Bio 22 exam 3
1/90
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
91 Terms
what are inducible operons/systems
when the presence of a specific molecule induces gene expression/ such as the lac operon
what is a repressible operon/system
when the presence of a specific molecules inhibits gene expression such as an abundance of end product in environment/ such as the Trp operon
Parts of the lac operon
Repressor gene (lacI), promoter, operator, lacZ, lacY, lacA
what does lacZ code for/ product do
beta-galactosidase, allows the conversion of lactose into glucose or galactose
what does lacY code for/protein do
Permease, facilitates entry of lactose into bacterial cells
what does lacA code for/protein do
Transacetylase, removes toxic by-products of lactose digestion
Polycistronic mRNA
type of mRNA transcribed from the lac and Top operon, one mRNA that codes for multiple products/proteins
lac operon with and without lactose present
When lactose is present, allolactose will bind to the repressor protein so the repressor protein cannot bind to the operator. Then RNA polymerase can come in and translate the genes needed to digest lactose
When lactose is not present the repressor protein will be able to bind to the operator and stop transcription from occurring
CAP?
Catabolite-activating protein. It is part of the lac operon. It binds to cAMP and then it is able to bind to the promoter of the lac operon which facilitates RNA polymerase to bind and transcribe. It is only active when bound to cAMP. Adenylyl cyclase is what turns ATP into cAMP. Glucose inhibits adenylyl cyclase which inhibits cAMP production which makes it so CAP or RNA polymerase can bind. Glucose present means no cAMP, no CAP binding, and no transcription. No Glucose present means plenty of cAMP for CAP which can then bind and RNA polymerase can transcribe
Trp operon components
trpR-repressor, trpP-promoter, trpO-operator, trpE,D,C,B,A-structural genes
Understand how the presence of tryptophan affects transcription of the trp operon
Tryptophan present: The repressor binds to Tryptophan which then causes a conformational change allowing for Tryptophan to then bind to the operator and stop transcription
No Tryptophan present: The repressor has not Trp to bind to, and it cannot bind to the operator so transcription of the genes occurs making Tryptophan
Leader region of the trp operon. Secondary structure of mRNA, and hairpins
Leader region is the part that is always being transcribed (attenuation) as a way to tell if Trp if or isn’t present.\
When the leader region is transcribed into mRNA it will form a different secondary structure depending on if Trp is or isn’t present and that will determine if transcription of the genes for Trp synthesis will occur
Trp present: terminator hairpin, transcription does not occur and RNA Pol falls off, ribosomes can continue translation
Trp not present: anti-terminator hairpin, ribosome stalls while RNA Pol transcribes the genes needed to make more Trp
Describe a riboswitch.
When mRNA/RNA can take up their own shape or secondary structure after being transcribed. They bind with small ligands that cause a conformational change and induce a second RNA domain
Chromosome territory
a chromosomes distinct location in the nucleus
interchromatin compartment
channels between chromosomes that contain little to no DNA but can have RNA Pol
Transcription Factory
The edges of the chromosome territories where there are many active RNA Polymerase molecules (4-30)
Explain how histone modifications lead to changes in transcription. What enzymes control acetylation of histones
all modifications to histones lead to the DNA being less tightly wrapped which makes it more open for transcription. For example acetylation decreases the positive charge which makes the affinity between the histone and DNA less strong. Acetylation of histones is done by histone acetyltransferase enzymes (HATs)
SWI/SNF mechanisms
alteration of the DNA-histone contacts, alteration of the DNA path (how DNA is wrapped), and remodeling of the nucleosome core particle (switch out a histone in the nucleosome core)
Describe the role of DNA Methylation in the control of transcription. Which base becomes methylated? At what location on the chemical structure?
