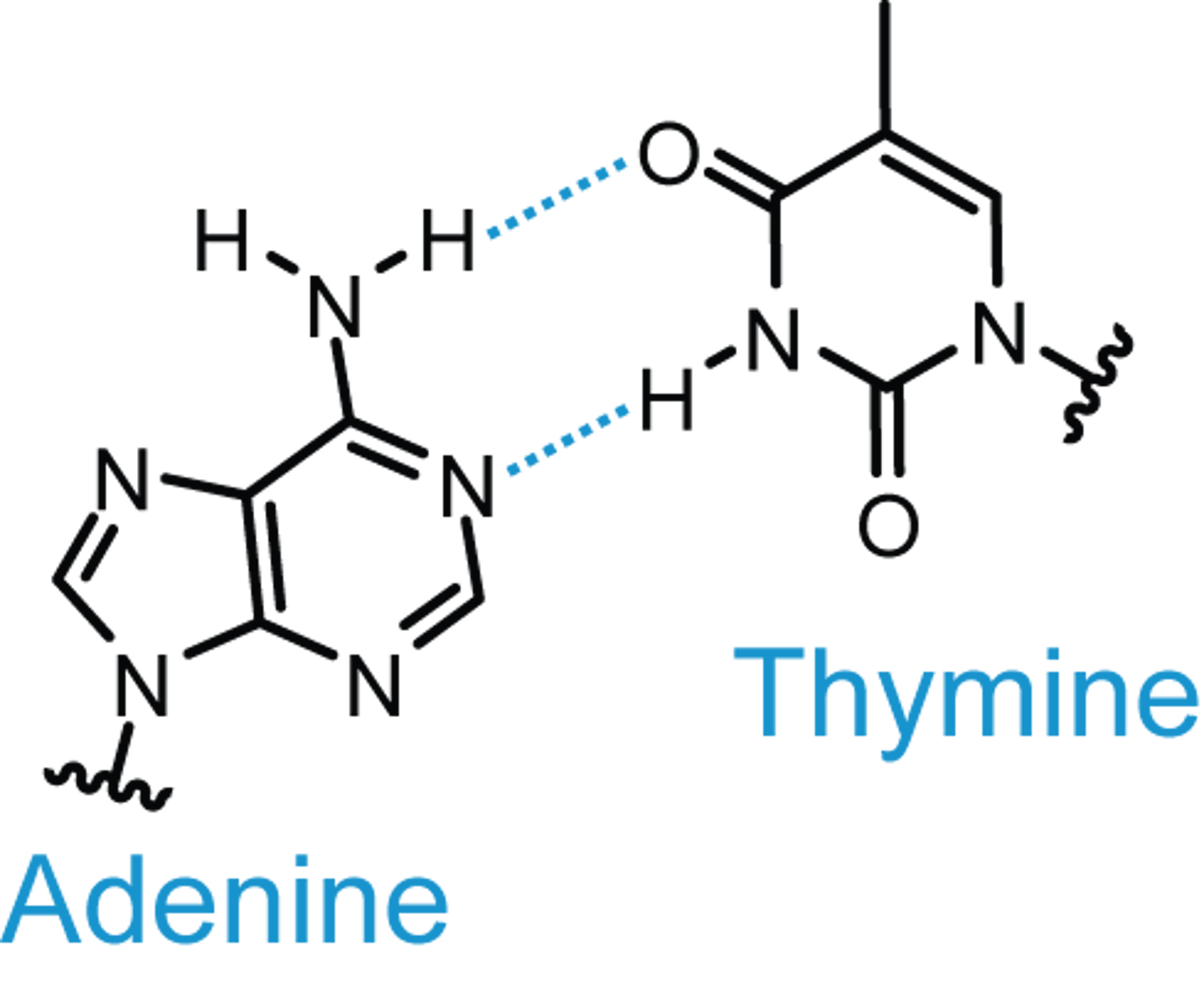
How many hydrogen bonds bind an adenine from one strand of DNA to a thymine on another DNA strand?
Two
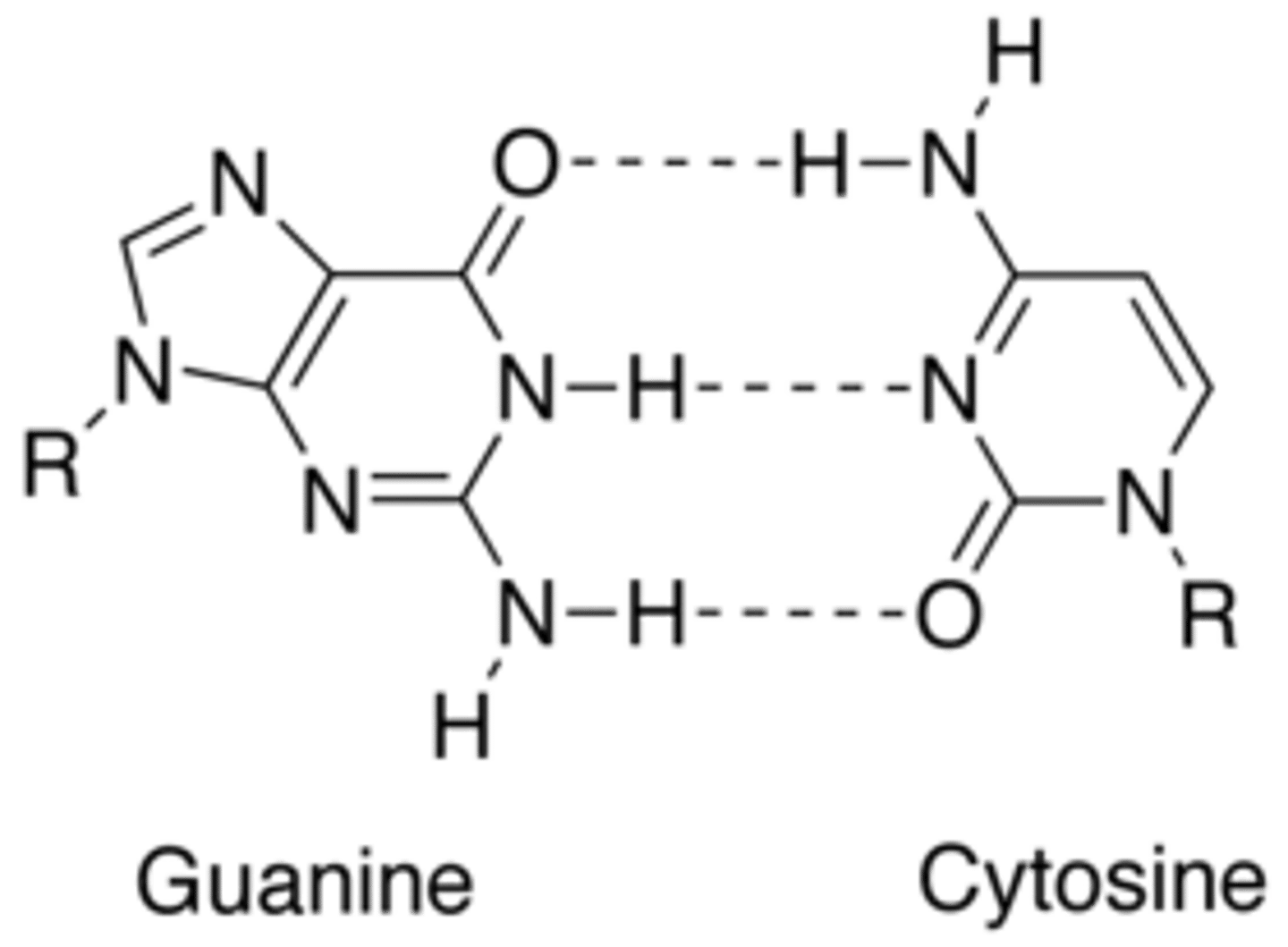
How many hydrogen bonds bind a cytosine from one strand of DNA to a guanine on another DNA strand?
Three
Functional group is found on the 5’ carbon of nucleotides.
Phosphate group or -PO3 group
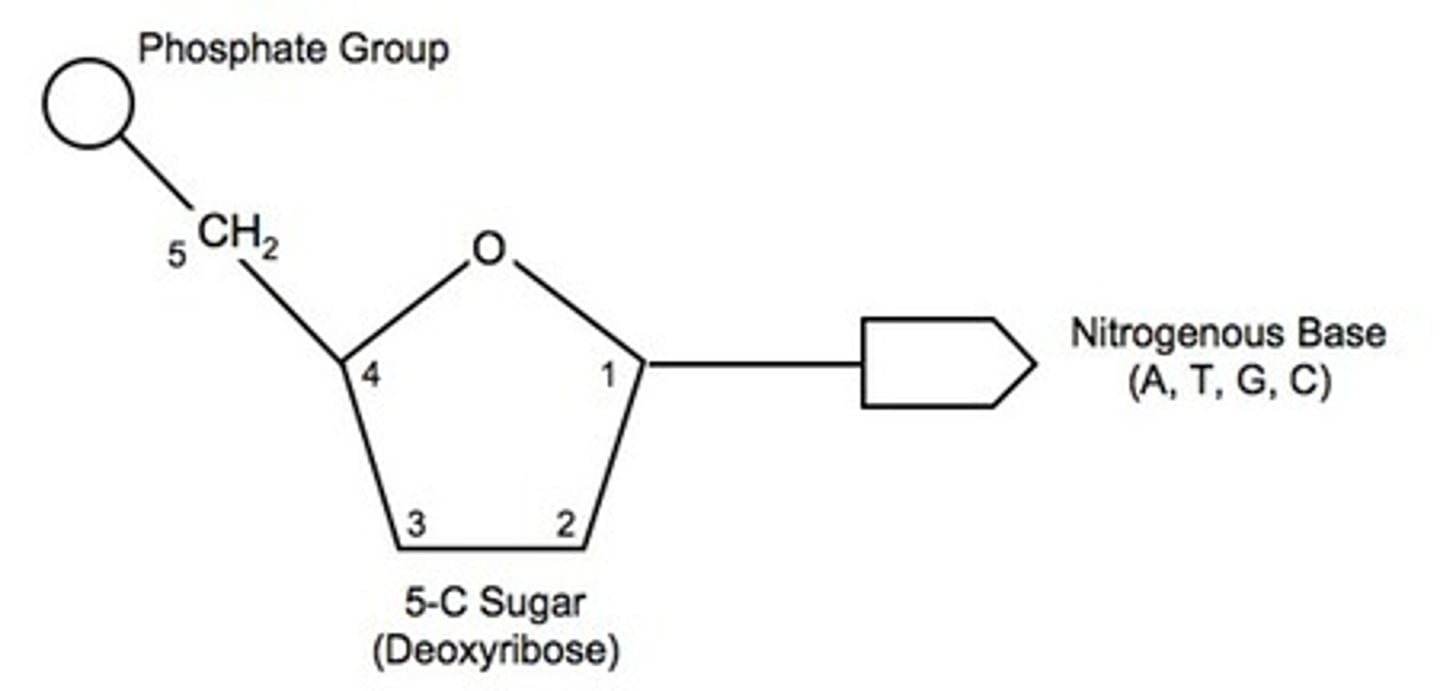
Functional group is found on the 3' carbon of nucleotides.
Hydroxyl group or -OH group
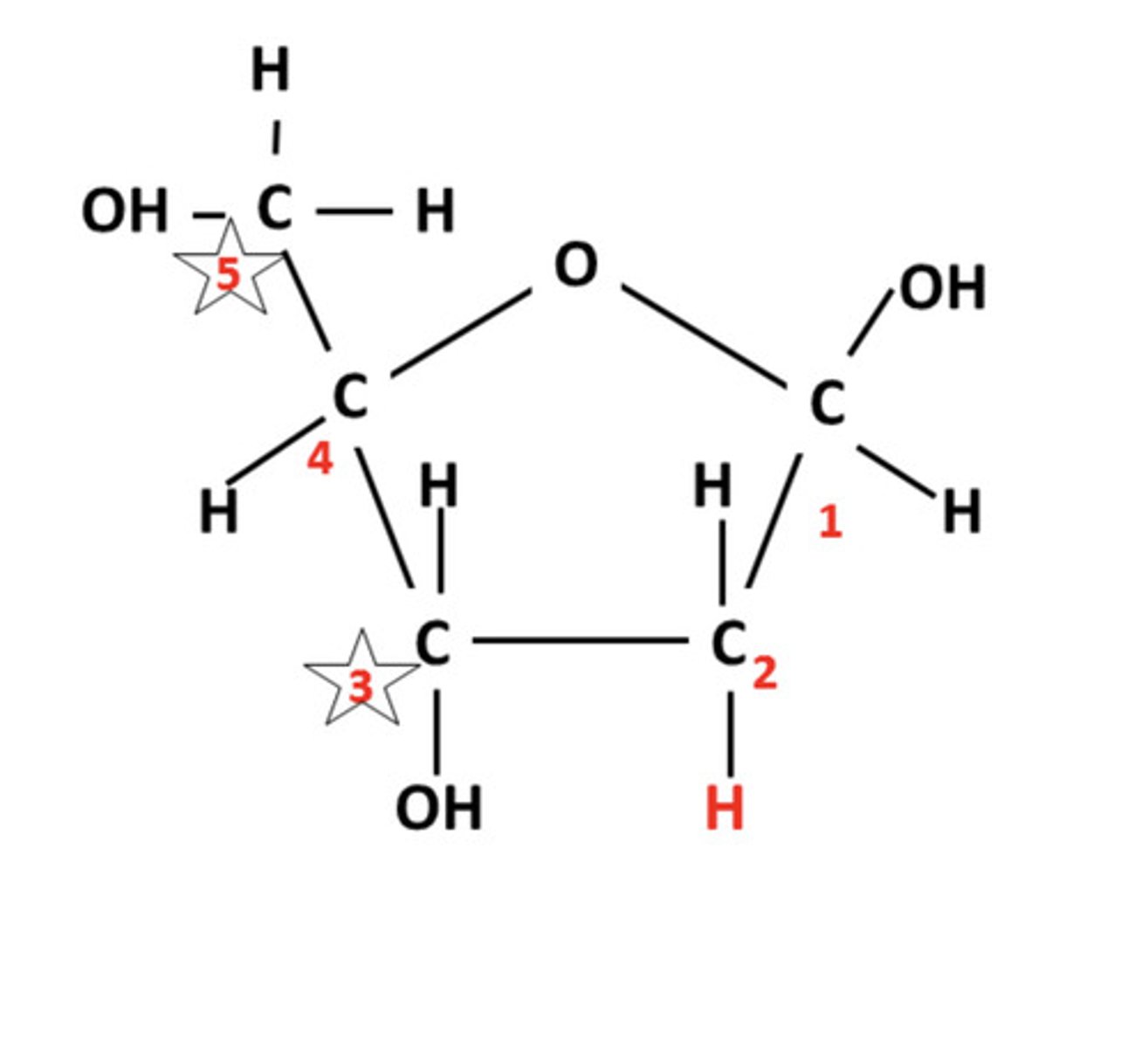
Do DNA strands run parallel or antiparallel (do they go in the same direction or opposite direction)?
Antiparallel
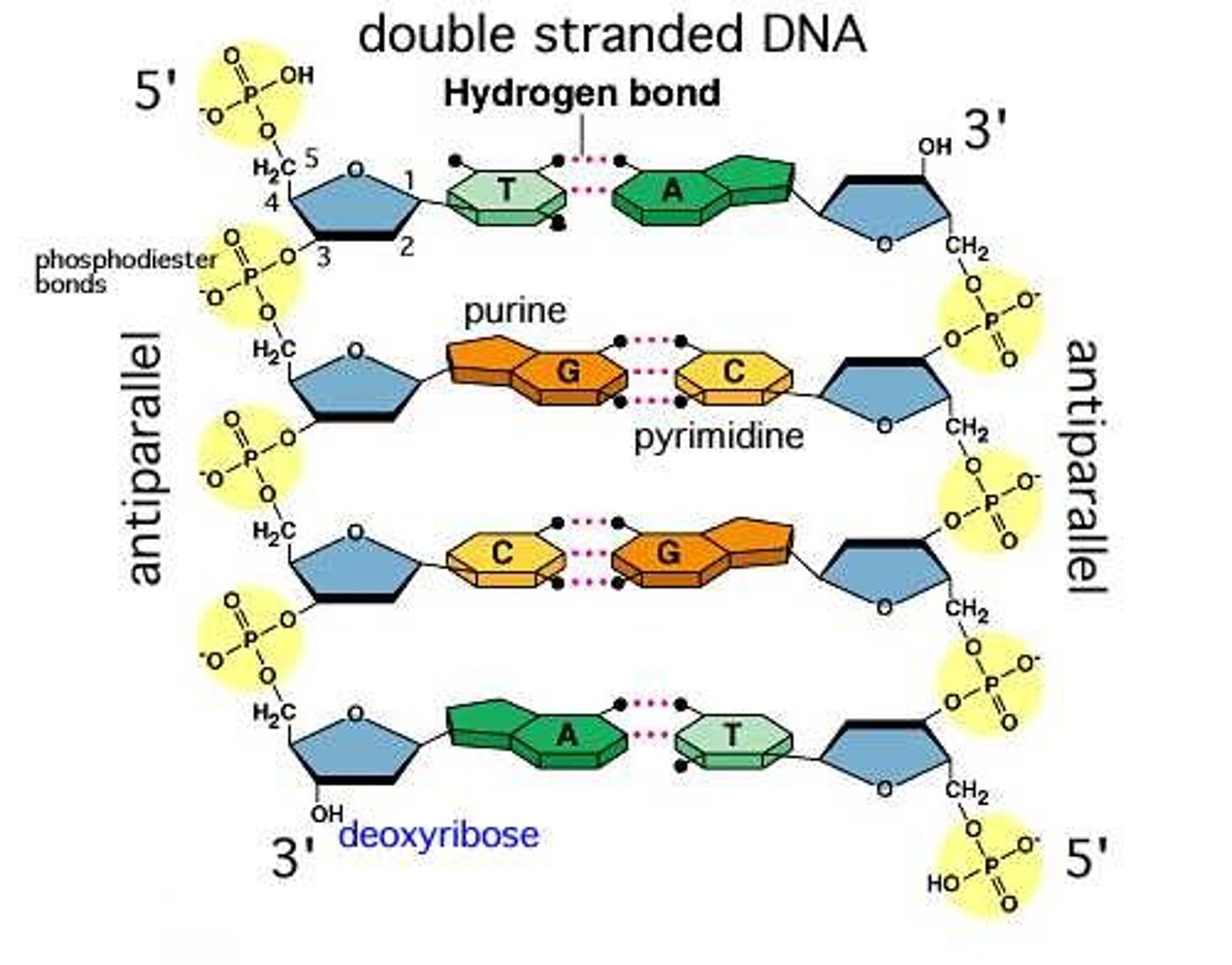
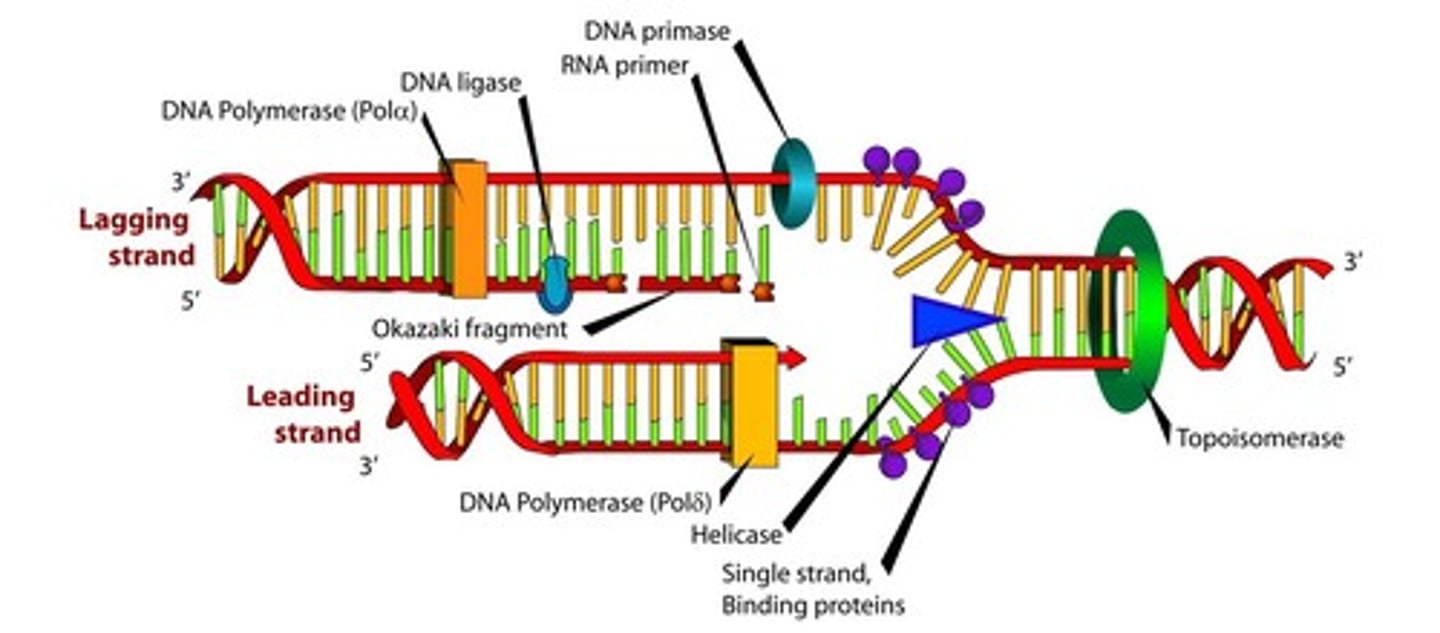
Can both DNA strands be used as a template to make new DNA?
Yes both strands can be used as templates
Although there are only four types of nucleotide bases, there seems to be an unlimited number of genes. Why is this the case?
Nucleotides can be arranged in different ways to generate different amino acid sequences.
•Each amino acid for protein encoded by three nucleotides or triplet on DNA
•20 amino acids are represented by at least one triplet (64 codons total)
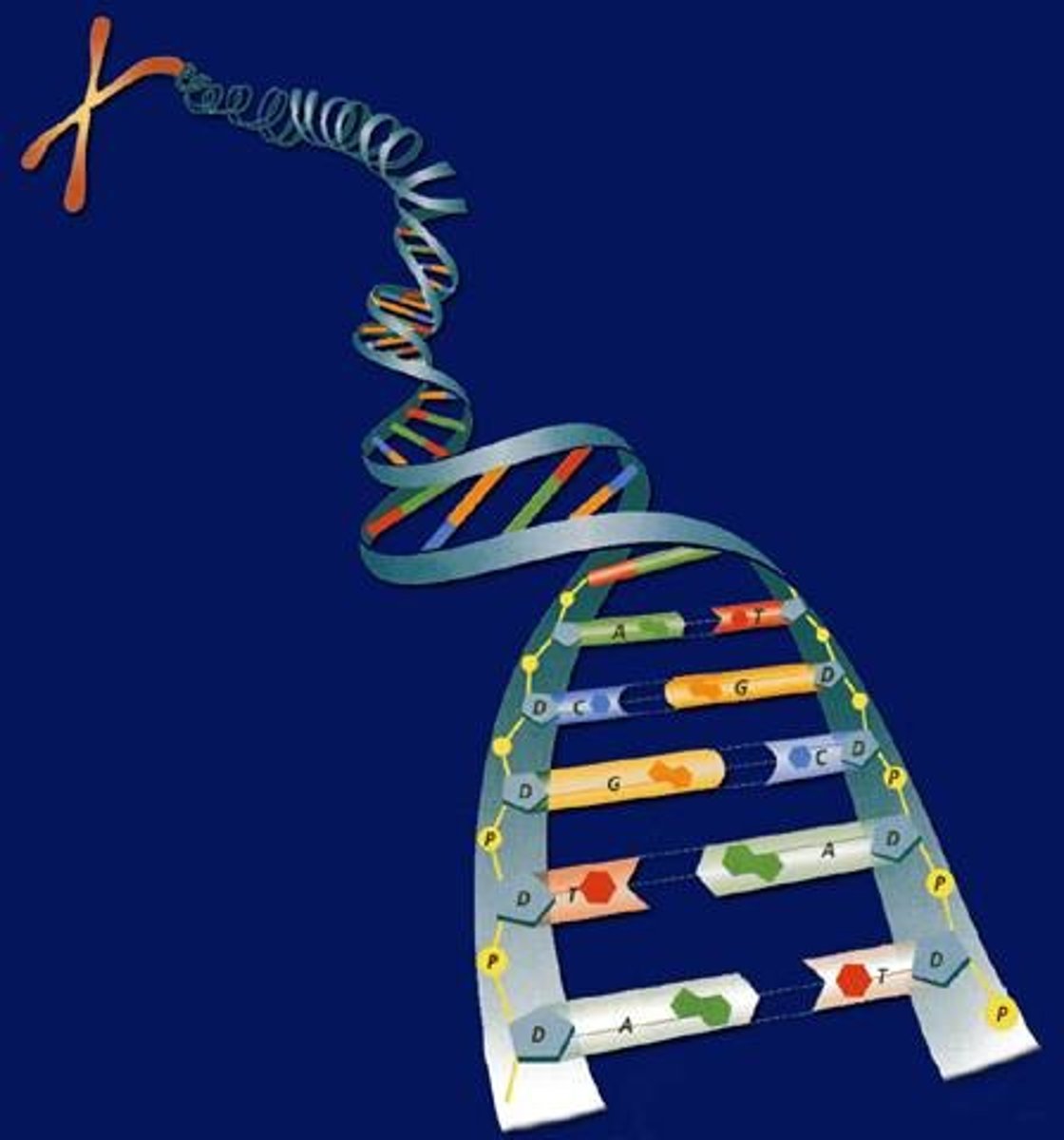
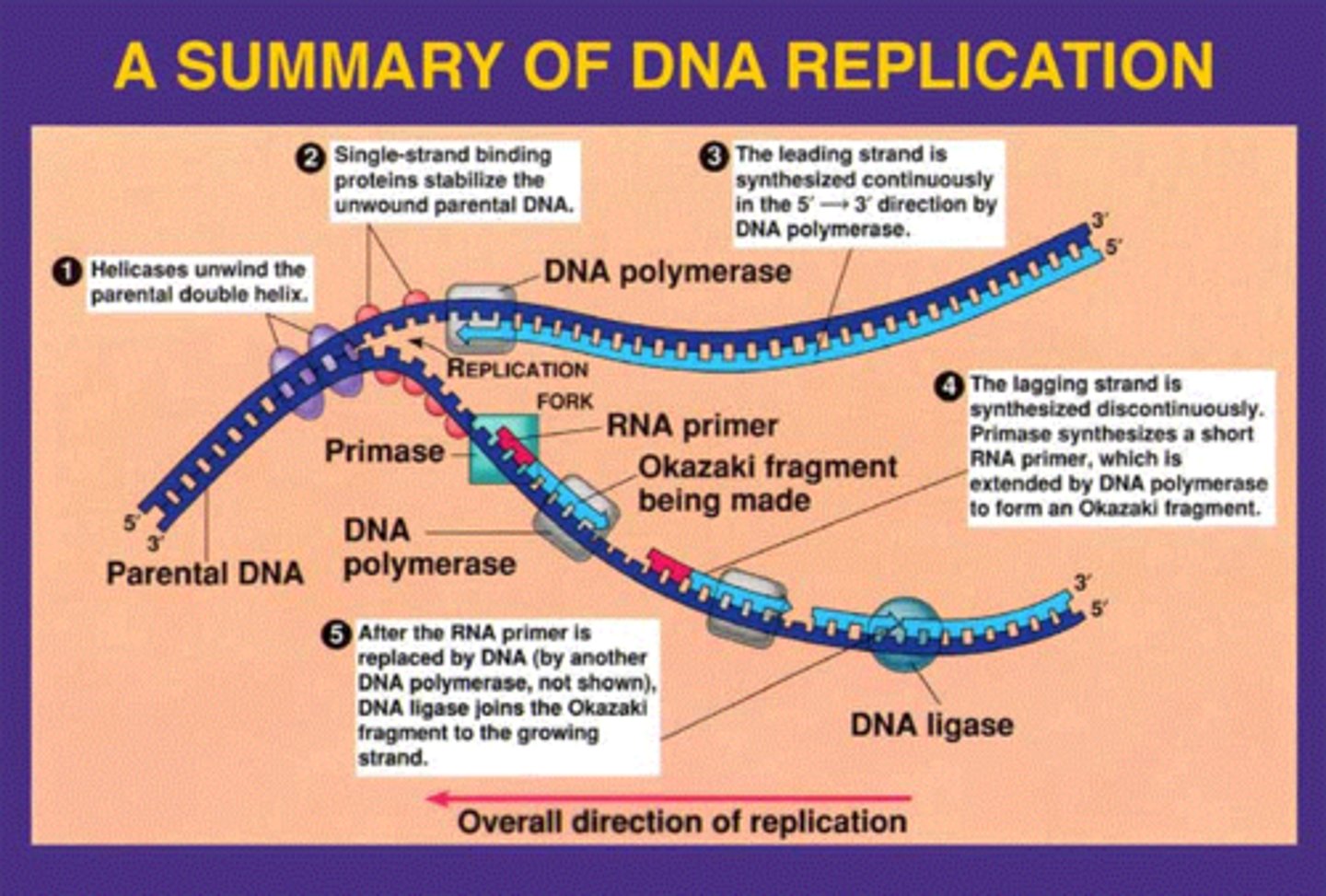
Fundamentals of DNA replication
Helicase opens up the DNA at the A-T rich origin of replication.
Once opened, the SSB proteins hold the DNA open and topoisomerase untwists the DNA as it opens so the DNA does not break.
Primase was add RNA nucleotide primers to the DNA template strands to start replication.
DNA polymerase III will bind the primers and start adding DNA nucleotides on the 3' end of the newly formed DNA strand. The strand that is made away from the fork is known as the lagging strand and the strand that is made toward the fork is known as the leading strand.
DNA polymerase I will remove the RNA primers and fill in the spaces DNA nucleotides.
DNA ligase will seal all the fragments together.
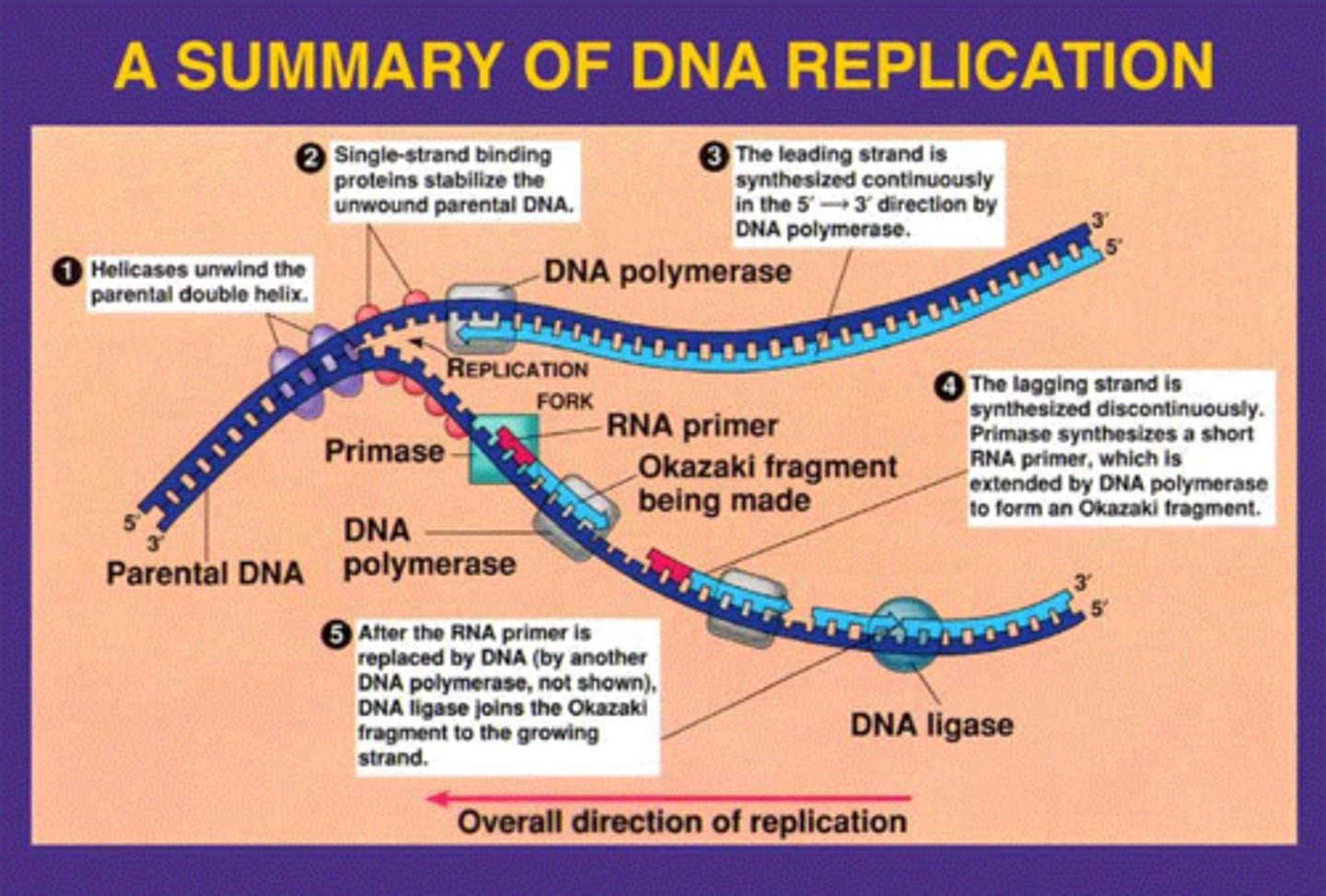
Why are origins of replication made with many A-T pairs instead of G-C pairs?
