Chapter 22 - Nucleotides and Nucleic Acids
1/55
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
56 Terms
energy for metabolism (ATP)
enzyme cofactors (NAD+)
signal transduction (cAMP)P
3 functions of nucleotides
storage of genetic info (DNA)
transmission of genetic info (mRNA)
processing of genetic info (ribozymes)
protein synthesis (tRNA and rRNA)
4 nucleic acid functions
nucleotide = nitrogenous base + pentose + phosphate
nucleoside = nitrogenous base + pentose
nucleobase = nitrogenous base
nucleotide vs nucleoside vs nucleobase
negative charge
what charge is the phosphate group at neutral pH
5’ position
what position does phosphate group normally attach to
built using 5’-triphosphates but each nucleotide has 1 phosphate per nucleic acid
nucleic acids are built using what phosphates (mono, di, tri) and how many phosphates do they contain per nucleotide
RNA : β-D-ribofuranose
DNA : β-2’-deoxy-D-ribofuranose
pentose in RNA vs DNA nucleotides
the straight-chain (aldehyde) and ring (β-Furanose) forms
in solution what forms of free ribose are in equilibrium
pyrimidines or purine
what are nucleobases derivatives of
nitrogen-containing heteroaromatic molecules and planar or almost planar
what type of molecules are nucleobases
250-270 nm
what UV light do nucleobases absorb
cytosine → DNA and RNA
thymine → only DNA
uracil → only RNA
pyrimidine bases and what they are found in (DNA/RNA)
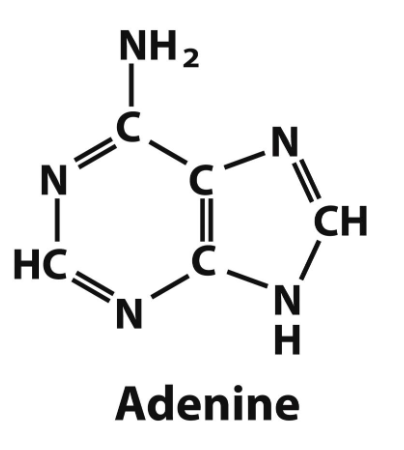
adenine structure
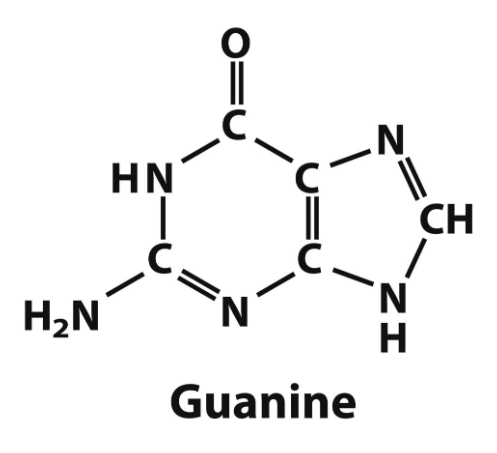
guanine structure
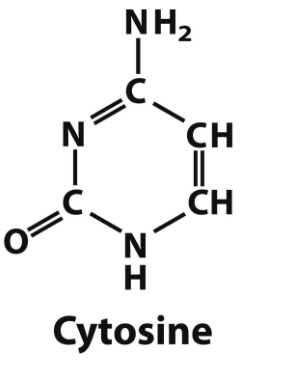
cytosine structure
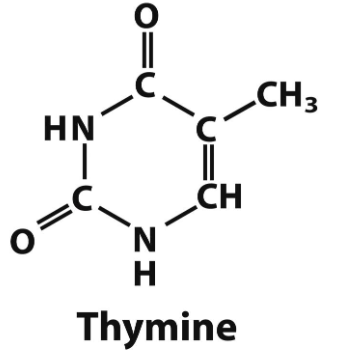
thymine structure
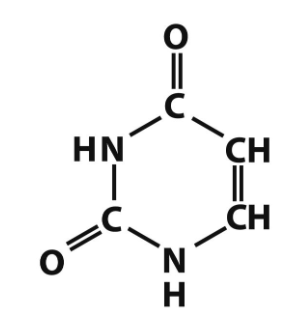
uracil structure
adenine → DNA and RNA
guanine → DNA and RNA
purine bases and what they are found in (DNA/RNA)
N-glycosidic bond
to position N1 in pyrimidines
to position N9 in purines
what type of bond is the pentose ring attached to the nucleobase via in nucleotides and to what position in pyrimidines vs purines
acid
what is N-glycosidic bond cleavage catalyzed by
purines: -sine (ex: adenosine)
pyrimidines: -dine (ex: cytidine)
what does nucleoside nomenclature end in in purines and pyrimidines
-ylate (ex: adenylate)
what does nucleotide nomenclature end in
eukaryotes (but also found in bacteria)
what organisms is 5-methylcytosine modification common in
bacteria (NOT found in eukaryotes)
what organism is N6-methyladenosine modification common in
prokaryotes → way to mark own DNA so that cells can degrade foreign DNA
eukaryotes → way to mark which genes should be active
purpose of epigenetic marker (such as methylation) in eukaryotes and prokaryotes
they are degraded by sequence-specific restriction enzymes and cleaved
what happens to foreign DNAs (not methylated) that are introduced into the cell