DNA methylation is a way to silence genes, it leads to decreased transcription and gene expression by binding to transcription factors of DNA. It occurs on Cytosine bases at CpG islands (many C's and G’s next to each other on the same stand). The methylation specifically it on C5 on cytosine’s nitrogen ring
Methylated sequences are concentrated in which regions? Where are these regions often found
in CpG islands typically near or in the promoter region
Define cis-acting elements and trans-acting factors. Give examples of each
Cis-acting elements are regulatory elements located on the same chromosome as the gene that it regulates. EX: promoters, enhancers, and silencers
Trans-acting elements are regulatory proteins that bind to cis-acting elements. EX: transcription factors that can activate or repress
Promoter parts
Core-promoter: absolutely critical and determines accurate initiation of transcription
Proximal-promoter elements: control how frequently RNA polymerase can bind.EX: CCAAT and GC boxes
Differentiate focused and dispersed promoters
Focused promoter: one spot for RNA polymerase to bind to, specific transcription initiation at the start site, majority for lower eukaryotes
Dispersed promoter: RNA polymerase has multiple sites to bind to, direct initiation from several weak transcriptional start sites. Gives higher levels of transcription and proteins
core-promoter elements
TFIIB recognition element (BRE), TATA-box, BRE, initiator, motif ten element (MTE), downstream promoter element (DPE)
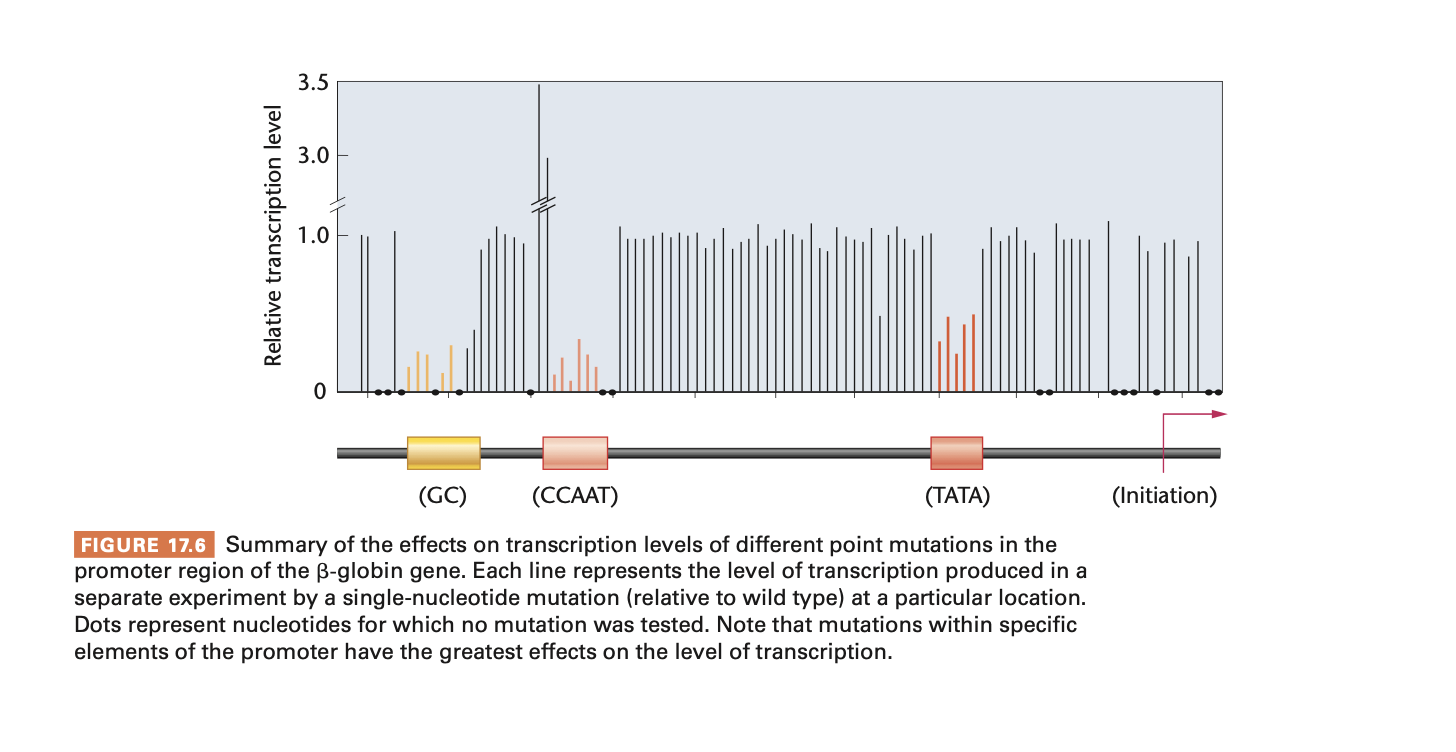

know what this graph shows
mutations in the proximal promoter elements (CCAAT and GC box) majorly decrease transcription
enhancer
regulate/enhance transcription of eukaryotic genes and important for reaching maximum level of transcription. Located anywhere is relation to the gene (upstream, downstream, in the gene)
Insulator
found between an enhancer and a promoter, it allows for some enhancer-promoter interactions while blocking others, stops enhancers from enhancing the wrong part
silencer
repress the level of transcription initiation
two functional domains of eukaryotic transcription factors?
