L24: Cytoskeletal Motor Proteins
1/23
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
24 Terms
Myosin
Myosin = actin-based motor proteins
Couple energy from ATP hydrolysis
Mechanochemical enzymes = convert chemical energy → mechanical energy
(+) end directed motors = move toward (+) end of actin
EXCEPTION: Myosin VI (6) moves toward (-) end of actin
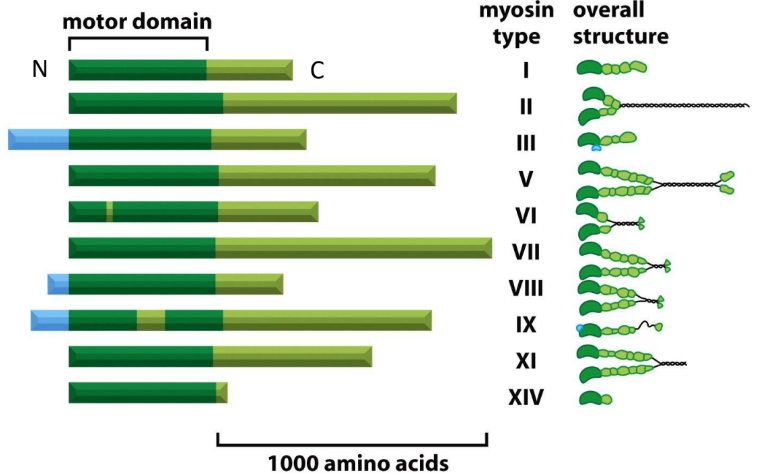
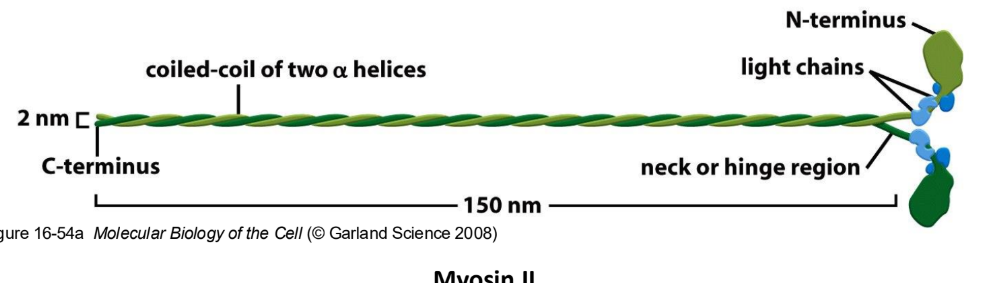
Explain the structure of Myosin complexes
Myosin = 2 heavy chains + 4 light chains
Myosin heavy chain = Head, neck, tail domain
Head domain = actin-binding + ATPase activities
Determines direction of movement
Neck domain = myosin light chain binding site
Structural + regulatory roles
Tail domain = different b/w different myosin
Determines specific properties
Myosin light chain = small polypeptide that binds to + stabilizes “neck” domain of myosin heavy chains
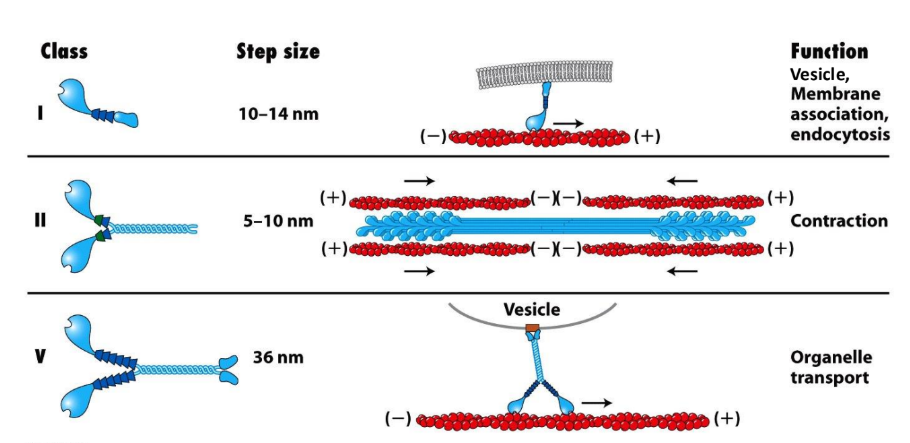
What do myosin tail domains define? Explain functions of Myosin I, Myosin II, and Myosin V
Specific properties of myosin determined by tail domains
Myosin I = bound to membrane ≠ dimerize, single-headed
Tails = different
Some have 2nd binding site → sliding
Some have membrane binding sites → bind to vesicles/organelles
Vesicle, membrane association, endocytosis
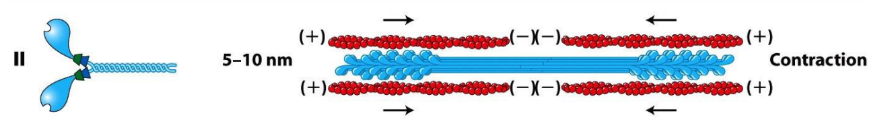
Myosin II = assembled into bipolar filaments = contraction (muscle contraction, contractile ring)
Myosin V = dimerized, organelle transport
Oddball = does processive movement (single motor molecule to take multiple steps along filament w/o detaching)
Which myosin have functions that are not as well understood?
