Unit 2 - Gene Interactions
1/127
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
128 Terms
Nucleosome
Repeating subunit of chromatin in eukaryotic cells
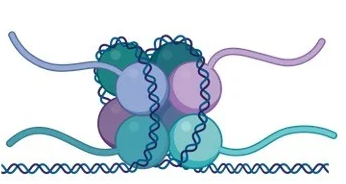
Heterochromatin
DNA tightly bound to nucleosome
F Plasmid
Has genes that assist with transfer of that plasmid to another host bacterial cell
R Plasmid
Has genes that confer antibiotic resistance
Lateral or Horizontal Gene Transfer
Transfer of genetic material between bacteria, archaea, other organisms
Conjugation
Transfer of replicated DNA from donor to recipient through pilus, single stranded plasmid is replicated as it leaves/enters
Transformation
Uptake DNA from environment
Transduction
Transfer of host DNA via a virus accidentally incorporating host genome
tra_
Genes that help transfer F plasmid
oriT
Origin of transfer
traA
Structural subunit of F pilus
Conjugation of F plasmid
F+ assembles pilus
Relaxosomes binds at oriT to cleave the T strand of DNA
Relaxosomes partially degrade, leaving relatase bound at 5’
Exporter move relaxase-T complex & recipient and replication begins
High Frequency Recombination
Bacterium with F plasmid integrated into main chromosome
Conversion of F to Hfr
Insertion sequence lines up and integrates into a giant chomosome
Hfr with F-
Donor bacterial chromosomal genes and maybe partial F plasmid genes are transferred to the recipient because pilus degrades
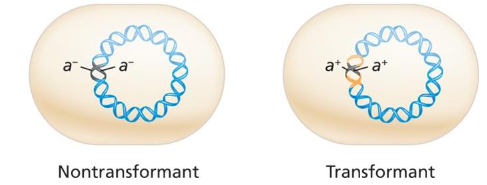
Transformation Steps
dsDNA enters at receptor where a strand is degraded
Transforming strand pairs with homologous region
Cell division makes a transformant and 1 nontransformant
Competent
Cell capable of being transformed
Transductant
Donor DNA integrated into recipient’s chromosome forms
Prophage
Bacteriophage integrated into a bacterial host chromosome
Genetic Complementation Analysis
Analyzing mutated bacteriophages with plaques to determine if it’s on the same or different genes
Even Plaques from Genetic Complementation Analysis
Mutation is on different genes that the bacteriophages complement each other
No Plaques from Genetic Complementation Analysis
Mutations are on the same genes, so no complementation
Abnormal Plaques from Genetic Complementation Analysis
Homologous recombination causes a copy of the gene to be normal
Major Grooves in DNA allows
Base pairs to be exposed from larger spacing
Conversative Replication
Synthesized DNA has no parental DNA
Semiconservative Replication
Synthesized strand has half parental and new
Dispersive Replication
Synthesizes strand has sections of parental
Messelson-Stahl Experiment
Replicated DNA in N15
Allowed 1 cycle of replication in N14 twice
Ultracentrifuged tubes and observed where the DNA suspended
Replisome
Complex of proteins at replication fork
Helicase
Unwinds double helix
DNA Topoisomerase
Relaxes supercoiling
SSB
Prevents strands from resealing
Primase
Synthesizes DNA primers
DNA Polymerase 3
Synthesizes DNA
DNA Polymerase 1
Removes and replaces RNA primers with DNA
DNA Ligase
Joins DNA segments
DNA Replication
Helicase breaks DNA and topoisomerase binds
SSB binds and primase synthesizes RNA primers
DNA poly 3 makes new strand
DNA poly 1 removes and replaces RNA priemrs
DNA ligase joins okazaki fragments
Base Pair Mismatch
DNA strand swings down and 3’ to 5’ exonuclease cleaves off the base pairs at the 3’ end
Telomeres
Repetitive sequences that ensures important information isn’t lost
Chromatin Memory
Turns off genes are immediately remembered to be turned off once replicated finishes
Telomere Synthesis
Telomerase attaches to empty strand where primers were removed
RNA template grows leading strand in a repetitive sequence
DNA poly extends other strand
Hanging strand knots upon itself
Ribonucleoprotein
Enzymatic complex that has protein and ribonucleic acid
Hayflict Limit
Number of times somatic cells can divide, around 50-70
PCR
Uses temperatures to denature DNA, DNA primers and Taq polymerase replicates DNA
Thermus Aquaticus
Bacteria that tolerates high temperatures and has Taq polymerase
PCR Ingredients
DNA template
dNTPs
Taq polymerase
2 DNA primers
Buffer
Sanger Sequencing
Determines nucleotide sequence by separately using a single type of ddNA and combining them together
Dideoxynucleotide
Lacks 3’ OH, which stops DNA synthesis
Sanger Sequencing Integredients
