Dynamics and disorder
1/23
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
24 Terms
What timescale is bond stretching and angle bending?
ps-fs
what timescale is side chain/loop motion?
ns-ps
What timescale is folding of small peptides?
microseconds to nanoseconds
What timescale is protein folding and domain motion?
ms
How do disorder affect the protein?
sequence conservation, faster mutation rate and tolerates more mutations, enriched in eukaryotes vs prokaryotes
What is gibbs free energy determined by?
hydrophobic effect, hydrogen bonds, electrostatic interactions and conformational entropy
What is the order of least to most disorder protmoting amino acids?
W, F, Y, I, M, L, V, N, C, T, A, G, R, D, H, Q, K, S, E, P
What are properties of disordered protein-ligand binding?
lacks secondary or tertiary structure, highly flexible, conserved sequence, MoRFs and fly-casting as model, favours hydrophobic interactions, high specificity low affinity, enriched in eukaryotes, faster mutation rate
What are the properties of ordered protein ligand binding?
secondary, tertiary, Quaternary structure, favour polar interactions, segmented interface, small number of partners, high specificity, high affinity, no major structural change
what are MORFs?
molecular recognition features
how do disordered proteins flycast?
since greater capture radii, and chain flexibility, so disordered protein partially weakly binds to partner from large distance and reel in bindin partner while completing folding simultaneously enhancing binding speed
What is one-to-many signalling?
one protein is responsible for multiple transcriptional regulations
What is many-to-one signalling?
many amino acid polymers can bind to the same enzyme
How is binding driven in folded proteins?
enthalpy driven (H-bonds and electrostatic interactions)
why is binding weaker in disordered proteins?
They have to fold to entropy apposes enthalpy so low gibbs, so interactions are weaker
Why is specificity still high if binding is weaker?
since they are uncoupled
What are the mechanistic consequences of disordered-ligand binding?
multiple conformers, so different rates, the domination can also change with temperature
How can you detect disorder?
NMR, crystallography, circular dichroism, fluorescence spectroscopy, single molecule detection
How can crystallography detect disorder?
through lack of density, and a high B-factor indicates mobility and dynamics
How does circular dichroism detect disorder?
random coil shows disorder AKA CD signal vs wavelength plot, goes down then up and then plateaus
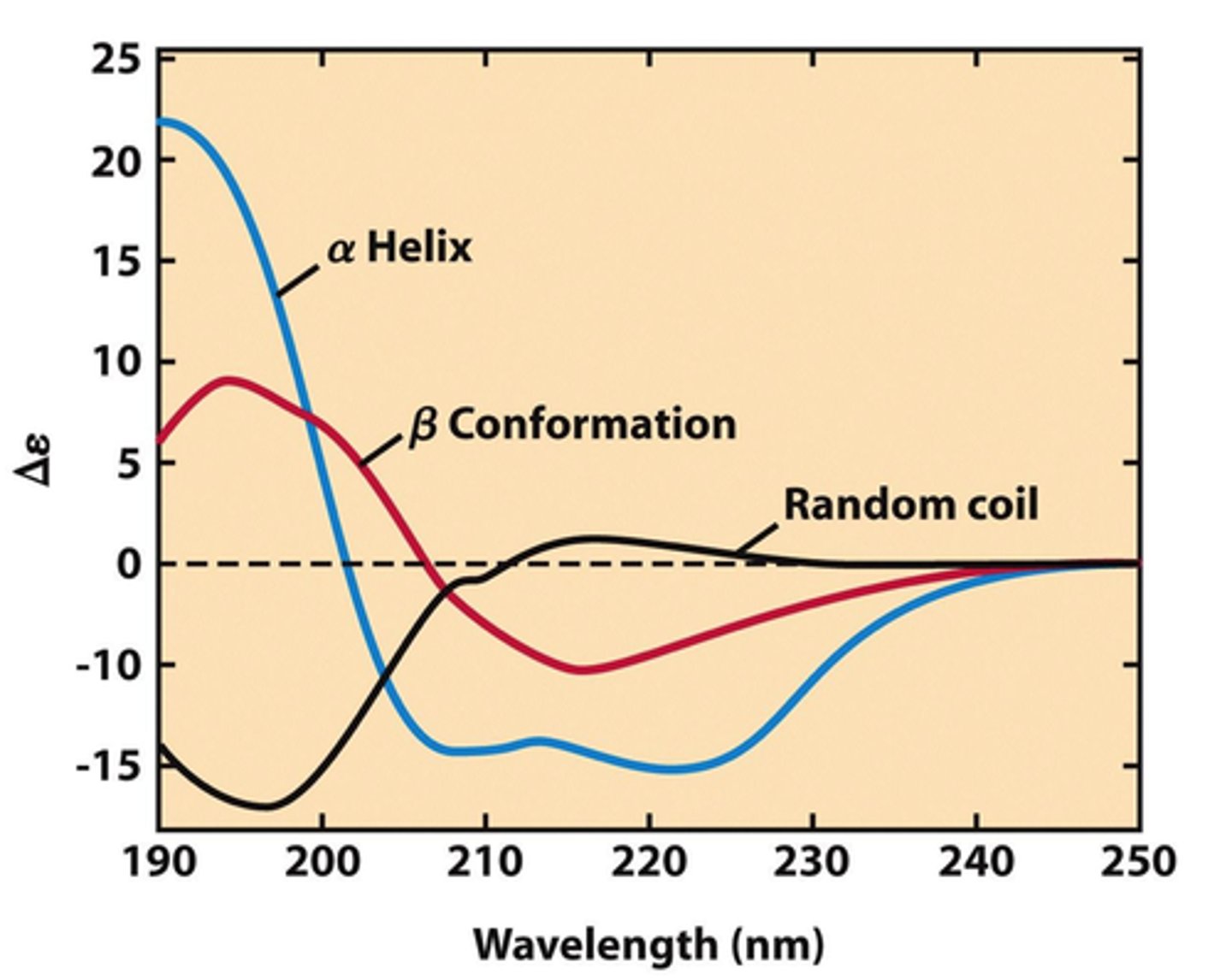
How does fluorescence spectroscopy detect disorder?
Use Trp fluorescence emission by looking for red shifts and intensity changes upon denaturation. REES can be used to report on equilibrium distribution of conformational states
How does NMR work for detecting disorder?
can label proteins with different isotopic labels and pick out specific interactions from specific residues to see how protein has changed
How does single molecule detection detect disorder?
low resolution so it doesn't tell you which parts are folded, but does show how many conformational states it can occupy and the interconversions of different energy levels.
What is a downside of using NMR for protein disorder detection?
isotropic labelling is very expensive and requires technical capability