A-T only has 2 hydrogen bonds, so it's much easier to open
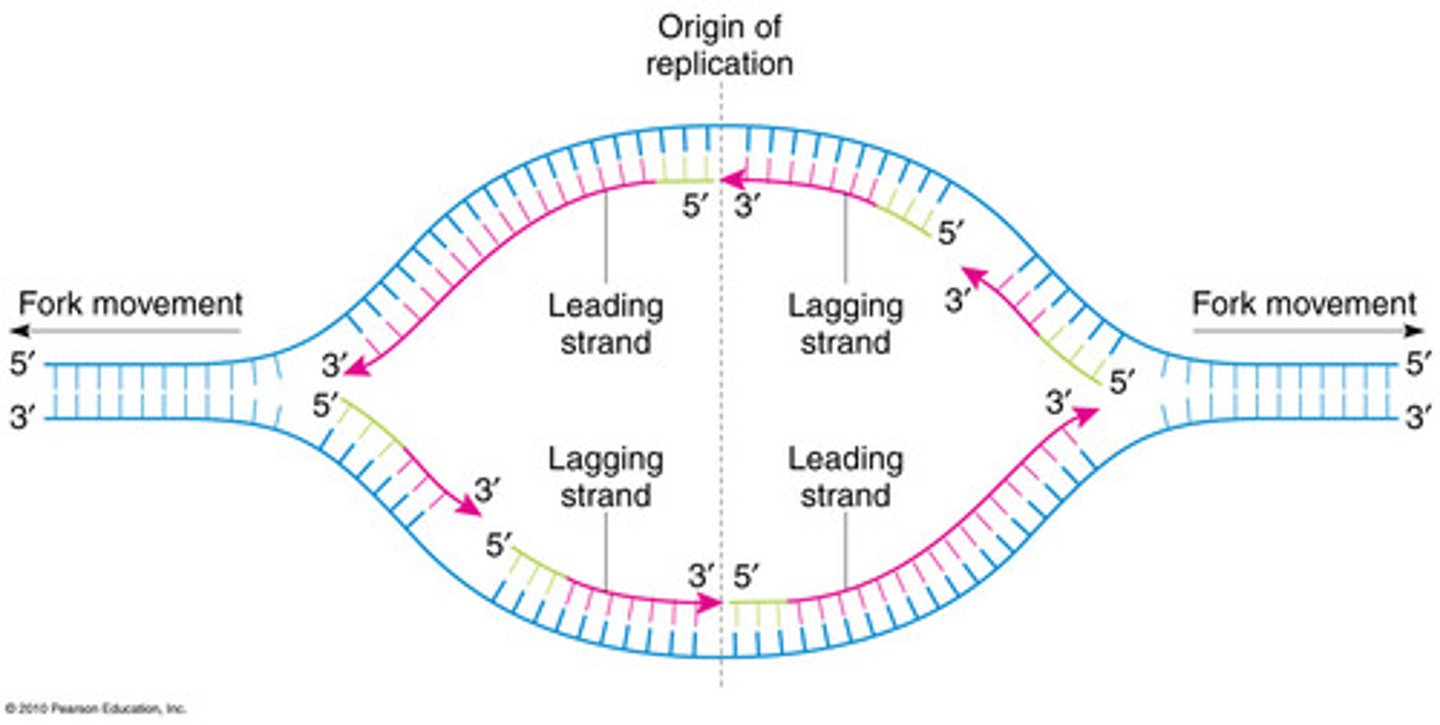
Why does a RNA primer need to be added?
DNA polyamerase III needs 3' end (-OH) to add nucleotides
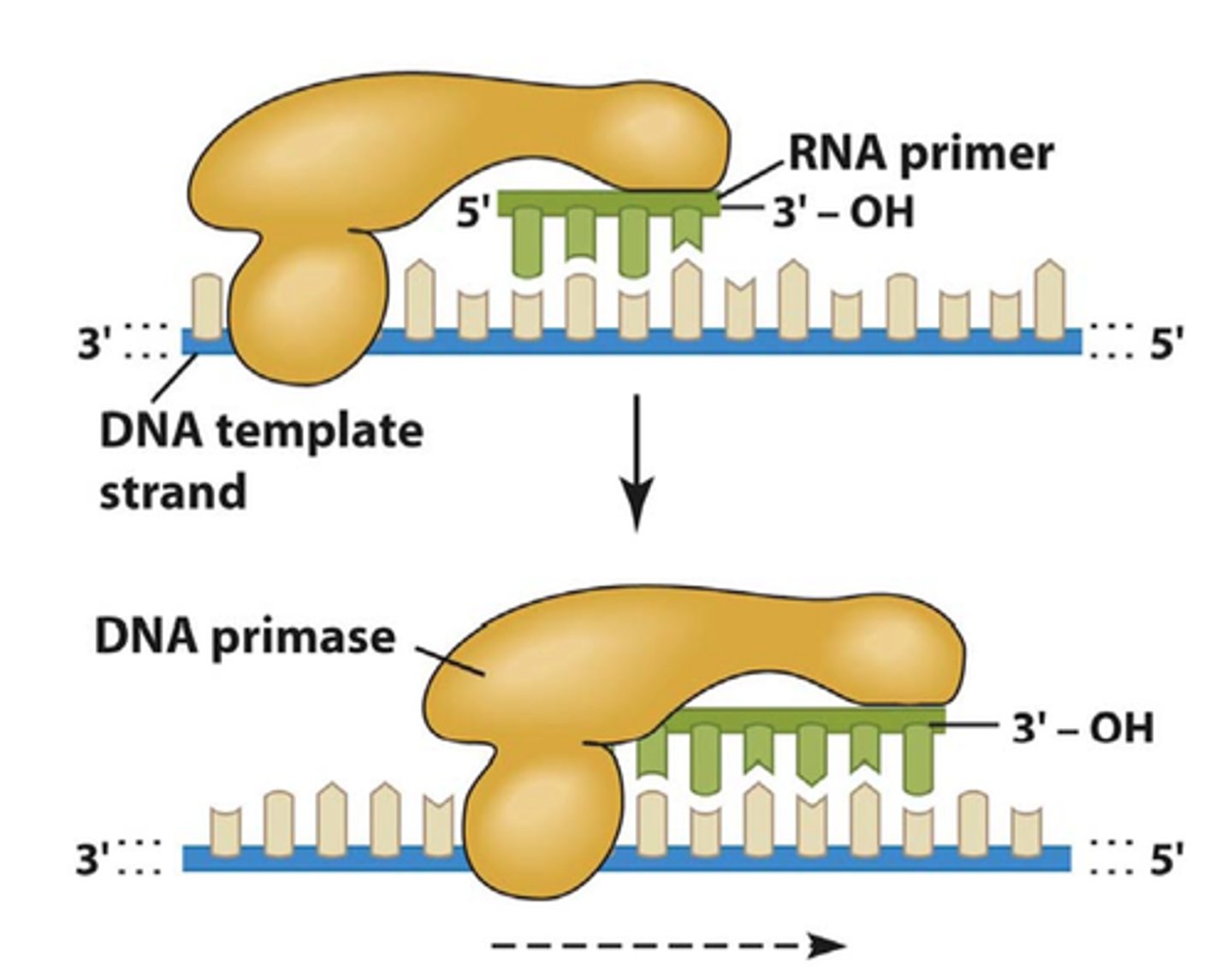
Does DNA polymerase III add nucleotides in a 5' to 3' direction, 3' to 5' direction or both?
DNA polymerase III adds nucleotides in a 5′ to 3′ direction
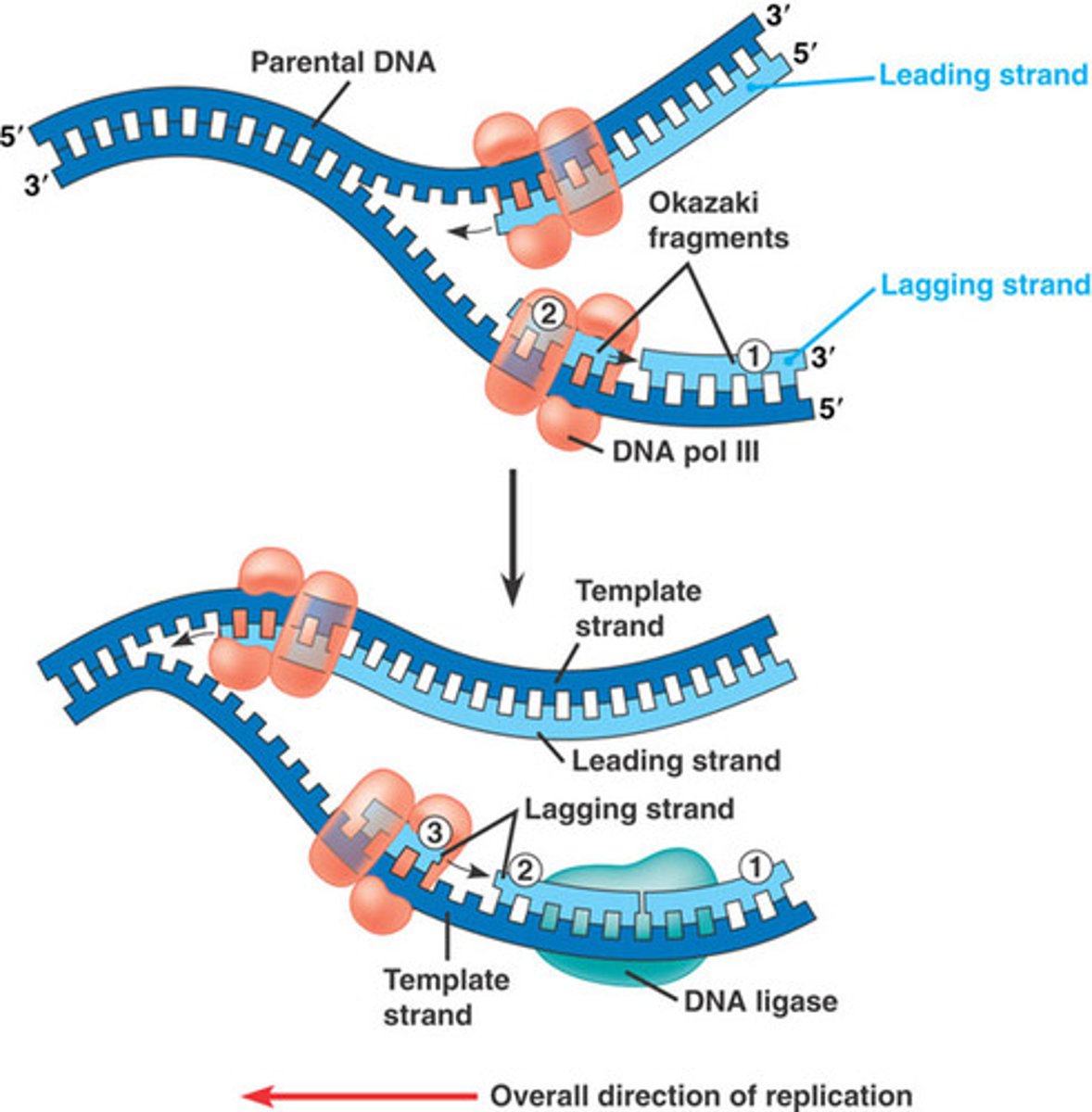
During DNA replication, do both DNA strands have Okazaki fragments, only the leading strand, only the lagging strand?
Only lagging strand has Okazaki fragments
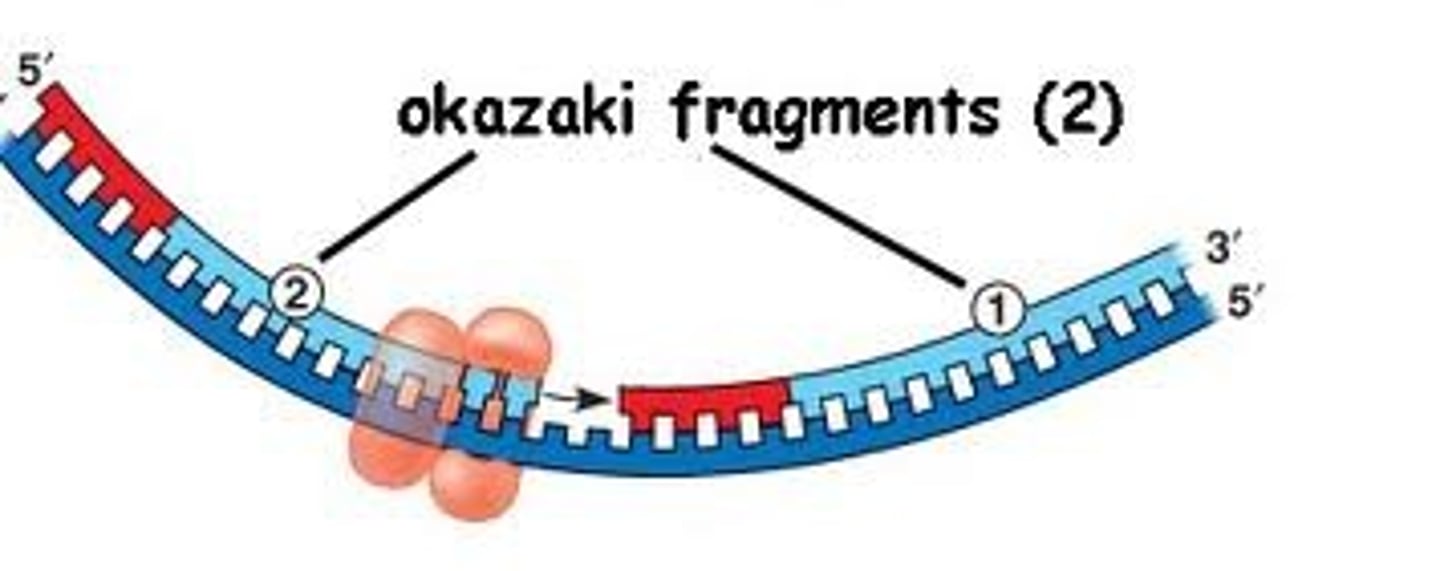
Why do the leading and lagging strands form during replication?
The leading or continuous strand forms since the 3' end of the newly form strand where nucleotides are added is made toward the fork.
The lagging or discontinuous strand forms since the 3' end of the newly form strand is made away from the fork and new primers must be made every time the fork opens in order to add more nucleotides to the new strand. These short fragments are okazaki fragments.
Similarities between prokaryotic and eukaryotic DNA replication
Bi-directional, semiconservative replication
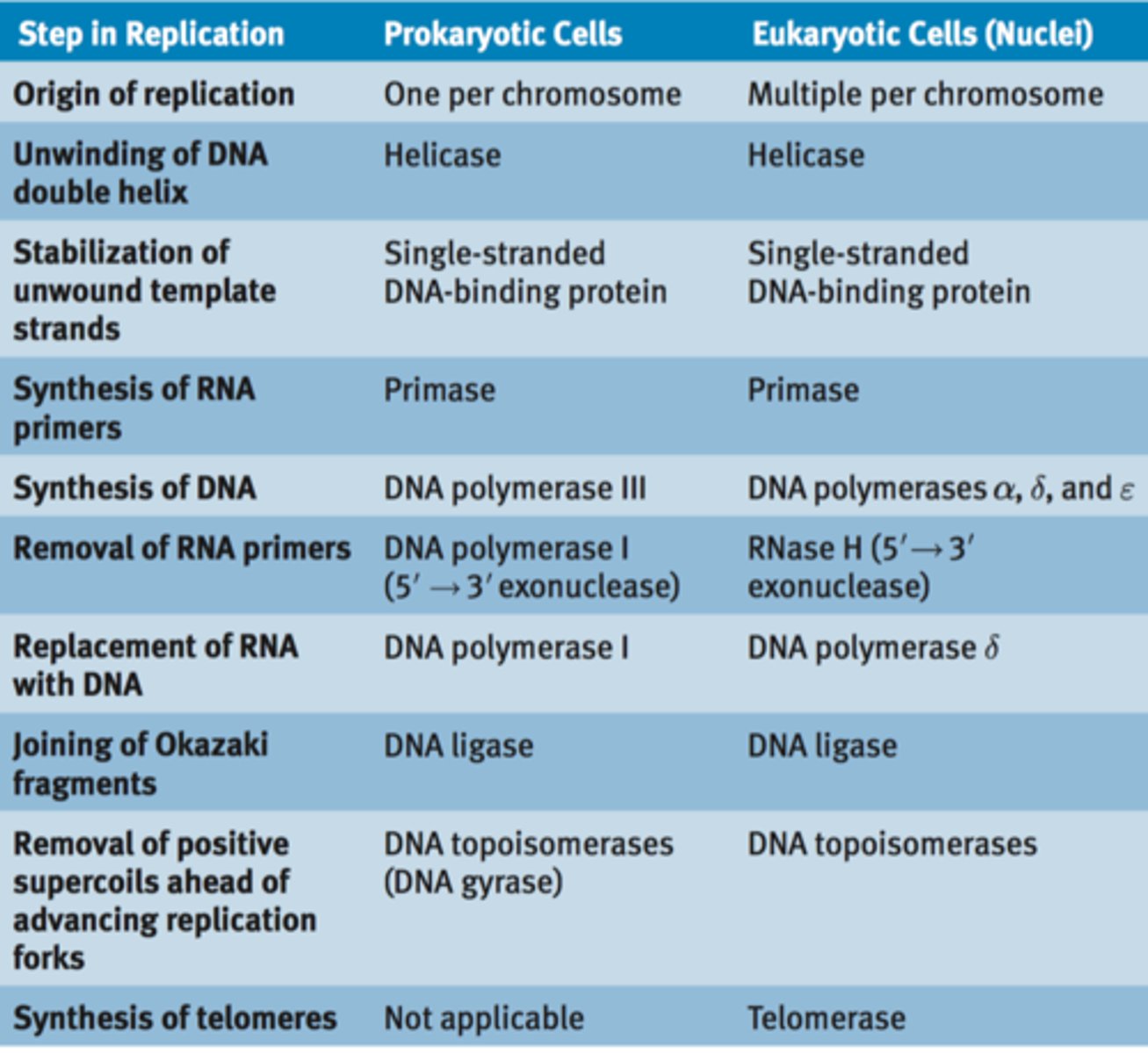
Differences between prokaryotic and eukaryotic DNA replication: Number of chromosomes
Prokaryotic: Have 1 chromosome
Eukaryotic: Have 46 chromosome
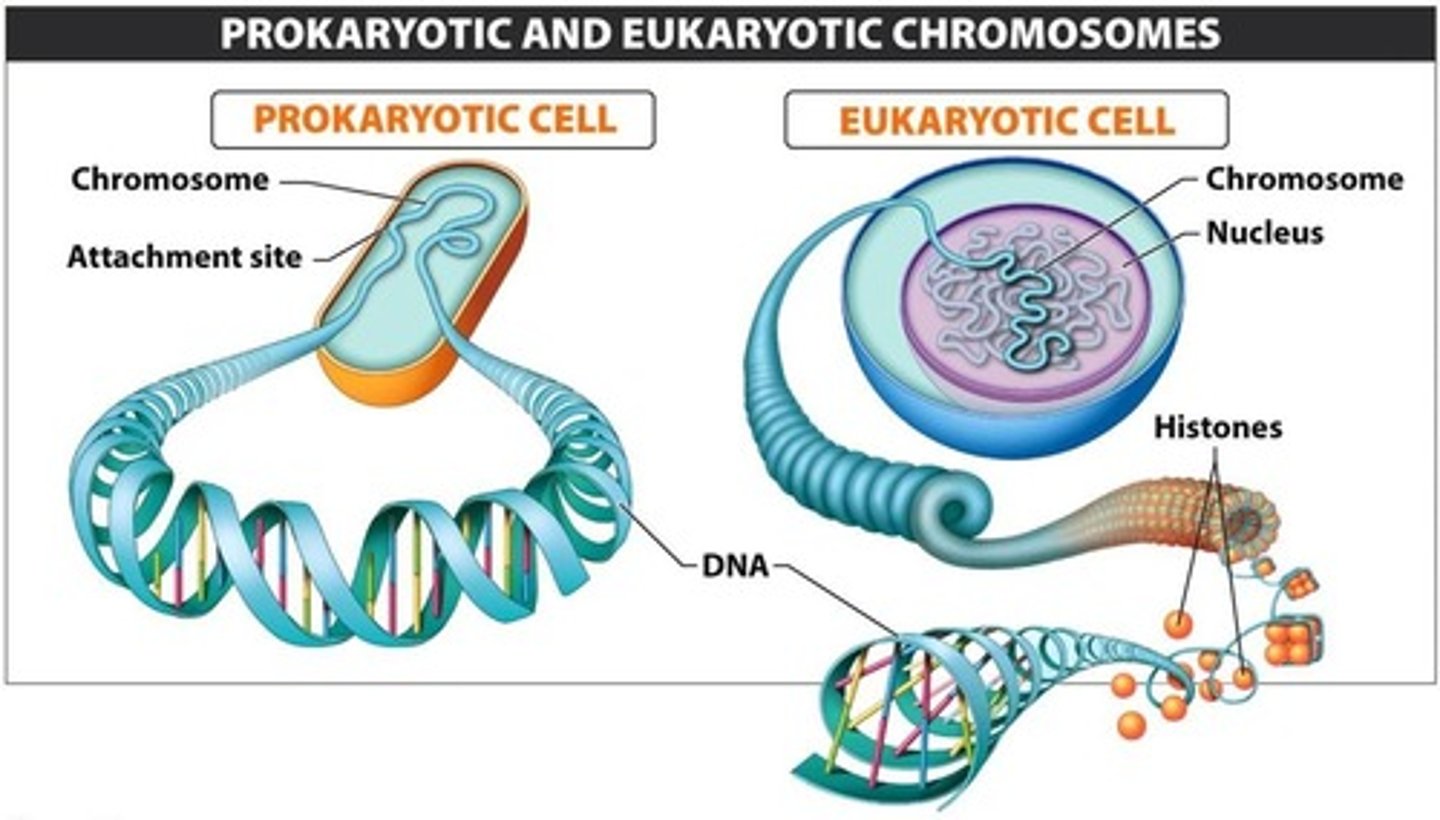
Differences between prokaryotic and eukaryotic DNA replication: Number of origins
Prokaryotic: Single origin
Eukaryotic: Multiple origin
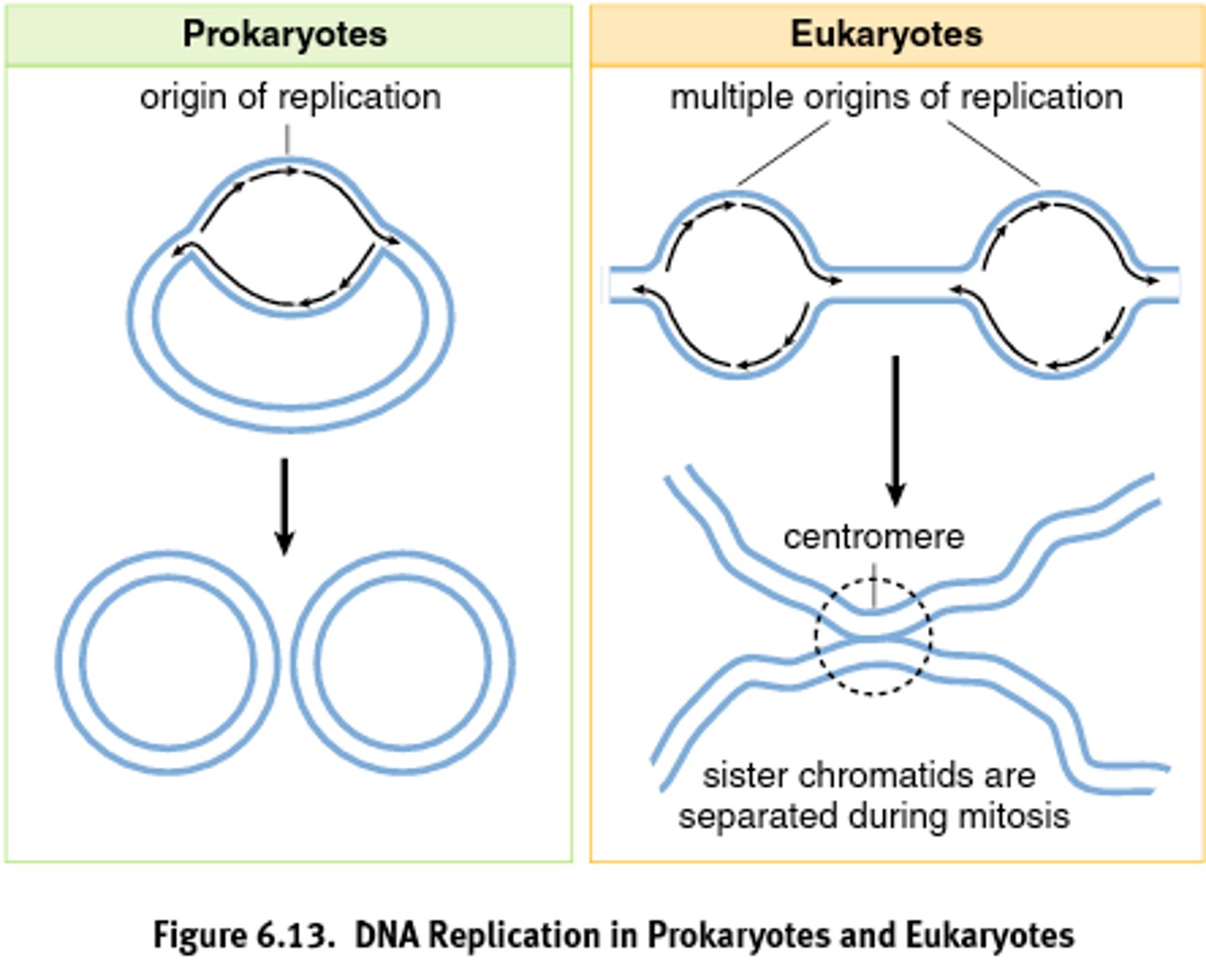
When does DNA replication occur at what stage of the cell cycle?
S phase
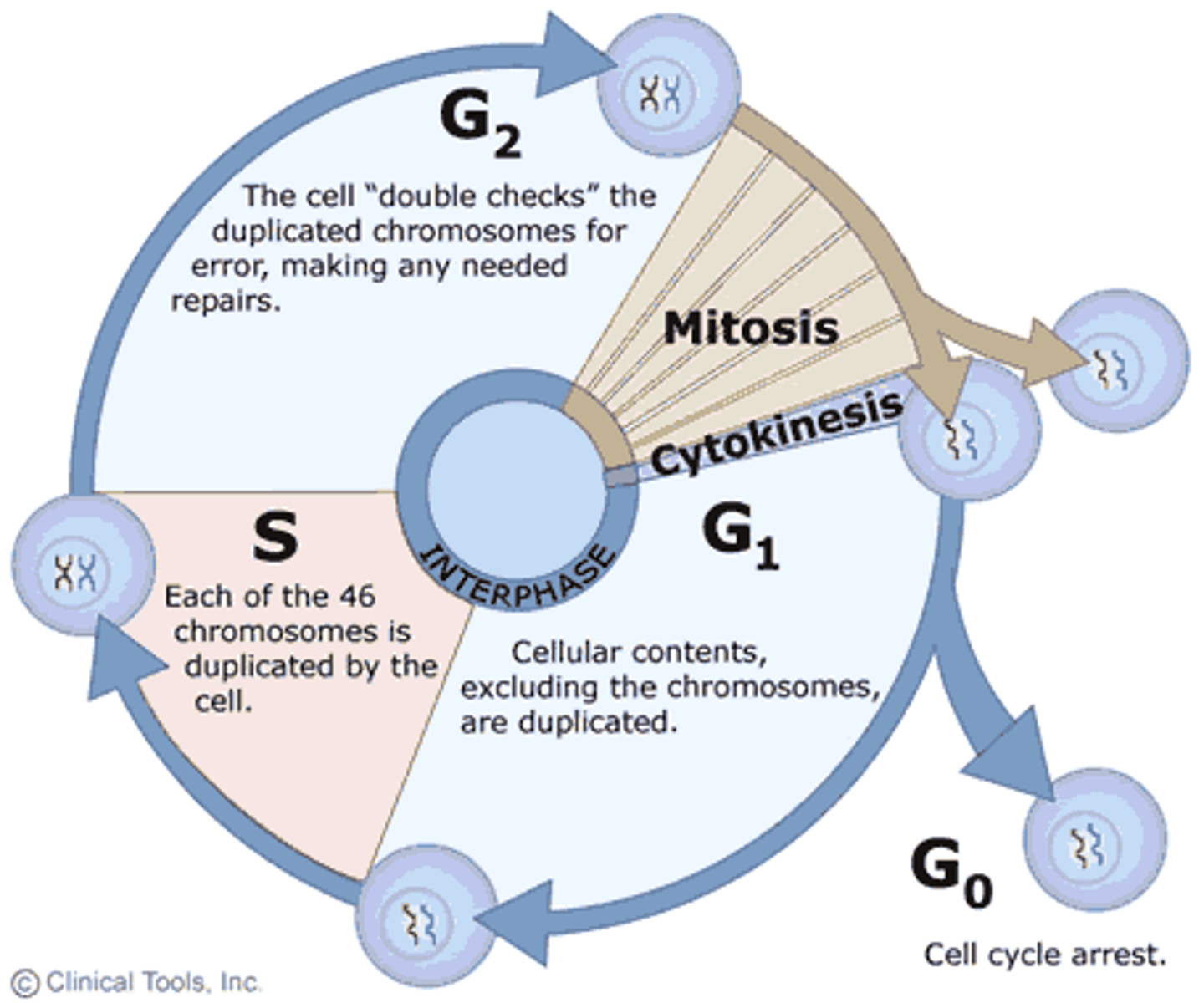
Name the three main processes that involve DNA.
Transcription (with translation), Replication, Recombination
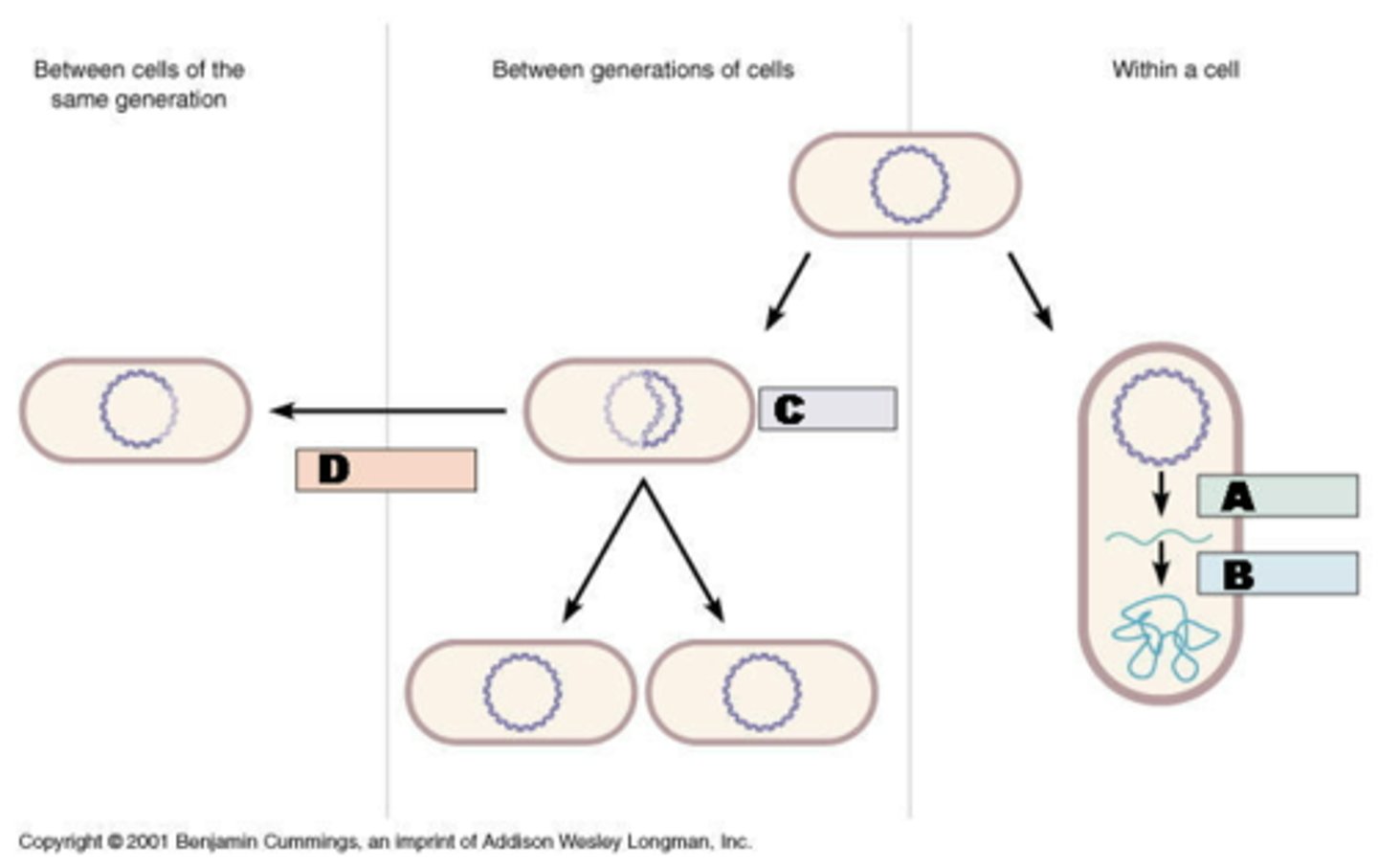
Does transcription always result in a protein being made from the RNA?
no tRNA and rRNA (primers too) do not make proteins directly from their RNA sequence
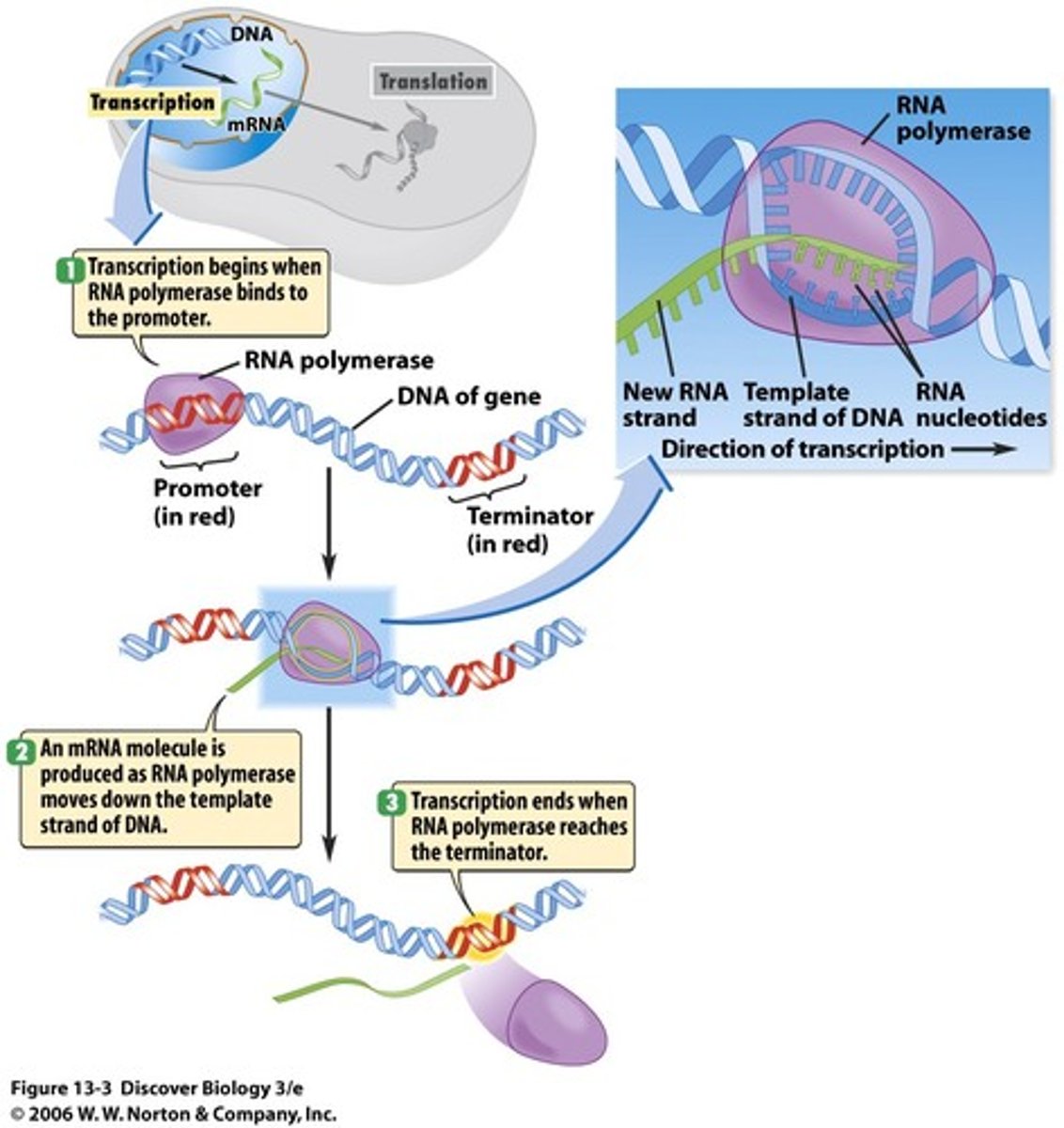
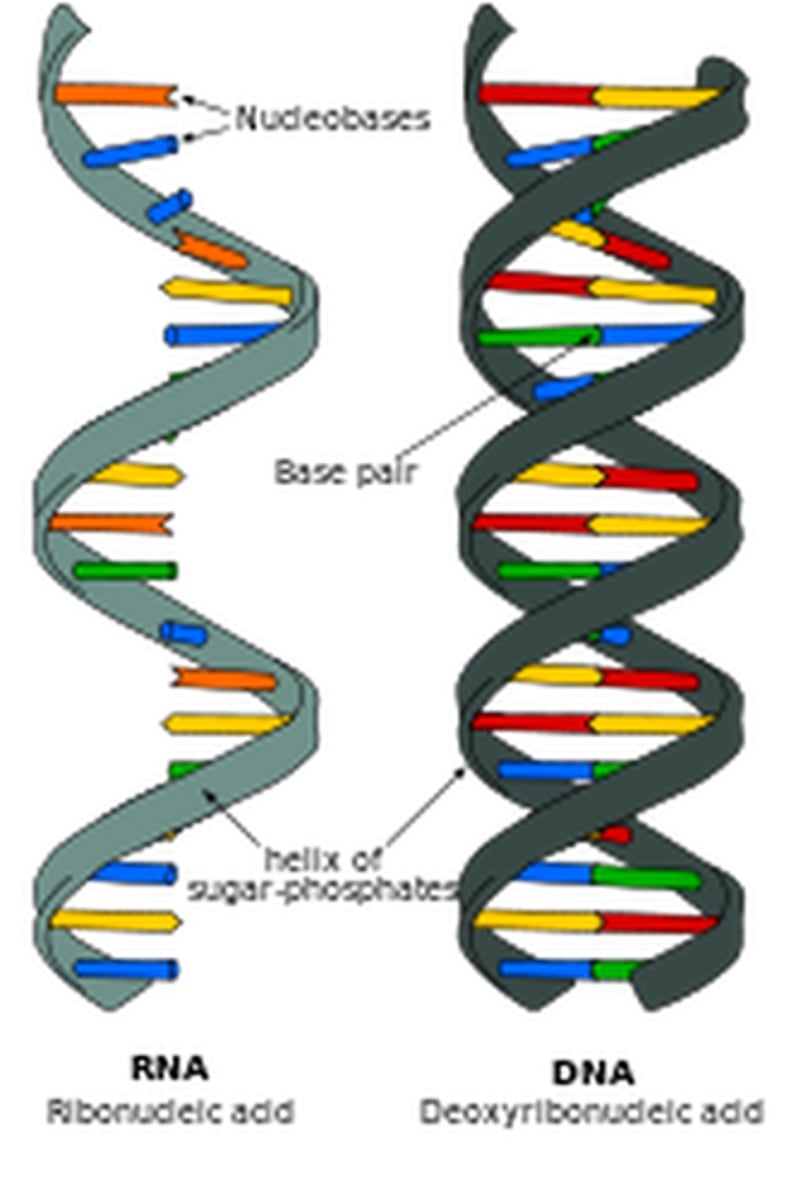
Differences between the structure of DNA & RNA
DNA:
Double stranded
Sugar: Deoxyribose
Bases: A,T,C,G
RNA:
Single stranded
Sugar: Ribose.
