Non-Homologous End Joining
1/14
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
15 Terms
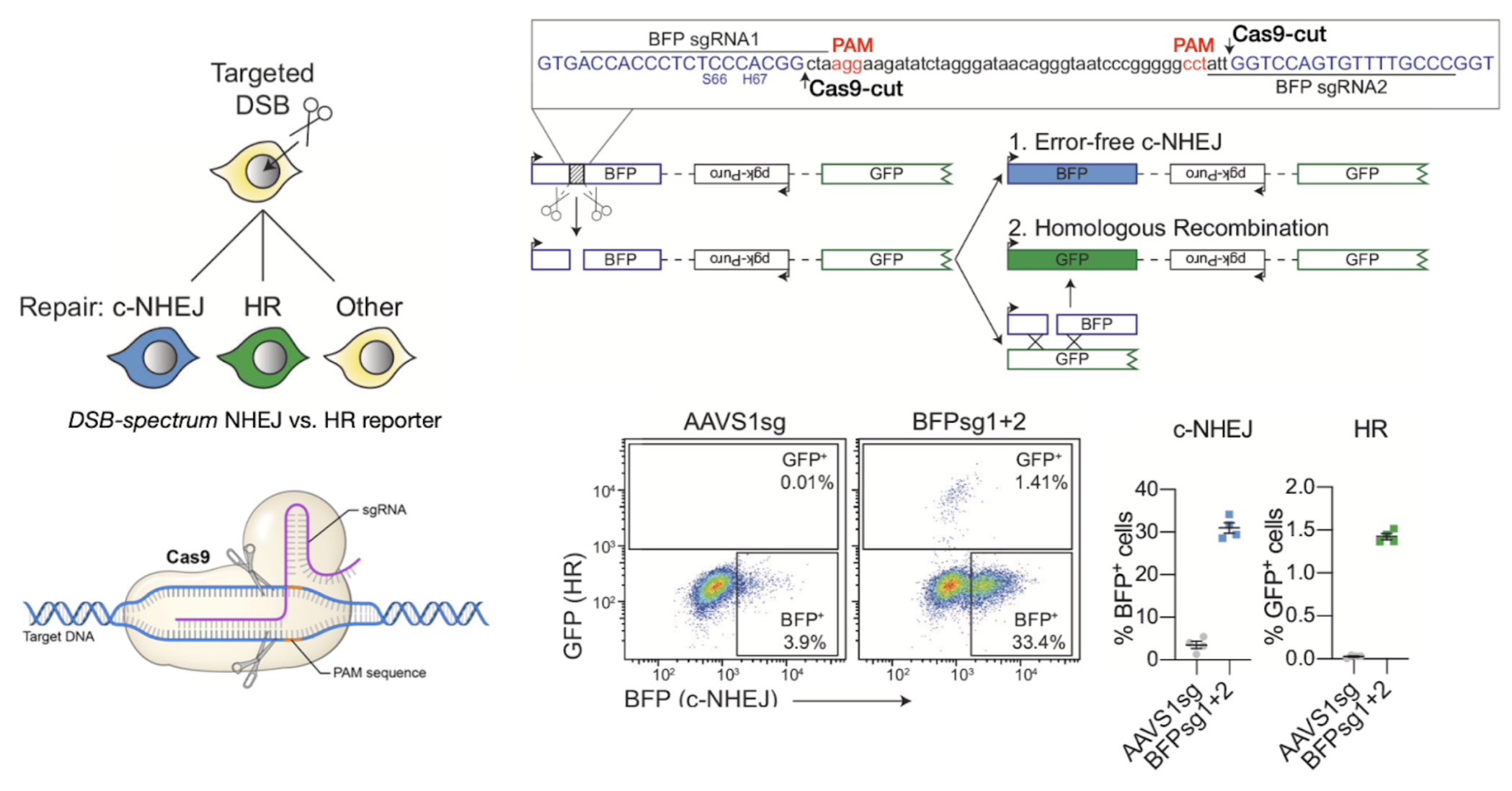
How did a reporter assay confirm the presence of NHEJ in cells with CRISPR induced DSBs?
Genomically integrated reporter contracts allow disrimination between NHEJ and HR repair of DSBs.
Large proportion of error-free, blue NHEJ cells observed
confirmed a secondary DSB repair mechanism alongside HR
Error-prone NHEJ not examined
How was the presence of NHEJ confirmed in the 80s? What did individual NHEJ/HR deficient cell experiments show about these pathways interactions?
