Cell Biology- Final Exam
1/88
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
89 Terms
Describe cleavage furrow formation in animal cells.
cleavage furrow is formed by a belt-like bundle of actin filaments called the contractile ring
contractile ring forms just beneath the plasma membrane (aka in the cortex) during anaphase
cleavage furrow → cytokinesis (cytoplasm division)
as cleavage progresses, the ring of actin tightens around the cytoplasm eventually pinching the cell in 2
entire contractile ring is dismantled shortly after cytokinesis is complete
During cleavage furrow formation in animal cells a ring of actin tightens around the cytoplasm and pinches the cell in 2- how is the actin ring tightened?
tightening of the actin ring involves interactions between actin filaments and myosin II
movement of myosin along the actin causes the contraction and tightening of actin filaments
Describe cell plate formation.
plant cell cytokinesis
microtubule associated
vesicles travel on microtubules to midline in cell division
during late anaphase a group of small vesicles derived from the golgi complex align themselves across the equatorial region of the spindle
vesicles contain polysaccharides and glycoproteins required for cell wall formation
vesicles contain polysaccharides and glycoproteins required for cell wall formation
vesicles are guided to the spindle equator by an array of microtubules and associated motor proteins
microtubule motor proteins: dynein, kinesin, myosin
vesicles fuse together to produce a cell plate which represents the cell wall in the process of formation
vesicles fuse together to produce a cell plate which represents the cell wall in the process of formation
plant cell cytokinesis DOES NOT INVOLVE ACTIN
What role does microtubules play in the formation of the cell plate.
vesicles travel on microtubules to midline in cell division
What cytoskeletal filaments are needed for cytokinesis in animal cells and plant cells?
animal cells: actin and microtubules
plant cells: ONLY microtubules
How do cells move? What are the steps of movement?
cells move by protruding a portion of the cytoplasm
steps in movement:
1. protrusion
2. adhesion
3. contraction
What types of cells need to move?
amoeba, cells of immune system, embryonic cells, macrophages, fibroblasts during development
Do cilia and flagella use actin, microtubules, or both?
only microtubules
Describe the first step of cell locomotion.
1. protrusion- front of cell pushes out an extension called a lamellipodium
lamellipodium filled with actin filaments that have (+) end facing towards plasma membrane
formation of lamellipodium:
actin assembly is required for protrusion (assembly pushes on leading edge of cell membrane to form protrusion)
myosin I moves along actin filaments causing them to slide past one another
Describe the second step of cell locomotion.
2. adhesion
cells must adhere to substrate in order to move
adhesion is mediated by integrin protein
integrin: transmembrane adhesion protein
bound to an intracellular bundle of actin filaments
bound to extracellular matrix
focal adhesion
Describe the third step in cell locomotion.
3. contraction
back of cell contracts to move the front of the cell forward
maybe myosin based since myosin II has been localized to the rear end of the cell
myosin II in rear of cell interacts with actin filaments to contract the filaments (propels cell forward)
Describe the cell motility of disease-causing organisms and the role of actin in them.
disease causing microorganisms can use the cells normal cell adhesion and cell motility systems to enter a cell
gram (+) bacteria - Lysteria monocytogenes
infects cells by binding to a cell adhesion molecule on the cell surface and enters the cell (phagocytosis) [not destroyed by the lysosome]
once inside the cell, the bacterium moves in the cytoplasm where it can divide rapidly
the bacterium moves inside the cell by producing a “comet tail” of polymerized actin [using the cytoplasmic g-actin of the host cell]
Describe microtubule associated motor proteins.
motor proteins: kinesin, dynein, myosin I
kinesin: (+) end directed, movement mediated by ATP hydrolysis
ex: movement of chromosomes during mitosis; intracellular vesicle transport
dynein: (-) end directed, movement by ATP hydrolysis
ex: movement of chromosomes during mitosis, movement of sperm flagella
myosin I: (+) end directed, movement by ATP hydrolysis
functions: form mitotic spindle, chromosome segregation in mitosis, intracellular transport, anchor organelles
What is the role of microtubules in cell divison?
form mitotic spindle
microtubules radiate from centrosome [(-) embedded in centrosome]
centrosome: forms poles of mitotic spindle
centromere
kinetochore
microtubule types: astral, polar, kinetochore
How do chromosomes move in cell division?
microtubules can change length while attached to the kinetochore
tubulin subunits add to the (+) end
tubulin subunits can be removed at the kinetochore or at the pole
Describe microtubule length change in cell divison.
microtubule length change:
microtubule lengthens as subunits are added to the (+) end
lengthens from the pole to capture the kinetochore
microtubule lengthens to find a target (chromosome)
Describe the role microtubule motor proteins play in cell divison.