Bases: A,U,C,G
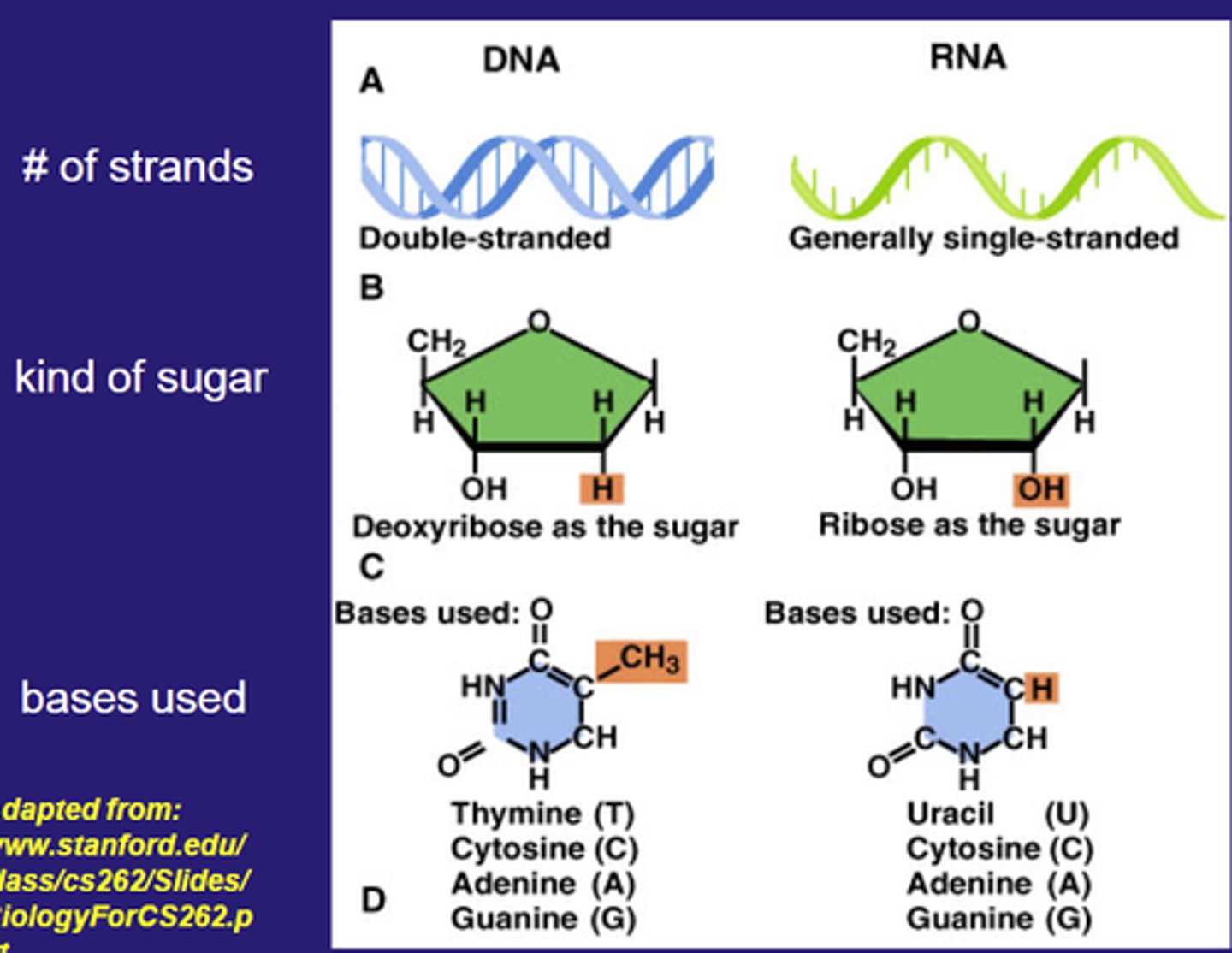
Three components of a RNA nucleotide
5 carbon sugar: Ribose
Nitrogen bases: Adenine, uracil, guanine, cytosine
Phosphate group
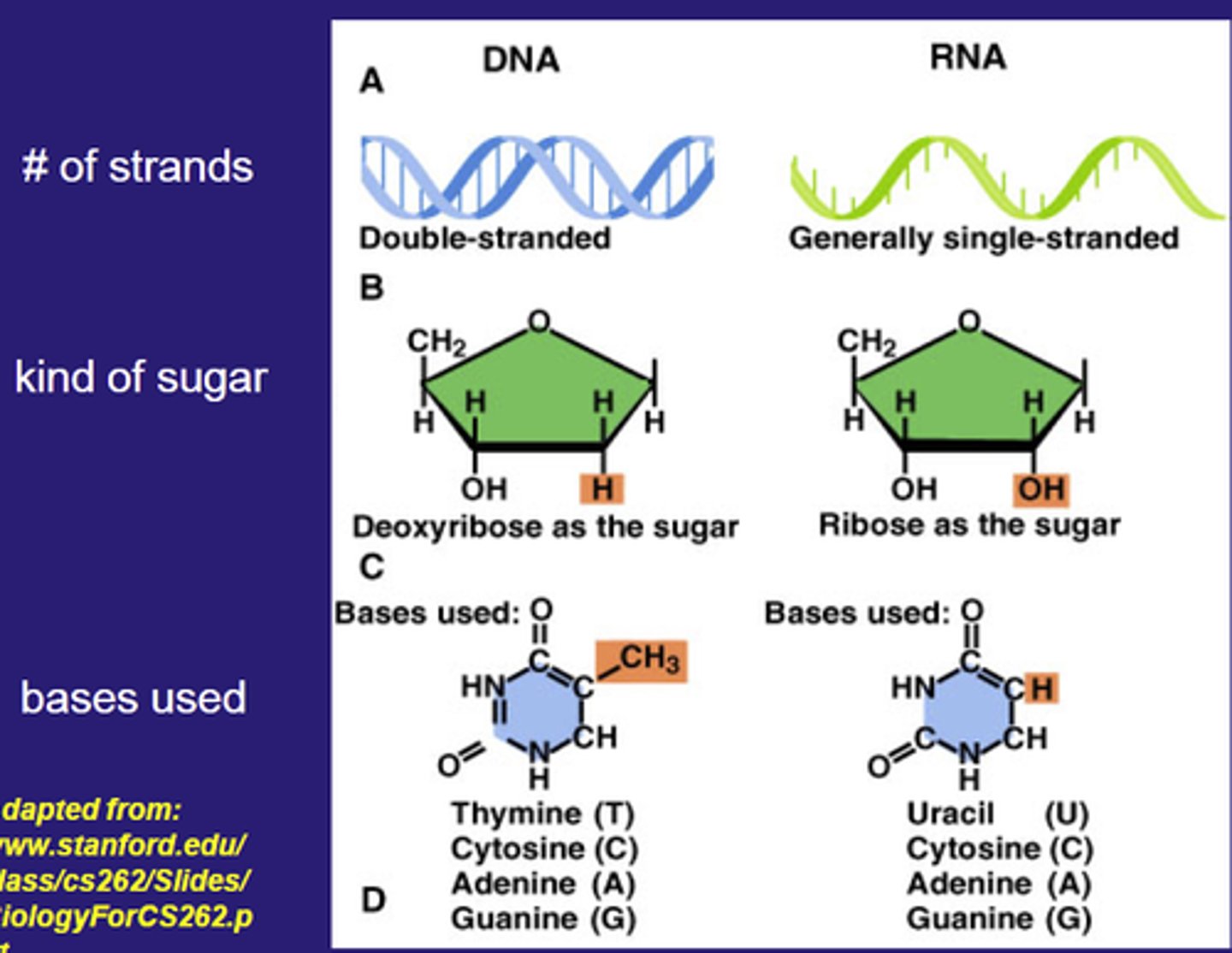
What does a codon stand for?
A codon is a three nucleotides or triplet that represents one amino acid or tells the ribosome to start (start codon) or stop (stop codons) translation
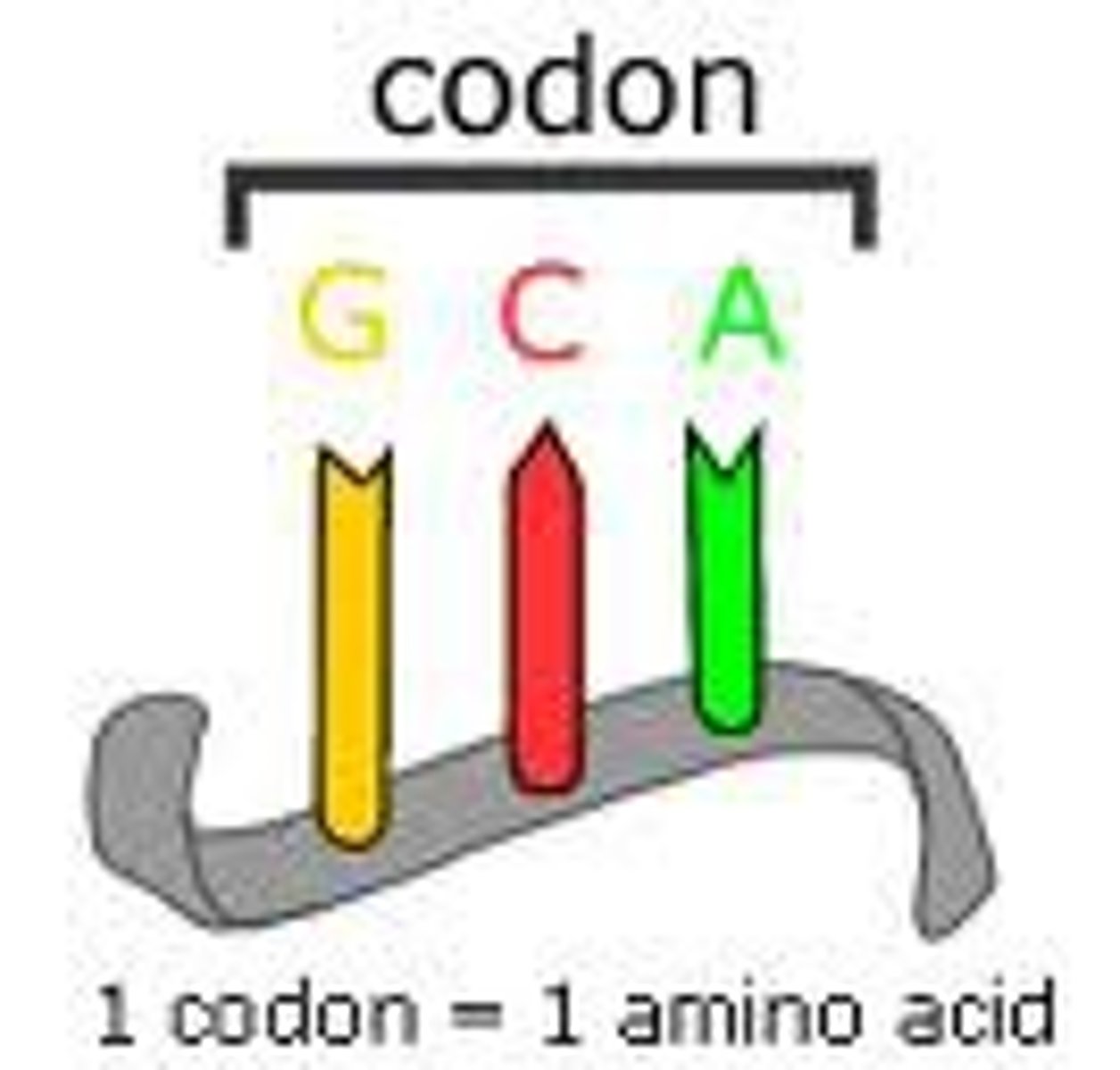
Is every amino acid represented by only one codon?
No, most amino acids are represented by multiple codons, 64 codons for 20 amino acids.
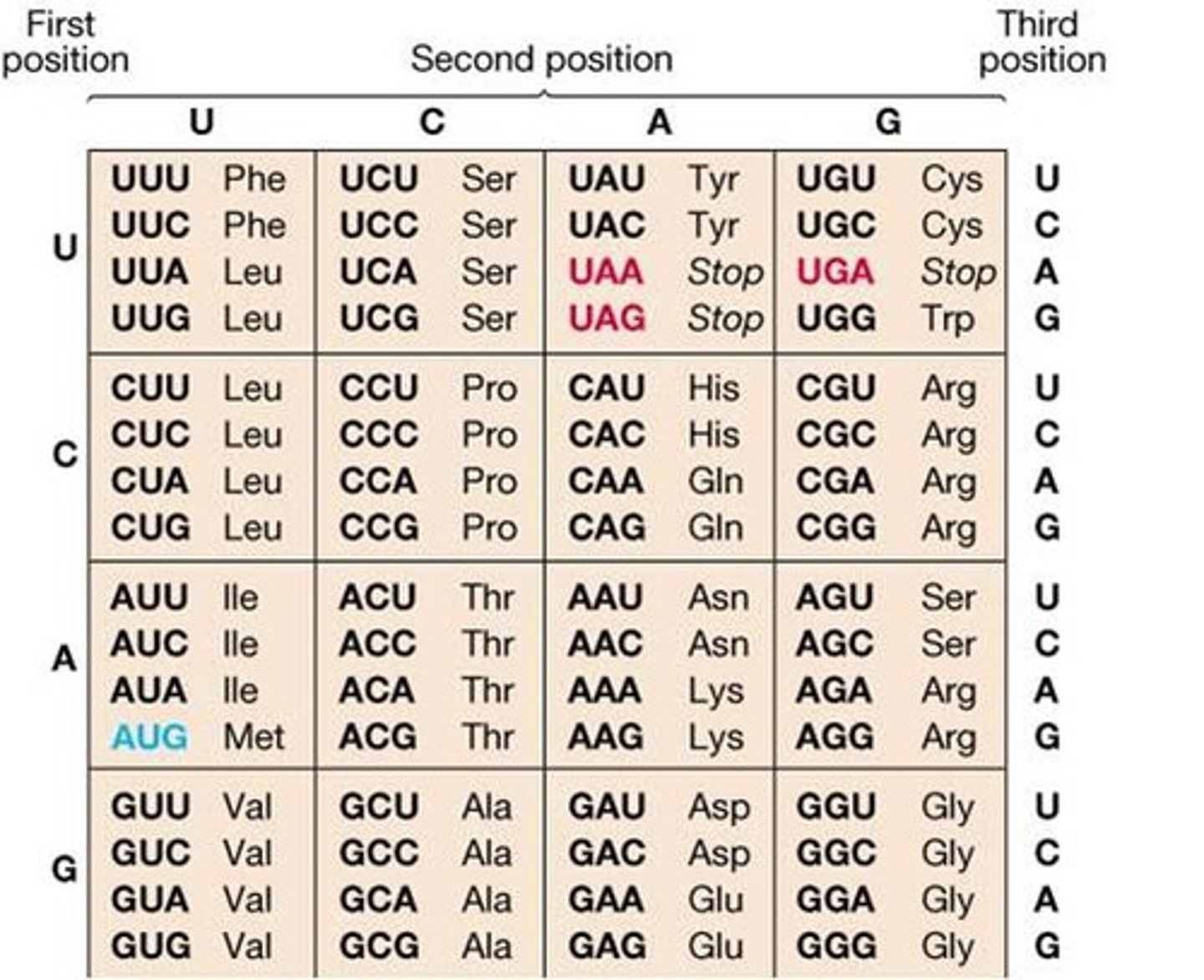
Why would it be bad if most amino acids were only represented by one codon?
This is needed in case of mutation. The more codons that represent each amino acid, the increase the chance that the mutation will not change which amino acid is added
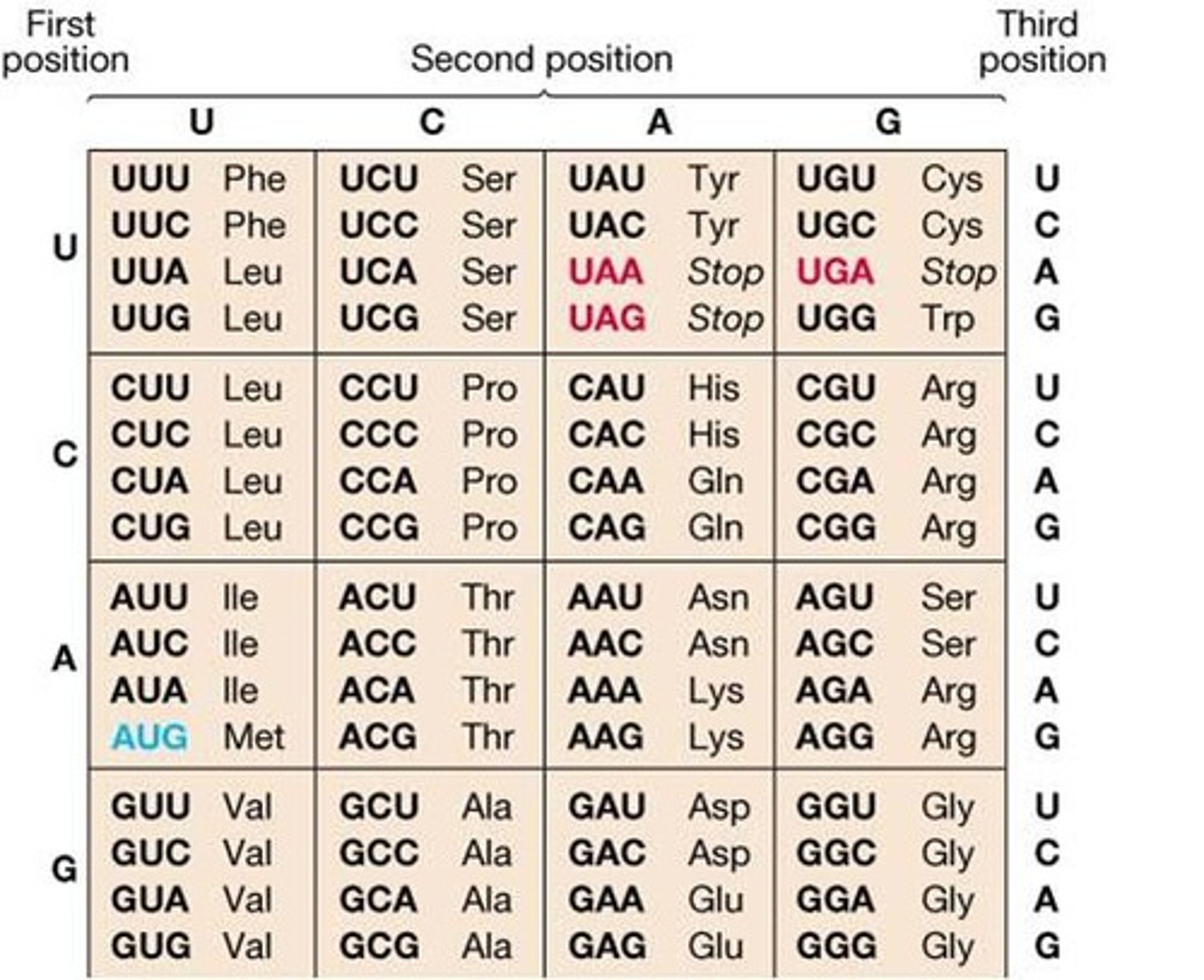
Three types of RNA
Messenger RNA (mRNA), Transfer RNA (tRNA), Ribosomal RNA (rRNA)
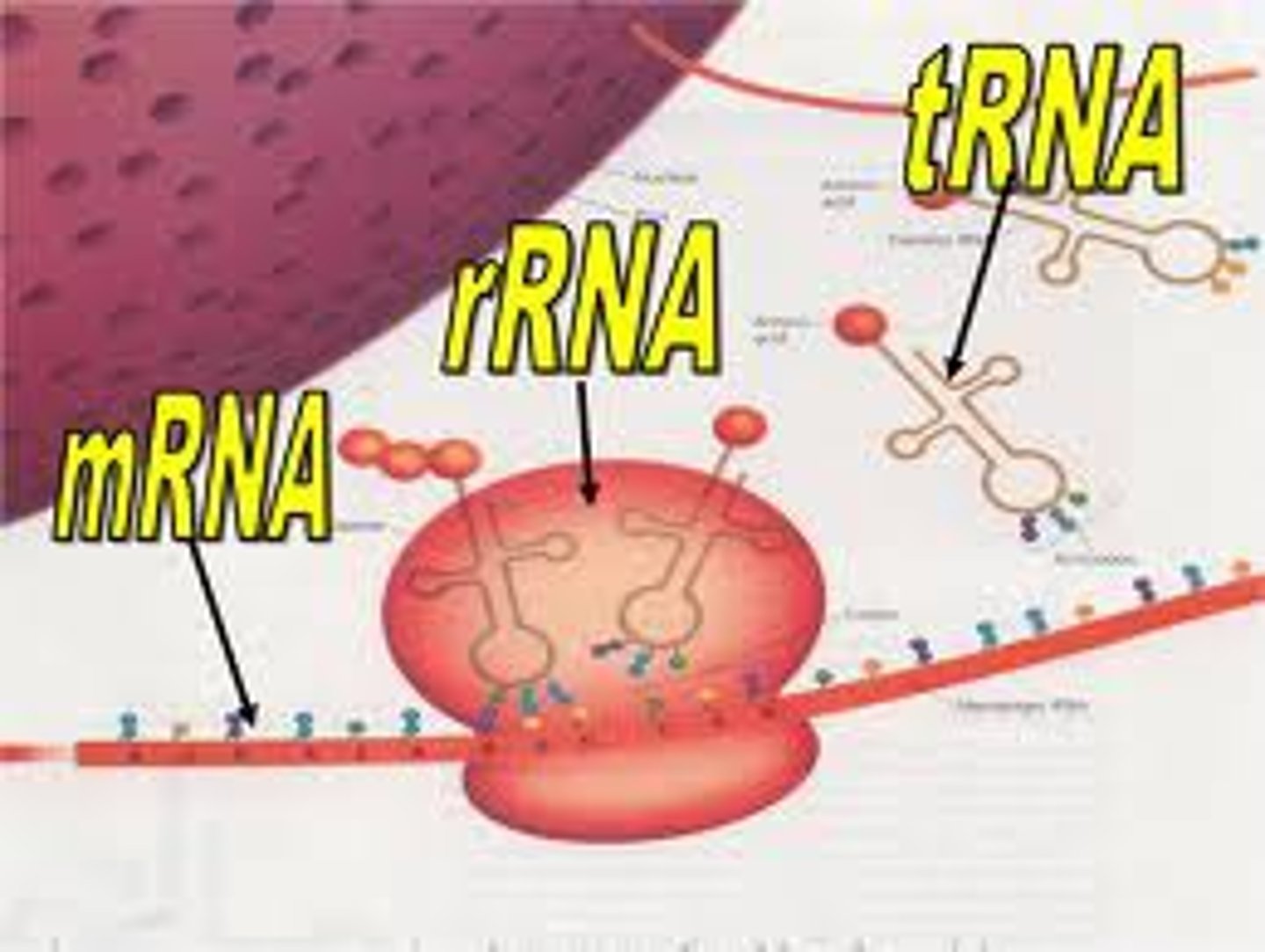
mRNA basics
Copy of structural gene, carries DNA message through complementary copy; translated by ribosomes, message is in triplets called codons
tRNA basics
Made from DNA; bottom loop exposes a triplet of nucleotides called anticodon which designates specificity and complements mRNA; carries specific amino acids to ribosomes
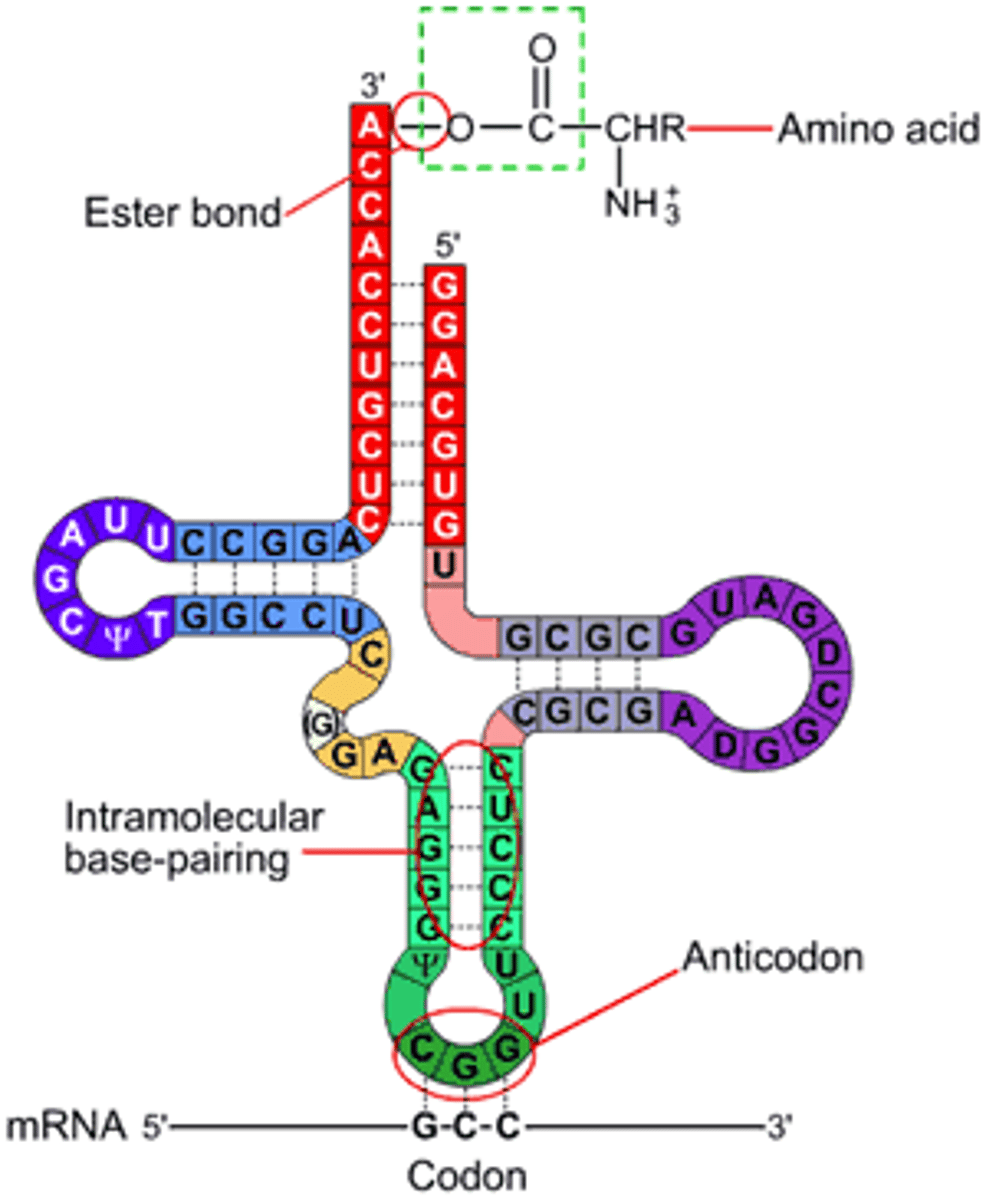
rRNA basics
Made from DNA, Component of ribosomes where protein synthesis occurs
Used to classify bacteria (16S) since it does not mutate very often
Which RNA is used as a means of classify bacteria? Why is this RNA used?
16S rRNA because does not mutate very often
Does RNA polymerase add nucleotides in a 5' to 3' direction, 3' to 5' direction or both?
Adds RNA nucleotides complementary to the template strand of a segment of DNA in the 5′ to 3′ direction
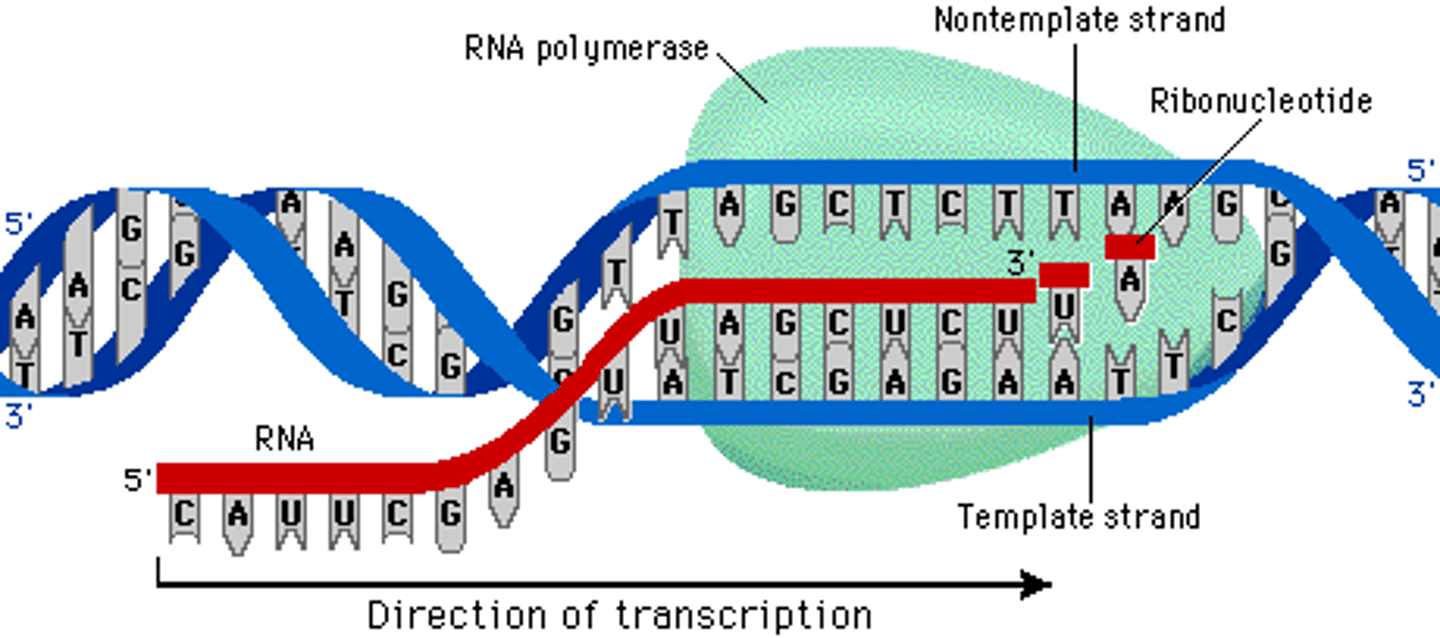
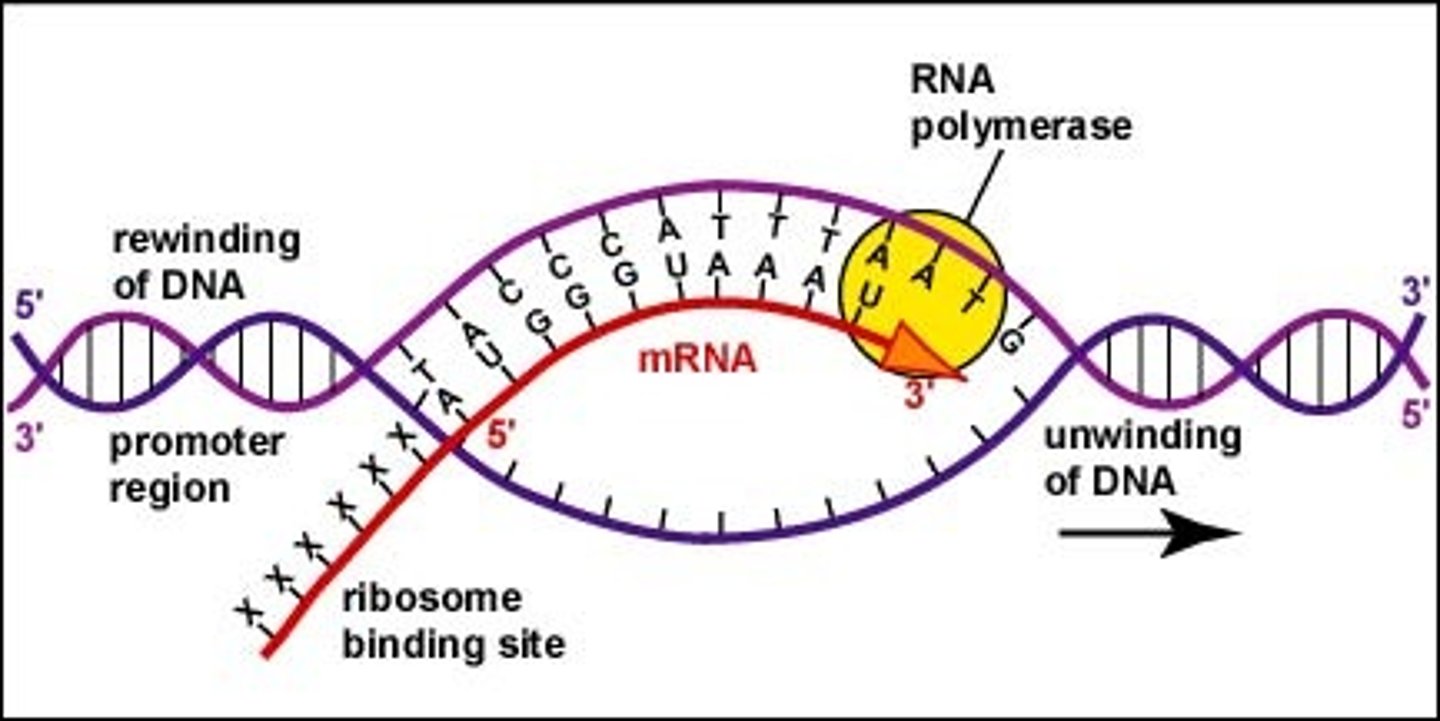
Fundamentals of transcription
Process of making an RNA copy of a gene sequence from DNA
RNA polymerase binds to a A-T region on DNA (promoter) and will use one strand as the template to make the RNA copy.