DNA-binding domain which binds to specific DNA sequences in the cis-acting regulatory site
Trans-activating domain which does the work/ activates or represses transcription by binding to other transcription factors or RNA polymerase
three examples of DNA-binding domains
helix-turn-helix, Zinc-finger, basic leucine zipper
What is the PIC and parts
Pre-Initiation Complex and it is a complex of proteins. First TBP (TATA-box binding protein) binds to the TATA box. Then TAFs (TBP-associated factors) binds to TBP creating TFIID. Then the other TFII’s join. RNA polymerase then comes and a mediator goes on top. An enhanceosome facilitates this process.
What is an enhanceosome
a complex formed by activator and coactivator proteins that direct transcriptional activity
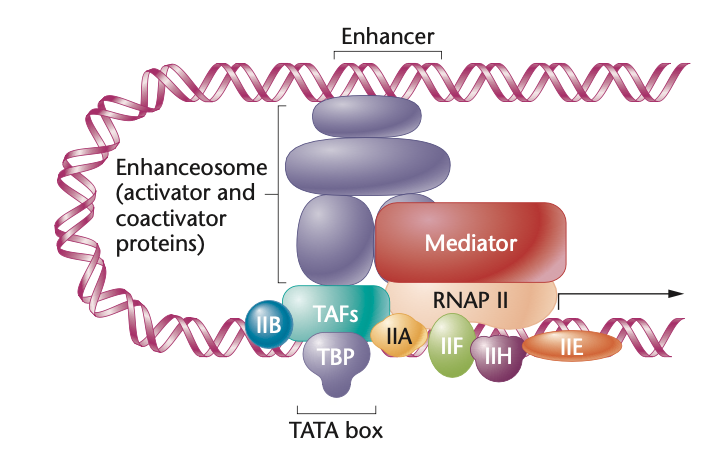
pic is enhanceosome and PIC

How are methylated promoters different from unmethylated promoters
When a promoter is unmethylated the gene can be transcribed and when a promoter is methylated the gene is silenced so transcription cannot occur
Where on the DNA structure are cytosines methylated and by what
The cytosines are methylated on C5 of their nitrogenous bases resulting in 5-methylcytosine. In CpG islands near the promoter. Catalyzed by DNA methyltransferases
Histone acetylation description and effect on transcription
Acetyl groups added to histone tails which reduce positive charge and therefore affinity between DNA and histone making DNA easier to transcribe. Done by histone acetyltransferase enzymes, lead to increased transcription
DNA methylation description and effect on transcription
Adding a methyl group to C5 of Cytosines nitrogenous ring, at CpG islands. Done by DNA methyltransferases, leads to decreased transcription
Chromatin remodeling description and effect on transcription
the process of changing how tightly DNA is wrapped around histones and exposing or hiding promoter regions, less wrapped is more transcription and more wrapped is less transcription
types of histone modifications and where do they occur
on the N-Terminal tail. Acetylation, methylation, phosphorylation, ubiquination, isomerization
Types of noncoding RNA
lncRNA (long noncoding RNA): do not encode proteins because they lack a functional open reading frame. They regulate gene expression at the transcriptional and chromatin level by interacting with DNA, RNA, or proteins, including chromatin-modifying complexes, to influence whether genes are turned on or off.