microtubule motors
dynein located at kinetochore is trying to move along the filaments towards the (-) end
chromosomes move toward pole
tubulin subunits lost where filaments connect to kinetochore
kinesin is located at poles and is trying to move toward (+) end
subunits removed at pole region (overall shortening of microtubule)
combo of dynein and kinesin moves chromosomes during anaphase
role of katanin:
microtubule associated protein
severing protein promotes removal of tubulin
localized to pole region
Describe role of microtubules in the metaphase stage of cell divison.
balance between forces exerted by kinesin at poles and dynein at kinetochore
no overall loss of subunits results in chromosome alignment
process not well understood
Describe loss of function experiments associated with the cell cytoskeleton.
knock-out or loss of function (kinesin/dynein)
knock-out or block protein function
look at position of the chromosomes
look for a chromosome shift or absence of chromatid segregation
stain chromosomes following loss of function (ex: DAPI)
stain microtubules using immunofluorescence and look for shortening or abnormalities
What are some cell cycle controls?
p53
transcription factors
cell cycle checkpoints
apoptosis
What is p53? What is its function?
p53: tumor suppressor gene that encodes a transcription factor
functions:
activate DNA repair enzymes
can halt a cell at G1/s (regulation point- halts growth and DNA replication)
activate apoptosis (programmed cell death)
What role does the p53 gene play in HPV and cervical cancer?
HPV: human papilloma virus
upon infection, HPV produces a viral protein called E6
E6 can bind to p53 protein and inactivate it
What role does transcription factors play in the cell cycle? What would a mutation in transcription factor do?
needed to activate transcription
bind to enhancer sequences and other DNA regulatory sequences to loosen interaction of DNA with histones and allow RNA polymerase to bind
mutation in the transcription factor can:
prevent activation of gene expression (DNA repair enzymes or proteins involved in cell cycle checkpoints)
What is the purpose of cell cycle checkpoints?
allow for cells to progress through cell cycle stage without errors
genes encode for cell cycle checkpoint proteins
oncogene is a mutated form of ones of these genes
What is the role of apoptosis in the cell cycle?
aka programmed cell death
biochemical events lead to changes in cell morphology and cell death
changes in cell shrinkage, nuclear fragmentation, chromatin condensation, and chromosomal DNA fragmentation
dependent on family of proteins called caspases
caspases:
cleave nuclear lamins causing the break down of the nuclear envelope cleave inactive forms of DNase to create an active form
DNase: degrades DNA
How do elephants and other large mammals crush cancer?
have more cellular DNA (more mutations)
numerous rounds of cell division (more chances for DNA errors)
yet large mammals can survive up to 60 years in the wild
elephants don’t develop cancer:
evolved to have 40 copies of the p53 gene (humans have 2 copies)
cells with damaged DNA are required or apoptosis is triggered
What are anti-mitotic drugs? What are some examples?
anti-mitotic drugs: used to treat cancer (don’t target only cancer cells)
Vinblastine: binds to tubulin and prevents microtubule formation
tubulin is a dimer (α and β tubulin)
α and β subunits join to form protofilaments
protofilaments form wall of microtubule
Taxol: binds to microtubule and prevents disassembly
What is taxol and how does it stop cell divison or mitosis?
taxol: binds to microtubule and prevents disassembly
the microtubules can’t disassemble during anaphase to search for chromosomes
chromosome movement during cell division requires disassembly of microtubules
if microtubules cannot disassemble, chromosomes can’t segregate anaphase
taxol-treated cells divide abnormally into more than 2 cells
cells have jumbled chromosomes and are destroyed by apoptosis
What is metastasis?
migration of cancer cells
mechanism is unknown (actin mechanism)
cells break cell adhesion
What role does the cellular cytoskeleton play in neurodegenerative diseases (such as alzheimers and parkinsons)?
cause a change in the shape of the axon of neurons
axon microtubules are stable- do not undergo dynamic instability
Tau protein (MAP): binds to and stabilizes microtubules
abundant in neurons of CNS (also found in other cells)
microtubules collapse into a constricted structure (not well understood)
abundant abnormal aggregates of cytoskeletal proteins in neurons and glial cells of CNS (ex: tau protein)
tau protein becomes phosphorylated
phosphorylated tau forms aggregates (clumps)
causes constriction of microtubule bundles
phosphorylated tau can leave an affected nonfunctioning neuron and attach to a healthy neuron
What is the function of microtubules (with associated proteins)?
guide intracellular transport
anchor intracellular organelles
form mitotic spindle
function in chromosome segregation during cell divison
Define centrosome, centromere, and kinetochore in terms of mitosis.
centrosome: poles
centromere: holds chromosomes/chromatids together and connects chromosome to spindle fibers via the kinetochore
kinetochore: complex of proteins and RNA molecules attached to the centromere
What is intracellular transport?
proteins made in the cytoplasm must be transported to different compartments in the cell
proteins carried in vesicles
vesicles attach to cytoskeletal filaments (ex: microtubules, actin)
movement of vesicles is mediated by motor proteins
Regarding intracellular transport, how do proteins find their correct target?