In 80s, irradiated, HR-deficient cells still exhibited repair - confirms presence of an alternative DSB repair pathway
NHEJ and HR deficient independent cell lines did not show comparable survival to control - these pathways cannot fully cover for one another
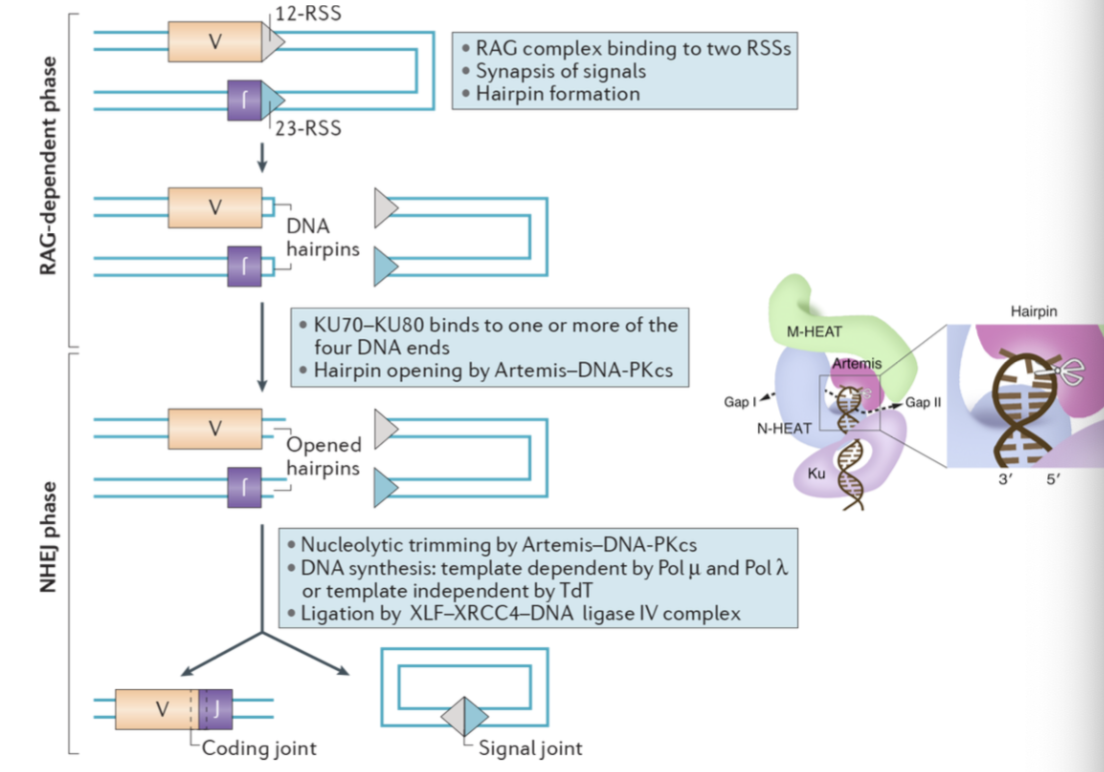
What are the three steps of NHEJ, and the main associated proteins? Which step has multiple, varying protein requirements? Give an example of one of these proteins.
End binding - Ku70/80
End processing - DNAPKcs
End-ligation - XRCC4-Lig4-XLF
Depending on the constitution of DNA ends, various end processing molecules may be required - based on repair needs
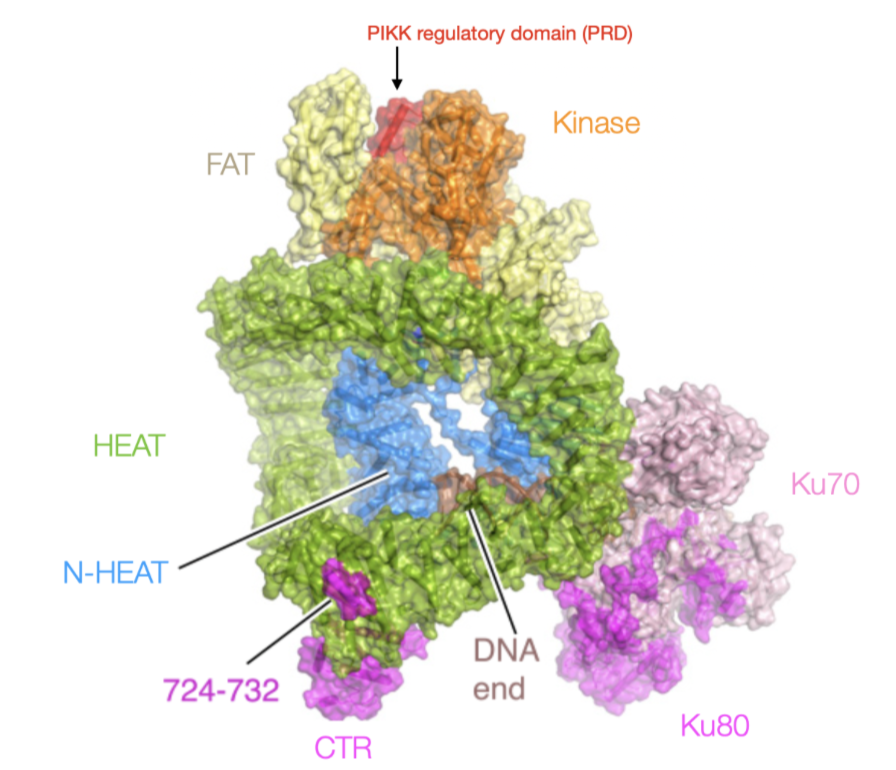
How does Ku binding initiate NHEJ? How is DNA-PKcs activated to promote end processing?