sncRNA (short noncoding RNA): are small RNA molecules that do not encode proteins because they lack an open reading frame. They regulate gene expression after transcription by base-pairing with target mRNA, which leads to mRNA degradation or inhibition of translation.
mechanism of action for lncRNA
Decoy: lncRNA binds to the protein instead of DNA
Adapter: brings multiple protein together
Guide: brings proteins to a specific sequence of DNA
Enhancer: brings proteins to activate transcription
Three classes of MAE
Monoallelic expression: Imprinting, random inactivation of one X-chromosome in women, random monoallelic expression of autosome genes
What is genetic imprinting?
A phenomenon in which genes show expression of only one allele (paternal or maternal) while the other is silenced via methylation
Most known imprinted genes encode what types of genes
growth factors or other growth-regulating genes
examples of imprinting diseases
Prader-Willi syndrome, Beckwith-Weidemann Syndrome, Angolan Syndrome
Explain Beckwith-Weidemann Syndrome
typically a person gets two alleles, one form each parent. Both alleles have a gene for H19 and IGF2 (insulin growth factor 2). In a normal imprinting pattern one allele’s promoter for H19 be methylated and nonfunctional and the other will encod a lcnRNA that will silence expression of the IGF2 of that same allele. So each allele will either code for H19 or it will code for IGF2. In BWS both H19 promoters are methylated so one IGF2 isn’t silenced and that person has double the amount of IGF2 expressed.
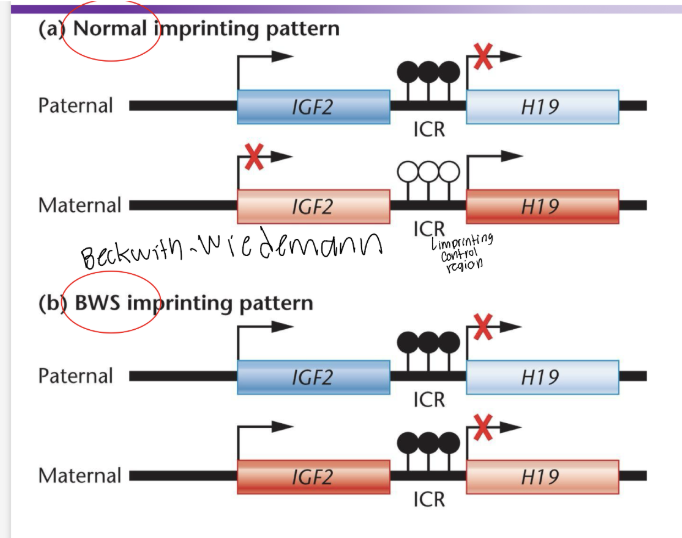
Describe random MAE as it relates to inactivation of the X-chromosome
What this means is that half of embryonic cells will randomly inactivate the paternal chromosome while the other half will randomly inactivate the maternal X chromosome via XIST RNA. This silences all genes on the inactivated chromosome ensuring no double amount of gene expression
What are the risks associated with ART in terms of imprinting errors
assisted reproductive technologies. They have an increased risk of imprinting errors that can lead to Beckwith-Wiedemann Syndrome, Prader-Willi Syndrome, and Angelman Syndrome. IVF children tend to have reduced levels or a complete loss of maternal specific methylation
Describe the abnormal histone modifications seen in Rubinstein-Taybi Syndrome.