signal sequence: stretch of amino acids that specifies a destination
Describe the structure of the nucleus and the nuclear pore.
nuclear envelope: double membrane, contains nuclear pore complexes
nuclear pore structure:
octagonal structure of repeating subunits that forms a channel through the membrane
repeating subunits: nucleoporins
fibrils extend into the cytoplasm and nucleus: nuclear cage
How do small and large molecules transport across the nuclear envelope?
small molecules move through the aqueous channel by diffusion (ex: water, small ions)
large molecules:
move through the channel by active transport (require energy)
proteins to be imported require a nuclear localization signal
nuclear localization signal: stretch of amino acids in the protein sequence that directs the proteins to the nucleus
ex: histones, proteins, ribosomes, DNA replication enzymes, proteins needed for transcription, mRNA
Provide an example of a molecule that would be exported from the nucleus.
mRNA, ribosomes (produced by the nucleolus)
Describe the mechanism of transport for large molecules being imported into the nucleus.
active transport
nuclear import receptors:
soluble cytoplasmic proteins (importins)
soluble protein receptors found in cell cytoplasm
recognize the nuclear localization signal on the protein to be transported
recognize the nucleoporin structure (FG repeats)
FG repeats: short stretches of phenylalanine and glycine
importin proteins bind to FG repeats
in nucleoporin- proteins that form nuclear pore
guide the transport of proteins through the pore (post-translational)
how does transport occur?:
transport is powered by the hydrolysis of GTP: GTP binds to the import receptor and pore
hydrolysis of GTP causes a conformational change in pore structure
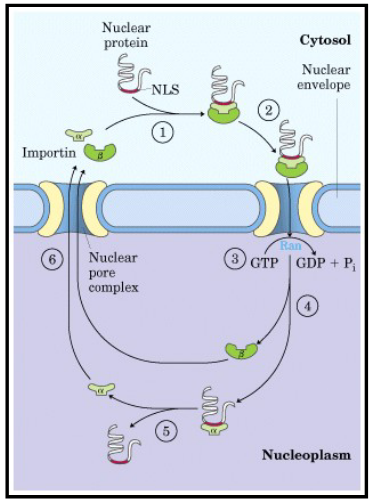
Describe the export of large molecules out of the nucleus.
same process that is used for import of molecules
nuclear export signals are located on molecules to be exported
nuclear export receptors: soluble proteins
export receptors bind to the export signal and the nucleoporin and guide the cargo through the pore to the cytoplasm
export is powered by the hydrolysis of GTP
Describe protein synthesis in different parts of the cell.
populations of ribosomes in:
1. cytoplasm: structural proteins (ex: actin and tubulin), enzymes needed for metabolism
2. associated with ER: soluble proteins that are destined for secretion (ex: hormones), resident ER proteins
3. mitochondria and chloroplasts: synthesize proteins needed to carry out their own function
Where are proteins destined for the ER synthesized?
proteins destined for the ER are initially synthesized on cystolic ribosomes
Describe protein transport to the ER.
ER signal sequence directs the protein to the ER
ER signal sequence: stretch of amino acids within the protein sequence
signal recognition particle (SRP) recognizes and binds to the signal sequence
SRP:
composed of 6 polypeptides and an RNA molecule
binds to signal sequence on protein being synthesized
binding of the SRP temporarily halts translation of the protein
SRP/ ribosome complex is transported to the ER membrane
SRP binds to a receptor on the ER membrane
SRP is released from the complex and translation resumes
as the protein is translated, it is threaded through a pore in the ER membrane (requires the hydrolysis of GTP)
co-translational import
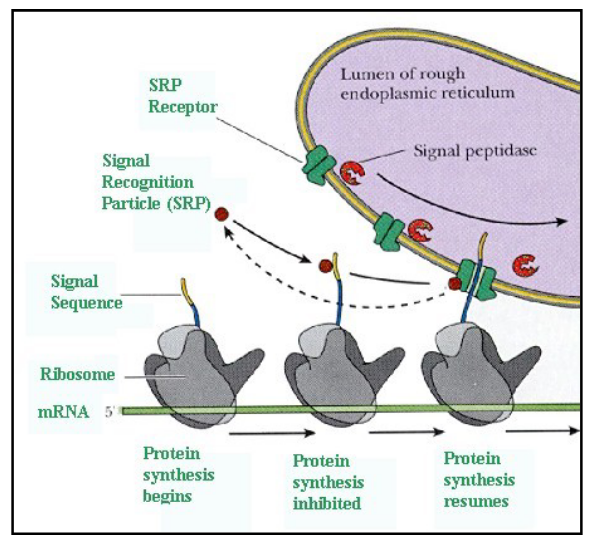
What is the main difference between nuclear protein transport and protein transport into the ER?
nuclear transport: post-translational
ER transport: SRP, co-translational
What types of proteins get transported to the ER?
soluble/ secretory proteins
single pass trans-membrane ER proteins
multi pass trans-membrane ER proteins
Describe transport of soluble/secretory proteins to the ER.