RNA polymerase will add complementary RNA nucleotide in a 5" to 3' direction until it reaches a terminator region on the DNA and this release the RNA polymerase and RNA copy.
mRNA copy will be read by ribosomes to make protein
tRNA copies will attach to amino acids and bring amino acids to the ribosomes
rRNA copies will become parts of ribosomes
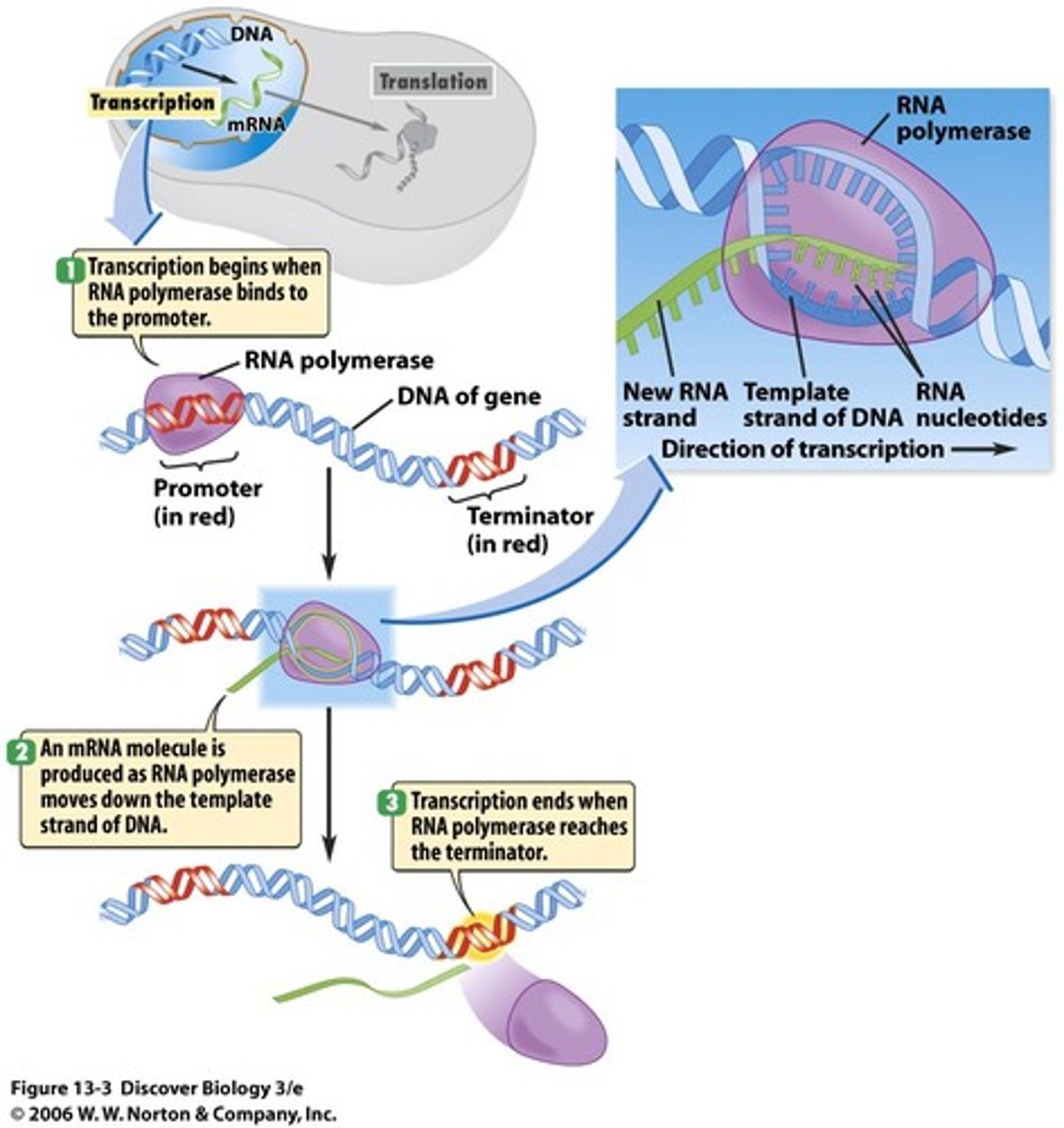
Fundamentals of translation
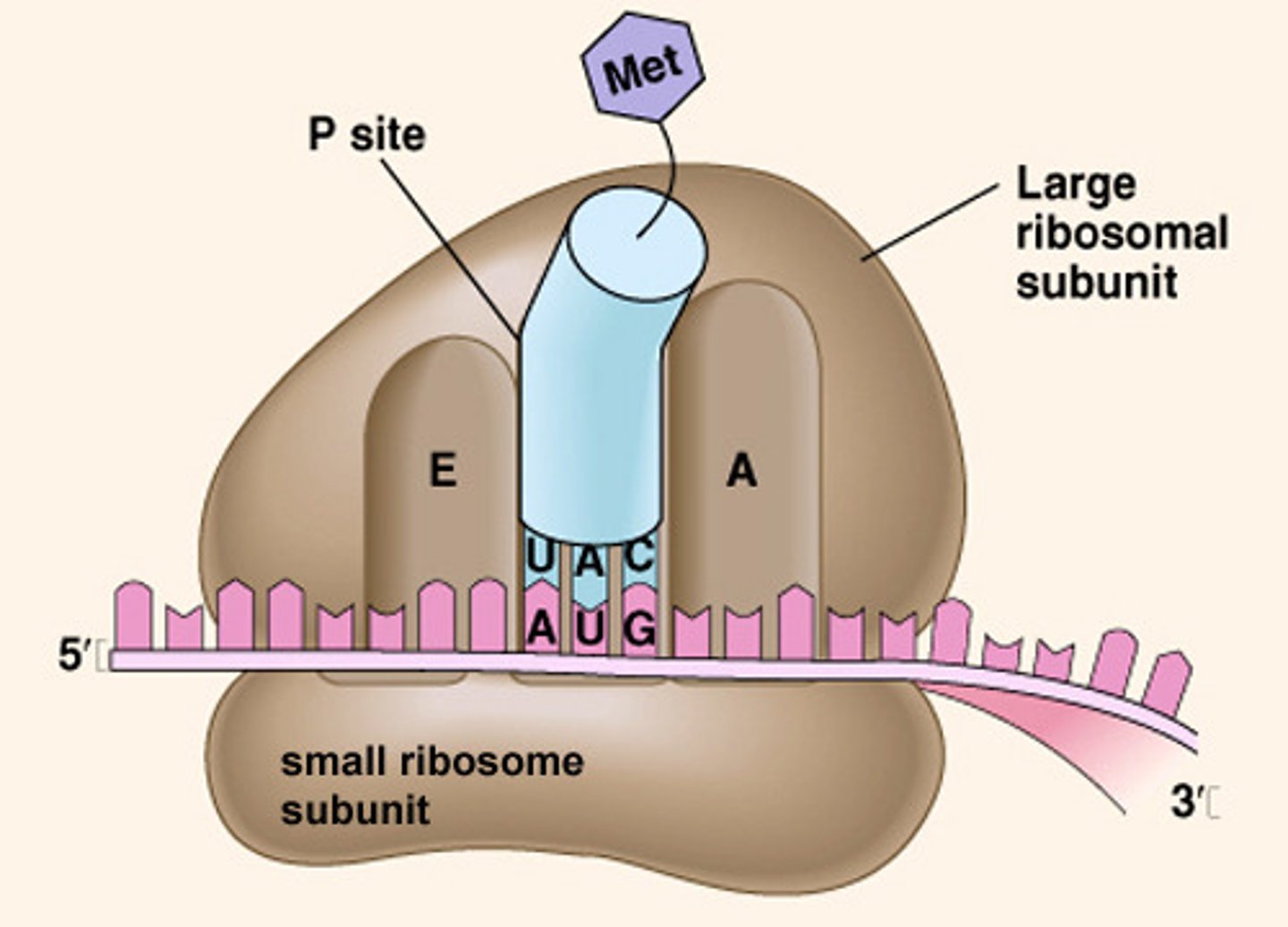
Process of translating the sequence of a messenger RNA (mRNA) molecule to a sequence of amino acids using ribosomes
The small subunit of the ribosome binds to mRNA and mRNA is read as three nucleotides/codon/triplet. The start codon on the mRNA will line up the P site of the ribosome. A tRNA with a complementary anticodon sequence will bring an amino acid (Met or f-MET) to the P site of the ribosome. A second tRNA will bring an amino acid to the A site of ribosome. The amino acids are covalently bonded together (peptide bond) by the ribozyme in the large subunit of ribosome and the growing peptide chain is transferred to the tRNA in the A site. The ribosome moves forward exposing the next codon and the tRNA with the growing chain goes to the P site. The tRNA without an amino acid leaves through the E site. The A site has the next codon to add the next amino acid.
This continues until a stop codon is reached by the ribosome in the A site. This codon does not code for an amino acid. A release factor enters the A site of the ribosome to separate the mRNA, ribosome and peptide.
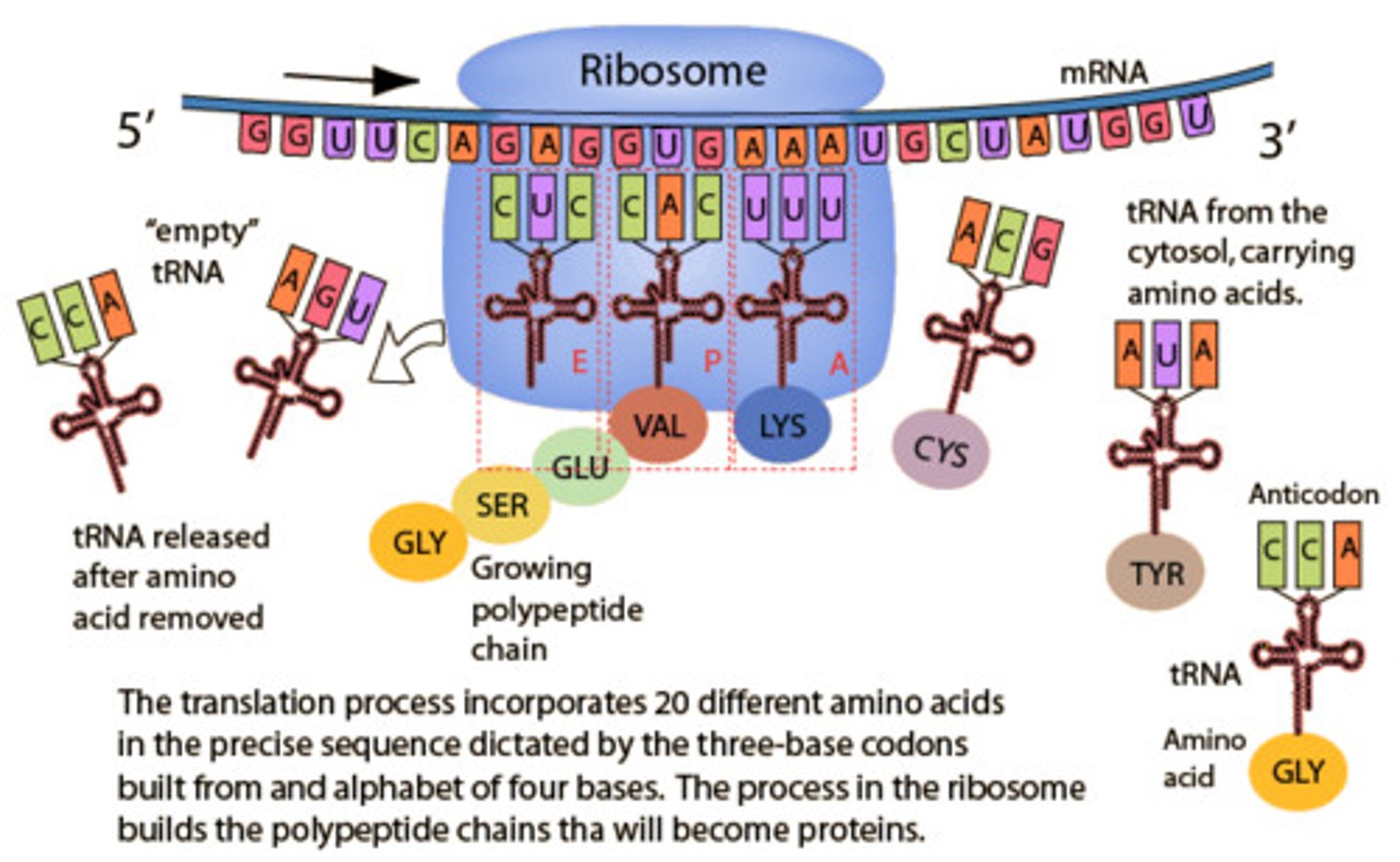
Why do you think the promoter region is made up of a lot of A-Ts?
There are only 2 hydrogens between A-T as compare to 3 hydrogen bonds for G-C so it takes less energy
Ribosomal subunit attaches to the mRNA
Small subunit
tRNAs enter which ribosomal subunit?
Large subunit
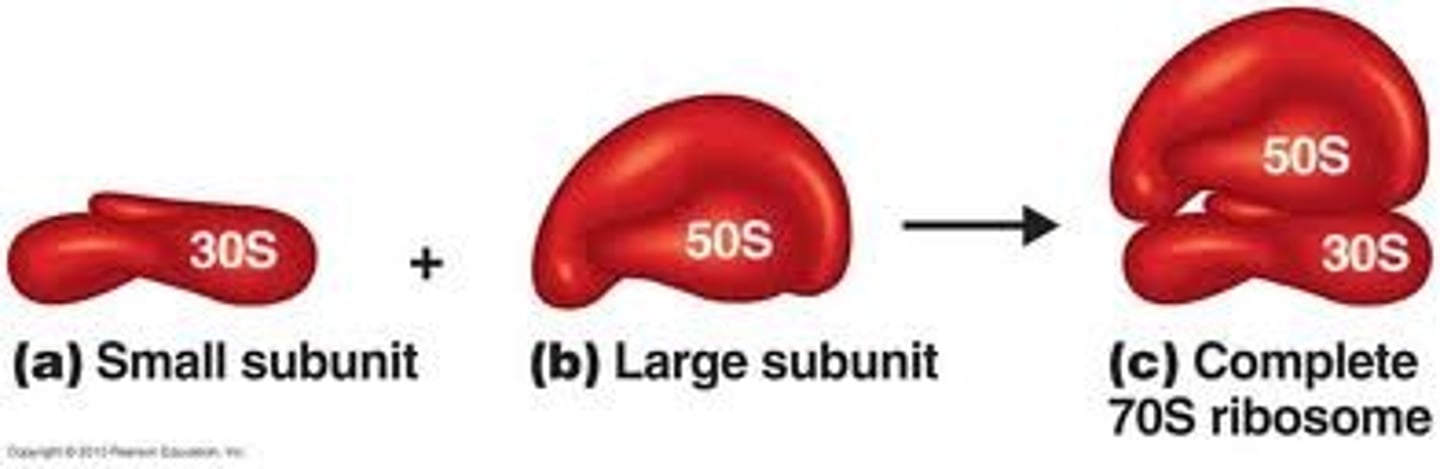
What unique amino acid is typically found as the first amino acid in bacterial proteins?
Bacteria usually have formyl methionine instead of methionine (eukaryotes) as its first amino acid
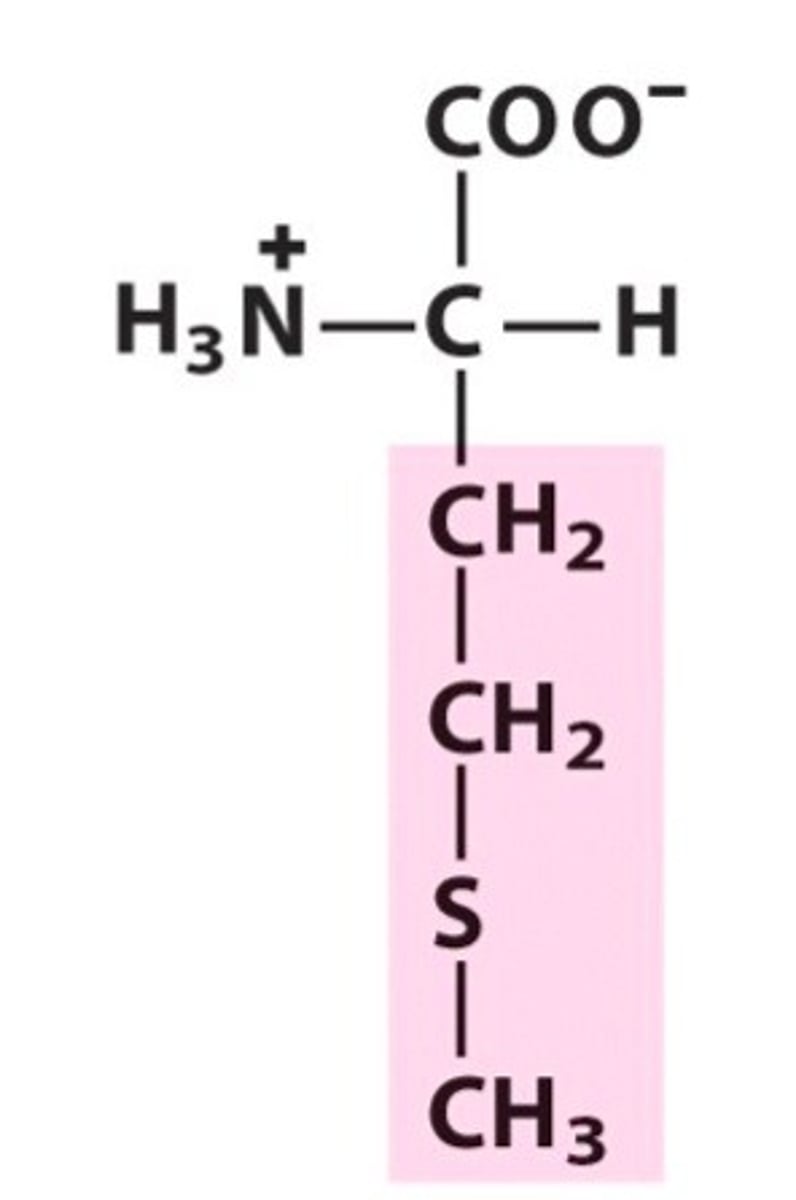
Is translation cost the cell a little bit of energy or is a lot of energy required for this process?
Uses 1200 ATP to make average size protein, so a lot
About 32 glucose molecules (through aerobic respiration) or 600 glucose molecules (through fermentation!)

Nucleotide sequence of the start codon
AUG
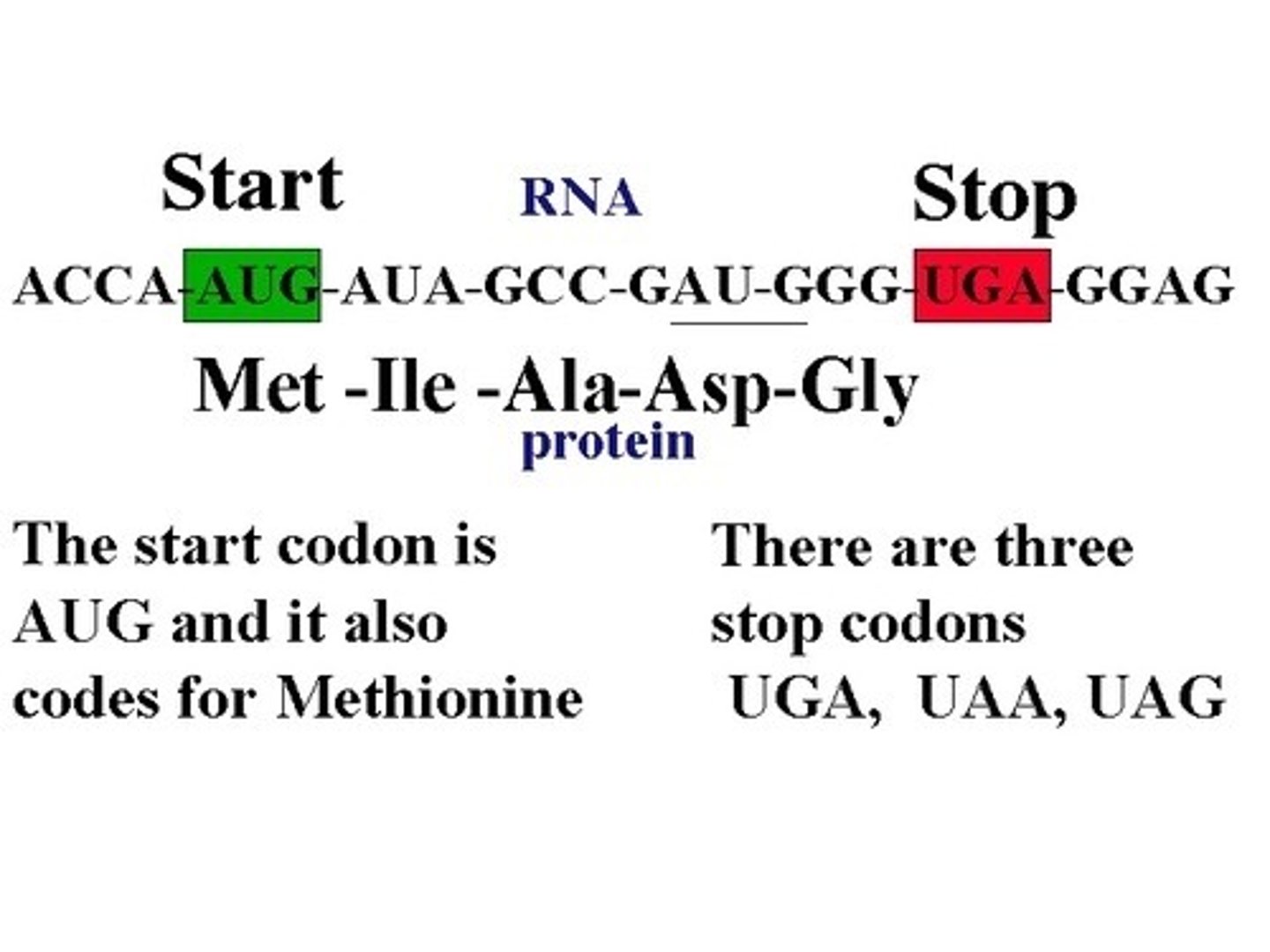

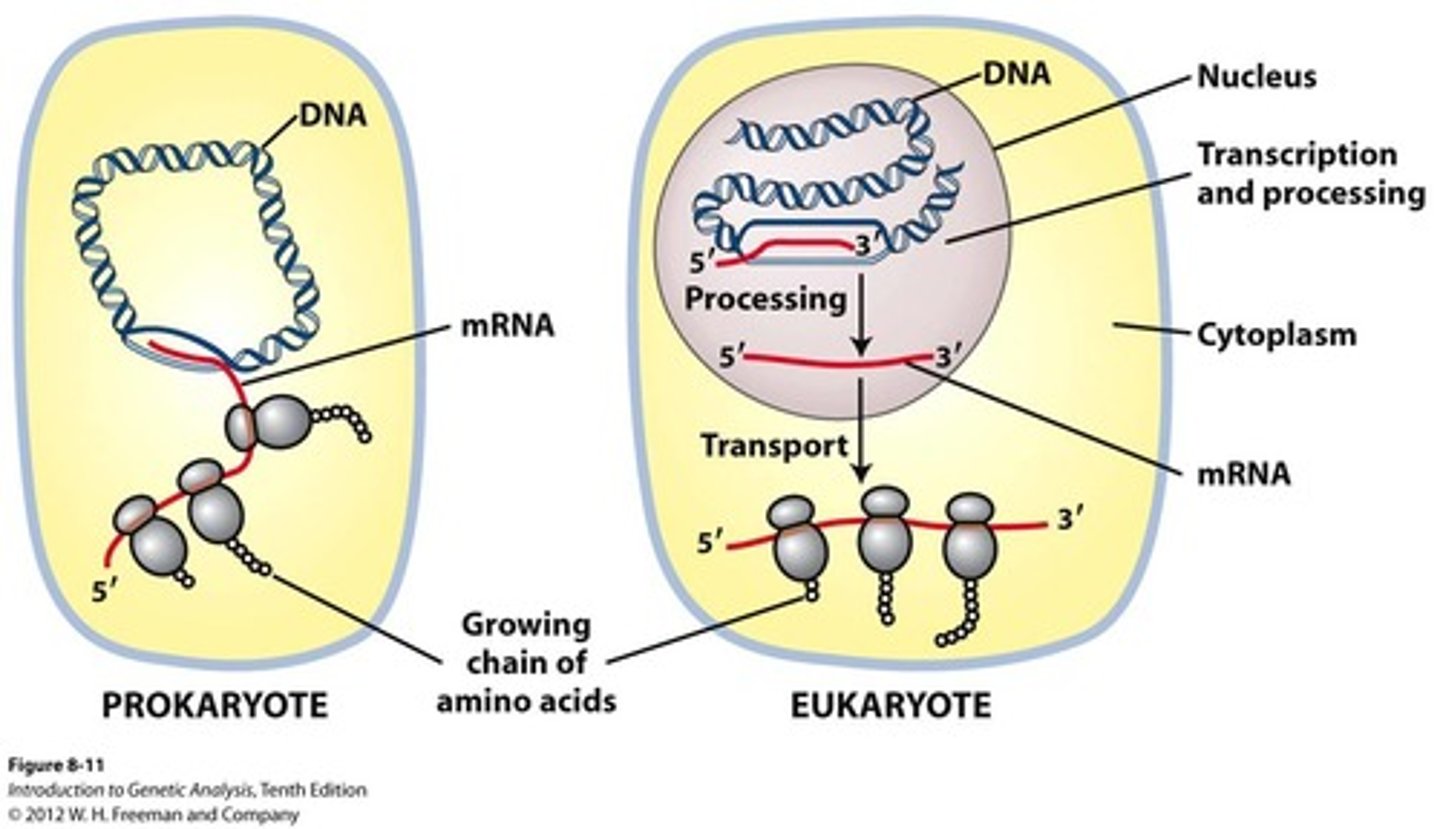
Basic differences between prokaryotic & eukaryotic transcription/translation: Cellular location
Eukaryotes: transcription and translation take place in different cellular compartments: transcription takes place in the membrane-bounded nucleus, whereas translation takes place outside the nucleus in the cytoplasm.