hydrogen bond
type of bond between two bases to form a base pair
A - T
C - G
Purines with pyrimidines
Watson-Crick base pairs in dsDNA
isolated “nuclein” from cell nuclei → hydrolysis of it produced phosphate, pentose, and a nucleobase → chemical analysis revealed phosphodiester linkages and pentose is ribofuranoside
what did Friedrich Miescher do
Watson, Crick, and Wilkins
who shared Nobel prize for discovering DNA double helix
5’ to 3’
what direction is DNA sequence read in
DNA polymerase
synthesis of new strand of DNA is catalyzed by what enzymes
one daughter strand and one parent strand
newly made DNA molecule has what 2 strands
mRNA
RNA that is the code carrier for the sequence of proteins
DNA template
what is mRNA synthesized from
contains ribose instead of deoxyribose
contains uracil instead of thymine
difference between mRNA and DNA template
yes
can one mRNA code for more than one protein
monocistronic has one promoter per gene → eukaryotes
polycistronic have one promoter for multiple genes → prokaryotes
monocistronic vs polycistronic
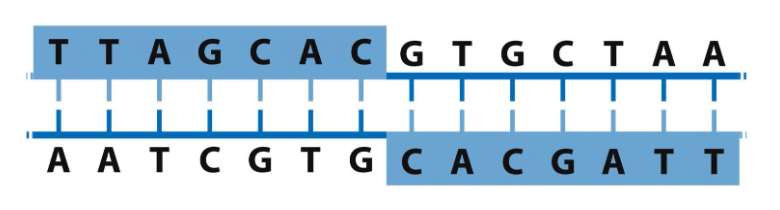
it’s a sequence that is the same forwards and backwards (on opposite strands) → can form hairpins and cruciforms
what is palindromic sequence and what can they form
1 strand of DNA or RNA
how many strands are involved in a hairpin structure
covalent bonds remain intact (along with genetic code)
hydrogen bonds are broken (strands separate)
base stacking is lost (UV absorbance increases)
what happens to DNA when denatured
high temperature or change in pH
what can induce DNA denaturation
annealing
name of the process of reversing denaturation
the polymerase chain reaction
what does the reversible thermal denaturation and annealing of DNA form the basis for
260 nm
DNA is commonly monitored by UV spectrophotometry at what wavelength
base composition (high CG increases it)
DNA length (longer DNA increases it)
pH and ionic strength (high salt increases it)
what 3 things does the midpoint o melting depend on
hybridize
what can 2 near-complementary DNA strands do
N-glycosidic bond
what bond is hydrolyzes in depurination
cytosine → uracil
5-methylcytosine —> thymine
adenine → hypoxanthine
guanine → xanthine
deamination results for cytosine, 5-methylcytosine, adenine, and guanine
guanine and apurinic residue
resulting 2 products of deamination of guanosine residue
hydroxylation of guanine → mitochondrial DNA is most susceptible
oxidative damage
methylation of guanine
chemical alkylation
nitrogen mustard, dimethylnitrosamine, and dimethylsulfate
3 alkylating agents
dimerization of pyrimidines
what mutation does UV light induce
ring opening and strand breaking
what mutation does ionizing radiation cause
aging and carcinogenesis
accumulation of mutations is linked to what