ER signal sequence acts as a start transport sequence
ER signal sequence is cleaved off by the signal peptidase enzyme
protein is released into the endoplasmic reticulum
co-translational transport
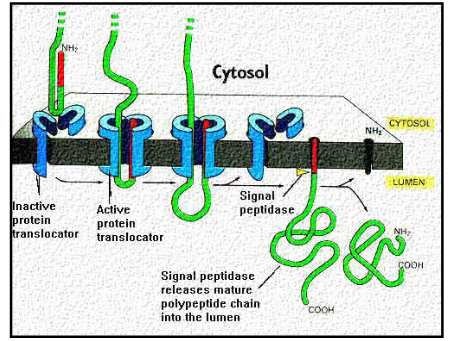
Describe transmembrane proteins in the ER and the two types.
transmembrane proteins:
span width of membrane
contain a domain that faces outside ER and a domain that faces ER lumen
single pass transmembrane proteins:
cross membrane one time
multi pass proteins:
cross membrane more than once
Describe transport of single-pass transmembrane proteins into the ER
ER protein: signal sequence initiates translocation (as described for soluble proteins)
signal sequence found at amino terminal end of protein or in middle of sequence
stop transfer signal:
anchors the protein in the membrane
middle of protein sequence
Describe transport if multi-pass transmembrane proteins to the ER.
crosses the membrane more than once
protein signal sequence initiates translocation (as previously described)
protein signal sequence is located in the middle of the polypeptide sequence (portein signal sequence serves as the start transfer sequence)
ALWAYS in the middle whereas for single pass the signal sequence can be at the end or the middle
protein is threaded through the membrane until the stop transfer sequence
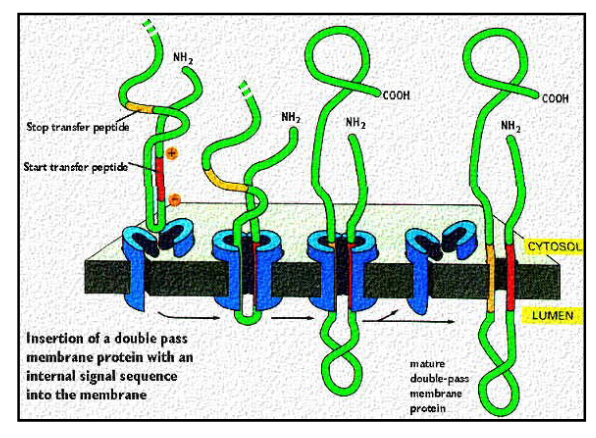
Why do proteins need to be folded in the ER?
ER soluble proteins ae passed through the membrane unfolded and must refold inside the lumen of the ER
What are binding proteins (BIP)?
specific to ER (resident ER protein)
class of proteins that bind to the incoming soluble proteins and aid in protein folding (ER)
chaperone protein
How do chaperone proteins work?
found in ER and cytoplasm
process is not well understood
some chaperone bind to proteins to be folded and protect the protein from degradation by forming a large aggregate with the protein
once the protein is properly folded, the chaperone releases
other chaperones: heat shock proteins (Hsp60, Hsp70)
disruption of the folding process can affect cell development and growth
abnormalities in chaperone proteins can cause: photoreceptor degeneration in the eye, CNS abnormalities, male infertility
What are Hsp chaperone proteins?
heat shock proteins
produced or activated in response to an increase in temperature
help prevent protein denaturation or protein unfolding
What are the two methods of protein degradation?
degradation by the lysosome
ubiquitin proteosome pathway (UPP)
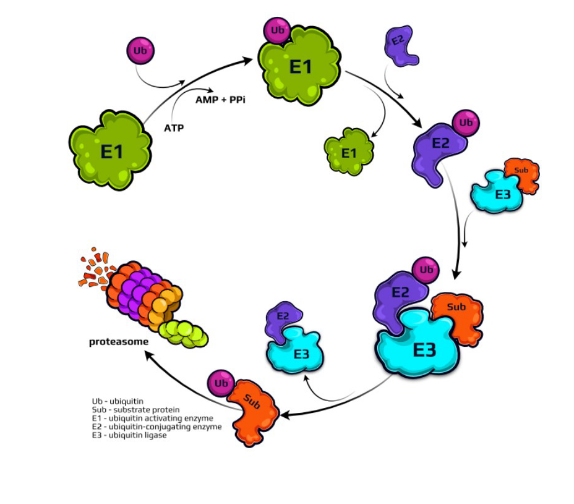
What is the ubiquitin proteosome pathway?