Prokaryotes: the two processes are closely coupled in the cytoplasm and occur simultaneously
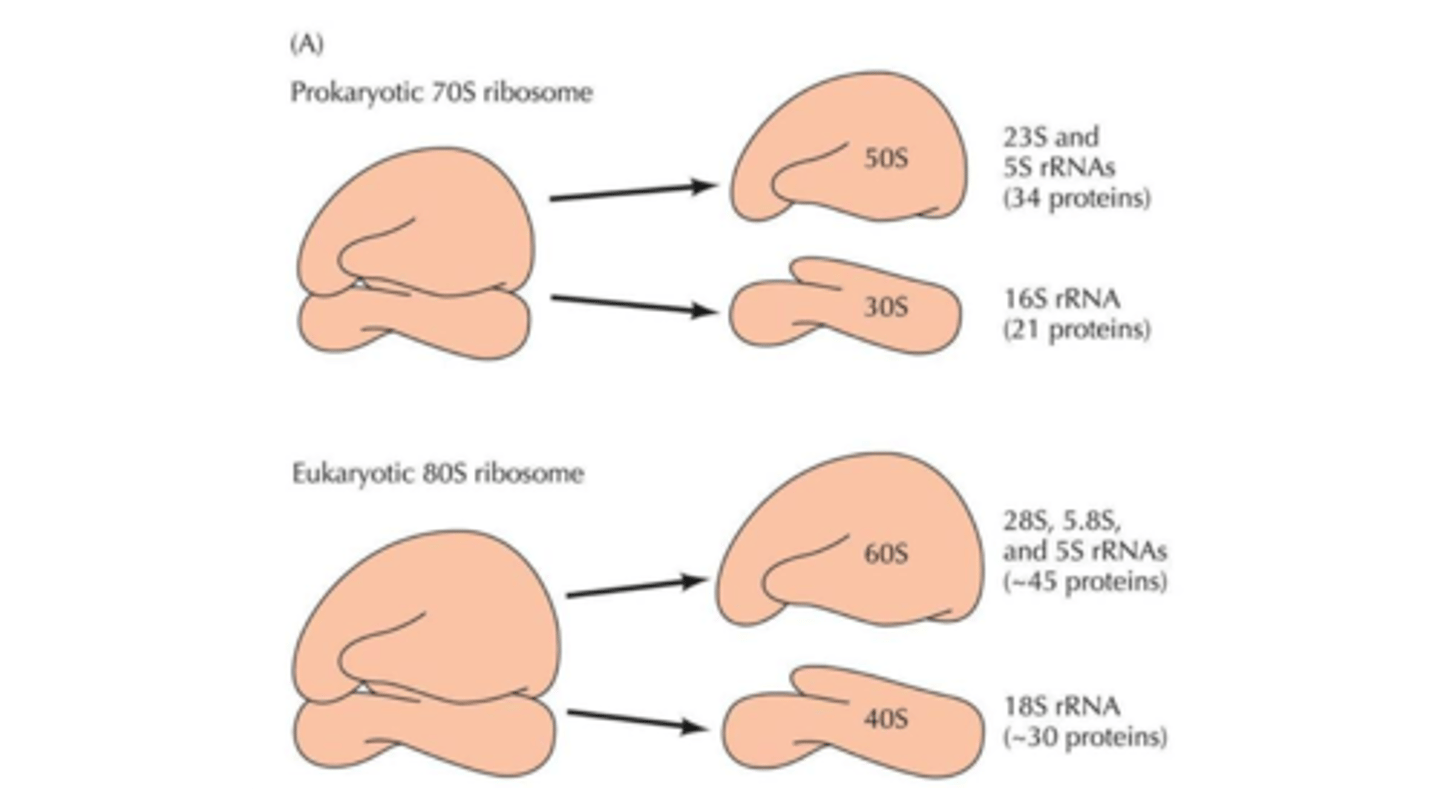
Basic differences between prokaryotic & eukaryotic ribosome size
Eukaryotes: 80S
Prokaryotes: 70S
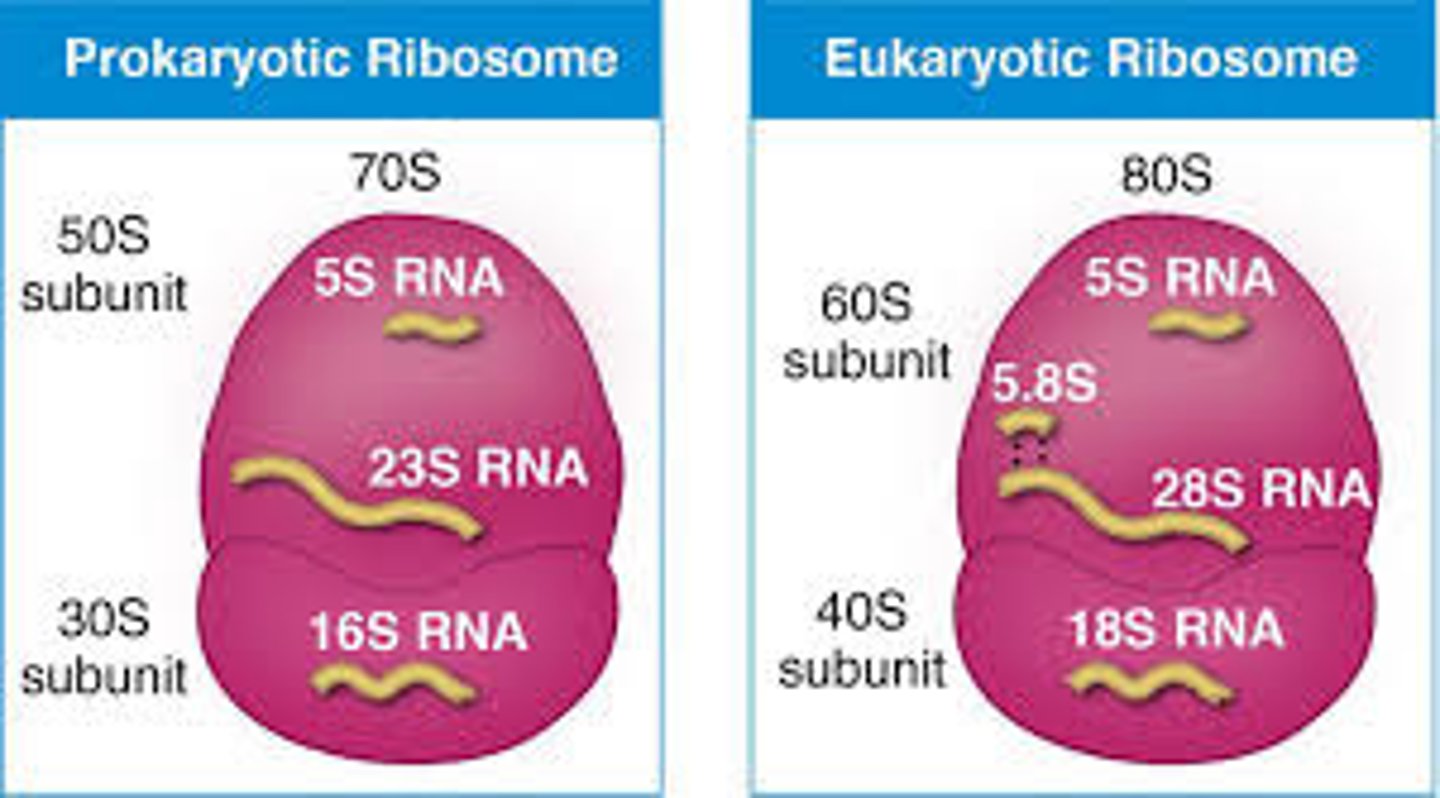
Basic differences between prokaryotic & eukaryotic mRNA processing
Prokaryotes: No special processing
Eukaryotes: Introns are removed from mRNA transcripts and exons are ligated together to form the final transcript using the spliceosome, caps and tails are added to the transcript
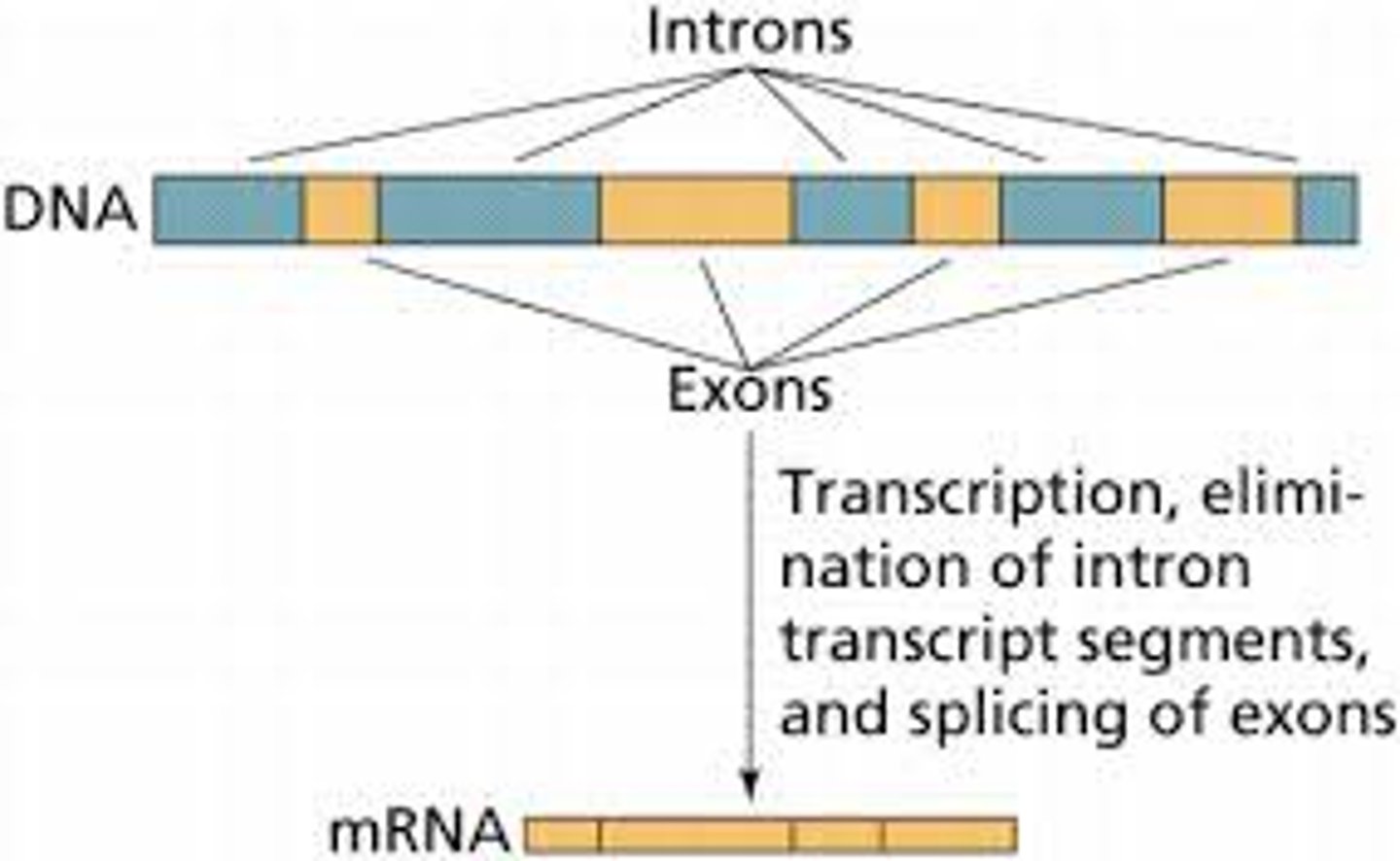
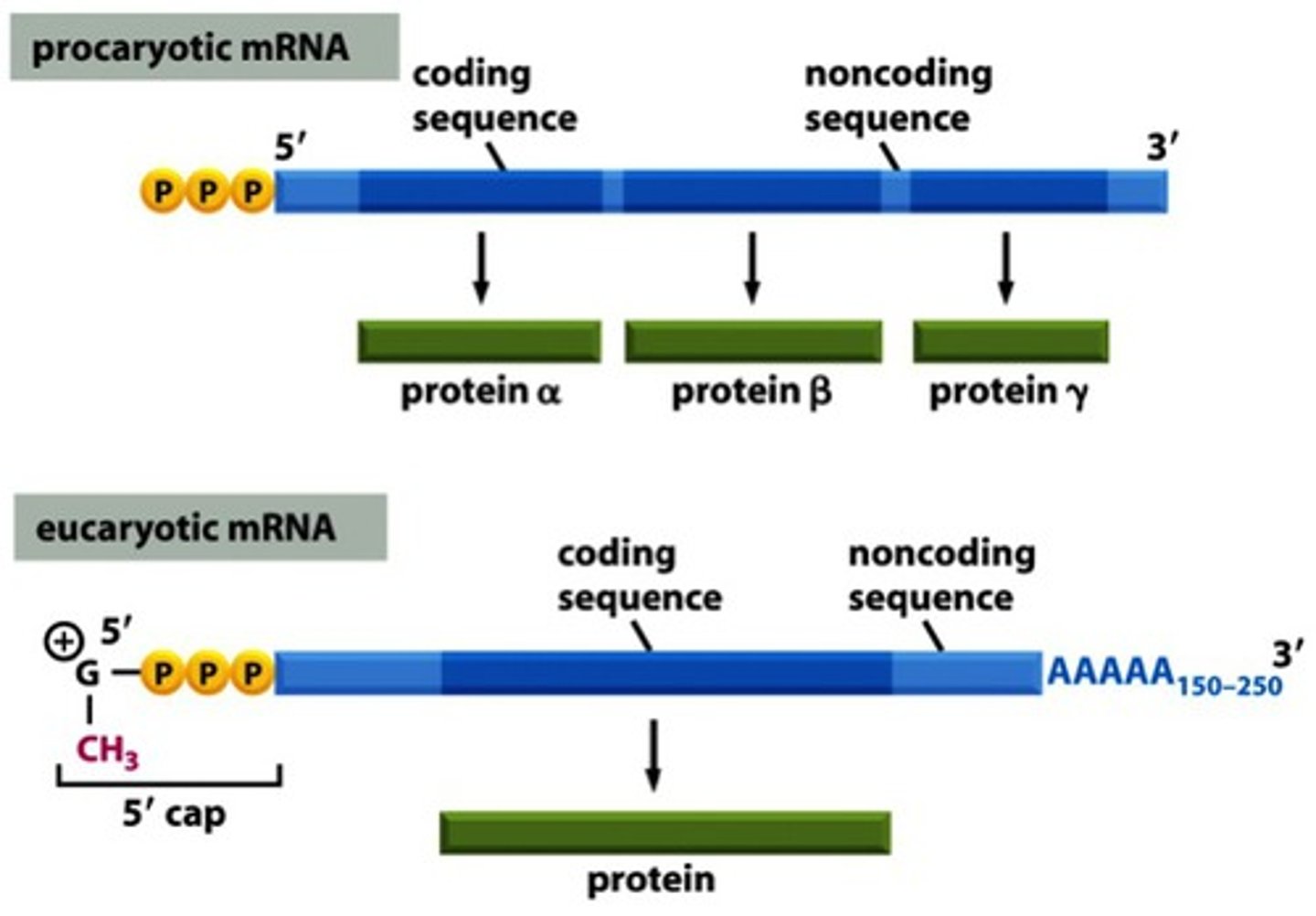
Basic differences between prokaryotic & eukaryotic gene numbers on transcripts.
Prokaryotes: Multiple genes per mRNA transcript
Eukaryotes: One gene per mRNA transcript
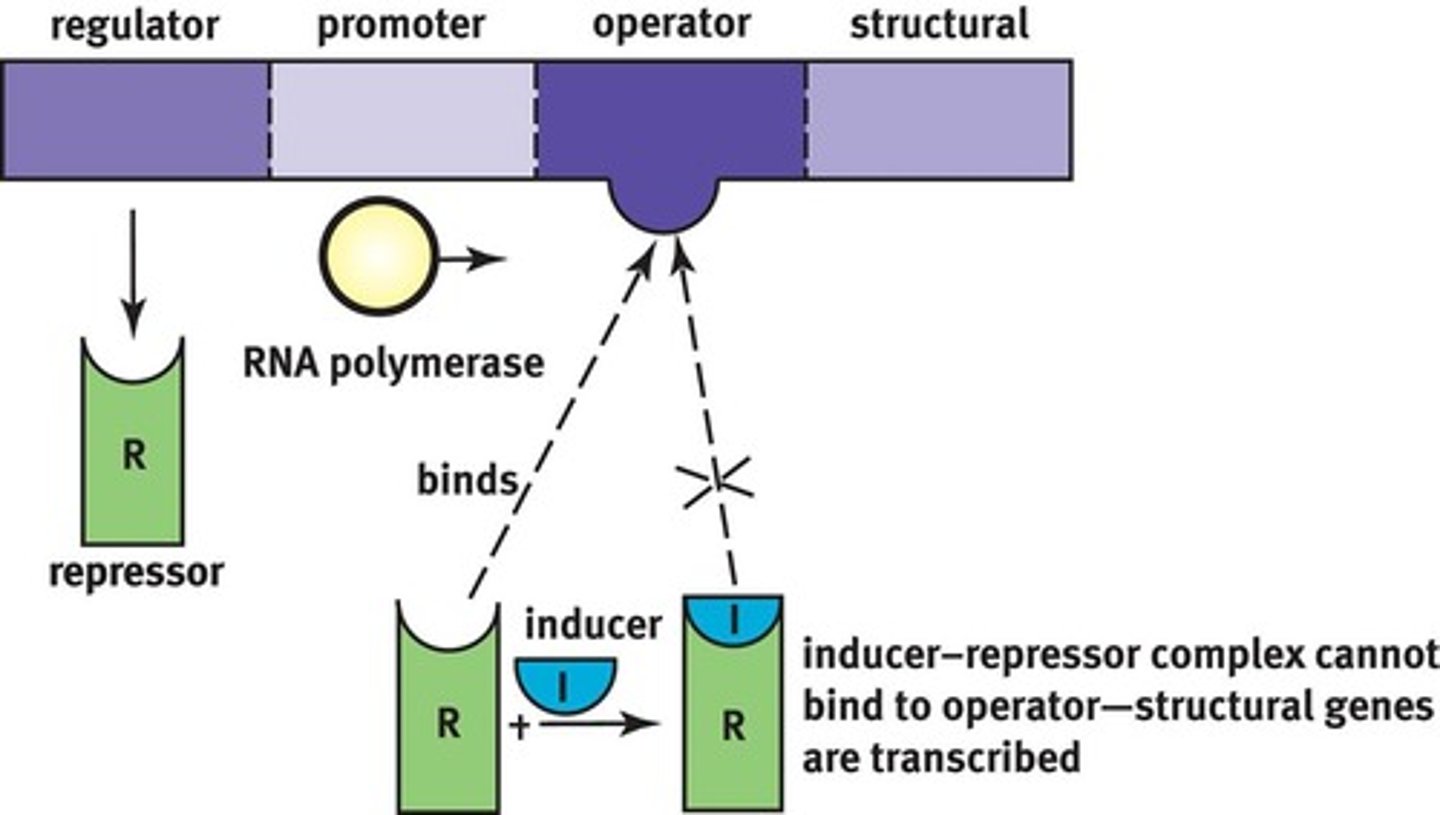
Two types of operons and examples of each
•Inducible operon: Catabolic operons (lactose metabolism)
•Repressible operon: Anabolic operons (make amino acids)
Are operons found in bacteria or eukaryotes?
Bacteria
If lactose is present and the bacterium has the lac operon, lactose can or cannot be used as an energy source.
Lactose can be used as an energy source.
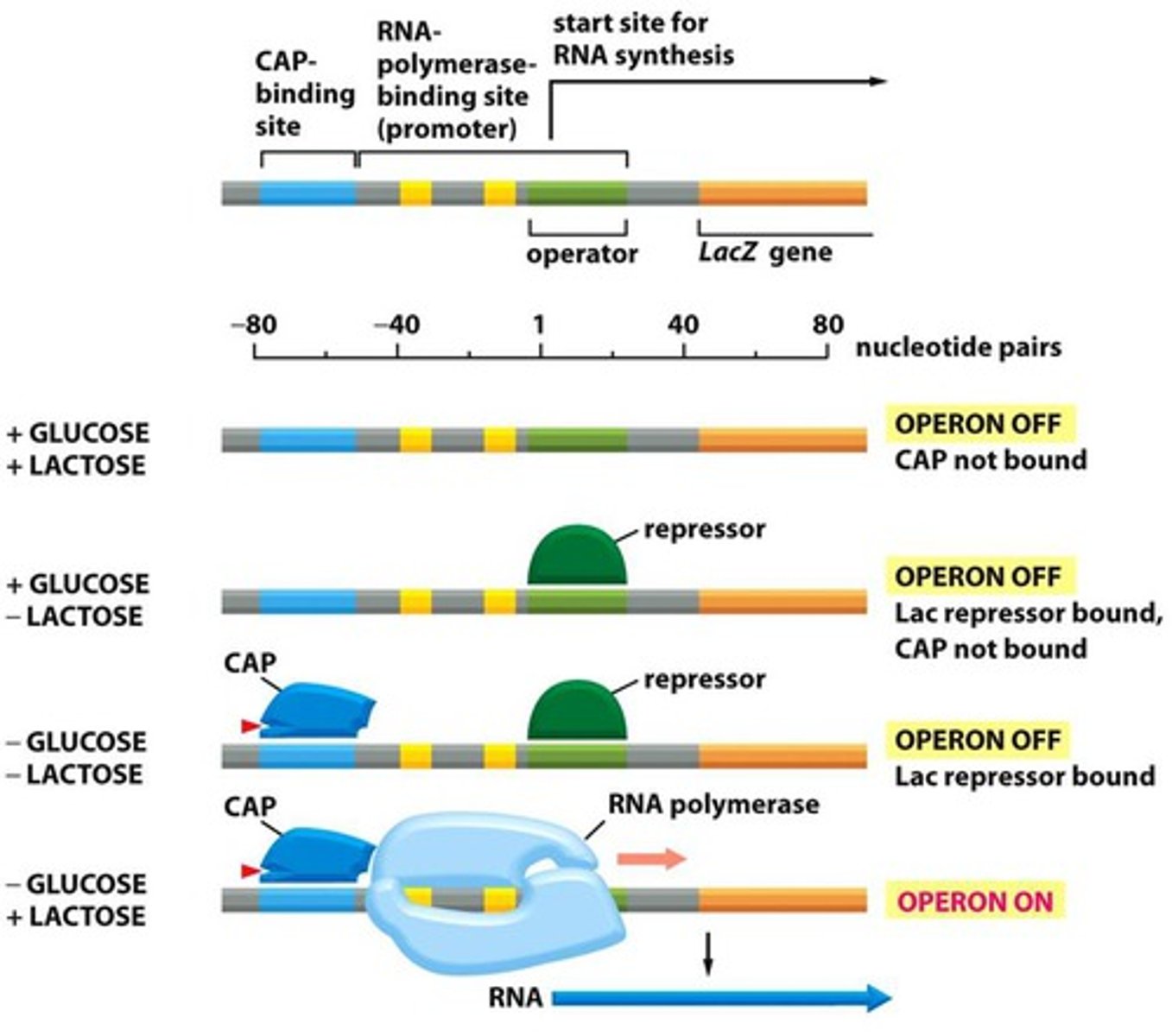
If lactose is present and the bacterium does not have the lac operon, lactose can or cannot be used as an energy source.
If a bacterium does not have the lac operon it cannot use lactose as energy source.
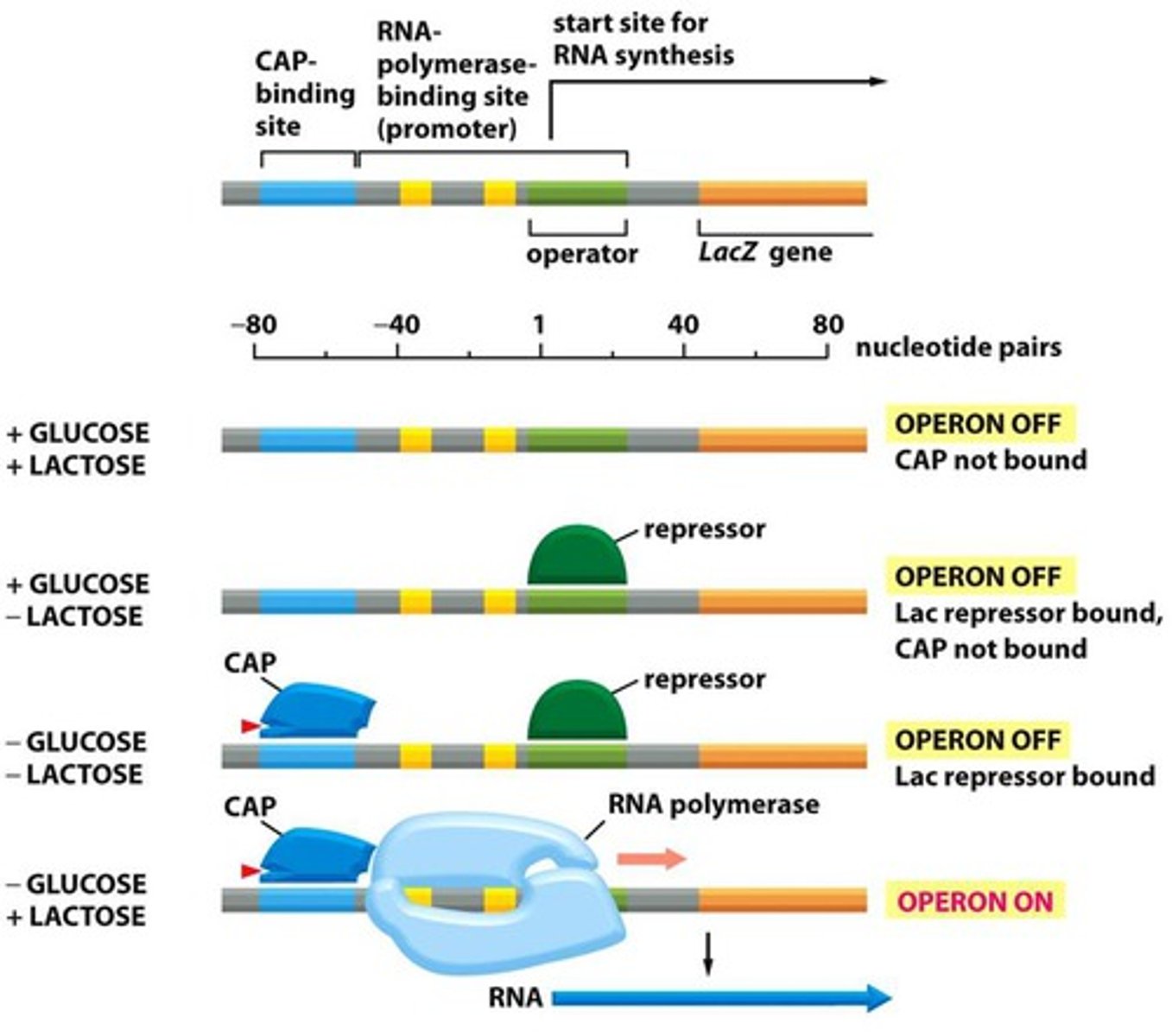
If lactose AND glucose are present and the bacterium has the lac operon, lactose can or cannot be used as an energy source.
If there is glucose available then glucose is used first then lactose.
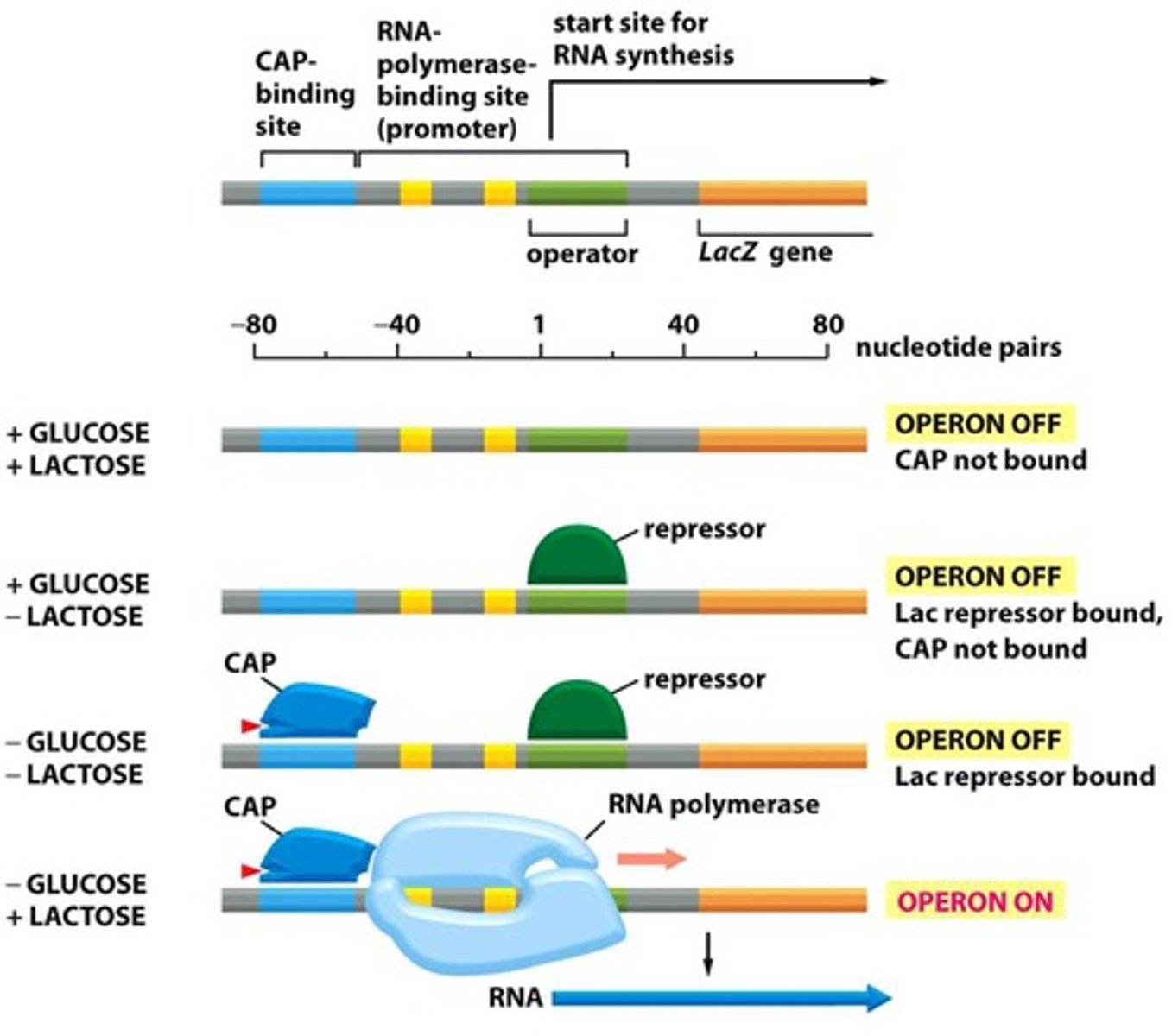
When are the lac proteins made: when lactose is absent or present?
Lac proteins are made when lactose is present

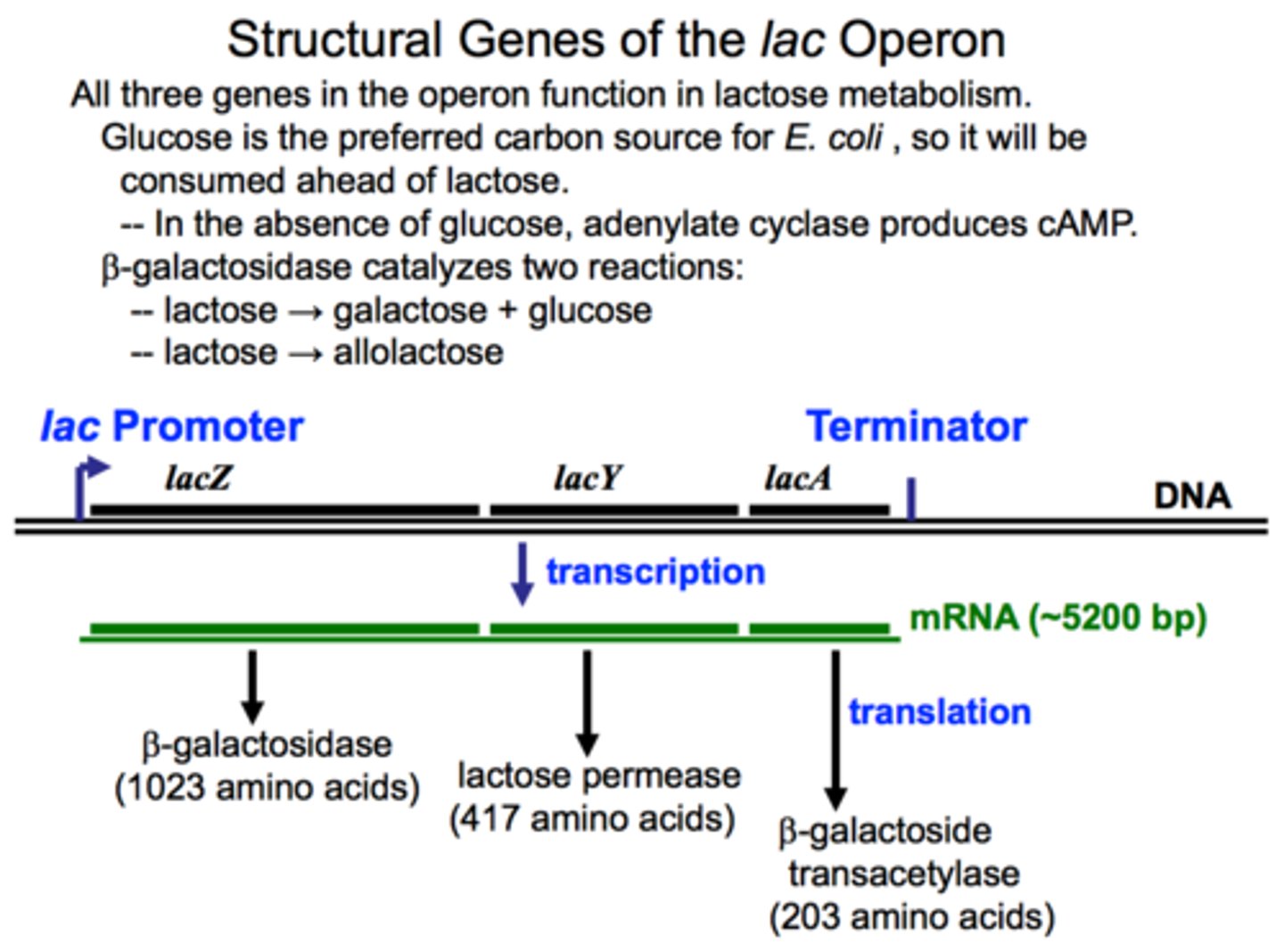
Understand the basics of how the lactose operon works.
Genes that allows microbes to use lactose for energy.
Operon for lactose metabolism is normally on "off mode" when lactose is absent.
The operon is on an "On mode" when lactose is present.
The genes for lactose metabolism will only be used if lactose is present and glucose is not present
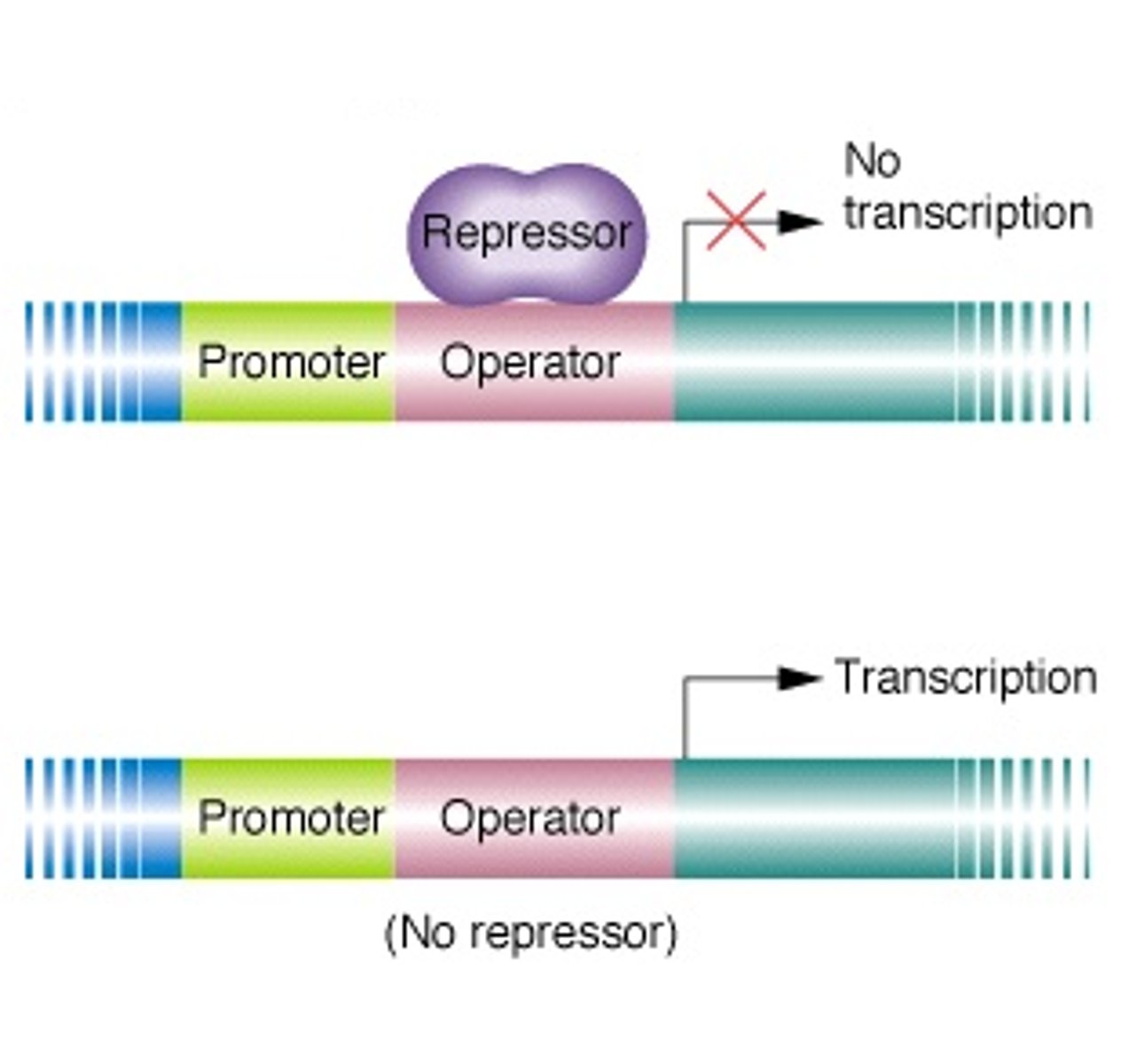
When there is no lactose, a protein repressor will bind the operator region on the DNA to block RNA polymerase from binding the DNA promoter thus stopping transcription of the lactose metabolism genes.
When there is lactose, the inducer allolactose, formed from lactose, will bind the protein repressor so it cannot bind DNA, then RNA polymerase can then bind the DNA promoter and start transcribing the genes needed to metabolize lactose.
