Unit 4 (Lessons 26-27)
1/40
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
41 Terms
Gregor Mendel and William Bateson
Gregor=founder
William=champion
What are Mendel’s 3 Laws?
Law of segregation
law of independent assortment
law of dominance
Law of Segregation
2 copies of each genetic factor segregate during gamete formation, ensures each parent’s offspring obtains one factor
Predict probabilities of inheritance
Law of Independent Assortment
segregation of alleles at one locus is independent of segregation of alleles at another locus
Law of Dominance
one allele dominates the other
why is the law of independent assortment an assumption rather than a law?
basic unit of transmission is the chromosome and genes on chromosome that are close together do not actually assort independently
this creates linked genes
How many chrossovers per chromosome in meiosis?
a finite number
Recombination Frequency (RF)
measure of how often genetic crossover occurs between 2 loci during meiosis
=(recombinant/total chromosomes)*100
Genes with independent assortment have RF= 50%
1% RF= 1m.u. (1 centiMorgan)
how do genetic companies build genealogies?
tracking how chunks of alleles are inherited through generational time
leads to haplotypes
Explain the concept of “Identical By Descent”
certain chunks of DNA don’t recombine very often (b/c they are linked) so they are inherited in blocks
genotyping SNPs all along chromosome lets you infer relationships between distantly related people
how does “Identical By Descent” compare to “Identical By Choice”?
minimum segment size thresholds were built into the matching algorithm
false matches can still appear!
Neomorphic
GOF but completely new function
Hypermorphic
GOF with higher activity levels
Amorphic
LOF but complete loss of function
Hypomorphic
LOF but only partially decreases function, still works to certain extent
How do we know if 2 genes interact genetically? (as opposed to physical interaction of products)
by counting phenotypic classes among offspring of Dihybrid Cross (AaBb x AaBb)
expected ratios are 9:3:3:1
What are 4 types of gene interactions with inheritance?
Redundancy- 2 genes make same product
Additivity- 2 genes generate 1 phenotype (ex: amount of pigment adds to darker shades)
Epistasis- 1 gene masks phenotype of another (AKA Gatekeeper effect)
Modification- 1 gene changes how another gene behaves (but doesn’t have its own phenotype) (AKA called modifiers)
What role do Epistasis and Redundancy play in SMA (Spinal Muscular Atrophy)?
Epistasis: SMN1=Gatekeeper, master gene and is 100% functional in normal conditions
Redundancy: SMN2=Backup gene, duplicate in case SMN1 is broken, which then follows the logic of additivity
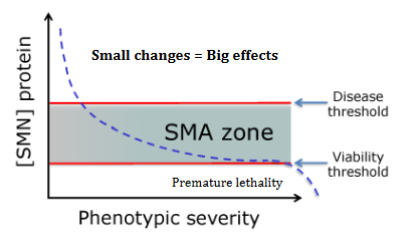
Explain genetics behind SMA (spinal muscular atrophy)
null mutant (complete LOF) causes embryonic lethality
SMA zone=hympomorphic state
2 thresholds=more than 1 SMN function
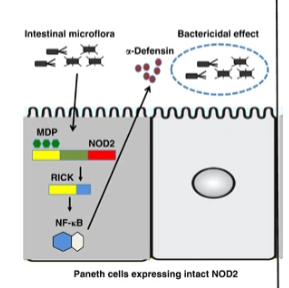
NOD2
encodes intracellular “sensor” that detects small molecules associated with bacterial pathogens and activates the immune response
when lost or reduced, bacteria overgrow and inflammation results
mutations in NOD2 not completely penetrant
What does penetrant mean with reference to disease?
everyone with mutation develops the disease
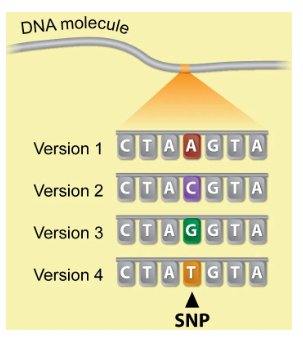
What are the “DNA markers” that allowed scientists to fine map locus and identify causal gene?
SNPs (Single nucleotide polymorphisms)
main difference between people
places in genome where one person has one bp and a different person has another
AKA SNPs are a type of genetic mutation that creates different alleles
to qualify as SNP, base change has to be found in at least 1% of human population
most SNP differences have no effect
True or false: any disease-causing mutation has many neutral SNPs linked to them
true!