There are mutations in the genes that encode for Histone Acetyltransferase (HAT) and Histone Deactylase (HDAC). What this means is that genes are way more closed since histones can’t be acetylated properly
Cancerous cells show what type of methylation
Typically hypomethylation. Less methylation of DNA means it can be transcribed more. However some genes can be hypermethylated (such as genes that fix DNA/errors) so that they get transcribed less
Promoter hypermethylation has what effect on tumor-suppressor genes
When tumor suppressor genes are hypermethylated at the promoter it prevents their transcription. This allows for the tumors to keep growing without suppression which is what causes cancer.
Global hypomethylation has what effect on chromosomal stability
Global hypomethylation decreases chromosomal stability because it leaves DNA more susceptible to mutations and can cause gene overexpression and promoter cancer progression
Know the examples of heritable epigenetic traits described in class
Mice coat color depending on diet, rats and MN and heritable stress, Dutch hunger winter famine
mice and epigenetics
Mice had a different colored coat depending on what they ate even though they were genetically identical. A diet that is rich in methyl donors is more likely capable of silencing the Agouti gene (and yield brown offspring) than a diet that is poor in methyl donors. Coat colors range from yellow (unmethylated promoter) to pseudoagouti (highly methylated promoter)
Rats and Epigenetics
Stress-Induced Behavior is Heritable. Newborn rats raised with low maternal nurturing in turn has low glucocorticoid receptors. This means that there are less receptor to bind to cortisol which lead the rats to be more stressed. Opposite for rats with high maternal nurturing. This was due to differences in histone acetylation and DNA methylation levels. Low-MN rats have higher levels of DNA promoter methylation and less histone acetylation for glucocorticoid receptors. High-MN rats had less DNA promoter methylation and more histone acetylation for glucocorticoid receptors
Dutch Hunger Winter
many people starved to death, Their generations now have heart disease, obesity, increased stress. Due to their IGF2 methylation being deregulated
Explain restriction enzymes and the sequences they recognize.
Restriction enzymes are protein endonucleases that recognize and cut double stranded DNA at specific sequences. Produced by bacteria as a defense mechanism against bacteriophages
Which enzyme is needed to seal the complementary fragments together after restriction digestion
DNA ligase
Cloning vectors purpose
act as a mean of transporting/inserting a desired sequence of DNA into an organism via transformation and they have a selectable gene marker like antibiotic resistance
plasmid vector
grows in Bacterial cells such as E. coli, They replicate individually from chromosomes, they have a number of convenient restriction sites, and a marker gene to select for presence in the host cell
phage vectors
grow in Bacterial cells such as E. coli, It is larger than plasmids (carry up to 45 kb of DNA)
YAC
Yeast artificial chromosomes, grow in Saccharomyces cerevisiae (yeast), Can be easily grown and manipulated, has posttranslational modification eukaryotic proteins
Define bacterial transformation
the process of inserting foreign DNA into a bacterial cell for it to take up and then transcribe itself
Steps to transformation
add plasmid to competent cells and incubating on ice
heat shock at around 42°C then back to ice
then recovery phase at around 37°C
How are cDNA libraries obtained? Which enzyme is required
cDNA is obtained via reverse transcriptase on an mRNA strand
Describe PCR
Polymerase Chain Reaction is a technique used to amplify small amounts of DNA, can be used to amplify a specific gene sequence
PCR Steps
Denaturation
Annealing
Extension
Repeat
PCR components
Taq Polymerase (heat stable polymerase)
dNTPs
Buffer- has the Mg2+ for Taq
Forward and reverse primers (around 20nt)
The DNA sample
Briefly differentiate Southern, Northern, and Western Blotting techniques. Remember SNOW DROP.
SNOW DROP
Southern Blot- detects DNA- probes with DNA or RNA
Northern Blot- detects RNA- probes with DNA or RNA
Western Blot- detects proteins- probes with antibodies
Gene targeting and knockout
Gene targeting: manipulate specific allele or gene in a genome
Gene knockout: disrupt or eliminate a specific gene to see what happens (knockout mice)
Describe the CRISPR-Cas9 system.