*
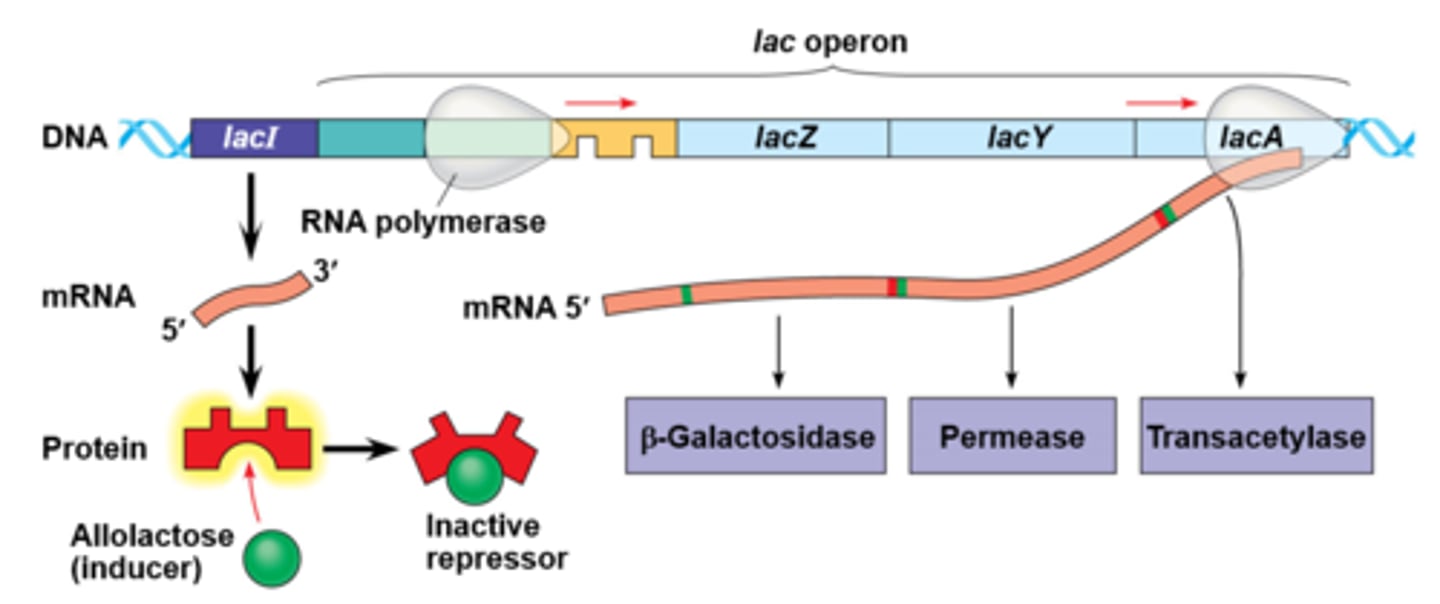
Structural genes for the lactose operon and their functions
LacZ: Enzyme that breaks down lactose into glucose and galactose
LacY: Permease or channel that allows lactose into the cell
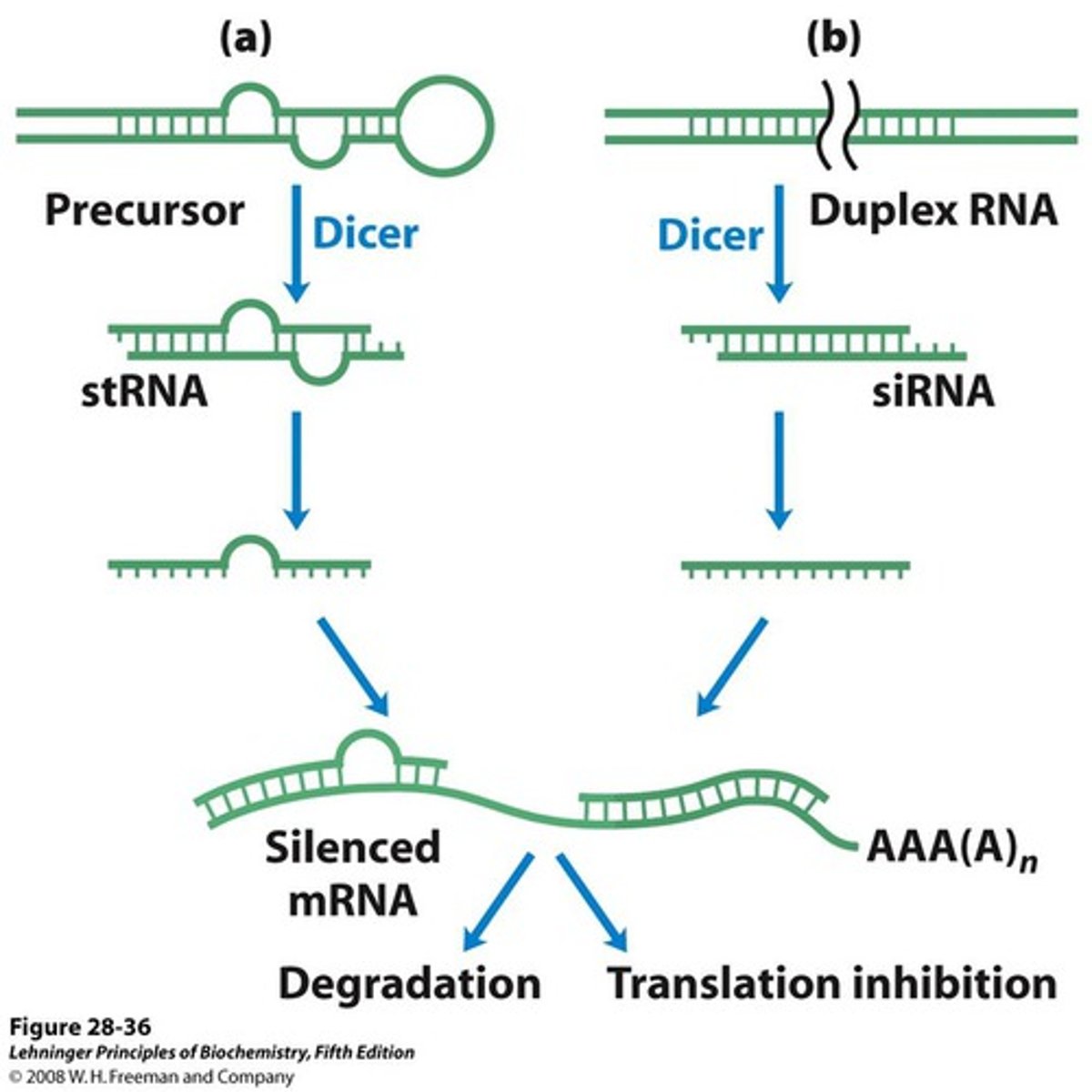
Why do cells form siRNA? What function do these nucleotides serve?
Cells form these siRNAs to control mRNA after transcription in the process of gene silencing.
These RNAs can be used to stop viral infections as siRNA will target viral genomic RNA so it can be degraded.
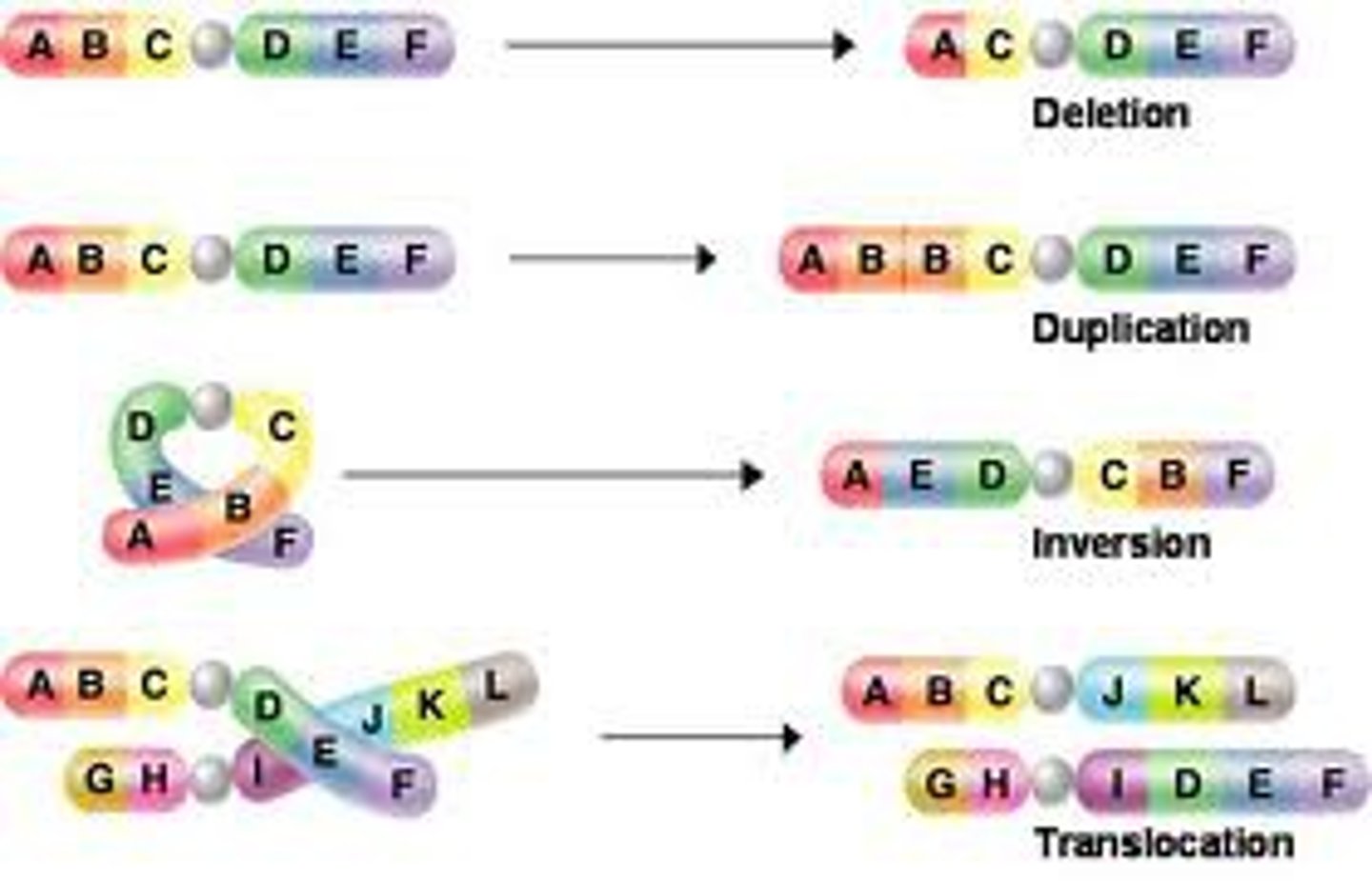
Are all mutations harmful to the host? Are there benefits and if so, what?
Some mutations are harmful/lethal.
Some are beneficial and can give organisms a evolutionary advantage (EX pepper moth transition from pepper color to brown) or can lead to a phenotype change like antibiotic resistance
Some mutations do not change the phenotype due to silent mutations (the mutation still leads to the production of the same amino acid)
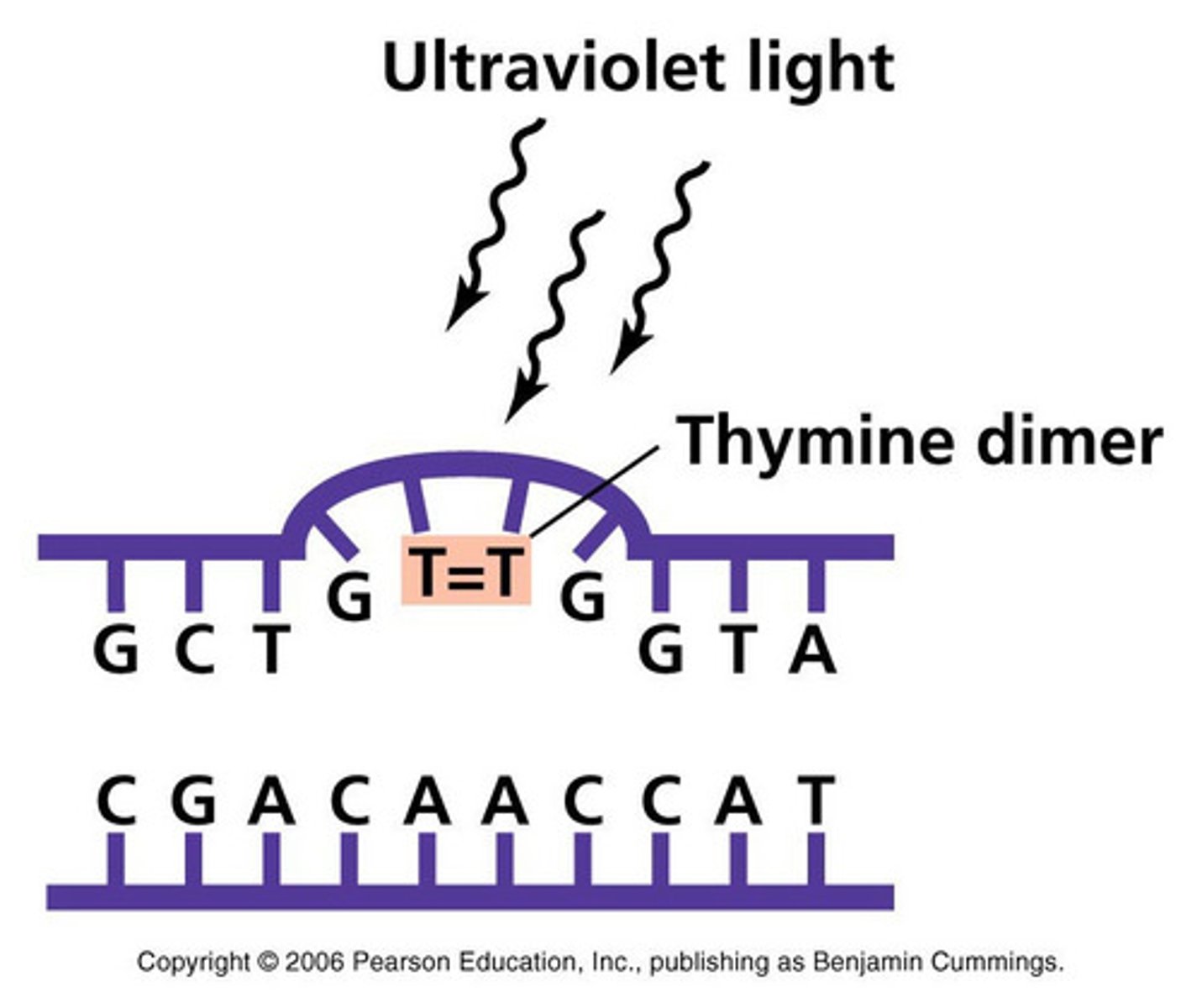
Examples of physical agent that cause mutations and what affect do these agents have on DNA?
Radiation
High energy gamma rays, X-rays can break DNA
UV can form abnormal bonds between adjacent pyrimidines/thymines to form dimers that prevents replication
Examples of chemical agents that cause mutations.
Many carcinogens
Nucleotide analogs like 5'-bromouracil (looks like thymine)
Intercalating agents: Ethidium bromide (insert between base pairs)

Resulting protein if there is a missense mutation
The protein is faulty, functions differently or functions the same
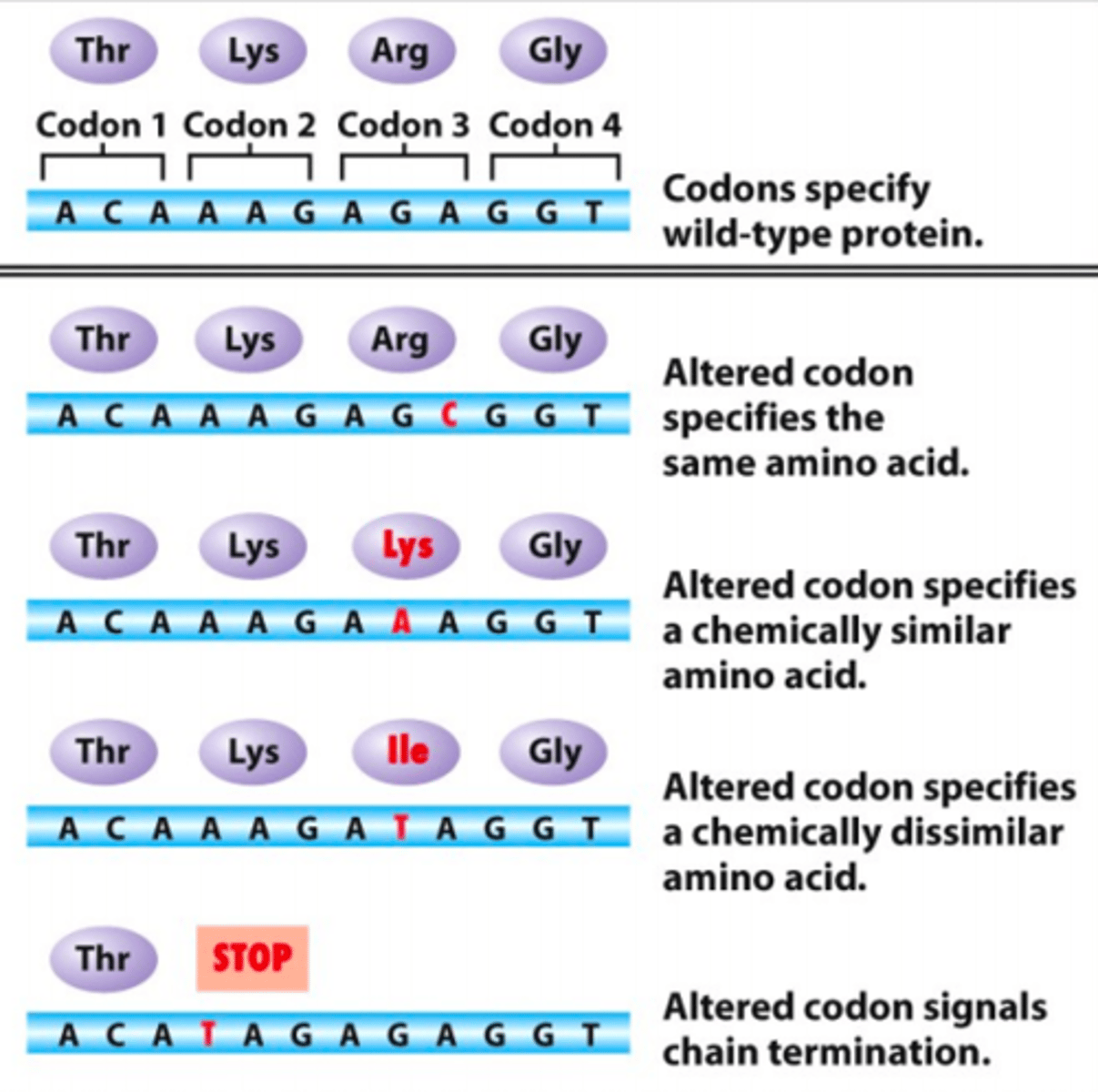
Resulting protein if there is a nonsense mutation
The protein is short or truncated, most nonfunctional

Resulting protein if there is a silent mutation
No effect on the proteins due to the wobble effect
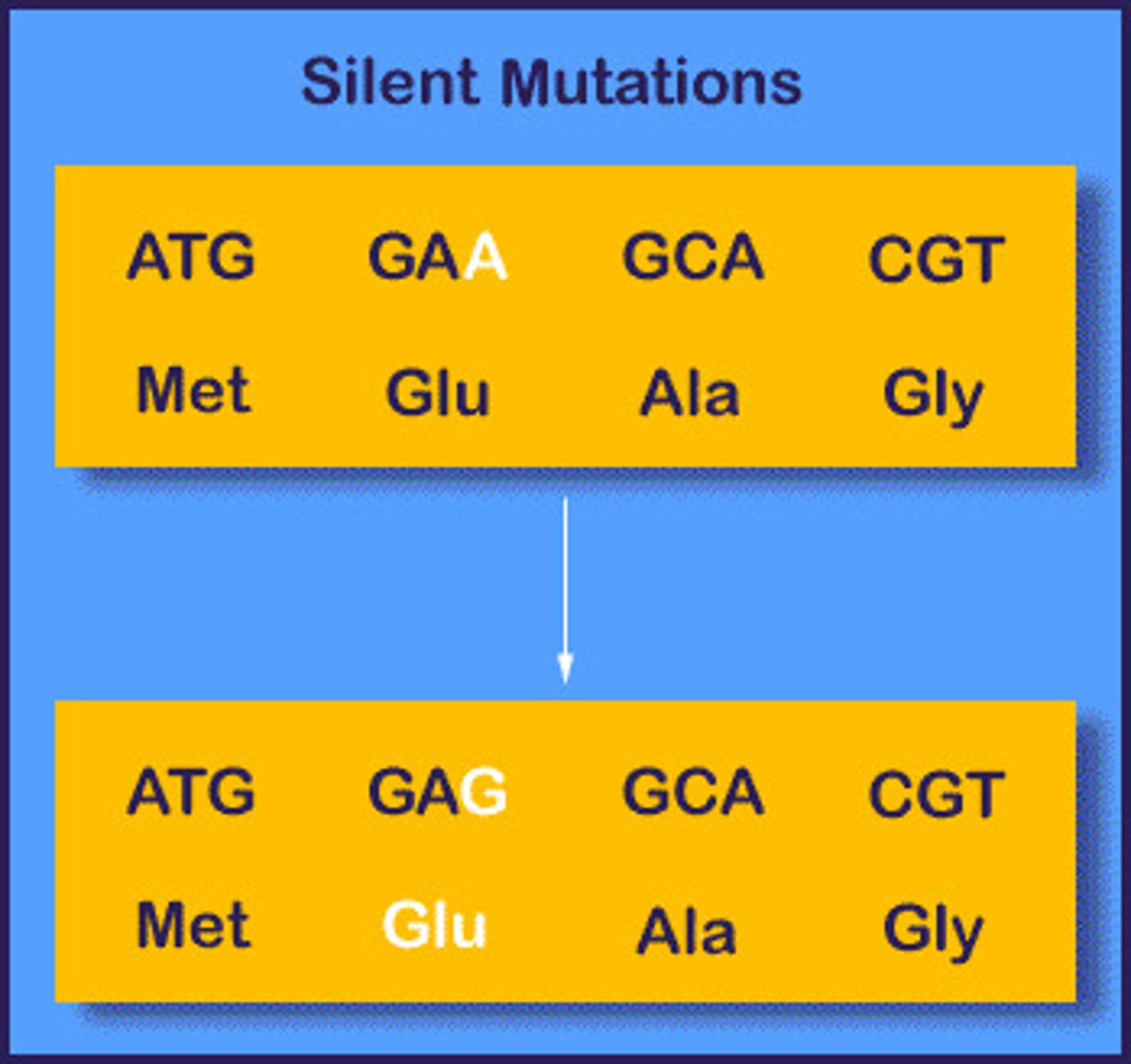
Resulting protein if there is a back mutation
The protein returns to its original shape
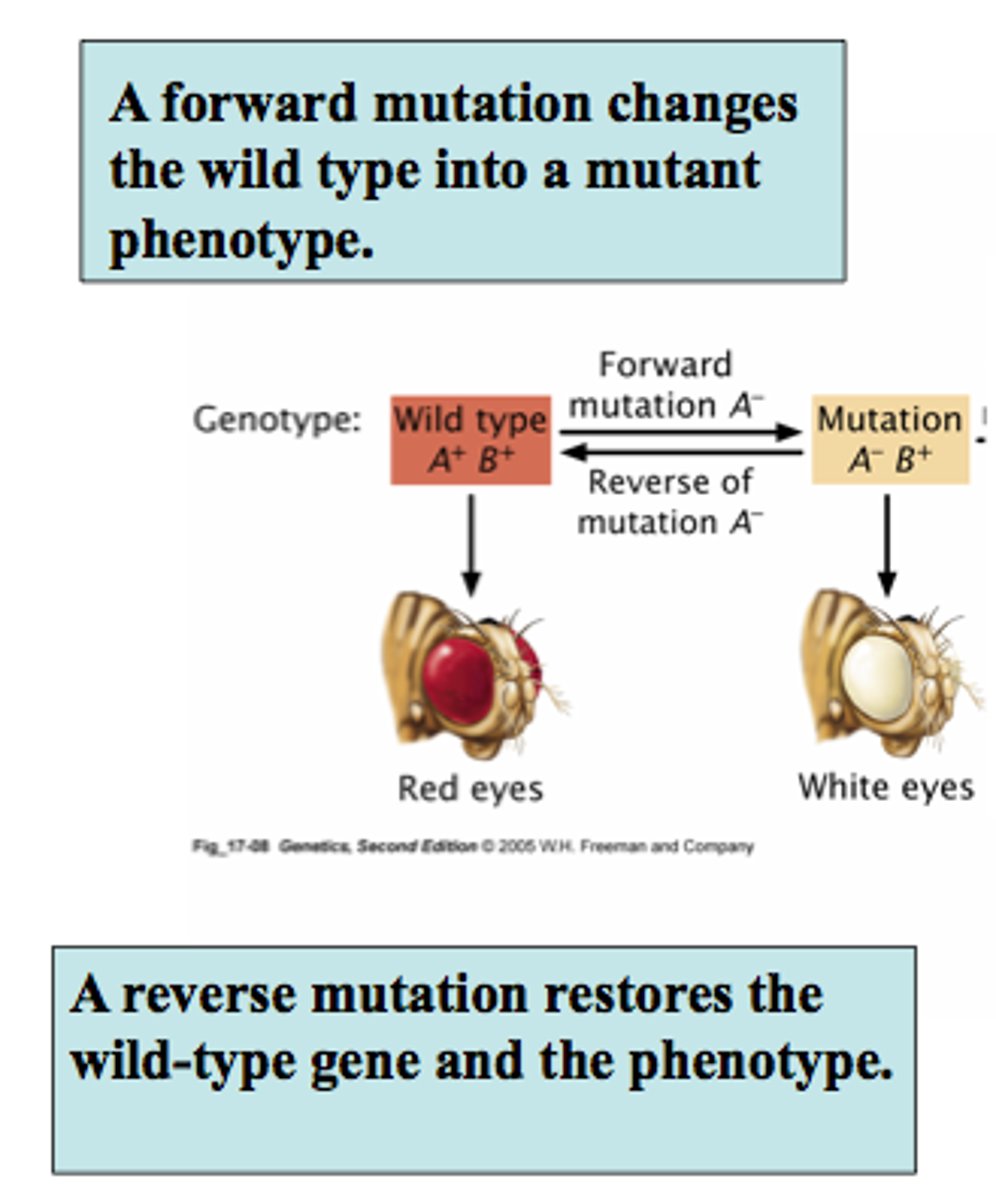
Resulting protein if there is a frameshift mutation
All amino acids added after the mutation differ, which may lead to an early stop, forming a nonfunctional protein
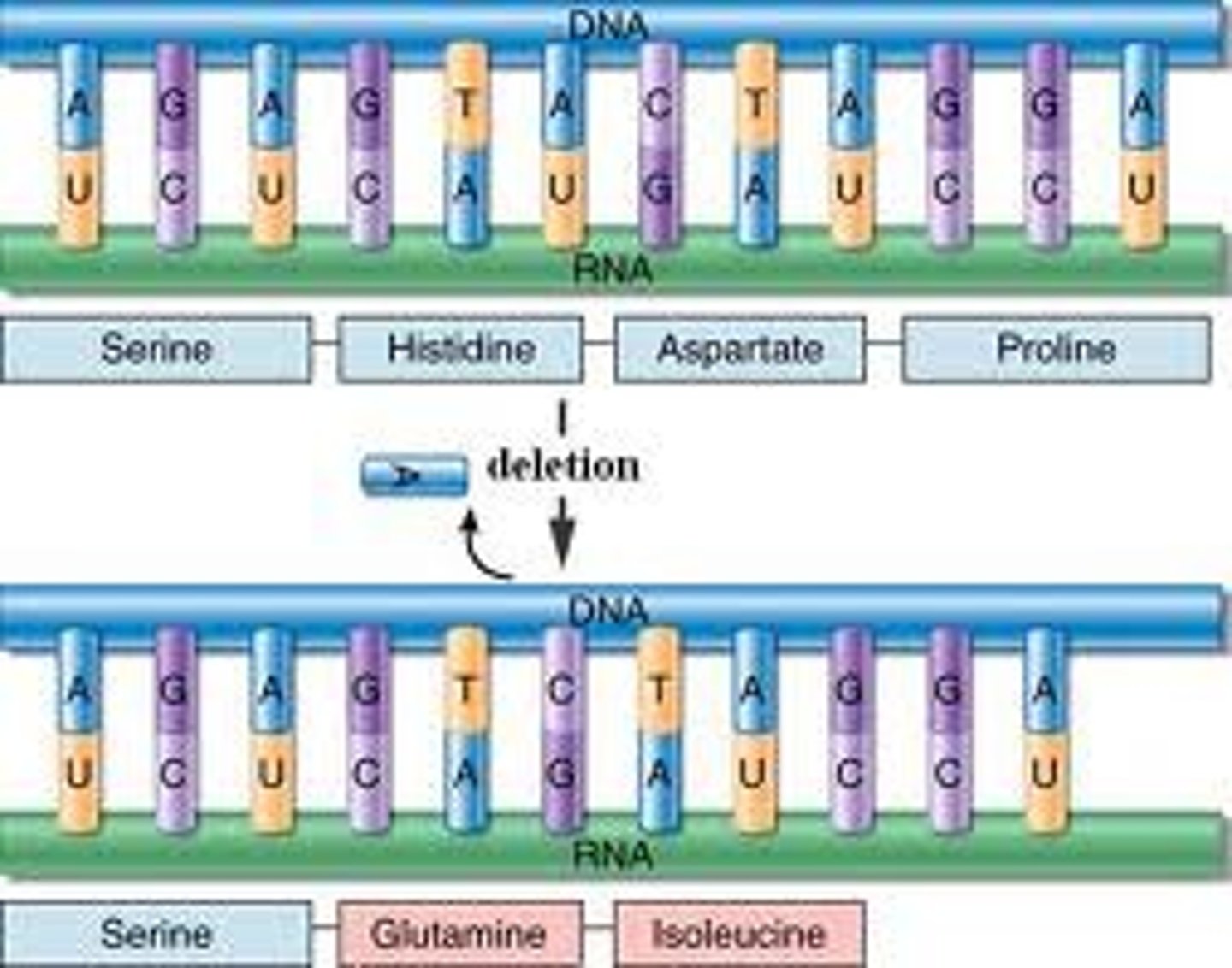
Four ways of repairing mutations in DNA
-DNA polymerase
-Mismatch repair
-Light repair
-Excision repair
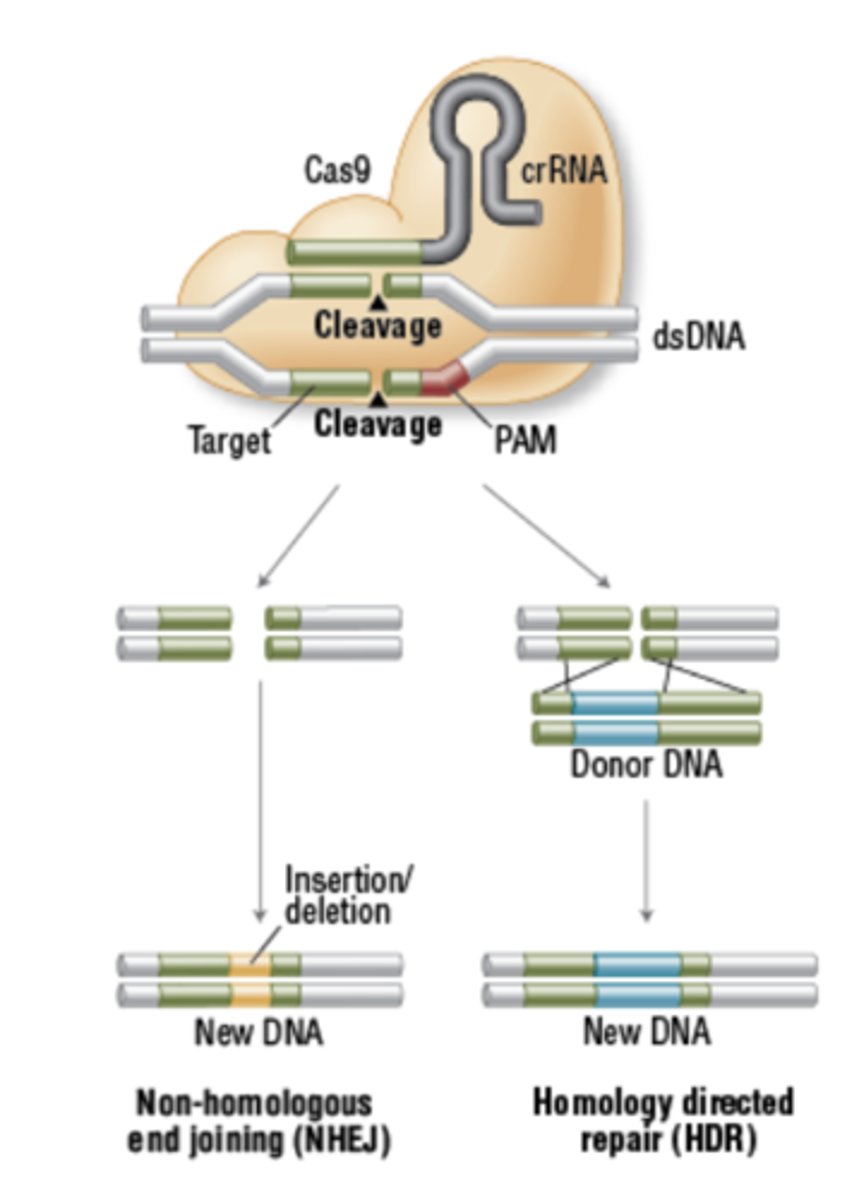
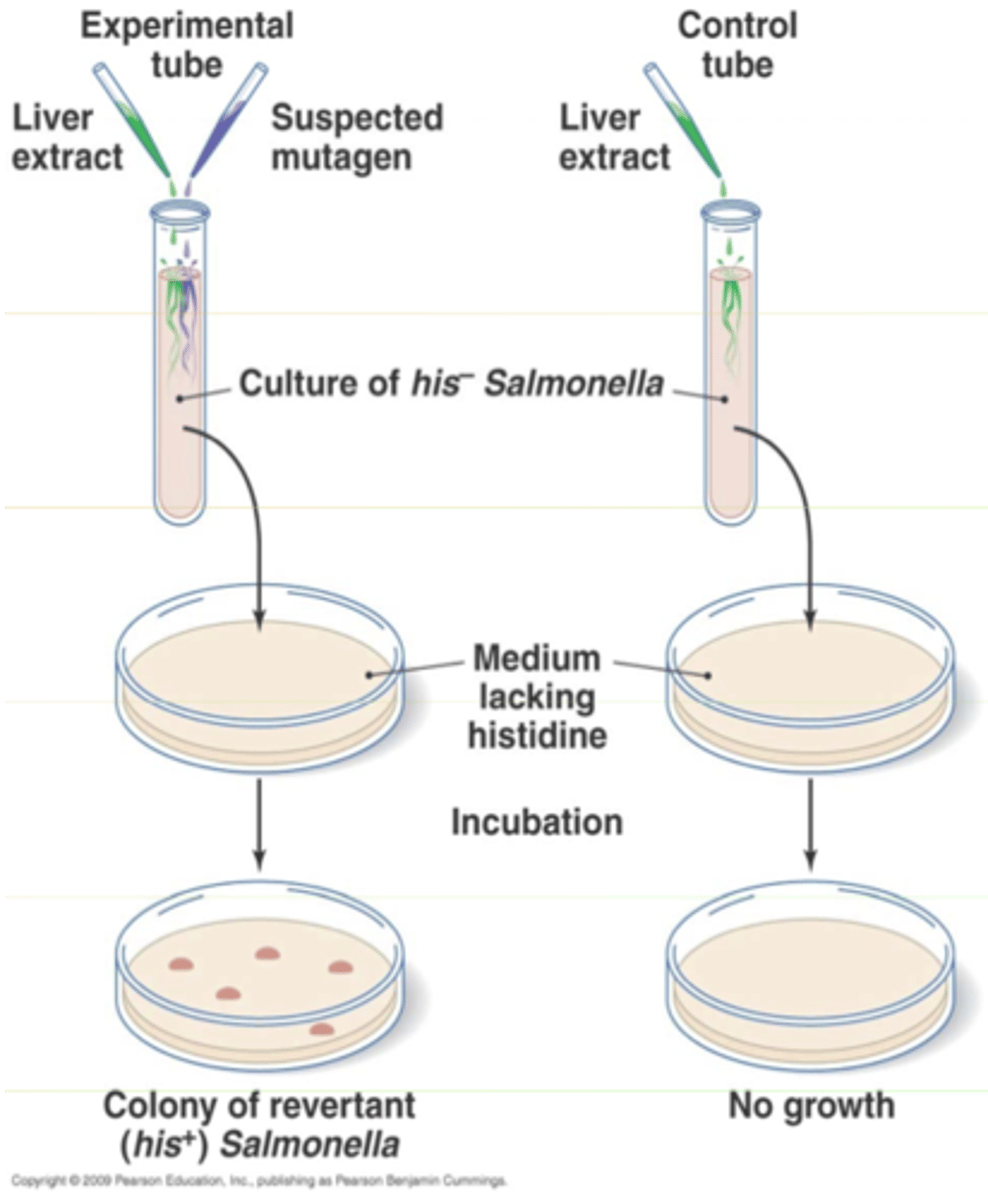
What is the purpose of the Ames test and how does it work? Be able to apply concepts.
The Ames test uses bacteria to test whether a given chemical can cause mutations in the DNA of the test organism.