Myosin III, IV, and VI-XV
Conserved head domains
Variable tail regions
Which myosin is the only known myosin to move towards the (-) end of F-actin?
Myosin VI (6)
Myosin II = conventional myosin
Myosin II = conventional myosin = first myosin discovered, most abundant type in muscle + non-muscle cells
2 heavy chains + 4 light chains = 6 proteins total
Tails = long α helices that mediate further polymerization of Myosin II → bipolar thick filaments
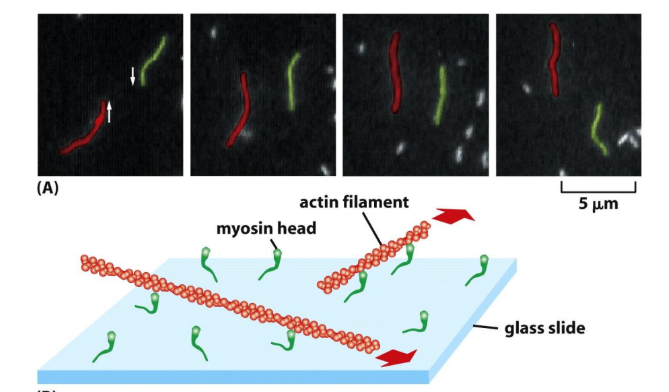
Explain how actin filament sliding assay works
How does ATP affect movement?
What 2 ideas does this sliding assay support?
Actin filament sliding assay = demonstrate motor function of Myosin
Myosin molecules stuck to glass (tails bound, heads free)
Fluorescent actin filaments added (labeled w/ phalloidin)
Filaments stick to Myosin on glass
ATP
No ATP → myosin binds tightly to actin
ATP → myosin moves actin
Myosin walk to (+) end & are bound to glass
(-) end is leading movement
MUST BE BOUND TO @ LEAST 2 MYOSIN (directionality)
ATP hydrolysis = coupled to Myosin motility
Myosin ATPase activity = actin activated
Actin absent = 4 ATP/hour
Actin present = 20 ATP/hour
Ensures myosin = active + using ATP w/ actin presence
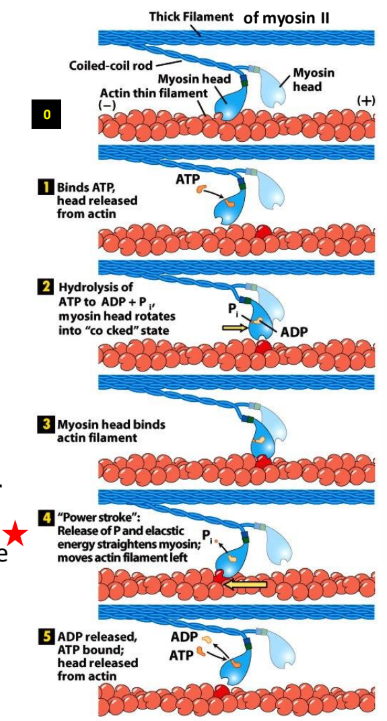
Explain the myosin-actin cross bridge cycle + 5 stages
Myosin-actin cross-bridge cycle = fundamental, repeating process of muscle contraction where myosin heads bind to actin filaments, undergo a “power stroke” to pull them inward + detach, driven by ATP
Only 1 head can be undergoing process @ one time
Rigor state
ATP binding site = empty
Myosin = tightly bound to actin
Rigor mortis of dead body = all muscles = tense (no ATP) → all myosin tightly bound to actin
ATP binding
ATP binding cleft = closes
Actin binding cleft = opens