DNA strand
DNA primers
DNA polymerase
DNA & dDNA
Capillary Electrophoresis
Uses a tube and a camera to capture fluorescently tagged dDNA as they cross a detector
Coding Strand
Non synthesized strands that’s identical to RNA made
mRNA
Encodes sequence of amino acids may be polycistronic in bacteria and archaea
rRNA
Helps large and small subunits form
small nuclear RNA
In eukaryotic nuclei where many snRNA joins with proteins to make spliceosomes
microRNA
Eukaryotic regulatory RNAs that base pair with mRNAs, changing stability and efficiency of translation
small interfering RNA
Eukaryotic regulatory RNA made from long ds molecules that are cut into smaller pieces to regulated stability and translation
Telomerase RNA
In telomerase, ribonucleoproteins complex that acts a template to maintain and elongate telomere length of chromosome
Consensus Sequence
Most common sequence (-35 and -10)
Pribnow Box
-10 consensus sequence AT rich in prokaryotes
RNA Poly Holoenzyme
Core RNAp and specific signma subunit
Sigma Subunit
Guides RNA poly to promoters, and different versions changes binding specificity
Synthesis is in the
5’ to 3’
Hairpin Loop
RNA folds back on itself from GC rich areas binding strongly, followed by weaker AU regions forcing polymerase off
Intrinsic Bacterial Transcription Termination
Hairpin loop forms
Rho-Dependent Bacterial Transcription Termination
Hairpin + ATP active protein forces unbinding
Euchromatin
Lightly packed, open chromatin for gene expression
RNA Polymerase 1
Several ribosomal RNA genes
RNA Polymerase 2
Protein coding genes and most small nuclear RNA genes
RNA Polymerase 3
tRNA genes, small nuclear RNA genes, ribosomal RNA gene
Saturation Mutagenesis
Individually mutate every base pair and areas where transcriptions decrease, it’s a promotoer
Band Shift Assay
Combine a section of DNA with a promotor binder, if a promotor binds, it’ll be slower in a gel and prolly a promotor
DNA footprint Protection Assay
DNA is incorporated with DNase that’ll randomly cut DNA but promotor bound with be protected causing a gap in gel
Enhancer Sequence
Increases level of transcription of specific genes by bringing certain gene promotors close to RNA polymerase to bind
Activator Protein
Binds to enhancer to bring complex to RNA polymerase II
5’ Capping
Methylated guanine added to 5’ end of the first 20-30 mRNA made to stop degradation and increases translation efficiency
3’ Polyadenylation
Poly A tail replaces some mRA at the 3’ to end transcription
Group I and II Introns
Self splicing, chloroplast, mitochondria related
Pre mRNA Introns
Requires splicesomes
5’ Splice Site
Beginning of intron with GU nucleotides
3’ Splice Site
End of introns with AG nucleotides
Branch Site
Conserved sequence near the end of introns connecting 5’ intron end to branch point A within branch site recognition sequence
Branch Point Adenine
A nucleotide within branch site
Polypyrimidine Tract
Sequence of pyrimindines near the end of to promote spliceosome aseembly
Intron Recognition and Cleavage
snRUP U1 binds to 5’ and U2 binds to branch site
snRUP U4, U5, U6 binds forming inactivate splicesome
U4 dissociates, activating complex to cleave 5’ end
2’ to 5’ bond form with branch point A stabilizing lariat intron
3’ splice creates a free 3’ OH on exon 1, freeing the lariat
Alternative Promoters
Different transcription start points are different for cell types
Alternative Polyadenylation
Different ending sites
Alternative pre-mRNA Splicing
Constitutive splicing
Exon skipping
Intron retention
Mutually exclusion exons
Constitutive Splicing
All introns removed
Exon Skipping
Some exons are skipped
Intron Retention
Some introns retained for regulation
Mutually Exclusive Exons
Different exons are combined
Calcitonin
Derived from CALCA with exon 4 retained
CGRP
Derived from CALCA without exon 4
Species with introns tends to have
Small variable breeding population (Ne), causing weak natural selection from stronger genetic drift
rRNA and tRNA are
Transcribed, but not translated
E. Coli
7 pre RNA genes, each with the same rRNA genes but different tRNA all in 1 transcript
rRNA and tRNA in Eukaryotes
From different pre RNA transcripts
ETS
External transcribed spacer
ITS
Internal transcribed spacer
tRNA Processing
5’ and 3’ ends trimmed
tRNA folds into 3D structure
Addition of bases