Tests to see if a chemical is a carcinogen (physical or chemical agent that causes cancer)
This test uses Salmonella that is mutated in histidine (amino acid) production so if the bacterium is grown on media lacking histidine it cannot grow. If the bacterium is treated with a chemical and the Salmonella can now grow on media without histidine this chemical has cause a back mutation in Salmonella gene for making histidine to its original form and is thus the chemical is a carcinogen
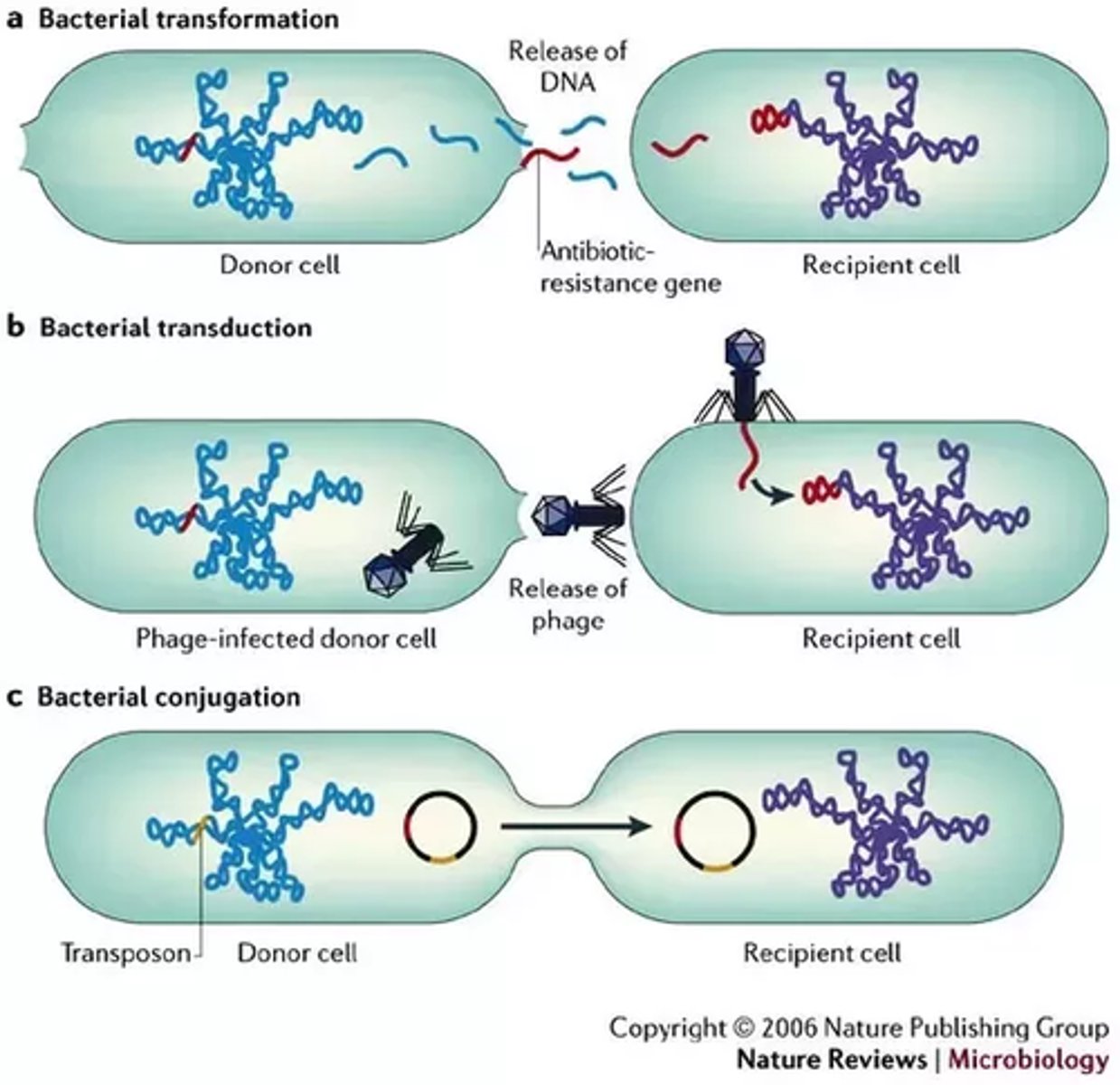
Basic means of genetic recombination in bacteria
•Conjugation
•Transformation
•Transduction
•Transposition
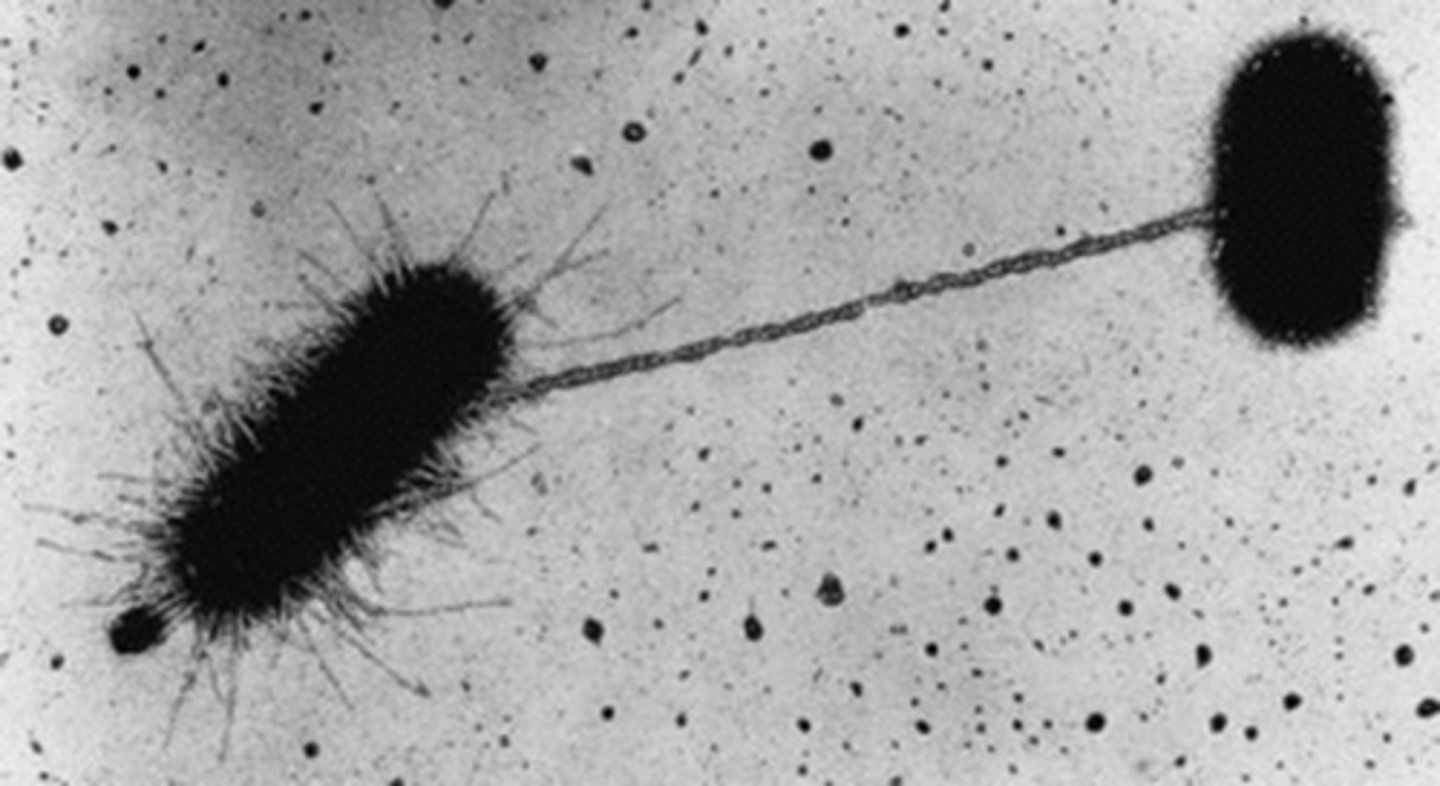
How are plasmids transferred from one bacterium to another in conjugation?
Via direct connection through a pilus
Difference between F plasmid transfer and Hfr conjugation
F plasmid transfer conjugation: the donor bacterium has a fertility plasmid that allows the synthesis of a conjugation (sex) pilus. Donors can transfer this plasmid to recipient cells of a related species or genus that do not have a fertility plasmid. Then the recipient cell can make the pilus.
Hfr conjugation: The donor's fertility plasmid has been integrated into the bacterial chromosome.
When conjugation occurs, a portion of the chromosome and a portion of the fertility plasmid are transferred to the recipient. This can lead to recombination of donor genes to the recipient cells but these cells cannot produce the pilus due to incomplete transfer of genes
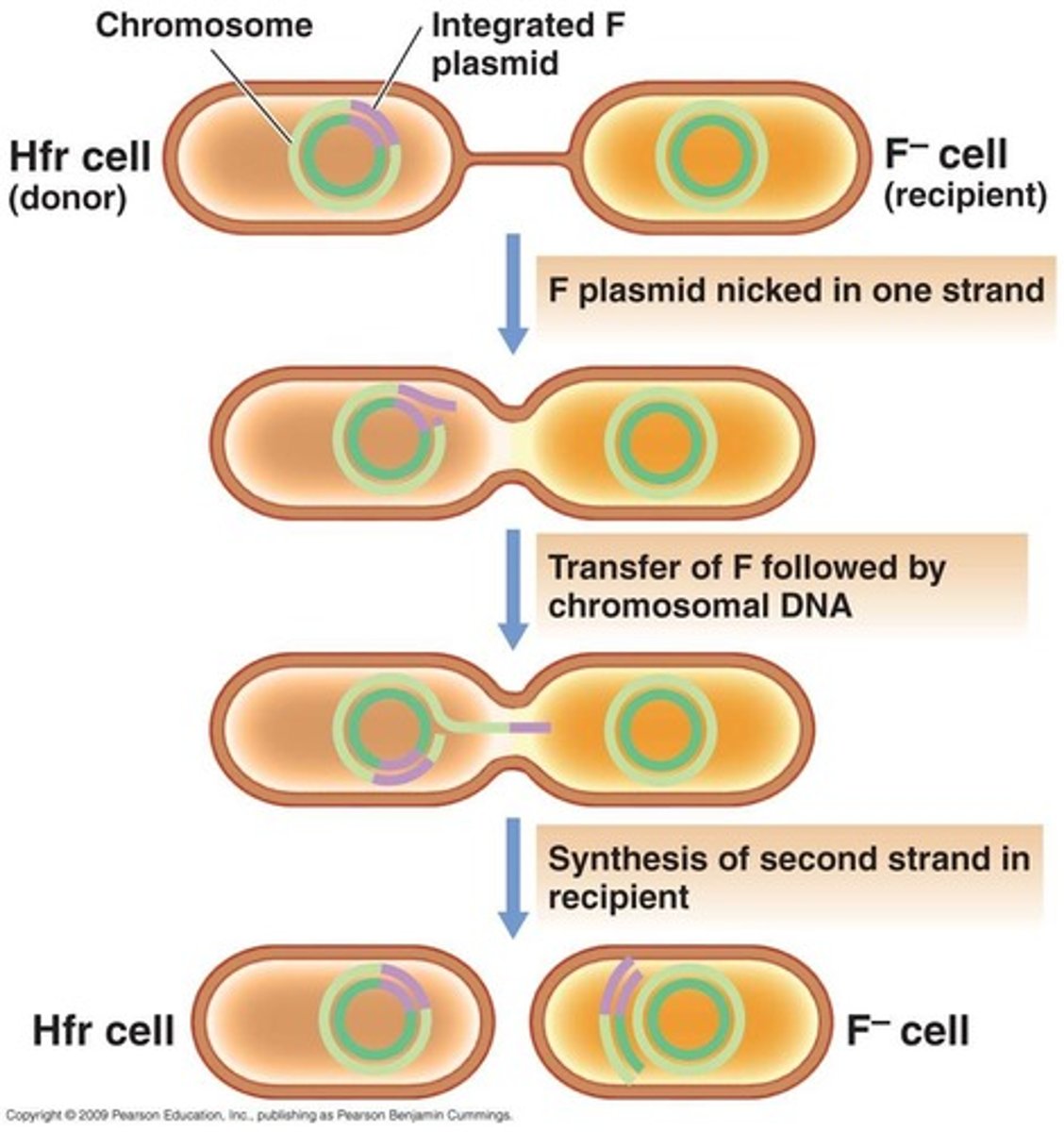
What is an advantage for a bacterium to carry the R factor?
This plasmid has the genes for resistance of multiple antibiotics , heavy metals, or virulence factors and this allows for the transfer of drug resistance between bacteria.
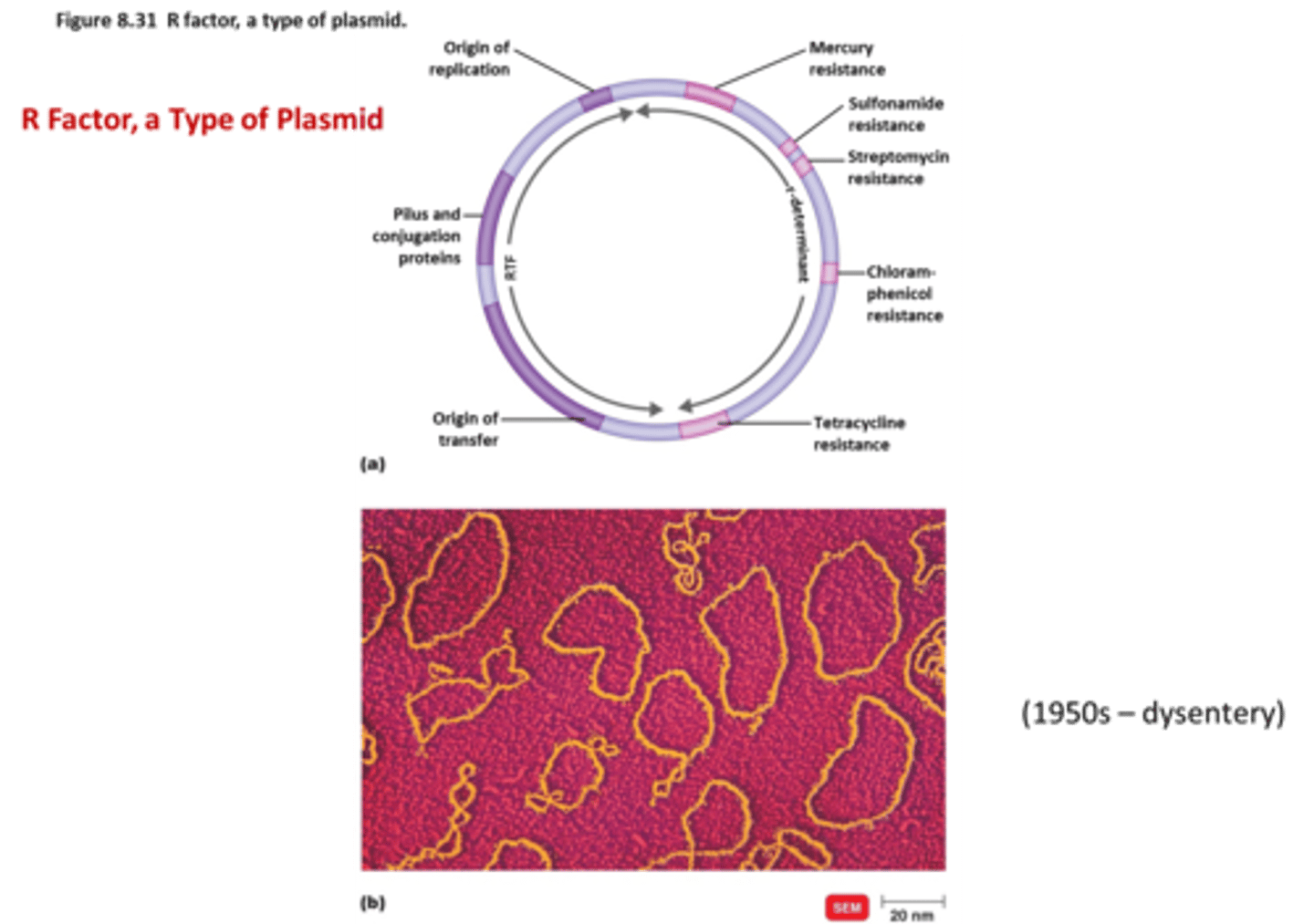
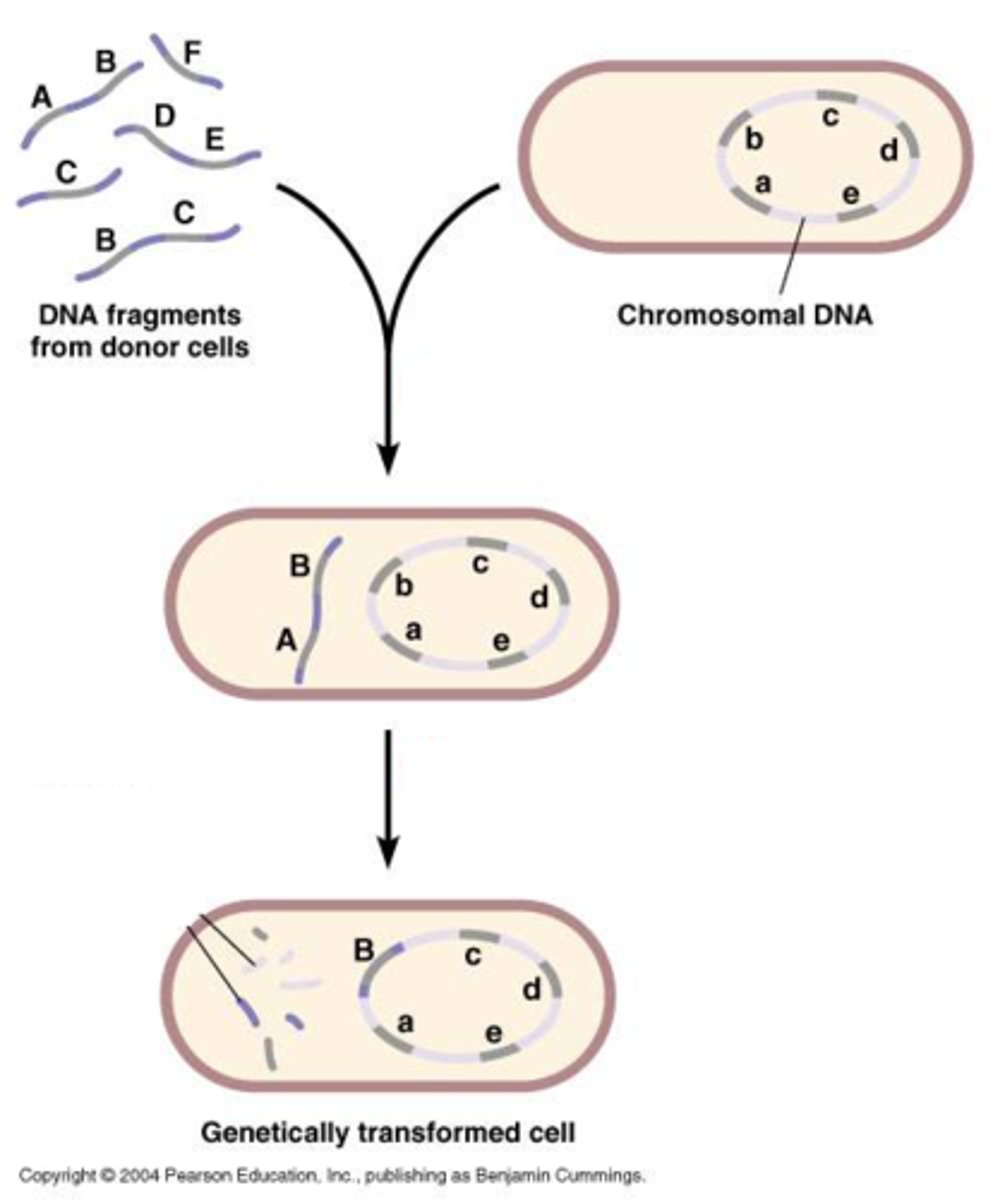
Can any bacterium take up DNA through transformation?
No only cells that are competent (have the proteins to allow DNA to cross the membrane) can transform DNA
Some bacteria can be treated in the lab to become competent but in nature only a few bacteria can take up DNA
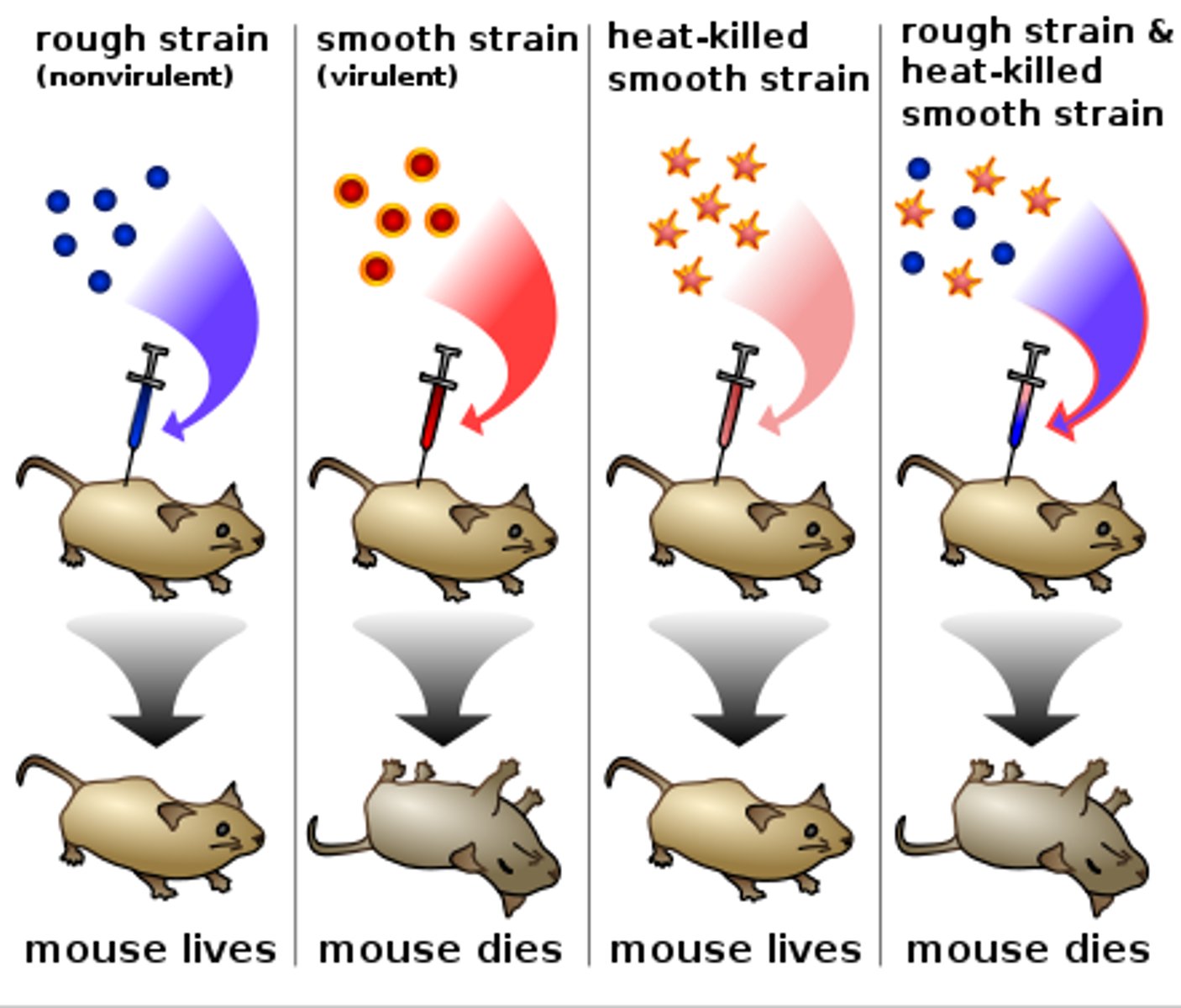
Understand the basics of the Griffith experiment.
Streptococcus pneumoniae in mice
Hypothesis: Dead matter (i.e. DNA) can transform living cells.
S type cells can kill mice, R type cells cannot kill mice
Sample containing heat killed S type bacterial cells (Have a capsule) was added to live R type bacterial cells (Have no capsule) resulted in S type cells that kill mice
This is due to R type cells using the dead S type cells DNA which includes genes for capsules
Two types of transduction
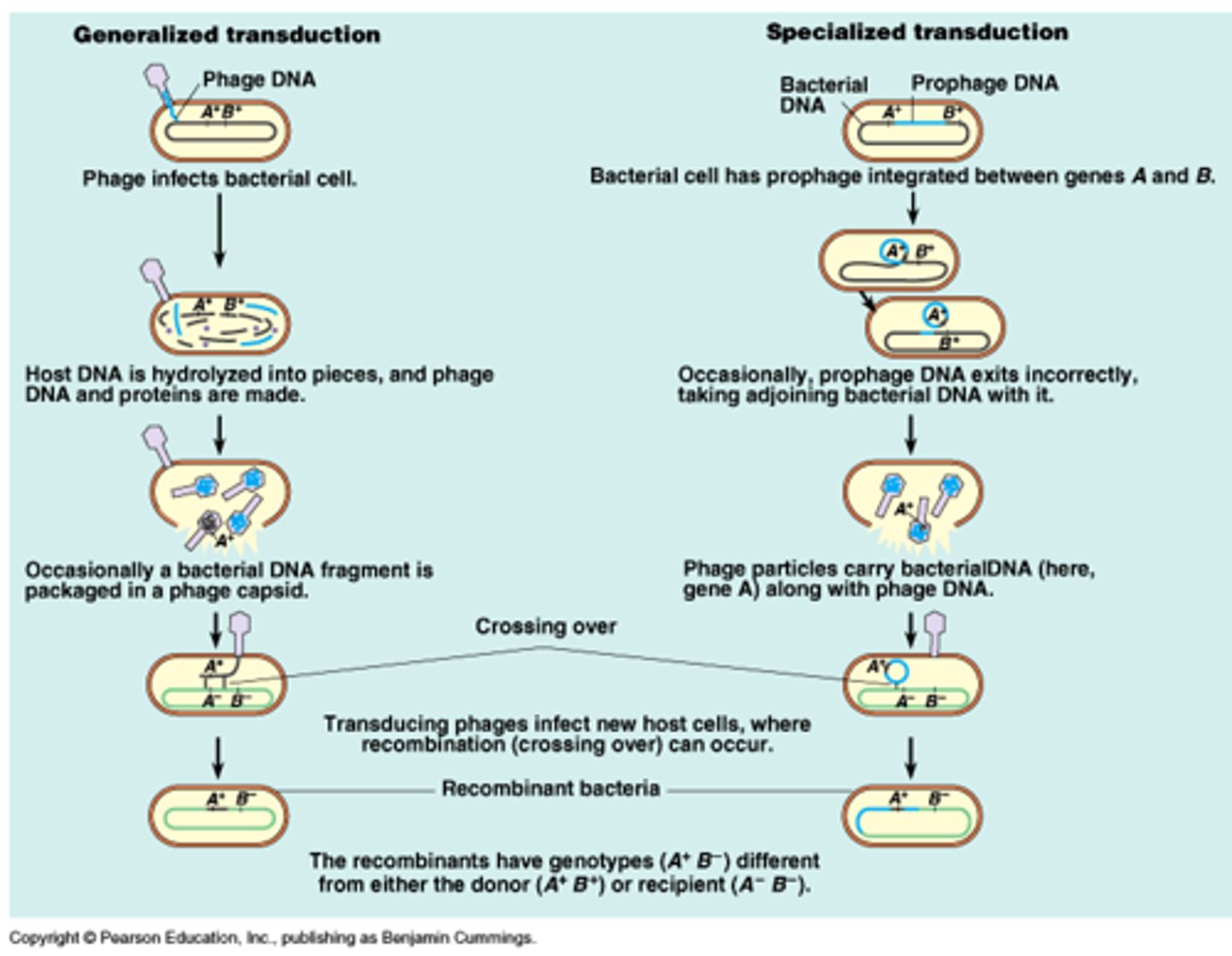
Generalized transduction and specialized transduction
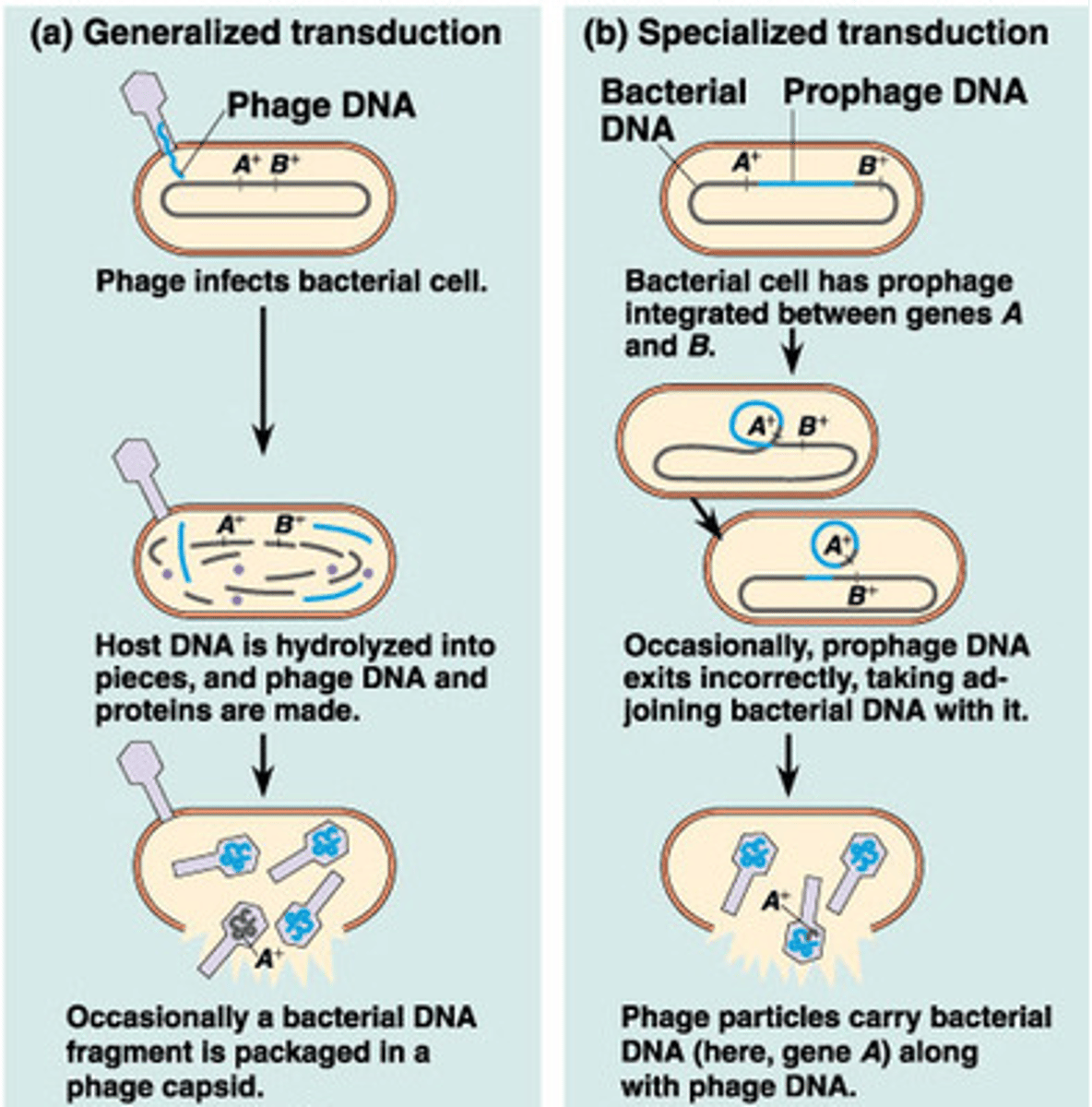
Differences between the two types of transduction
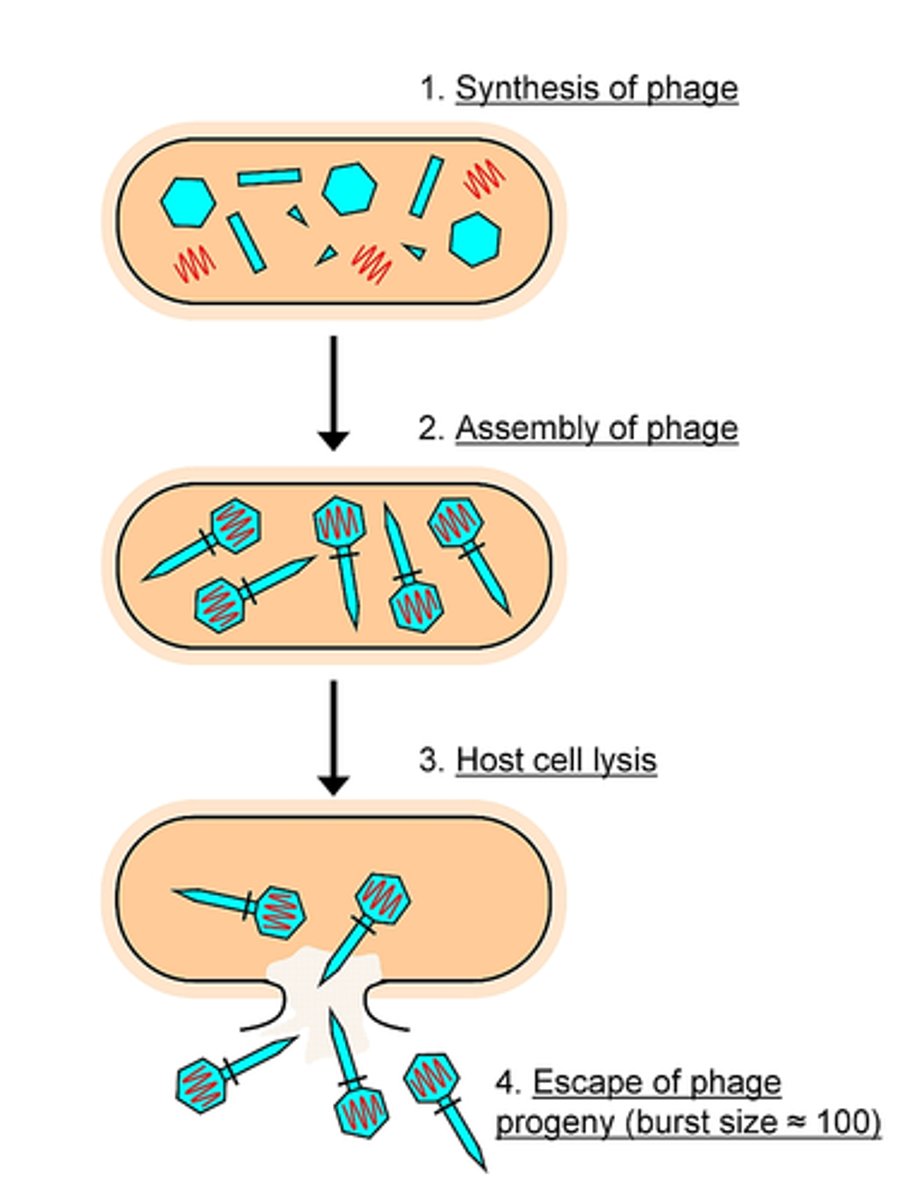
Generalized transduction: Lytic phages can pick up random fragments of disintegrating host DNA in their capsids during assembly; Any gene can be transmitted this way.