method of protein degradation for incorrectly folded proteins/ short lived proteins
two steps:
1. tagging of the substrate protein by the covalent attachment of multiple ubiquitin molecules
ubiquitin: small protein (76 amino acids in size)
role of enzymes (E1, E2, E3)
E1: ubiquitin activating enzyme
E2: ubiquitin conjugating enzyme
E3: transfers ubiquitin from E2 to protein to be degraded
protein destined for degradation must be tagged with multiple ubiquitin’s
poly ubiquitin protein vs. mono ubiquitin protein
2. subsequent degradation of the tagged protein by the proteasome complex
large proteolytic complex that breaks down the tagged portein
proteins targeted by this system are short-lived proteins
regulatory proteins (ex: cyclin = cell cycle regulator)
cyclin protein combines with CDK protein (cyclin dependent kinase) to regulate movement through cell cycle
quicker than lysosome degradation
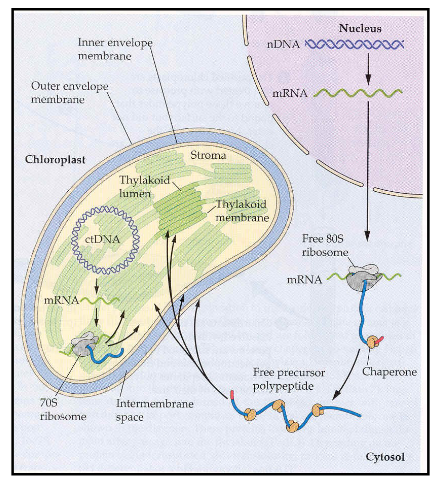
Similarities between protein import in mitochondria and chloroplasts?
proteins destined for mitochondria and chloroplasts are synthesized in the cytoplasm on cystolic ribosomes
contain a signal sequence that specifies their destination
transported into the organelles post-translationally
signal sequence is cleaved by peptidases following import
What are the functions of cell membranes?
form a cell boundary- confine biochemical reactions
sense external signals
protection
transport
cell shape
cell signalling
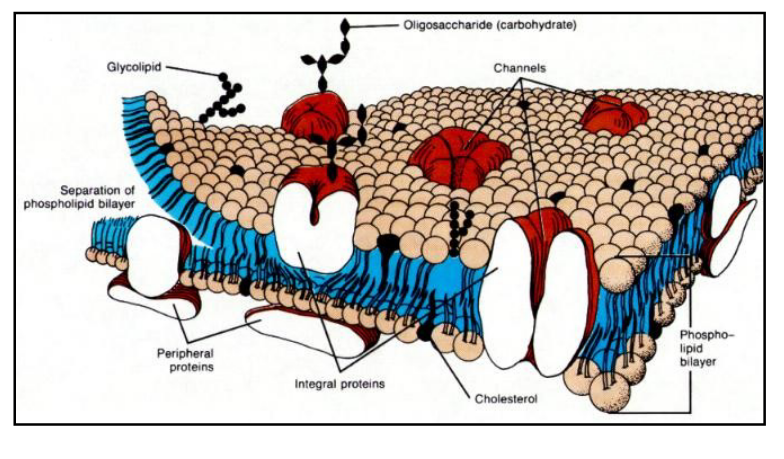
Describe the structure of the cell membrane.
composed of lipids and proteins
described by the fluid mosaic model:
membrane lipids are arranged in bilayer
proteins are embedded in the bilayer
fluidity:
lipid molecules able to move and behave more like a liquid than a solid
lipids can diffuse laterally in the membrane
cell membrane is a dynamic structure
molecules in cell membrane held together by noncovalent interactions
What are the reasons for fluidity in the cell membrane?
1. phospholipid tails are short: increases fluidity
because short tails reduce the tendency of the lipids to react with one another
2. phospholipid tails contain double bonds: increases fluidity
because double bonds create kinks in fatty acid tail and presence of kinks prevents packing together of phospholipids
What are the three types of cell membrane lipids?
phospholipids
glycolipids
cholesterol
Describe phospholipids.
membrane lipid
polar head (hydrophilic) and nonpolar tails (hydrophobic)
ex: phosphotidylcholine:
head group: glycerol, choline, phosphate group
tail region: fatty acid tails
amphipathic: contain hydrophilic and hydrophobic regions
most abundant lipid in the membrane
phospholipids can move laterally in the membrane
Based on what we have discussed in lecture, briefly describe two
different cellular mechanisms or processes that when defective can lead to
the appearance and accumulation of unfolded or misfolded proteins
within cells
chaperone proteins: mutated or missing chaperone proteins could lead
to misfolded proteins or unfolded proteins.lysosome: non-functioning organelle could lead to the accumulation of
misfolded or unfolded proteins in a cell.Ubiquitin Proteosome Pathway: pathway does not function correctly or
at all and could lead to the accumulation of misfolded or unfolded
proteins in a cell.