Define haplotype
particular combination of DNA variants in a region of chromosome
usually containing many linked SNPs
frequency of haplotypes varies in different human populations, reflecting different more recent common ancestors
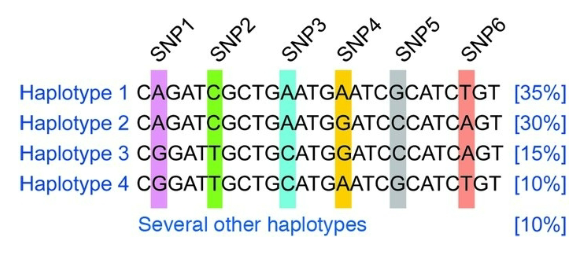
How would a missense mutation in an SNP in Haplotype 1 affect gene?
mutation would be associated with the other SNPs in Haplotype 1
AKA it is inherited by descendants
How can haplotypes be used to map disease causing mutations among multiple families with affected and unaffected individuals?
look for linked SNPs
look for haplotypes shared by affected individuals and NOT found in unaffected individuals
Shared haplotype likely contains disease mutation
Ex: TERT mutation in melanoma
When determining which haplotype is associated with a disease causing mutation, scientists found that only a fraction is more common, how did they determine if it actually caused the mutation?
calculated the likelihood ratio
AKA likelihood that it is ore frequent in the disease than expected by chance.
use logarithm score to yield simple number and set boundaries
What percent of people with Crohn’s disease have a first-degree relative with the disorder?
15%
Pleiotropy
one gene influences multiple traits
What % of disease-associated SNPs are located in gene coding sequences and have the potential to disrupt splicing or alter function of encoded proteins?
5%
Remaining 95% of disease-associated SNPs located in non-coding DNA sequences that make up 98% of genome
Forward Genetics
Mutagenesis
Mapping-by-sequencing
used to identify genetic basis responsible for observed phenotype, mapping done to identify underlying gene
Reverse Genetics
Ectopic expression
gene silencing
gene sequence is known but function is unknown
Gene then “knocked out” and phenotype is observed, then transgenic rescue or ectopic expression
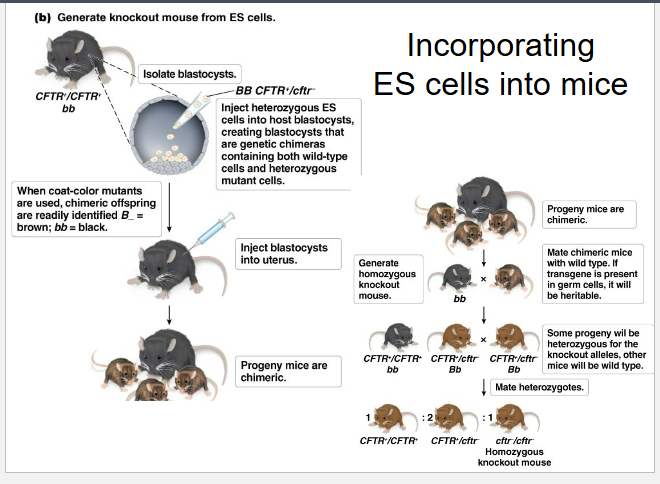
How are “Knock-out” mice made?
animal cells have less homologous recombination, must select for it
totipotency is not common so transgenes injected into eggs, embryos, or embryonic stem cells to give rise to gametes
develops genetic chimera
How are “knock outs” made in embryonic stem cells?
by homologous recombination
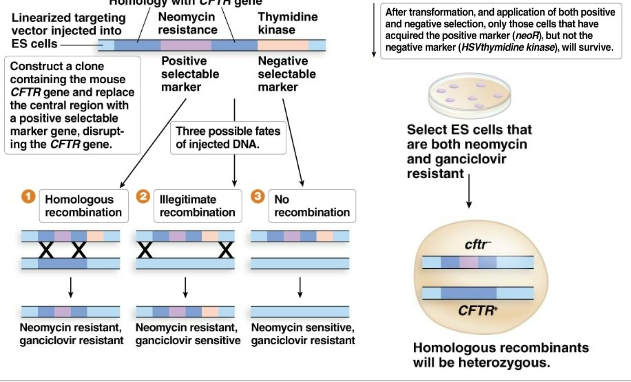
How are embryonic stem cells incorporated into mice?
How does CRISPR-Cas9 work as a gene editing tool?
naturally in bacteria to produce sgRNA
Natural bacterial immune system, leads to adaptation then expression then interference
utilizes guide RNA (crRNA), tracrRNA (helps Cas9 bind) and PAM
What is PAM?
Protospacer Adjacent Motif
short DNA sequence (2-3 bases) sits right next to target sequence
After you’ve made your knockout gene and analyzed the phenotype, how do you prove the observed phenotype is due to the mutation and not some genetic background effect?
complementation
Non-complementation
What is genetic copmlementation?
2 recessive mutants used as evidence that two genes operate within common pathway and not on different genes
Shows that defect generated is fixed by expression of transgene
If genes complement one another then they are in different genes
Fail to complement=same gene
What are histone PTMs?
histone post-translational modifications
chemical alterations to histone proteins (mostly on N-terminus)
AKA acetylation and methylation