Specialized transduction: When lysogenic phages are induced, some times this results in excising not only part of the viral genome but part of the host cell genome that can be inserted into the capsid. A new host can insert the part bacteria/part virus DNA into their chromosome (seen in E. coli, Salmonella, S. aureus)
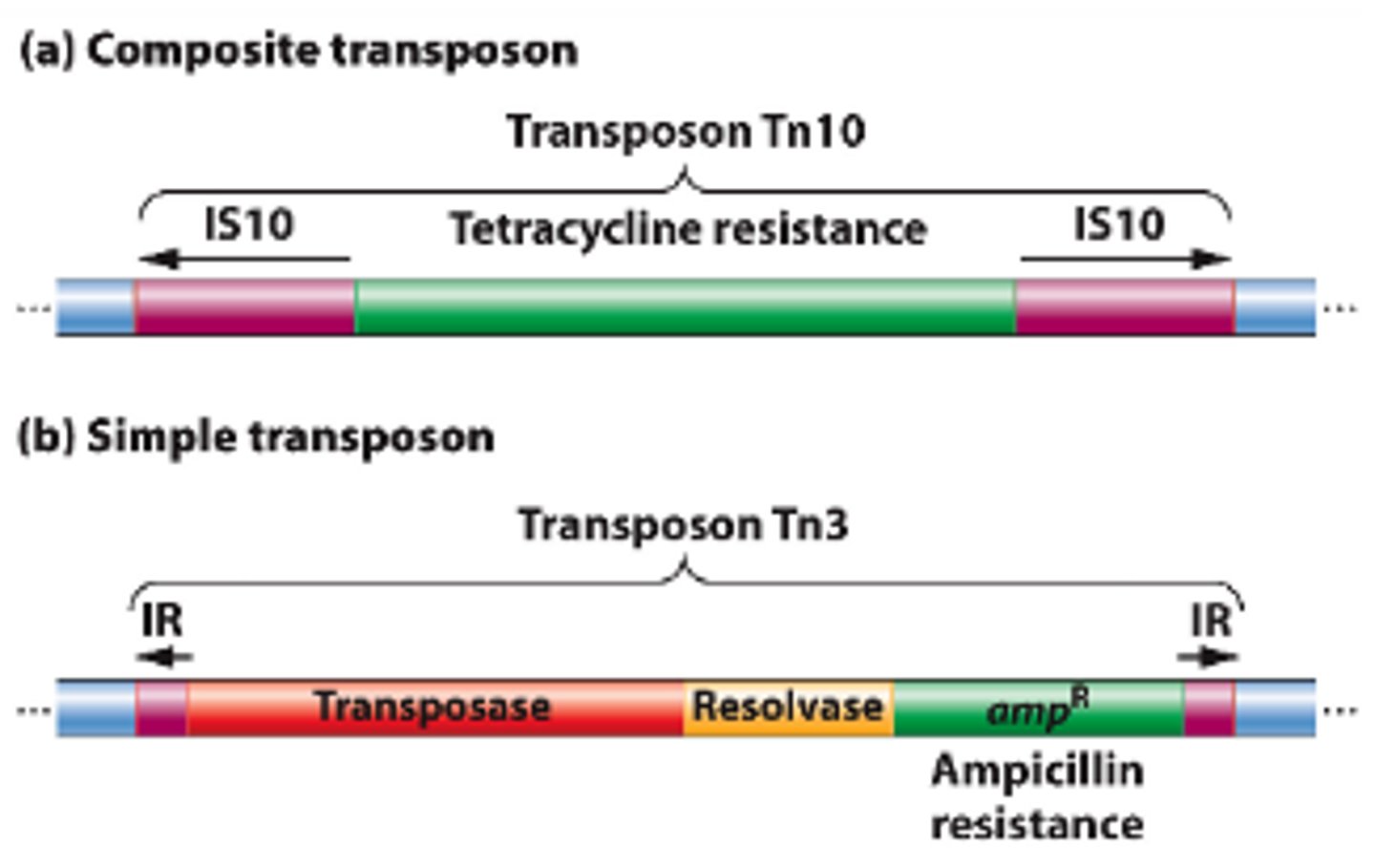
What are transposons?
Special DNA segments that have the capability of moving from one location in the genome to another - "jumping genes"
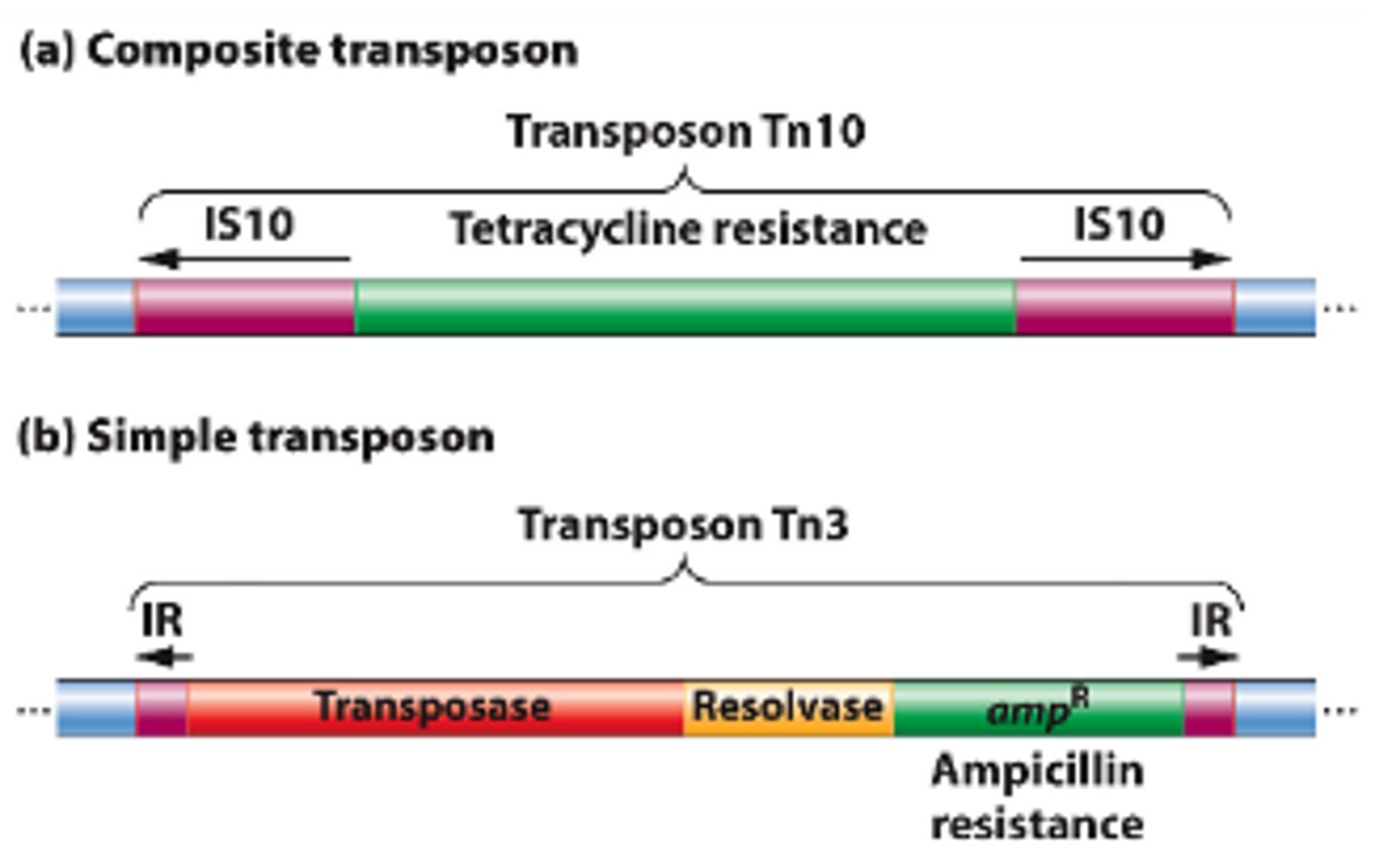
Why are transposon beneficial and harmful to bacteria?
Beneficial: Some transposons encode for antibiotic resistance and toxins for the bacterium
Harmful: Transposons can jump into genes and disrupt synthesis of those proteins
The case study: What made this virus different than other strains of the flu?
This strain had genetic material from pig, bird/avian and human flu
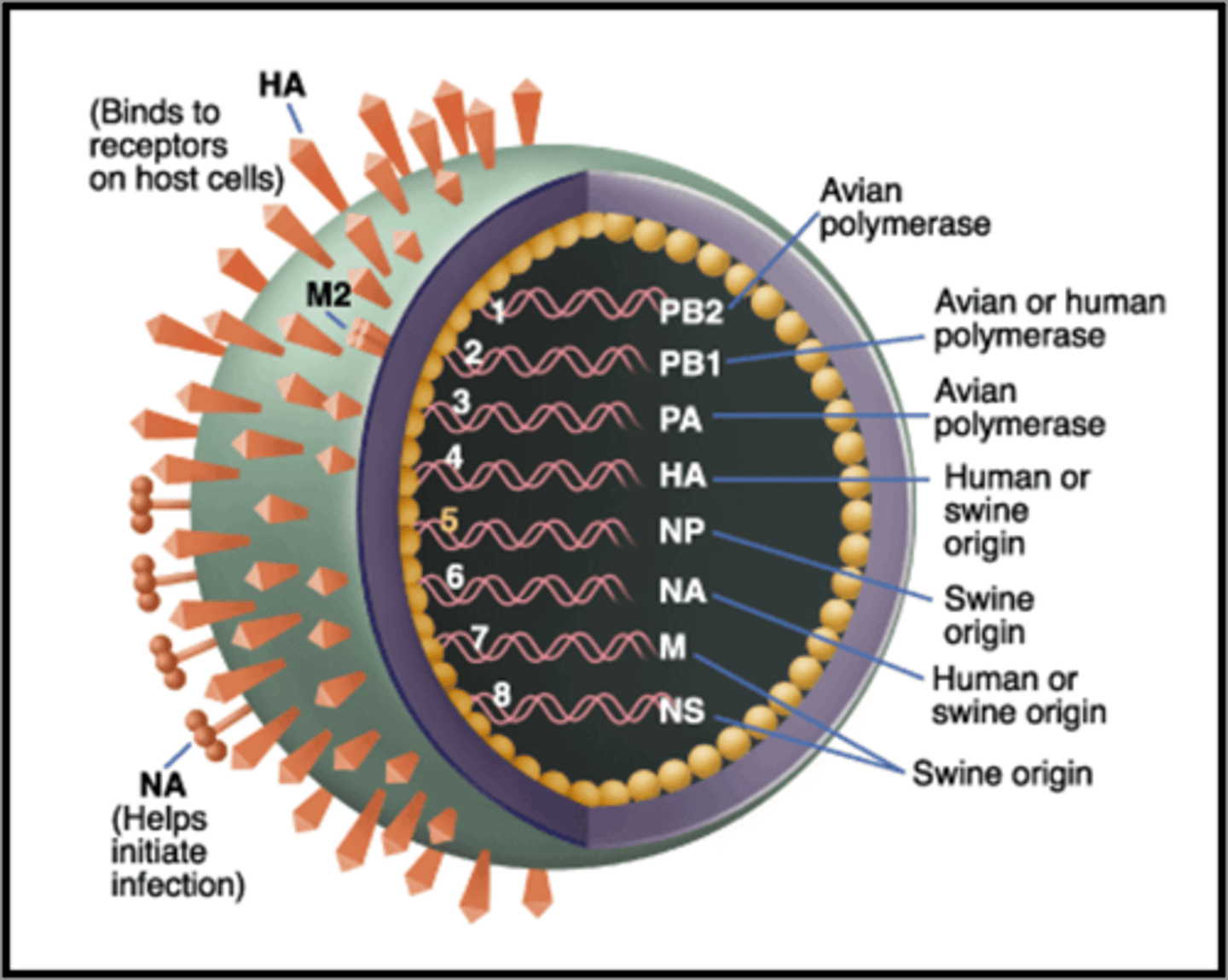
Why do we need a new flu vaccine every year?
The virus mutates so our immune system will only recognize previous flu strains.
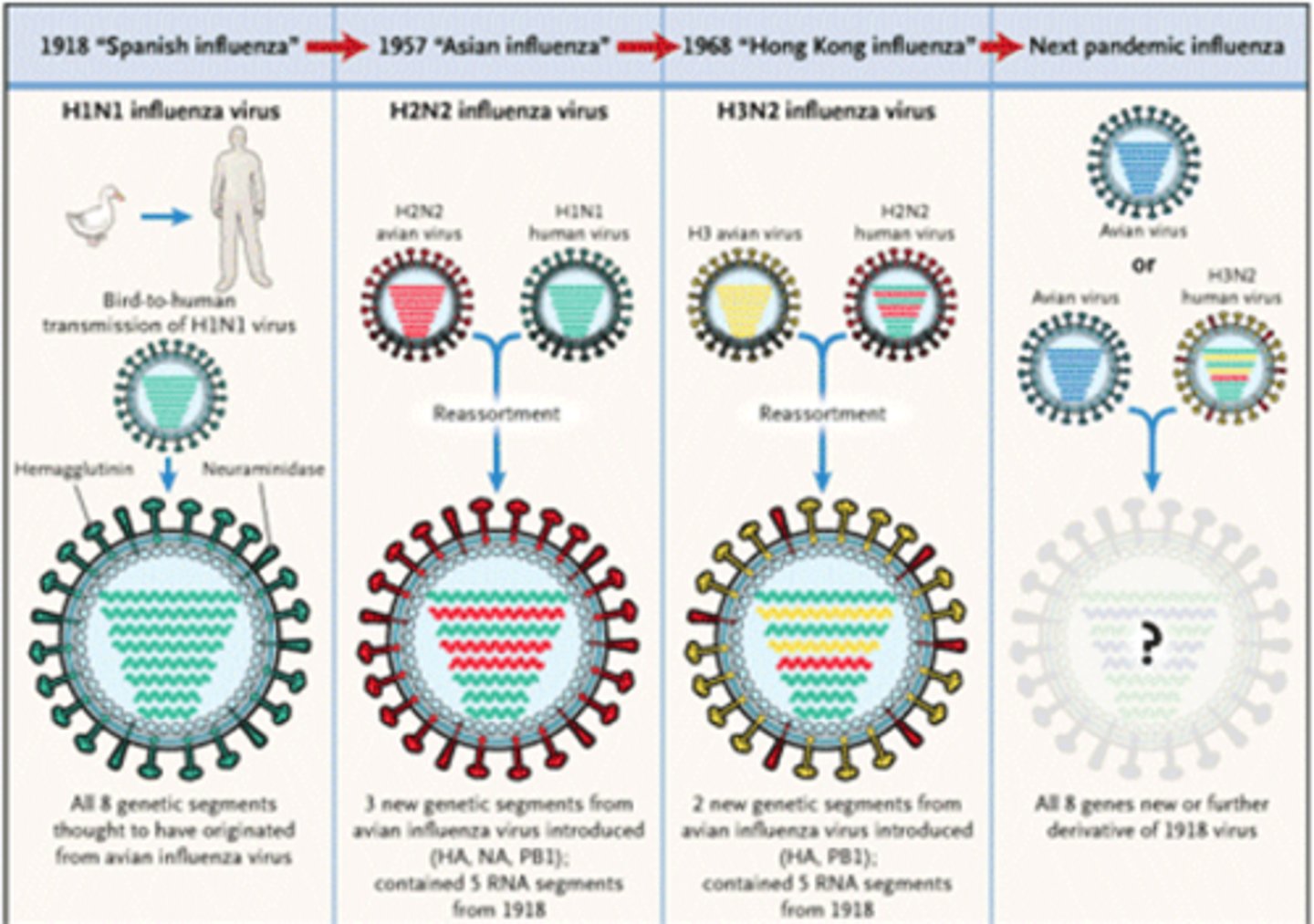
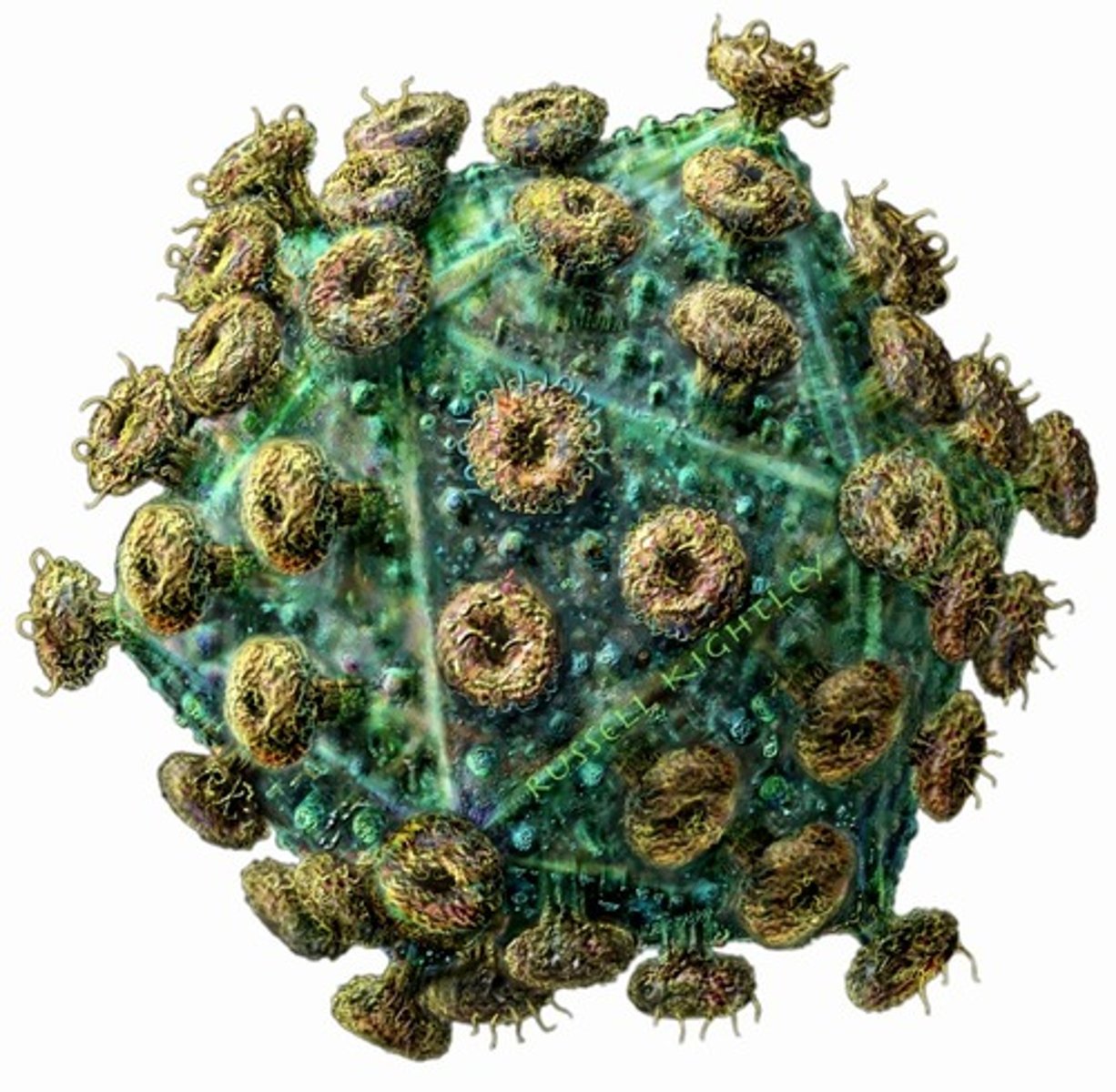
What role do the spike proteins play in the flu infection?
Viral spike proteins recognize particular proteins on the surface of certain host cells (including respiratory cells). These proteins help to determine which cells will be infected. The spikes serve as a means to attach to host cells.
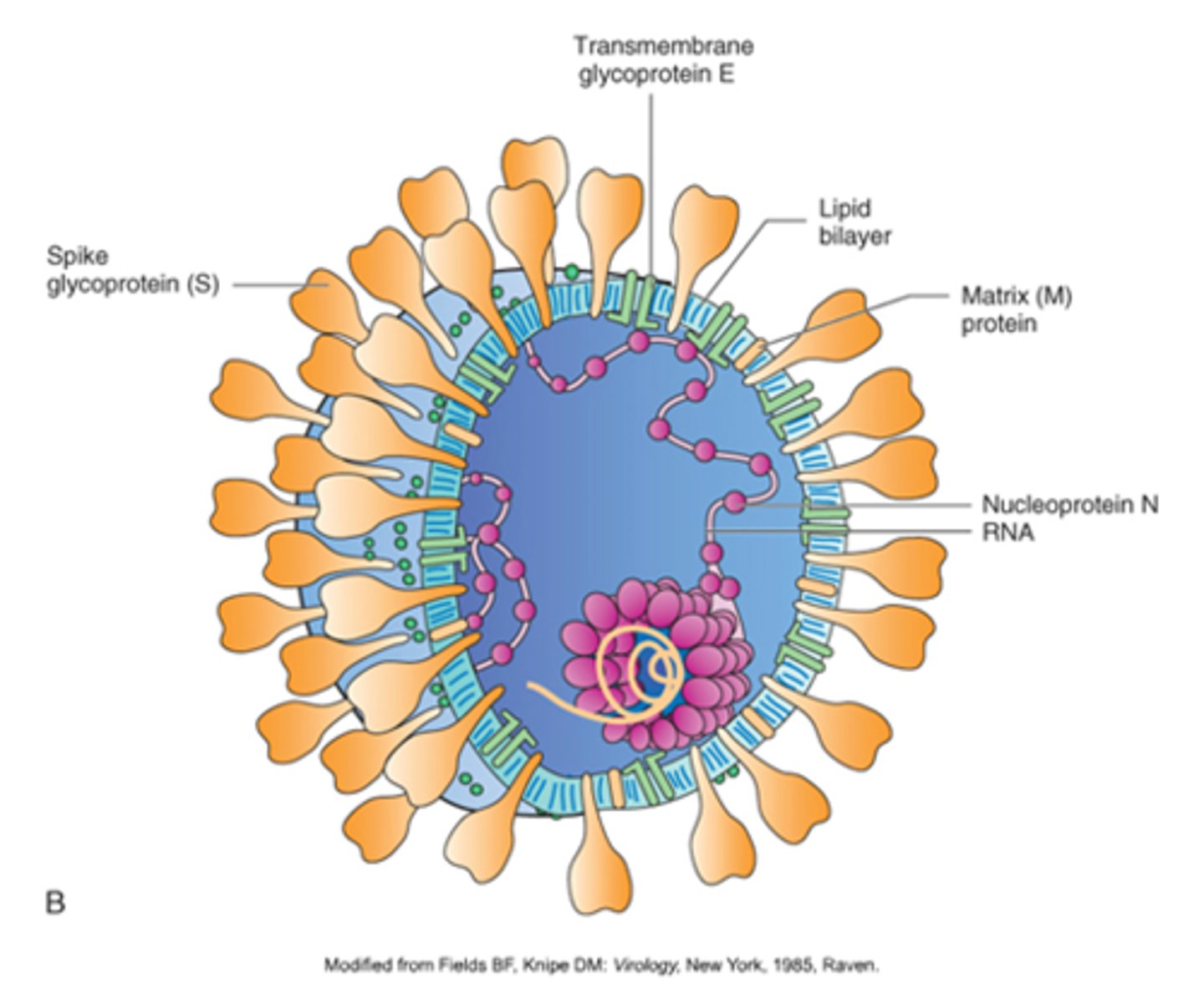
How can the flu be spread from animals to humans and vice versa?
The influenza virus recognizes proteins that are found in multiple animals like birds and pigs.

Why are viruses not considered "living things"?
Acellular (not made of cells)
Not able to exist without a host
What organisms cannot be infected by viruses?
None all organisms can be infected by viruses
Basic properties of viruses
Obligate intracellular parasites
Infect bacteria, protozoa, fungi, algae, plants, and animals
Ultramicroscopic size (nm)
Acellular
Can have DNA or RNA as genetic material not both

Why viruses are beneficial?
Vaccines, gene therapy, part of ecosystems
Why viruses are harmful?
Causes disease and cancers, gene mutations
Name the virus that is considered the link between viruses and cells.
Mimivirus

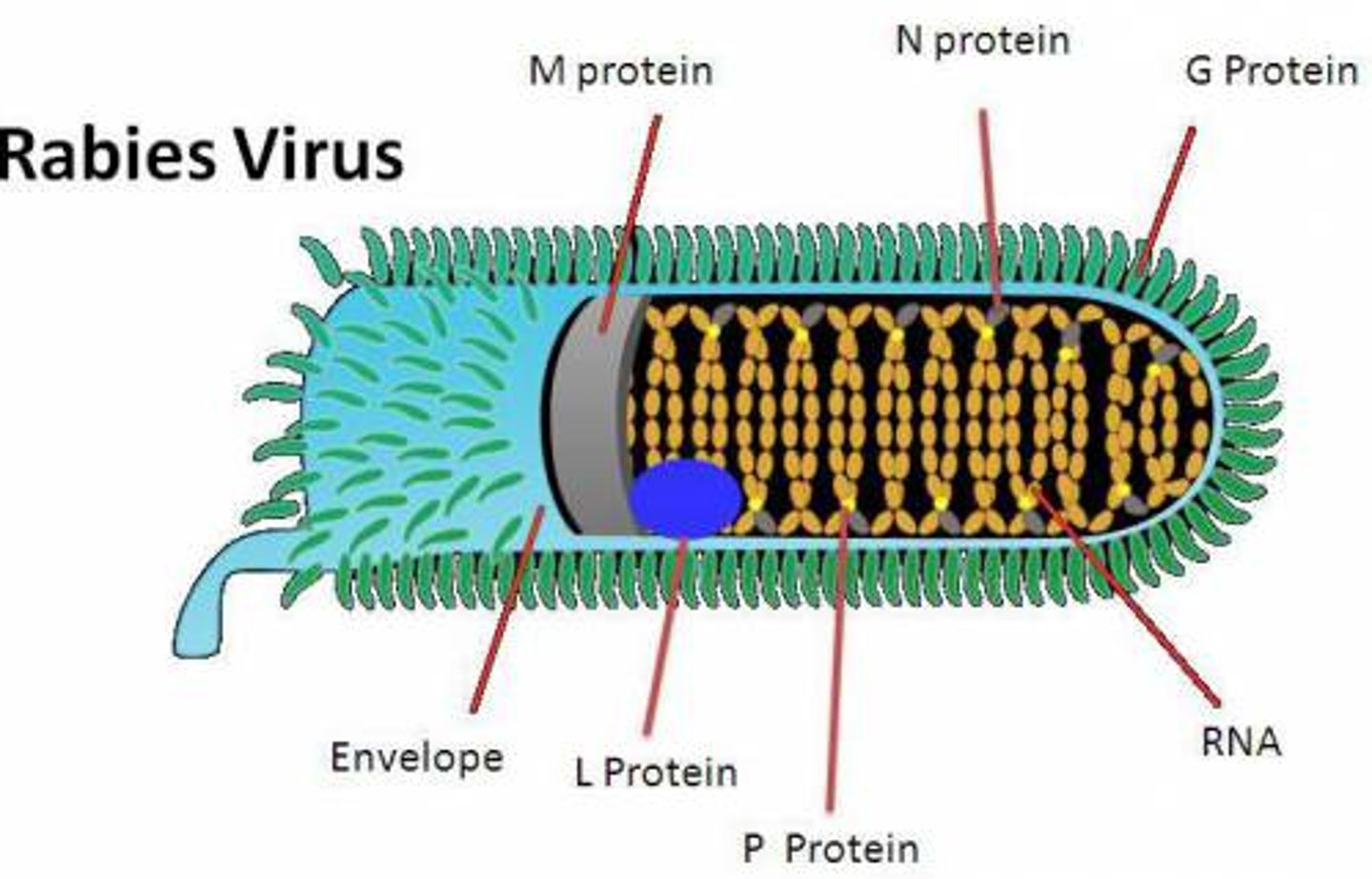
Name examples of enveloped helical viruses.
Avian flu, measles, rabies, Sars-CoV-2
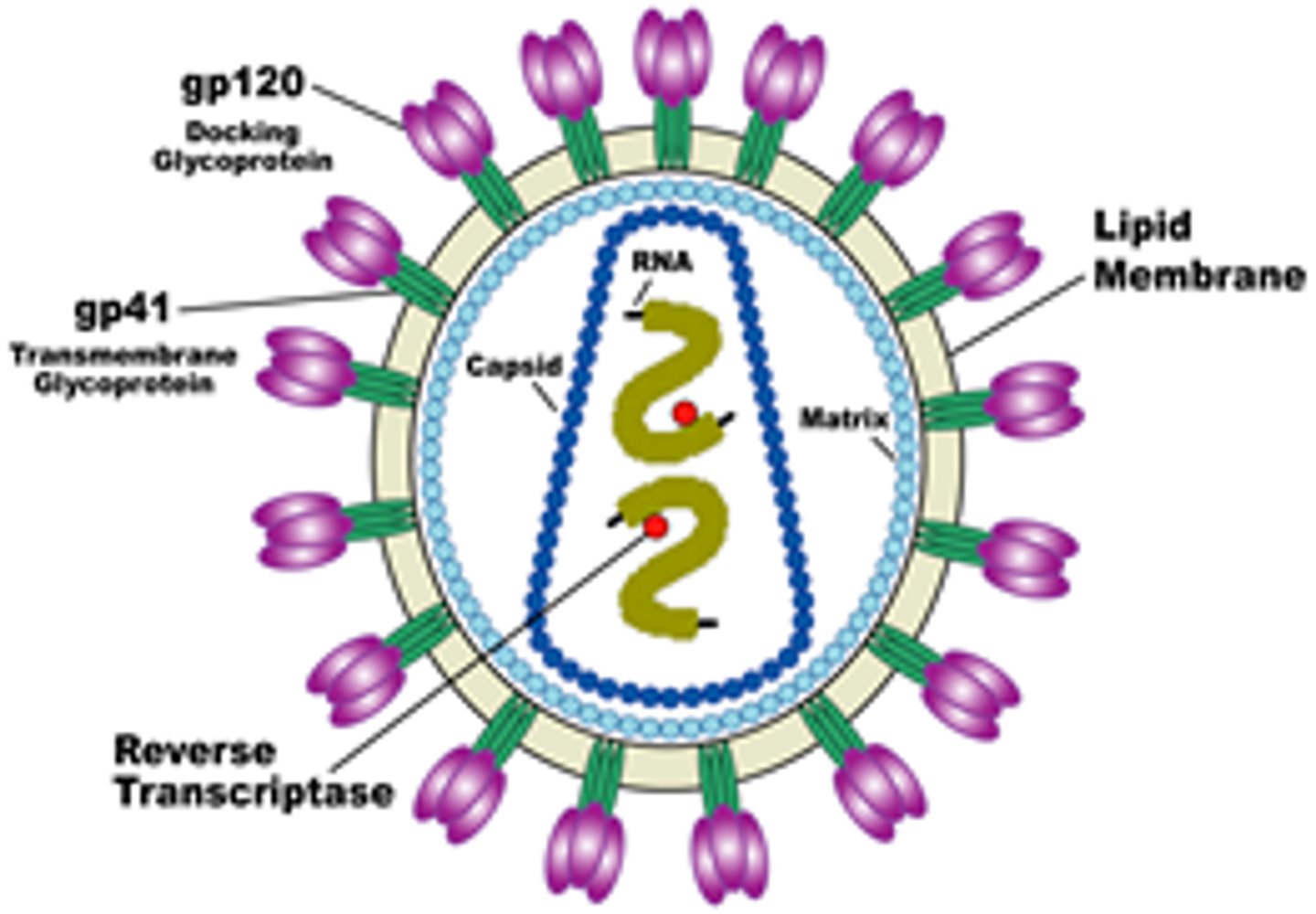
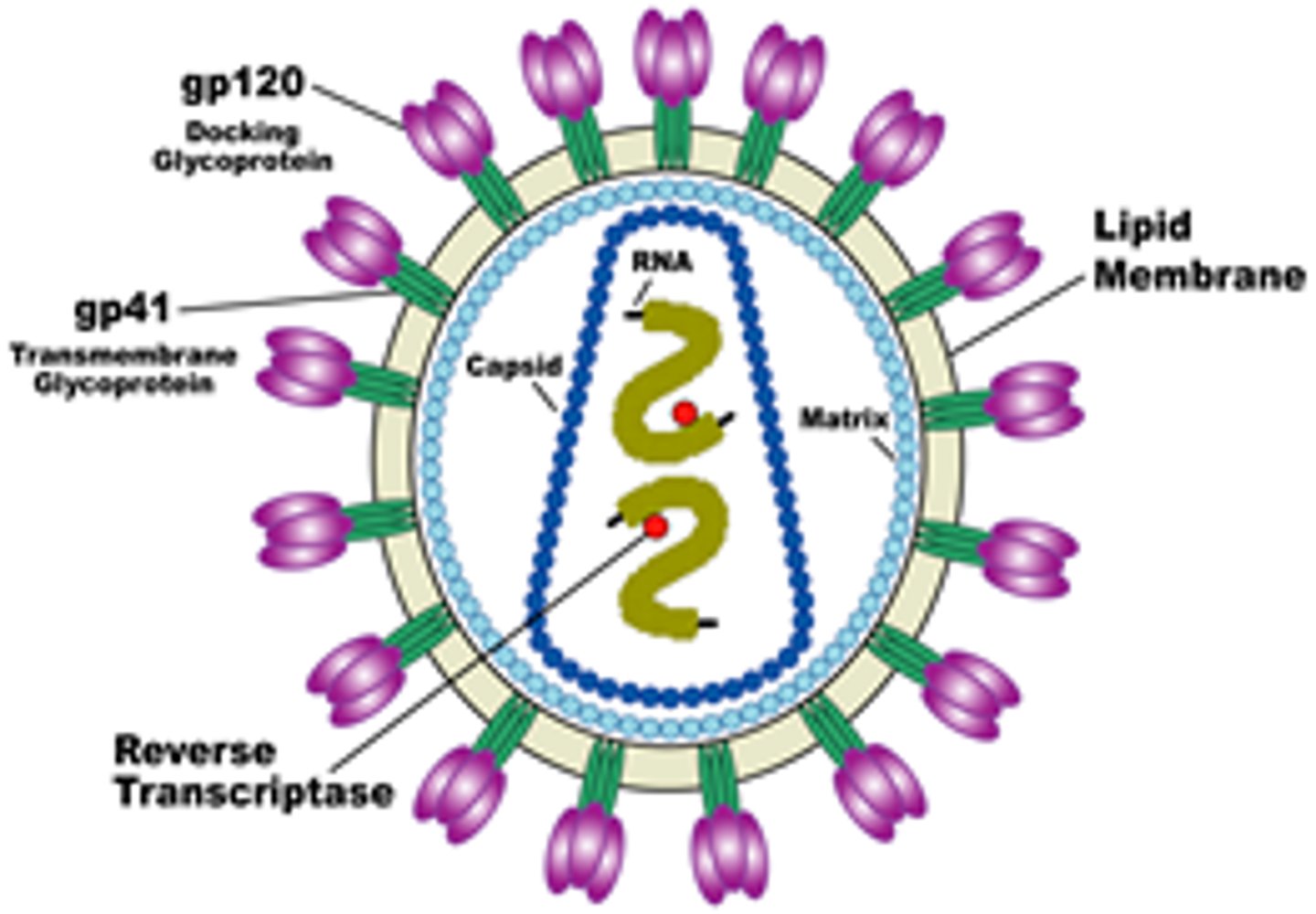
Name examples of enveloped icosahedral viruses.
Herpes, HIV