Interaction w/ actin = weak → myosin lets go of actin
ATP hydrolysis
ATP hydrolysis: ATP → ADP
Δ Conformation → head moves forward, head binds to actin
Head moves to new position BEFORE rebinding to filament
Pi release
Power stroke = myosin head pulls actin thin filament towards center of sarcomere (powered by release of Pi)
ADP release
Myosin = rigor state
ATP exchange releases head from actin
Mechanism = still debated
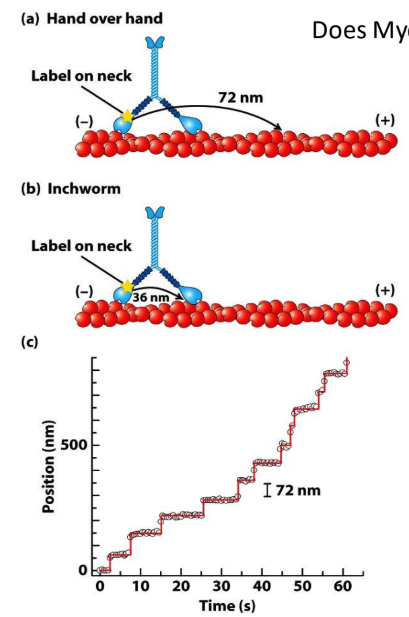
What is special about myosin V dimers?
Myosin V dimers = unconventional
Processive movement = 2 myosin V heads coordinated to interact w/ same actin filament
@ least 1 head always in contact w/ actin
Allows myosin V to carry cargo
Myosin II cannot move processively
2 heads act independently
What are the 2 models for Myosin V movement?
How can the step size of Myosin V be used to determine the correct model?
Processive movement = motor can walk on filament w/o detaching/falling off
Hand-over-hand = rear detaches → moves ahead of other head → alternate positions
Inchworm = one head consistently in front
Myosin V step size = 72 nm = 36 nm b/w 2 myosin heads x 2
Fluorescent tracking experiment = track position of1 myosin head over time
72 nm = Hand-over-hand model
What is the major difference b/w conventional and unconventional myosin?
Myosin II = conventional myosin = heads independently move
Myosin V = unconventional myosin = heads coordinate movement
What diseases can be caused by mutations in myosin genes?
MYH9 gene → Myosin IIA → bleeding problems, hearing loss, kidney disease, cataracts
Arg702 → decrease platelets, early-onset renal disease, hearing loss in infancy
Myosin VI ablation = deafness in mice
Myosin VII mutation → deafness + blindness in humans
What was used to isolate two classes of MT motor proteins?