Describe glycolipids.
membrane lipid
lipids with a carbohydrate side chain (sugar side chain is exposed at the cell surface)
function: protection of cell from harsh conditions, cell recognition and cell signaling
prominent in membranes of nerve cells (brain) and epithelial cells that line the intestine
What is an example of a bacterial infection interacting with glycolipids?
vibrio cholerae produces cholera toxin (enterotoxin)
V. cholerae- bacteria occurs in both marine and freshwater habitats
pathogenic to humans
the toxin acts on the mucosal epithelium of digestive tract
causes a sudden onset of massive diarrhea causing the individual to lose gallons of protein-free fluid and associated electrolytes, bicarbonates, and ions
enterotoxin binds to glycolipids on the surface of the intestinal cells
binding activates the adenylate cyclase enzyme in the cells converting the enzyme into a pump which extracts water and electrolytes from blood and tissues and pumps it into the lumen of the intestine (dehydration)
Describe cholesterol.
membrane lipid
found in some membranes
steroid: ring structure
ring interacts with tails of phospholipids
presence of cholesterol decreases the fluidity of the membrane
What are the types of membrane proteins?
integral
peripheral
lipid-anchored
Describe integral membrane proteins.
contain one or more hydrophobic regions embedded in the lipid bilayer
most integral membrane proteins are transmembrane proteins
ex: transmembrane protein
membrane protein that extends through the lipid bilayer
single pass and multi-pass proteins
multi-subunit proteins- composed of several polypeptide chains
Describe peripheral membrane proteins.
found on the periphery of the membrane attached to phospholipid head groups or other adjacent proteins
attach to the membrane by electrostatic interactions and H bonds
How would you purify transmembrane proteins and peripheral proteins?
1. transmembrane proteins: use detergent to break membrane
2. peripheral proteins (attached to membrane with weak interactions): can shift pH to break interactions
Describe lipid anchored membrane proteins.
peripheral protein
covalently bound to lipid molecules that are embedded in the lipid bilayer
linked to the outer membrane surface
attached by GPI anchor
linked to the inner membrane surface
attached by a fatty acid group
Describe the fluid property of membrane proteins.
fluidity
proteins can rotate in the membrane
proteins can diffuse laterally in the membrane
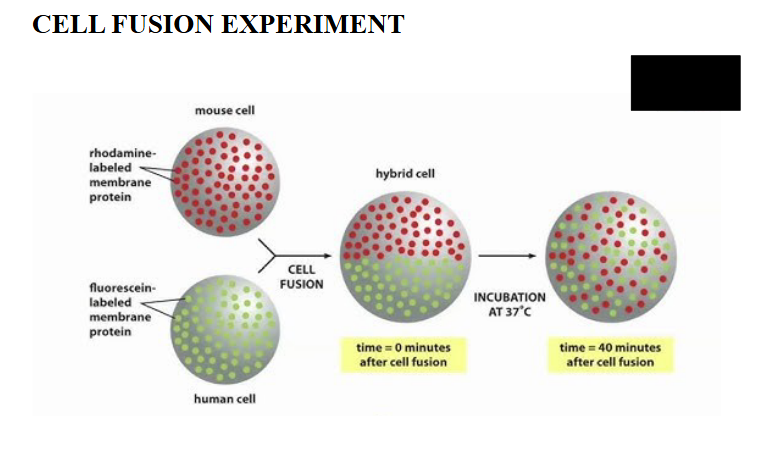
How was membrane protein fluidity discovered?
two experiments:
1. mouse/human cell fusion
mixing of mouse cell membrane proteins and human cell membrane proteins after cell fusion
membrane proteins diffuse over time, can be seen by mixing of cells
2. FRAP (fluorescent redistribution after photo-bleaching)
label a membrane protein with a fluorescent antibody
bleach the fluorescent molecules in a small area using a high intensity laser beam
fluorescence intensity recovers as the bleached molecules diffuse away and unbleached molecules diffuse into the irradiated area
What are factors that might prevent the movement of membrane proteins within the membrane?
proteins are linked to extracellular matrix, membrane components, cellular junctions (cell-cell interaction)
proteins can be attached to phospholipids, cholesterol, other proteins
cell junctions limit membrane protein mobility- proteins can migrate up to the junction but not past it
cells within a tissue layer have two surfaces:
apical surface
basal surface- attaches cells to ecm
What are the types of cell junctions?
tight junctions, gap junctions, anchoring junctions
Describe tight junctions.
bind cells in a tissue layer
prevent the “leaking” of material between cells
Describe gap junctions.
tunnels that connect cells
allow for exchange of materials between cells
allow for easy exchange of materials between cells- especially important in cell signaling and exchange of cell signal products between cells
opening and closing of gap junctions is a regulated process (controlled by levels of intracellular calcium)
high levels of intracellular calcium = closes cell junctions
PIP2 pathway (release calcium into the cell)
tested by labeling molecules with tag and looked to see if calcium molecules moved between cells
Describe anchoring junctions.
connect cells to each other and to extracellular matrix
can be connected to the cytoskeleton
focal adhesions
Difference between isolating peripheral and integral proteins?