Ku70-80 dimer binds DSB ends - facilitates assembly of the repairosome - all other cofactors in NHEJ have Ku binding domains
no enzymatic activity, recruits enzymes
DNA-PKcs form a DNA dependent kinase with Ku70-80 interactions (activated in presence of DSBs and DNA-bound Ku)
Activated by outward folding of PRD, allowing open ATP binding groove of kinase domain - autophosphorylates ABCDE cluster to lock in active state
Active DNA-PKcs loosens DNA binding, allows end-processing factors and ligases to access ends.
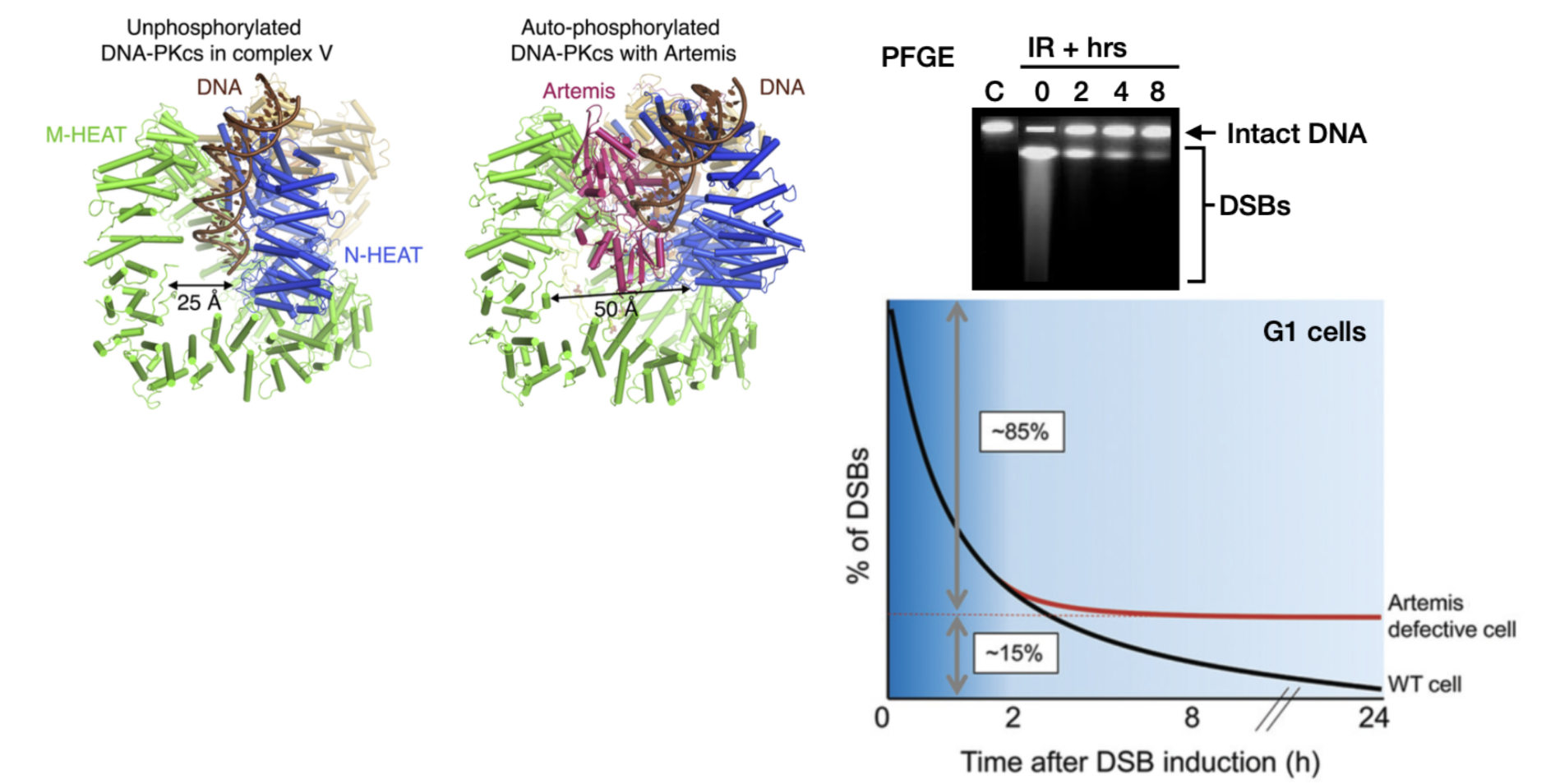
What is the role of Artemis in end processing? What % of DSBs rely on Artemis?
