Gene-L8-DNA replication III
1/13
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
14 Terms
what types of errors occur during DNA replication and what are the consequences? (2)
mismatch
incorporating the wrong base and substitutions can lead to mutations
protein alterations can alter amino acid residue and can cause change in gene expression
slippage
in repitive DNA usually- where there’s polymerase switching from DNA pol alpha and then to DNA delta or epsilon- misalignment for polymerases
can cause insertions or deletions
how is high fidelity maintained during DNA replication? 3
complementary base pairing- ensures the correct watson-crick
proofreading- 3’ to 5’ exonuclease- DNA polymerase checks the last nucleotide. if incorrect, it pauses and removes it
post replication mismatch repair- occurs immediately after replication and detects and fixes remaining mismatches
how does complimentary base pairing ensure accuracy during DNA replication? what is it based on? what is the mechanism?
based on the correct geometry of the Watson-crick base pairs
correct pairs fit precisely into DNA polymerase activity site- A-G mismatch makes a wrong shape
mechanism
polymerase active site closes around the correct base pairing
it uses the amino acid residues to for a hydrogen bond
if mistached- the active site cannot close properly and there is not proper Hydrogen bonding and the polymerase pauses
it unwinds DNA about 8bp and strand shifts to 3’ to 5’ exonuclease site and removes the incorrect nucleotide by hydrolysis
error goes from 10^-3 to 10^-8 error rate
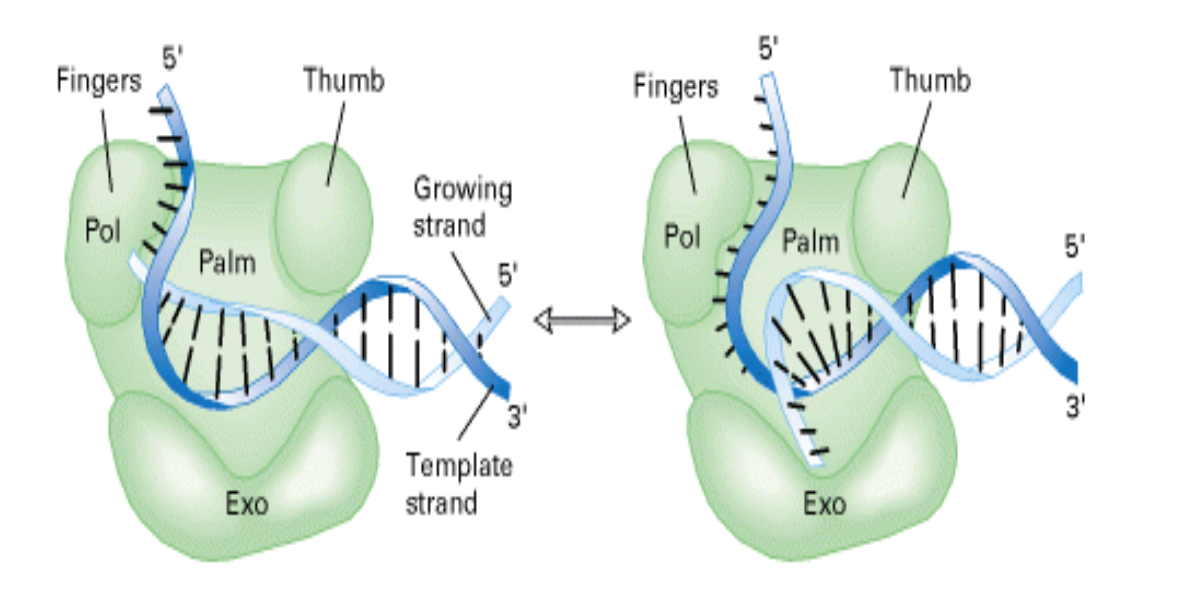
what happens if a mismatch isnt repaired after DNA replication? what happens if a C should have been added instead of a G
next round of replication can fix the error as a permanent mutation
outcome depends on which strand is used as a template though
if new strand- mutation propagates
if parental stand was used correctly- original sequence is retained
- If the old strand is repaired- then mutate persists
- If the new strand (with mutation) is repaired- the mutation stops
what is the mutHLS system? where is it present and what does it do? name the 3 components
mismatch repair system in e.coli
MutS: scans DNA for mismatches- distortions or bulges and recognises and binds to this error
mutL: is a linker or mediator and connects must to mutH
mutH- endonuclease and cleaves the newly synthesised strand
mismatch repair always targets the NEW strand
how does the MutHLS system identify the newly synthesised strand?
based on the DNA methylation at GATC strand
adenine is methylated in the parental strand
after replication the old strand is methylated and the new strand is temporarily unmethylated
mutH recognises hemimethylated DNA and costs the non methylated strand
leaves the parental strand intact
what are the steps of mismatch repair in e.coli? 8 steps- 4 sections
recognition:
mutS detects the hemimethylated DNA
mutL recruits mutH to the nearest GATC site
nick formation
mutH nicks the new unmethylated strand
excision
helicase unwinds the DNA
exonuclease 1 removes the section of new strand with the error
SSB proteins stabilise ssDNA
resynthesis and ligation
DNA pol III fills in the gap correctly
DNA ligase seals the remaining nice
what are the key components of the mismatch repair system in humans?
recognition by mutS homologous: MSH2, MSH3, MSH6 which form heterdimers and recognises different mismatches
mutL homologous- MLH/PMS proteins- MLH1/MLH3/PMS1/PMS2- act as mediators and regulators of repairs
haven’t found mutH homologs yet
how does the human MMR recognise the new strand?