1. peripheral proteins
extracted from the membrane by changes in pH or ionic strength
2. integral proteins
removed by detergents which disrupt the hydrophobic interactions between lipids in the bilayer
detergent: Triton X, NP-40
What are the steps of separating isolated proteins?
SDS-PAGE
Protein detection
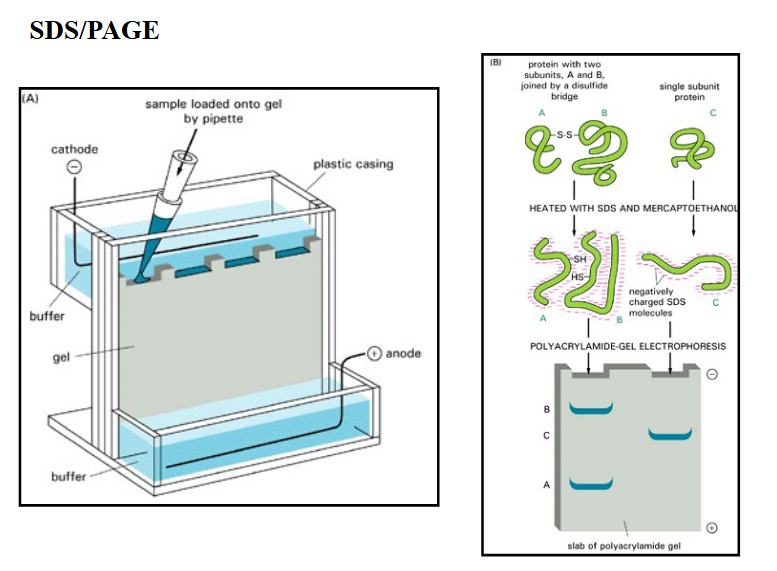
What is an SDS-PAGE?
separating isolated proteins
SDS-PAGE: SDS polyacrylamide gel electrophoresis
SDS (sodium dodecyl sulfate)
denatures proteins and coats them with a negative charge
electrophoresis separates proteins based on size
polyacrylamide gel: PAGE
negatively charged proteins migrate in the gel toward the (+) electrode
migration rate is dependent on size of the protein
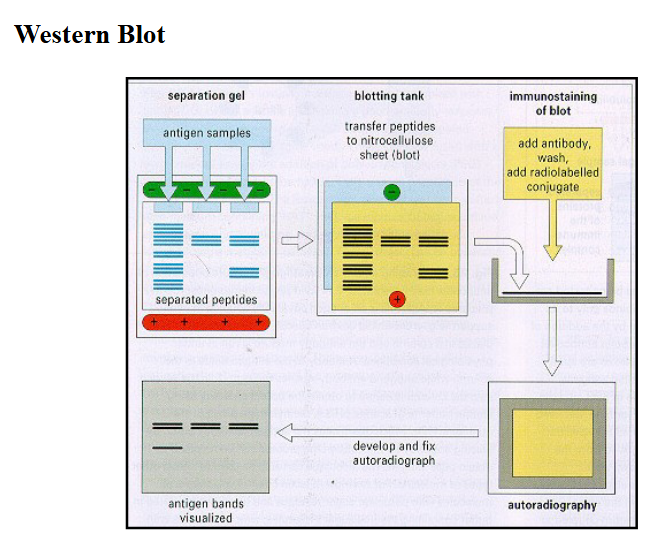
How would you detect proteins once you separate them?
Western blot
ELISA (enzyme linked immunosorbent assay)
indirect ELISA → HRP
microtiter plates
apply antigen to the surface of the plate
add antibody specific for antigen (use 1 or 2 antibodies)
antibody contains enzyme tag (HRP)
add substrate (peroxidase)
monitor color change
Describe cell membrane transport.
membrane transport is affected by size and polarity of the molecule
small nonpolar molecules can diffuse rapidly across the lipid bilayer
ex: oxygen, carbon dioxide
small uncharged, polar molecules such as water and urea will slowly diffuse across the lipid bilayer
ex: water (osmosis)
lipid bilayers are impermeable to charged molecules
the charge on the molecule and the molecule’s degree of hydration prevent it from entering the bilayer
ex: Na+, Ca2+, K+
membrane transport is mediated by membrane porteins
two main classes of membrane transport proteins: carrier proteins and channel proteins
Describe carrier proteins.
bind to the molecule to be transported
undergo a series of conformational changes to transfer the bound molecule across the membrane
carrier mediated transport:
1. uniport: transport of a single molecule (ex: Ca2+ transport [ER membrane])
2. symport: transport of 2 molecules (coupled transport), occurs in a single direction (ex: glucose, Na+)
3. antiport: transport of 2 molecules (coupled), occurs in opposite directions (ex: Na+/K+ pump)
![<ul><li><p>bind to the molecule to be transported</p></li><li><p>undergo a series of conformational changes to transfer the bound molecule across the membrane</p></li><li><p>carrier mediated transport:</p><ul><li><p>1. uniport: transport of a single molecule (ex: Ca<sup>2+</sup> transport [ER membrane])</p></li><li><p>2. symport: transport of 2 molecules (coupled transport), occurs in a single direction (ex: glucose, Na<sup>+</sup>)</p></li><li><p>3. antiport: transport of 2 molecules (coupled), occurs in opposite directions (ex: Na<sup>+</sup>/K<sup>+</sup> pump)</p></li></ul></li></ul><p></p>](https://assets.knowt.com/user-attachments/bde927cb-f1e2-4fa3-a629-1fd140637399.png)
Describe passive transport.