In vitro motility assays
Kinesins = similar ATP binding/hydrolytic core domain to myosin
Dynein = unique catalytic core
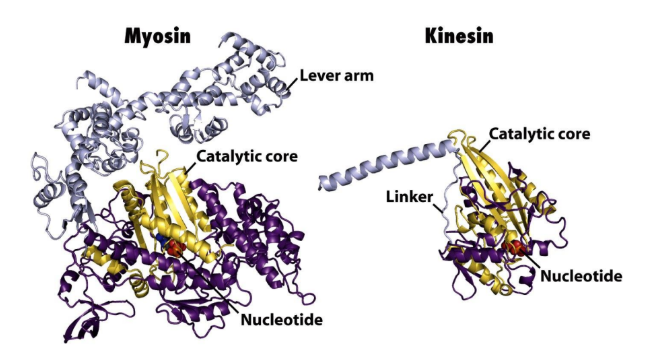
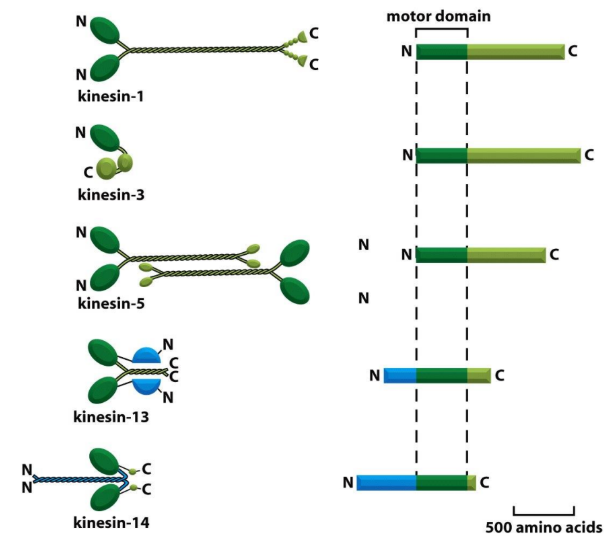
Kinesins definition + 3 classes
Kinesins = MT-activated mechanochemical ATPases
Kin-N Kinesin = motor domain @ N-terminus of protein
→ (+) ends
Conventional kinesins
Kin-C Kinesin = motor domain @ C-terminus of protein
→ (-) ends (rare)
Kin-I Kinesin = INTERNAL motor domain
Do not move along MTs
Bind MT ends → protofilament peeling
Kinesin-13
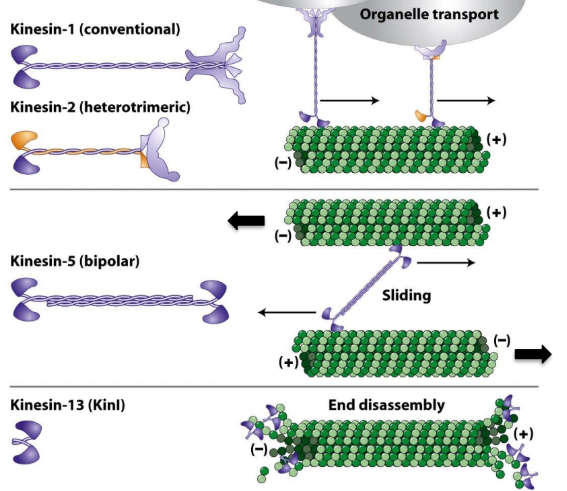
Describe structure of conventional kinesin
Conventional Kinesin = first Kinesin discovered + best characterized
2 heavy chains + 2 light chains
Heavy chains
Motor domain head @ N-terminus
Neck domain
Tail/stalk domain = α-helical coiled-coil
Light chains
@ C-terminus of tail region
Opposite to motor heads
Bind to cargo (organelles + vesicles)
Explain how the microtubule gliding assay works
MT Gliding Assay = demonstrate Kinesin motor function
Tails = attached to coverslips
Head = exposed
Fluorescent MTs moved around by head
Generally = Kin-N Kinesins (walk to (+) end)
(-) end MT = leading movement
Kin-C Kinesin (walk to (-) end)
(+) end MT = leading movement
Explain the kinesin cross-bridge cycle
ATP hydrolysis + Pi release → associated w/ Δ conformation of head/neck
ATP binds to forward head → Δ conformation in neck linker
Rear head swings forward past front head
New forward head release ADP
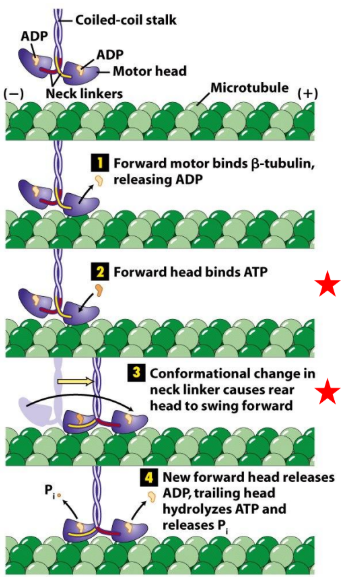
What kind of movement do kinesin motor proteins do?