Binds to phosphorylated (active) DNA-PK to cleave DNA hairpins and overhangs
Fast repair component is Artemis independent, but slow repair depends on Artemis (15% breaks persist in Artemis deficient cells)
heterochromatin/damaged ends with hairpin-like configurations may necessitate an involvement with Artemis
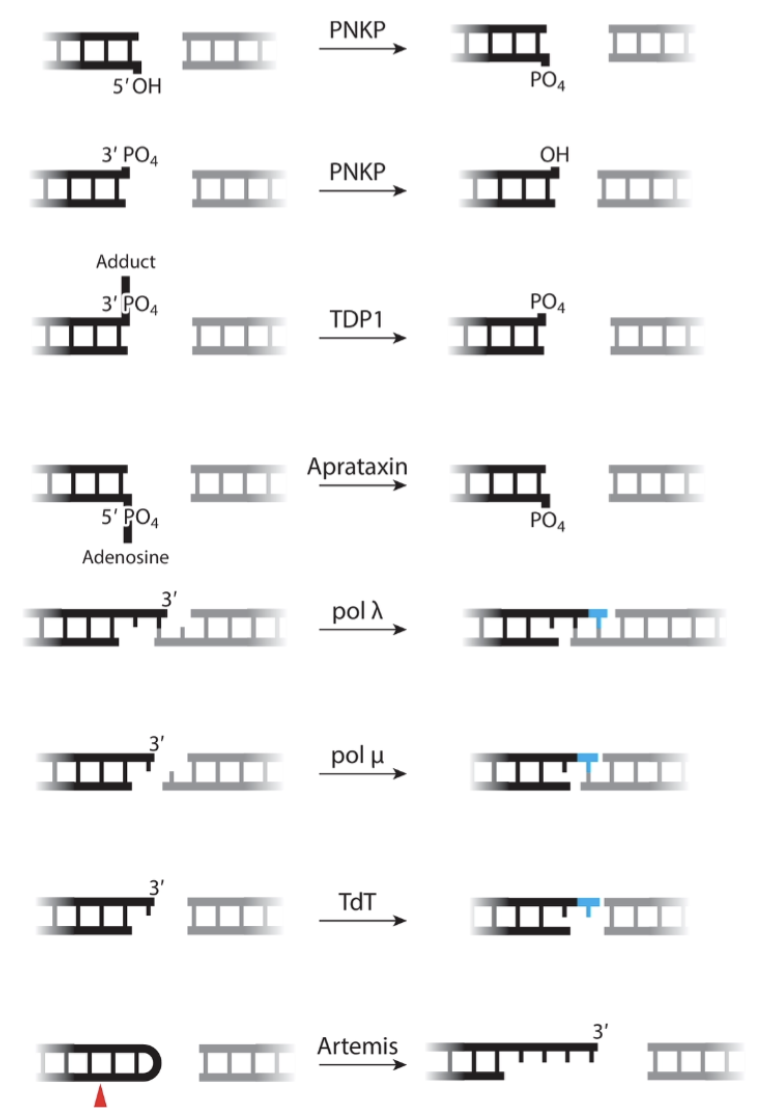
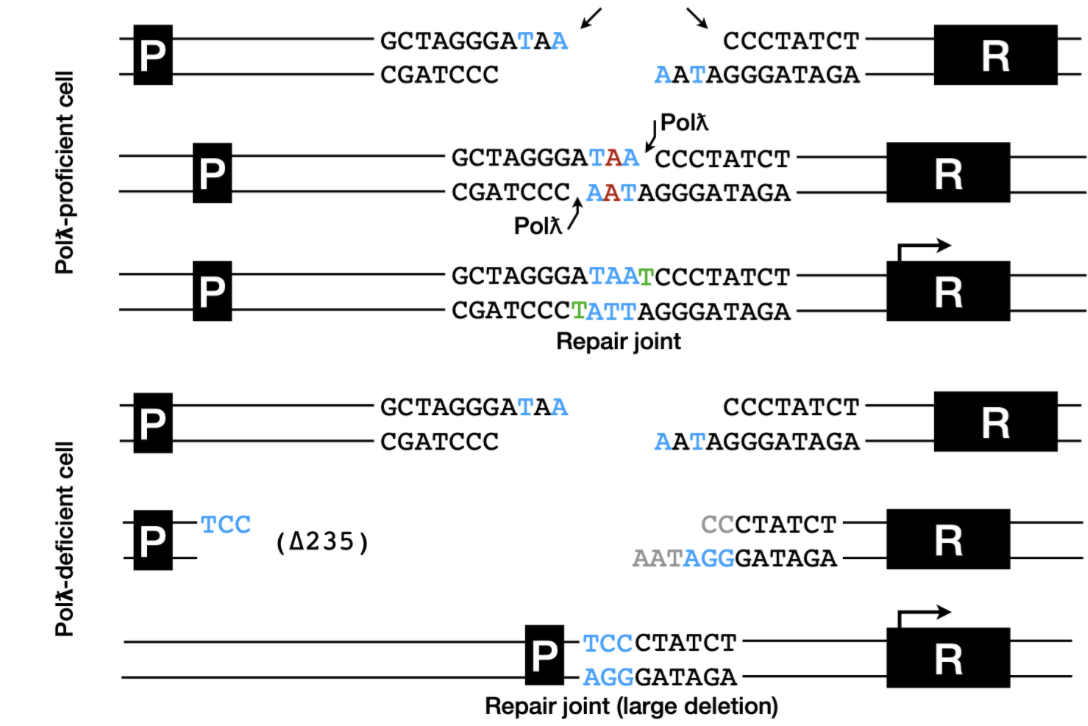