1. passive transport: occurs by diffusion
two types of diffusion:
simple:
concentration gradient drives transport
diffusion occurs from an area of high concentration to an area of low concentration
facilitated:
diffusion of molecules through the use of carriers of channels in the protein
What are two methods of driving membrane transport?
passive transport
active transport
Describe the methods of active transport.
1. coupled carriers
free energy released during the movement of one molecule down an electrochemical gradient is used as the force to pump the other molecule against the electrochemical gradient
energy stores in the electrochemical gradient of one molecule is used to drive the movement of the other molecule
ex: sodium and glucose
2. ATP driven pump
uses hydrolysis of ATP to pump molecules across the membrane
ex: Na+/K+ pump found in the plasma membrane of most animal cells
Na+ is pumped out of the cell against its electrochemical gradient
K+ is pumped into the cell
hydrolysis of ATP powers transport
3. light driven pump
found mainly in bacteria (halophilic bacteria)
couple transport against an electrochemical gradient to an input of light energy
carrier proteins are light-gated
absorption of light causes conformational change in the carrier protein
Describe channel proteins.
form aqueous pores that extend across the lipid bilayer
when pores open, molecules can pass
transport occurs quickly
can be channel protein or a gated-channel protein
gated channel protein requires stimuli to open the channel
What are the three stimuli that can open membrane channels?
1. voltage change across the membrane: dependent on membrane potential
voltage gated channels
ex: muscle contraction and nerve function
2. mechanical stress: connected to cell cytoskeleton
mechanically gated channels
many of the channels have cytoplasmic extensions that link the channel to the cytoskeleton
3. ligand-gated: connected to cell surface receptors
binding of a ligand opens the channel
ligand can be an extracellular mediator (ex: neurotransmitter)
ligand can be an intracellular mediator (ex: nucleotide-ATP)
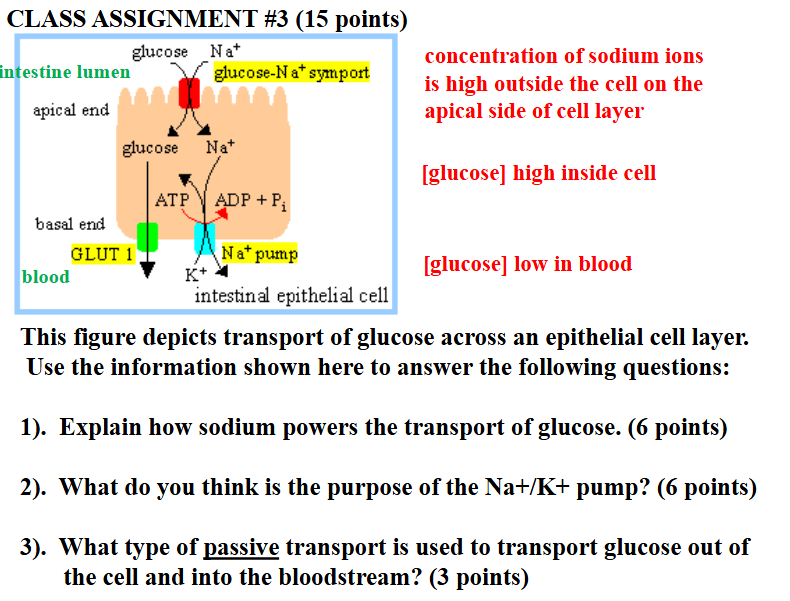
glucose is pumped into the cell by a sodium-powers glucose symport (coupled transport that occurs in the apical domain of the cell)
the sodium gradient driving the glucose symport is maintained by a sodium pump which keeps the internal concentration of sodium low
glucose passes out of the cell (down its concentration gradient) by passive transport mediated by a glucose carrier protein (basal domain of the cell) = facilitated diffusion
ESSAY QUESTION: POSSIBLE TOPICS:
COLOCALIZATION STAINING
ORGANELLE/PROTEIN ISOLATION AND IDENTIFICATION
CELL FUNCTION EXPERIMENTS (SHOW THE IMPORTANCE OF
A MOLECULE IN A SPECIFIC CELL FUNCTION)
EXOCYTOSIS (PROTEIN PRODUCTION TO SECRETION)
ISOLATION AND IDENTIFICATION OF MEMBRANE PROTEINS