Processive movement = movement over long distances w/o dissociating
Important for long-distance transport of vesicles + organelles
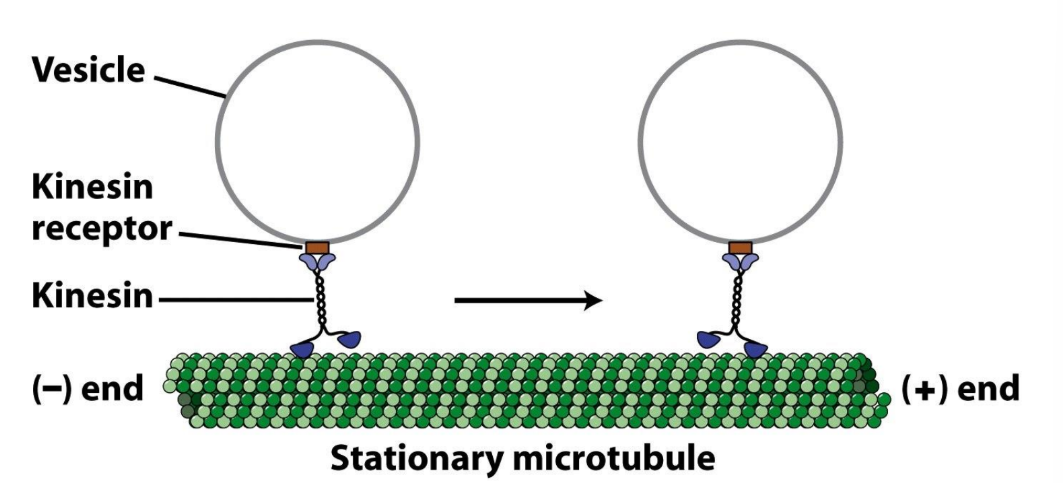
Define dyneins + describe its structure + 2 types
Dyneins = mechanochemical enzymes that use energy from ATP hydrolysis to (-) end directed movement along MTs
Structure = 2-3 heavy chains w/ unknown # of intermediate + light chains
Microtubule binding domain = 2 “feet” walking on MT
ATPase domain = bind + hydrolyze ATP → Δ conformation
Dynactin binding domain = bind to dynactin complex
Dynactin complex = intermediate b/w dynein + cargo
Cytoplasmic Dynein = most common form, many cellular functions
Ciliary/Flagellar Dynein = molecular motor → MT sliding → bending motions of cilia + flagella (microscopic hair-like appendages on eukaryotic cell surface → facilitate movement)
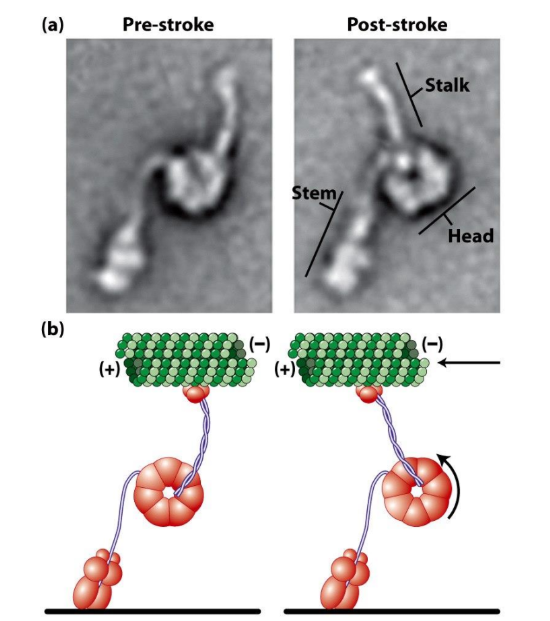
Describe power stroke of dynein + how it can be observed
Dynein power stroke = driven by ATP hydrolysis
Electron micrographs before + after power stroke
Δ angle b/w stem + stalk domains
Dynactin complex
Dynactin complex = multiunit protein complex = intermediate b/w Dynein and cargo
Dynein DOES NOT directly bind to cargo
Critical for cytoplasmic Dynein function
