Chapter 15: How damage to DNA that is not correctly repaired leads to mutations in DNA sequence that can alter the characteristics of organisms
1/73
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
74 Terms
Mutations definitiion
permanent change in the DNA sequence, therefore, a transient change to the DNA sequence is not considered a mutation
Does DNA damage always lead to mutation?
No, because if accurate DNA repair occurs before the DNA sequence is permanently changed, the mutation was avoided
do mutations always have to be in a gene?
No, although, we are more likely to notice them (see a phenotypic change) if the mutations are occurring in a gene
What are the categories of mutation sources?
Naturally occurring: spontaneous mutations
Caused by external agents: induced mutations
What are the causes of natural forms of mutations?
errors in replication
spontaneous chemical changes in the bases
naturally occurring recombination errors
What are the causes of induced mutations
chemical mutagens that cause base substitutions or indels
UV irradiation
X-rays or ionizing radiation
What do naturally occurring errors in replication lead to?
base substitutions and indels
what do spontaneous chemical changes in bases lead to?
depurination, deamination, or oxidation —> base substitutions
What do naturally-occurring recombination events lead to
chromosomal rearrangements
what do chemical mutagens result in?
base substitutions or indels
What does UV irradiation result in?
pyrimidine dimers → base substitutions
What do X-rays or ionizing radiation lead to?
chromosome breakage → chromosomal rearrangements; alternatively, base damage deamination, or oxidation → base substitutions
What are the two purines?
AG
What are the two pyrimidines
TC
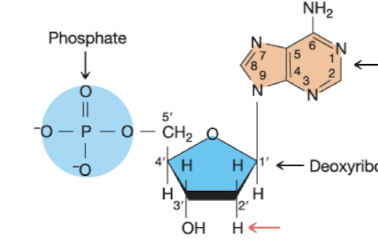
What nucleotide is this
Deoxyadenosine 5’ monophosphate
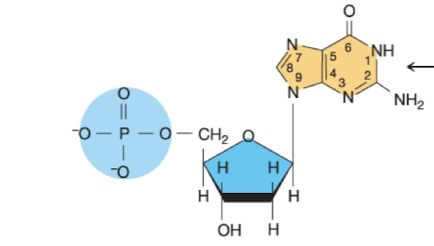
What nucleotide is this?
Deoxyguanosine 5’ monophosphate
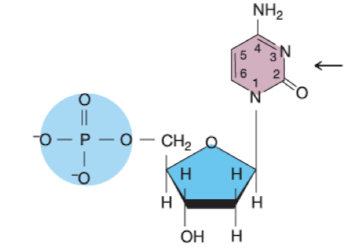
What nucleotide is this?
Deoxycytidine 5’ monophosphate
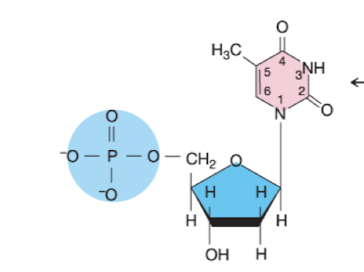
What nucleotide is this?
Deoxythymidine 5’ monophosphate
base substitutions
change one nucleotide to another without changing the number of nucleotides in the DNA
indels
insertions and/or deletions of nucleotides and by definition they are changing the number of nucleotides in the DNA
Transition
changes between purines or between pyrimidines
Is A ←>G a transition or a transversion
transition
Is C → T a transition or a transversion
transition
is T→ C a transition or a transversion
transition
Is A → T a transition or a transversion
transversion
Is T → A a transition or a transversion?
transversion
Is A → C a transition or a transversion
transversion
Is G → T a transition or a transversion
transversion
IS G→ C a transition or a transversion
transversion
transversions
changes in nucleotides between purines and pyrimidines
Are transitions or transversions more distorting?
transversions
What is the Cap on mRNA and what is its function (in eukaryotes)
7-methylguanosine attached to the 5’ end of the mRNA with three phosphate groups; protects the RNA from degradation and is required for translation in eukaryotes
silent mutation, AKA…
Change of the DNA that does not influence the gene’s function; synonymous mutation
Missense mutation, AKA…
nonsynonymous mutation; changes in the DNA that influences the genes function
nonsense
early addition of a stop codon
What do you cause a missense mutation that results in now encoding and amino acid that has similar properties as the original amino acid
conservative missense mutation
What do you cause a missense mutation that results in now encoding and amino acid that has different properties than the original amino acid
nonconservative
do mutations have to occur in a protein coding region to have a phenotypic outcome?