recognition occurs before replication is fully complete
relies on nicks in newly synthesised DNA- many nicks in the Okazaki fragments and occasional nicks in leading strand
repair machinery target the nicked new strands
likely involves PCNA- sliding clamp protein and increases the adherence of polymerase to DNA
what are defects in MMR associated with?
associated with a deposition to colon cancer
Lynch syndrome- mutations in MLH1 and MSH2
what causes DNA damage and what are the types? what are they fixed by? 3 types
sources: internal and external
internal- oxidative stress, ROS,
external- UV, x-rays, chemicals and sunlight
types:
small base damage from oxidation, xrays- BER- base excision repair
bulky lesions- UV damage- NER nucleotide excision repair
crosslinks/ds breaks- homologous recombination repair
mechanism of nucleotide excision repair?
recognition
-XPC complex scans DNA for distortions and bulges
unwinding
recruits TFIIH transcription factor IIH
has helicase activity and unwinds DNA
stabilisation
RPA ssDNA binding proteins stabilise the open DNA
25bp bubble is formed
XPG helps stabilise this
excision
endonuclease XPG and others cut both sides
remove 24-32nucleotides
DNA pol fills the gap and DNA ligase seals the nick
how does BER base excision repair work? 5 steps what does it repair?
repairs small base damage from oxidation or deamination- deamination of 5-methycytosine to thymine GT mismatch
DNA gkycosylase recognises GT and removes the damaged base by cutting glycosylic bond. leaves an AP site
AP endonuclease APE1 cuts DNA backbone at 5’ side of AP site
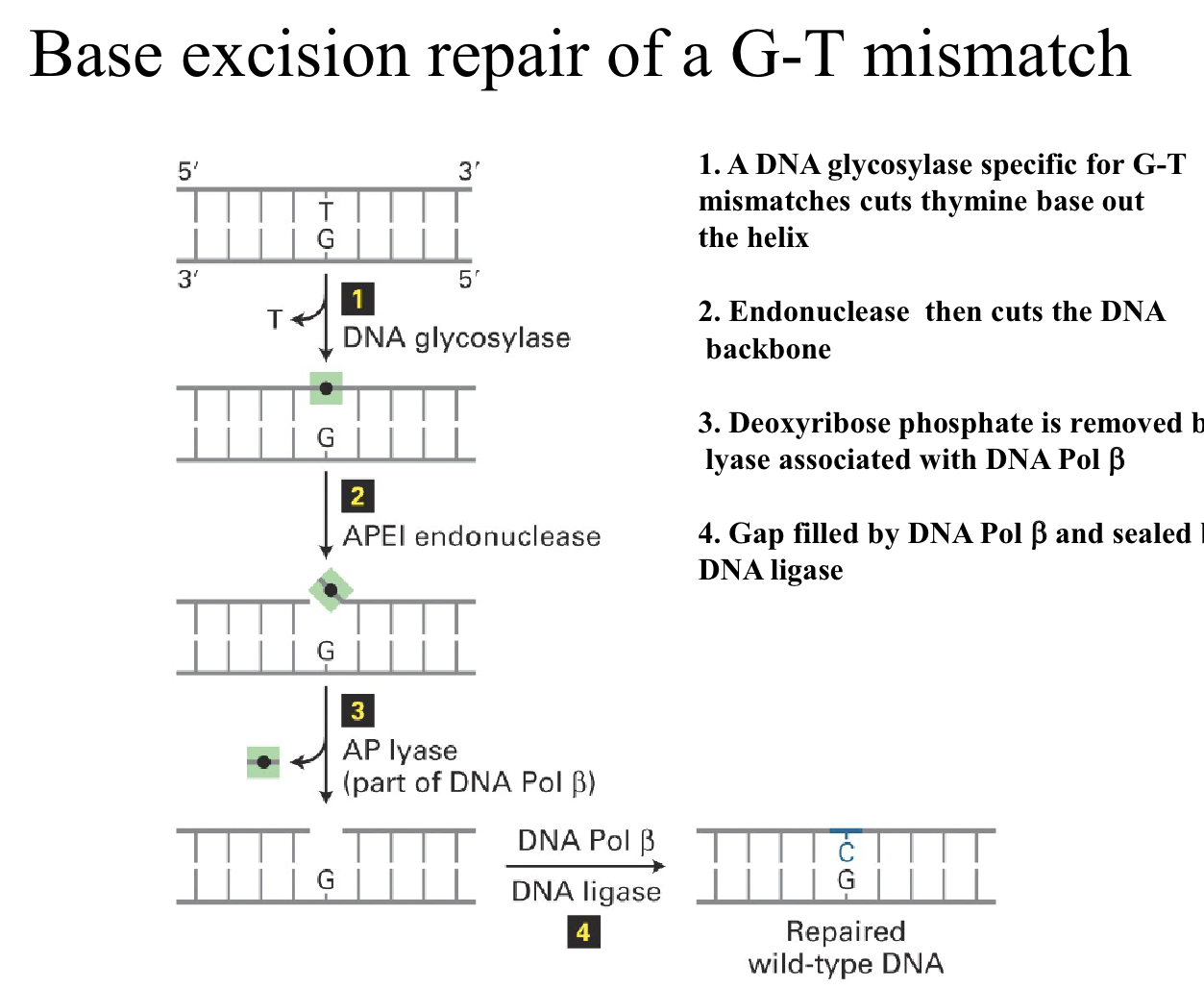
dRP removal- lyase activity with DNA pol beta removes sugar residue
DNA pol beta fills the gap
DNA ligase seals the nick
how do chemotherapy agents target DNA and why are cancer ells vulnerable? give an example
drugs induce DNA damage to kill cancer cells cells- cancer cells divide rapidly and have defective DNA repair pathways which makes them more sensitive to damage
cisplatin causes DNA crosslinks and nicks- disrupts replication and triggers apoptosis.