Mediate attachment of Dynein to cargo
~11 proteins
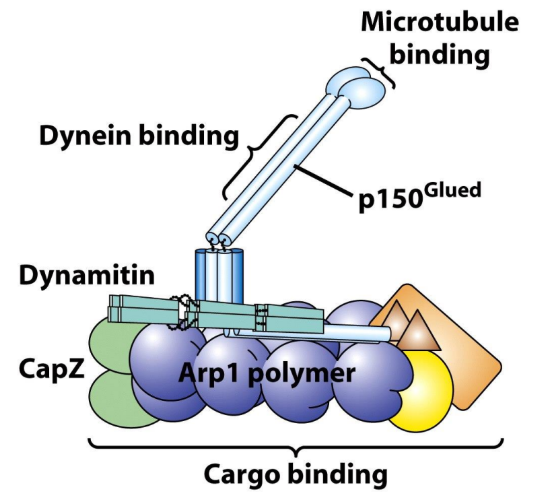
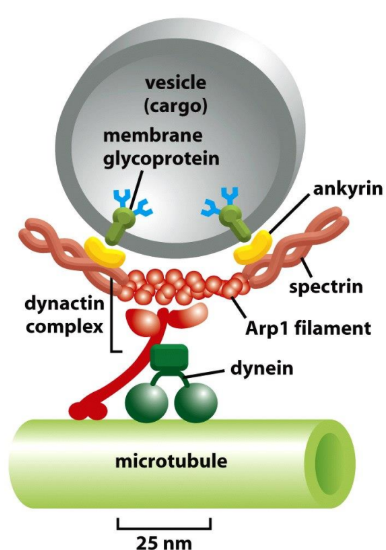
What part of dynactin complex is used to attach to surface of vesicle cargo?
Actin Related Protein 1 (Arp1) filament of dynactin complex → interact w/ Spectrin/Ankyrin membrane complexes of vesicle cargo membrane
Microtubule motors in disease
Kinesin deficiencies → Charcot-Marie-Tooth Disease, some kidney diseases
Dynein deficiencies → chronic respiratory tract infections (cilia ≠ function w/o dynein)
Dynactin Complex mutations (p150Glued subunit) → familial + sporadic ALS
Ciliary + Intraflagellar Transport defects → autosomal recessive polycystic kidney disease (ARPKD), retinal degeneration, other sensory disorders
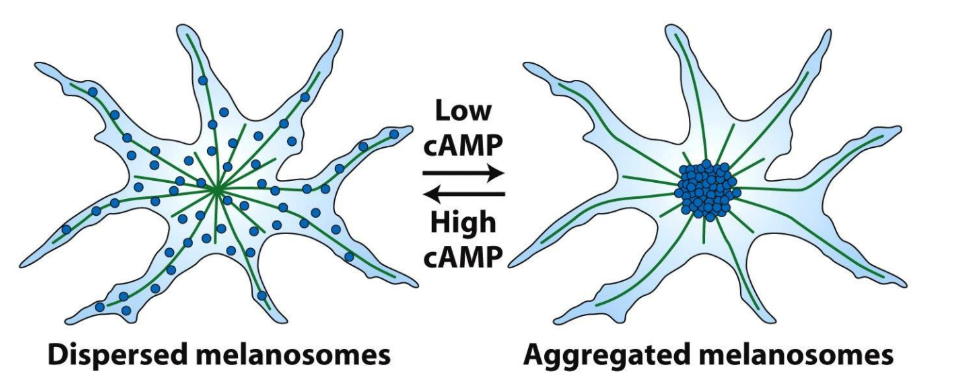
Explain melanosomes movement
A. Where are the microtubule (+) ends and (-) ends?
B. Are Kinesins or Dyneins important for melanosome localization in high cAMP?
C. Are Kinesins or Dyneins important for melanosome localization in low cAMP?
↑ cAMP → dispersed melanosomes
↓ cAMP → aggregated melanosomes
A. (-) ends = MTOC/centrosome, (+) ends cell periphery
B. ↑ cAMP → dispersed melanosomes → Kinesins
C. ↓ cAMP → aggregated melanosomes → Dyneins