Guide RNA binds to PAM then brings the Cas9 to the specific sequence of DNA that is to be cut. Different nucleases are what create breaks in the genome
Define whole-genome sequencing
shotgun cloning, strategy for sequencing a genome by cutting genomic DNA into fragments via multiple restriction enzymes then determine the DNA sequence
contigs?
refers to the fragments made when cutting genomic DNA for whole-genome sequencing with various restriction enzymes, they are continuous fragments that overlap and collectively make one continuous DNA
ChIP
chromatin immunoprecipitation
Crosslink proteins to DNA (freeze proteins onto DNA so interactions are preserved) and lyse the cells open via sound waves
The chromatin is then isolated and fragmented
A protein specific antibody is added and binds to the protein on the DNA
Purify/isolate the protein-DNA complexes via agarose beads
Reverse the crosslinks and isolate the DNA
Sequence the DNA and map to genome
Describe the human microbiome project
Human microbiome project: to determine if people have a shared core human microbiome which can help us understand any correlation between microbiome and health
Human Genome project
The creation of extensive maps developed for genes implicated in human disease conditions
nutrigenomics
to find genes associated with medical conditions or nutrient metabolism
stone-age genomics
to study evolutionary relatedness of various extinct and present day species using DNA from bone
metagenomics
(environmental), to sequence genomes from entire communities of microbes in environmental samples of water, air, and soil.
Describe the production of insulin in bacteria.
Mammalian proinsulin mRNA is gathered from pancreas cells
That proinsulin mRNA is then converted to cDNA via reverse transcriptase
cDNA is split into two sub units, A and B
Those cDNA subunit units are joined to a plasmid
Recombinant plasmid is transformed into E. coli
Extract and purify the proinsulin
Treat with cyanogen bromide to release A and B chains
Purify and mix/combine chains to make functional insulin
types of vaccines
inactivated, attenuated, subunit, nucleic acid
inactivated vaccines
prepared from killed samples of infectious virus or bacteria
attenuated vaccines
live viruses or bacteria that can no longer reproduce but can cause mild form of disease
subunit vaccine
surface proteins from virus or bacterium are developed by genetic engineering. EX: hepatitis B virus or HPV vaccines
nucleic acid vaccines
uses mRNA. EX: covid vaccines
GloFish and heavy metal detection?
The GloFish were genetically engineered to carry a fluorescent gene that is linked to an inducible promoter. This promoter is inducible via heavy metals. What this means is that when there is heavy metal contamination, the promoter will become active allowing transcription of the gene which causes fluoresce glowing. The fish would only glow in the presence of heavy metals
types of prenatal testing
amniocentesis (amniotic fluid), chorionic villus sampling (chorion tissue), and a blood draw (non-invasive 3-6% of blood has fetal DNA)
ASO
allele specific oligonucleotide
ASO how does it work
oligonucleotides are used a probes to identify alleles that can differ by a single oligonucleotide. DNA is extracted by the patients and denatured. Then it is probed with the ASO. One probe is for the normal sequence and one it for the mutant sequence. Then you see which probes bind and that shows you have that allele
tests for sickle cell gene
PGD= Preimplantation Genetic Diagnosis
Uses ASO method
Take one embryo at the 16 cell stage
DNA is isolated and amplified with primers via PCR for the B-globin gene
Then DNA is denatured and placed onto two membranes
One membrane is probed with the normal B-globin allele BA and one probed with the mutant B-globin allele BS
Results can show no mutant allele at all, only mutant alleles, or carrier with one mutant allele
what is personal genomics
individual genome sequencing utilized in medical clinics as a way to evaluate a person’s genetic information to give insight into the genetics of Alzheimer's disease, proteus disease, etc
What ethical, social, and legal questions arise from genetic engineering, genomics, and biotechnology
ELSI program: ethical, legal, and social implications program. Addresses issues about confidentiality of genetic information and implications for medical practice, genetic counseling, and reproductive decision making