no
examples of mutations in a non-protein coding region that results in a phenotype
point mutations in transcription enhancer or promoter elements can change the binding site of transcription factors or RNA pol complex, respectively; point mutations in the UTR can affect translation and or gene regulation by affecting the binding of miRNAs; point mutations in splice sites/A branch point can affect the binding of snRNPs and therefore affect splicing
Explain the basis of the Luria Delbruck fluctuation test
System: phage attacked bact, but some survived due to REM
Question: were the mutations induced by the phage or were they there before?
Dependent Variable: percent of the cultures that are resistant
In the luria delbruck fluctuation test, what result would you expect to see if the phage resistance mutation was induced?
the same number of resistant colonies in each assay
In the luria delbruck fluctuation test, what result would you expect to see if the phage resistance mutation was spontaneous?
You would see variation in the number of resistant colonies
In what number of occasions does DNA pol not use proofreading
very very rare, <1 in 10^9 occasions
Which nucleotides have keto groups involved in hydrogen bonds
G and T
Which nucleotides have amino groups involved in hydrogen bonds
A and C
What happens if you lose the keto groups of G/T or if you lose the amino groups of A/C
loss of hydrogen bonds, no base pairing
Tautomerization of DNA bases
the spontaneous isomerization of a ntirogen or oxygen base to an alternative hydrogen-bonding form
How does the conversion of a keto group to an enol group on Thymine change what the nucleotide can pair with?
now pairs with G (three hydrogen bonds)
How does changing the amino group on a cytosine to an imino group change what the nucleotide can pair with?
now pairs with A (two hydrogen bonds)
Does DNA pol recognize differences in bases that are caused by tautomerization?
no
What is the predominant form of A
amino
What is the predominant form of T
keto
What is the predominant form of G
keto
What is the predominant form of C
amino
What is the rare tautomer of A, and what does it pair with, # of hydrogen bonds
imino, C, 2
What is the rare tautomer of T, and what does it pair with, # of hydrogen bonds
enol, G, 3
What is the rare tautomer of G, and what does it pair with, # of hydrogen bonds
enol, T, 3
What is the rare tautomer of C, and what does it pair with, # of hydrogen bonds
imino, A, 2
Do tautomeric changes cause transitions or transitions
transitions because the nature of the base is changed, i.e., a purine stays a purine and a pyrimidine stays a pyrimidine
How can wobble contribute to DNA pol errors
In this process, there is no chemical change to the bases. The bases mispair due to a slight shift in their position
DNA pol wobble leads to the following atypical base pairs:
AC and TG
Does DNA pol wobble lead to transitions or transversions
transitions only
How do you get insertion in replication slippage
in the new strand, an extra base slips out, the loop is stabilized by repetitive DNA sequences, during the next round, a Pair is _________
How do you get deletion in replication slippage
in the template strand, extra bases loops out, the loop is stabilized by repetitive sequences, during the next round of replication, a base pair is ___________
What is an example of the impact of replication slippage
CCG expansion in fragile X that results in the most common form of an inherited intellectual disability
What is CCG expansion in Fragile X syndrome caused by molecularly?
the number of trinucleotide repeats positively correlates with methylation in the 5’ UTR of the FMR1 gene —> when methylation is high, FMR1 is silenced → this results in disease
Do all nucleotide expansions that cause disease lead to gene silencing?
No. Some result in disease because they affect coding sequence and created repeats of amino acids that are deleterious for the function of the protein
Depurination
spontaneous loss of G or A by breakage of glycosidic bond with deoxyribose → leading to random base substitution
Deamination
naturally-occurring conversion of a C to a U → transition mutation of GC to AT
Oxidation of bases
naturally-occurring oxygen radicals attack G&T and alter their structures —>results in transversions
Which bases can depurination occur on
A or G but are more common on G
Why does depurination lead to problems for DNA pol
DNA pol does not have a base to read on the template strand; often causes DNA pol to be able to pass through this area
Does depurination directly affect the DNA backbone
no