How do polymerases permit end ligation?
Fills uneven ends in DSBs to allow even ligation
Polymerase deficient cells result in deletion of bases - as ends must be even length for ligation
bases lost until repair joint can be formed - could be a severe level of base loss
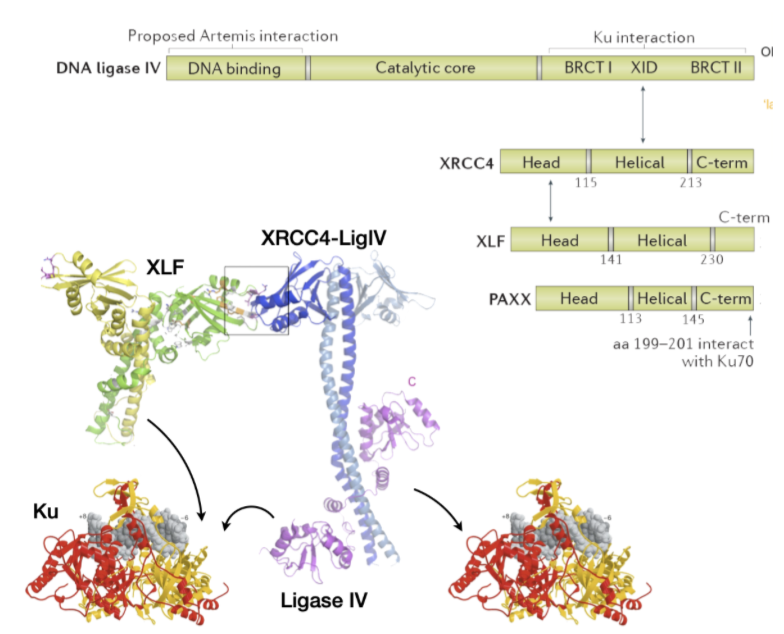
Which proteins form the Ligase IV complex involved in NHEJ?
