exam 3
1/89
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
90 Terms
polymorphism
common rare mutation within a population
variant
relatively rare mutation within a population
reversion
restoration of the original phenotype to a mutant
haploinsufficiency
single functional copy of a gene is not sufficient
pleotropic
single mutation that affects two or more seemingly unrelated traits
somatic mutations
mutations in non-reproductive cells - only affect a portion of an organism
germline mutations
mutations in reproductive cells - affect some of the offspring
silent mutation
no effect on protein
synonymous mutation
minimal effect on protein
evident mutation
clearly see a phenotypic change
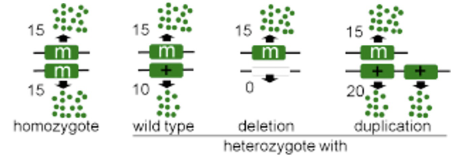
amorph
null allele, a complete loss of function, usually recessive
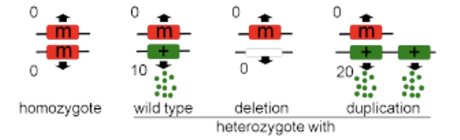
hypomorph
partial or reduced function, usually recessive
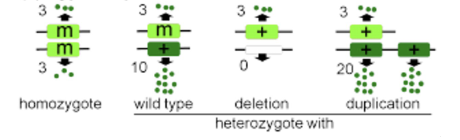
hypermorph
increased function or expression
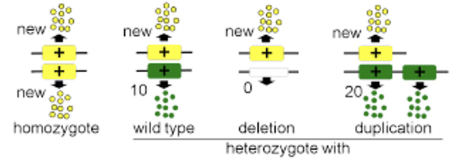
neomorph
protein has a new and different function
antimorph
activity that is dominant and opposite to the wild-type function; also known as dominant negative mutations
multimorphs
show multiple effects
non-conditional mutation
mutant phenotype present under all conditions
conditional mutation
mutant phenotype expressed only under certain conditions
spontaneous DNA damage
damage due to natural processes
induced DNA damage
damage from external sources
point mutation
change a base pair to a different base pair
transition mutation
purine → purine or pyrimidine → pyrmidine
transversion mutation
purine → pyrimidine or pyrimidine → purine
missense mutation
DNA change leads to change in amino acid
nonsense mutation
DNA change leads to formation of a transcriptional stop site
TAG
amber
TAA
ochre
TGA
opal
frameshift
insertion or deletion of 1 or more bases that is not a multiple of 3
deletion mutation
removal of a large section of a gene, entire gene, or multiple genes
insertion mutation
large amount of DNA added into gene - prevents normal function
inversion mutation
chromosomal rearrangement, affects multiple genes, not leaky, can revert
duplication mutation
have additional copies of a gene or region, due to unequal recombination events between daughter chromosomes
depurination
purine is removed by hydrolysis resulting in an apurinic site
depyrimidation
pyrimidine is removed by hydrolysis resulting in an apyrimidinic site
deamination
amine group on cytosine, or 5-methylcytosine can be removed
oxidation
reactive forms of oxygen produced during metabolism react with and alter DNA bases
base analogues
compounds similar in structure to a DNA base that can be incorporated into DNA during replication
nitrous acid
deaminates bases
autonomous transposable elements
encode all enzymes needed to move
non-autonomous transposable elements
can only move if supplied with enzyme from another transposable element
pericentric
inversion includes the centromere
paracentric
inversion does not include the centromere
translocation
joining of non-homologous chromosomes
true reversion
mutation exactly restores original DNA sequence
pseudo-reversion
second mutation restores the original phenotype
interaction suppressors
suppress mutations by restoring protein-protein interactions
allele specific
bypass suppressors
turn on a new pathway (or upregulate another pathway) that eliminates the need for a particular gene
physiological suppressors
general changes in cell physiology suppress the effects of the original mutation
informational suppressors
changes how the cell reads the information encoded in the mRNA
misreading the mutant mRNA allows the production of a functional protein from the mutant gene
micronucleus
particle in a cell that contains nuclear DNA; it might contain whole chromosomes, or broken pieces of chromosomes
photolyase
cleaves TT dimers in the presence of visible light
MutS
binds to G-T mismatch
MutL
helps recruit Vsr to T:G mismatch
Vsr
forms complex with MutS & MutL
endonuclease
cuts DNA backbone 5’ to the incorrect T
MutM
removes 8oxoG
MutY
removes A
MutT
degrades 8oxoGTP
synapse
point at which 2 DNA molecules held together by complementary base pairing between strands
heteroduplex
region of complementary base pairing in the synapse
translational coupling
mRNA secondary structure prevents binding of ribosomes unless previous region is being translated
catabolic operons
involved in the breakdown of sugars to provide energy for growth
anabolic (biosynthetic) operons
involved in the synthesis of various essential compounds, such as amino acids
effector molecules
bind a regulatory molecule and change its activity
inducers
bind to repressor or activator and allows initiation of transcription
corepressors
bind to a repressor molecule allowing it to block transcription
aporepressor
inactive repressor without bound corepressor
negative regulation
repressor must release the DNA to allow for transcription
may require the presence of a co-repressor to prevent transcription from occurring
positive regulation
activator must bind the DNA to allow for transcription
may or may not require the presence of an inducer molecule to activate activator protein
MacConkey lactose plates
pH sensitive dyes that turn colour if sugar is metabolized
X-gal
colorless compound that turns an intense blue colour when cleaved by β-galactosidase (product of lacZ)
lac operon
required for lactose metabolism
lacZ (β-galactosidase)
cleaves allolactose into useable sugar monomers
lacY (lactose permease)
allows lactose to enter the cell
lacA (transacetylase)
transfers acetyl group to galactose to prevent buildup in cell
lacI (repressor)
repressor protein binds lacO operator
CAP binding site
location where cAMP-CAP activator proteins at low glucose levels
promoter
location of RNA polymerase binding
lacO (operator region)
repressor binding location
glucose
cell’s preferred carbon source
Trp operon
encodes enzymes for synthesizing L-tryptophan
regulation by attenuation
transcription starts normally, but is terminated prior to transcription of the first structural gene of the operon