XRCC4, XLF, PAXX, ligase complex
multiprotein interactions with ligase and Ku
XLF, Ku and Ligase IV are necessary fro NHEJ and survival
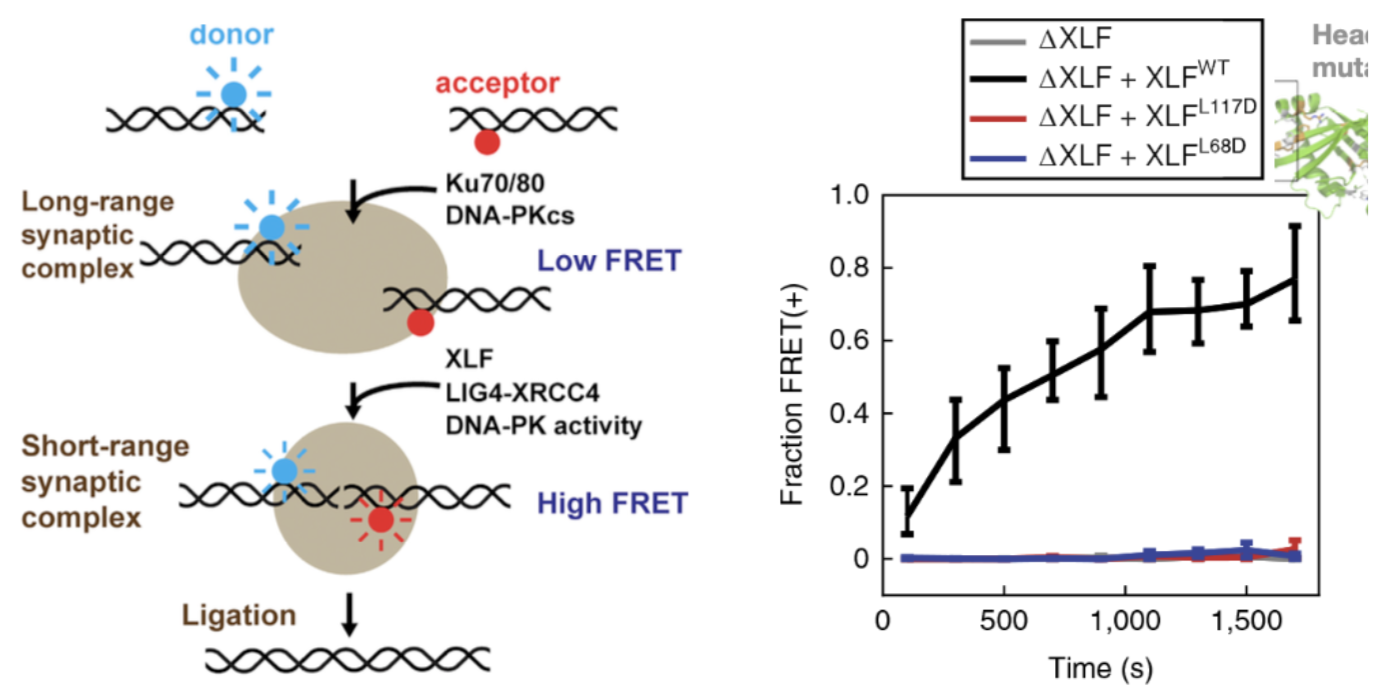
How is close-synapsis of DSB DNA ends induced?
XLF and LigIV-XRCC4 promote close synapsis of DNA ends
also requires DNA-PK activity for transition between long and short range synapsis
Tethered dimeric form of XLF with head mutations (disrupt binding site) in one subunit of XLF dimer exhibited inability to induce short-range synpasis
2 WT XLFs are required for close synapsis
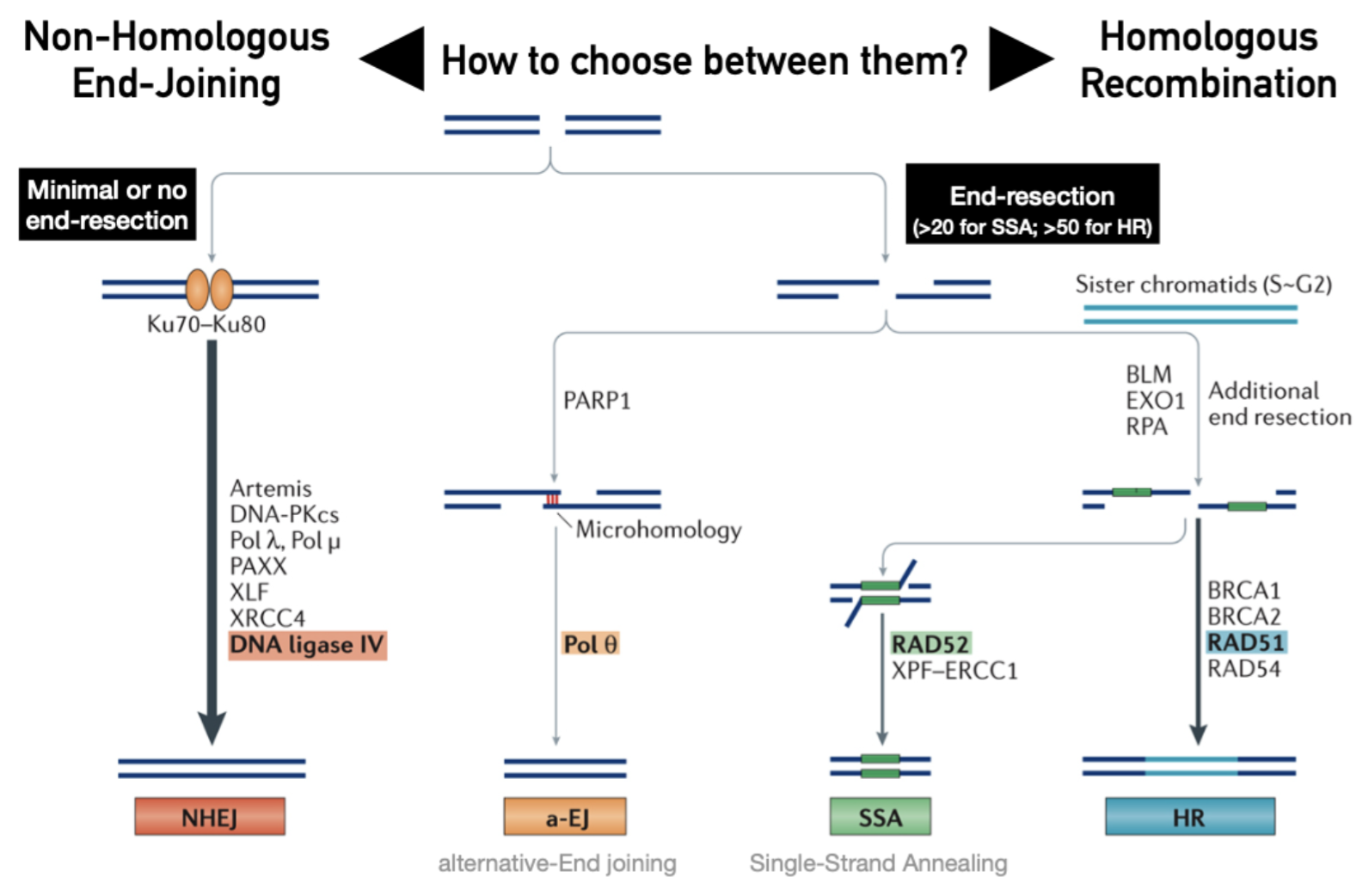
What is the proposed theory for a cells choice to induce NHEJ or HR?
End-resection: favours HR by providing ssDNA substrate for homology search (Rad51)
End-protection: favours NHEJ (resected ends disfavour Ku)
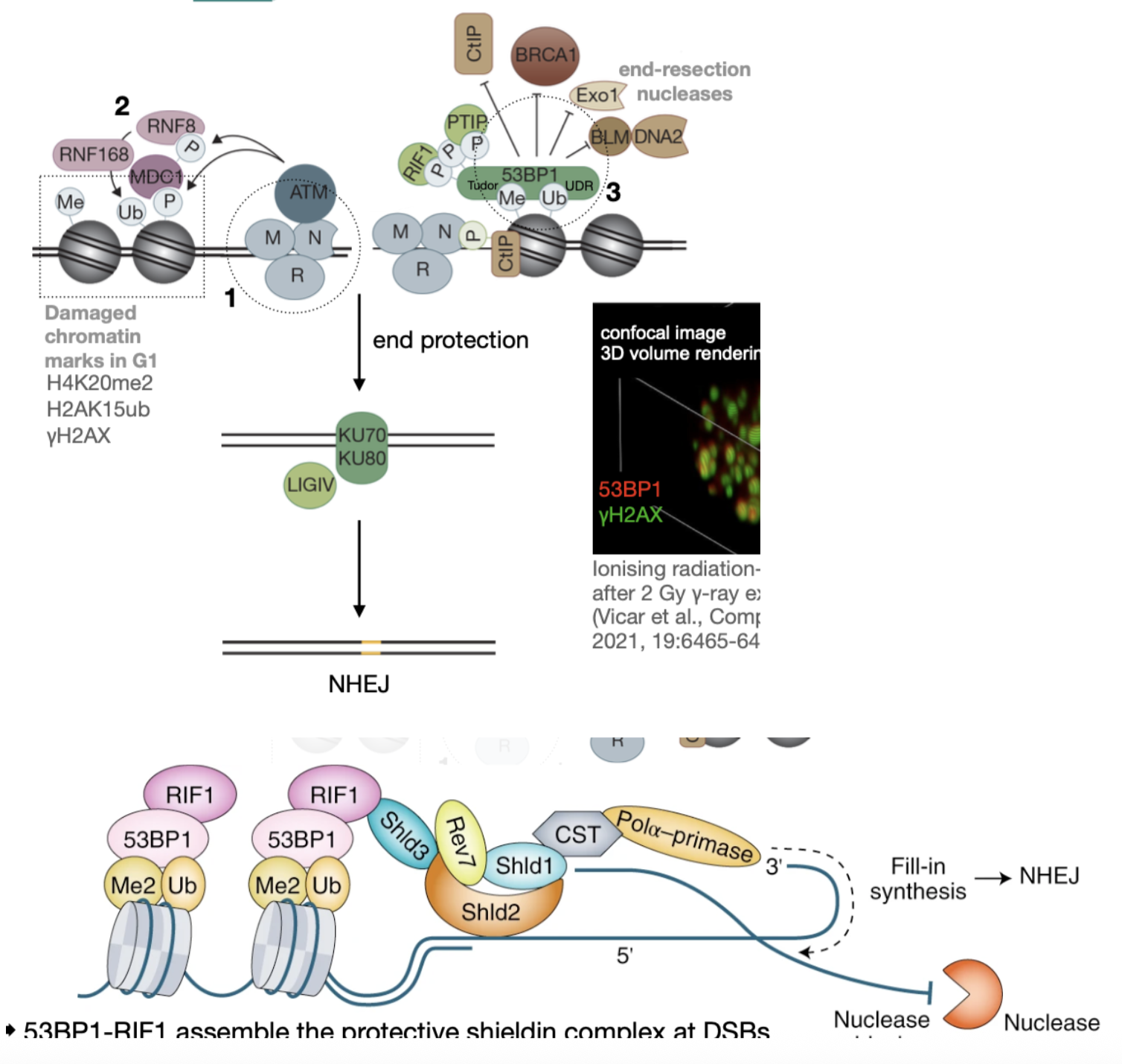
How is NHEJ initiated when breaks occur in G1?
In G1 (methylated histones), active ATM phosphorylates local histones with damaged DNA (marks as yH2AX)
phosphorylated histones recruit ubiquitin ligases (RNF8, then RNF168) to ubiquitinate the histone tail
Methyl and ubiquitin molecules in histone tails recruits 53BP1
Phosphorylated 53BP1, which recurs proteins to protect ends from resection nucleases (RIF1)
RIF1 recruits ‘sheilding complex’ - shields DNA from ER nucleases - Pola also refills resected ends
NHEJ is permitted
How is HR initiated when breaks occur in S/G2?
Since nascent sister chromatid is in immediate vicinity of break, template is easily accessible and HR is appropriate.
Same G1 ATM response with phosphorylation and ubiquitination of histones - but no methyl mark
No methylation leads to decreased 53BP1 binding
CDK activity (and ATR) phosphorylates DSCC1, which displaces RIF1 with PTIP1
BRCA1 binding partner BARD1 competes with 53BP1 for histone binding
BARD1 promotes end resection - and HR
Throughout the cell cycle, what is the spread of repair via NHEJ vs HR?
In G1, all repair is NHEJ
In G2, fast repair is NHEJ, slow is HR.
HR repairs endogenous DSBs from collapsed replication forks - sister strand nearby
How does NHEJ deficiency affect a patients immune system?
Defective NHEJ (due to mutations in DNA-PKcs, Artemis, Lig4, XLF and XRCC4) fail to repair DSBs induced by IR.
Immune cells require DSB repair in maturation (VDJ recombination), RAG1/2 induced breaks cannot be repaired - Ag receptors are not diverse - RS-SCID
What are the clinical features of LigIV deficiency?
Growth defects, microcephaly, malignancy
fast replicating cells are prone to perish due to defective end-ligation in NHEJ
How does excessive NHEJ provide a positive patient prognosis to cancer cells treated with PARPis?
PARP induces increased SSBs > DSBs
Imbalanced NHEJ vs HR can result in ‘BRCAness’ and PARPi sensitiviy
prostate cancer cells with DtX3L over expression has a negative regulatory effect on 53BP1, which induces HR deficient and chromosomal instability
High sensitivity to PARPis
53BP1 mutation restores HR and promotes PARPi resistance