Cytoskeleton
1/23
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
24 Terms
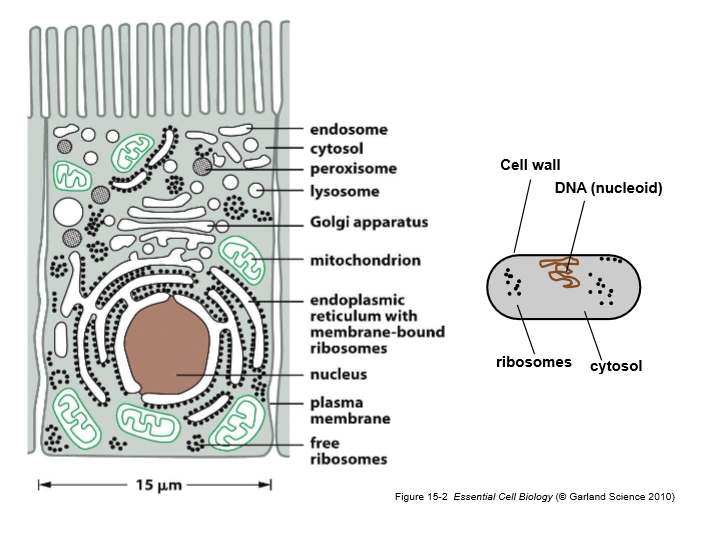
Label a prokaryotic and eukaryotic cell
How do cytoskeletal polymers assemble?
Non-covalent protein-protein interactions, driven by self-assembly and head-to-tail subunit addition
Assembly is regulated by nucleotide binding and hydrolysis
Intermediate filament assemble without nucleotides, relying on coiled-coil hydrophobic interactions and lateral contacts for strength.
What are the cytoskeletal polymers?
Actin filaments (microfilaments)
Microtubules
Intermediate filaments
Actin
Subunit: globular actin
Polymer structure: Head to tail → 2 stranded helix
Polarity: Yes +/- , growth occurs at + ends
Nucleotide: ATP
Microtubules
Subunit: αβ-tubulin heterodimer
Assembly: Head to tail protofilaments → 13-protofilament tube
Polarity: Yes +/-
Nucleotide: GTP (β-tubulin hydrolyses)
Intermediate filaments
Subunit: IF monomer (coiled coil)
Assembly: Dimer → Tetramer → unit-length filament → mature IF
Polarity: No
Nucleotide: None
Branched Actin network
ARP2/3 binds the side of an existing filament
Creates a 70° branch
Produces a dense meshwork at the leading edge
Plus ends face forward → protrusion
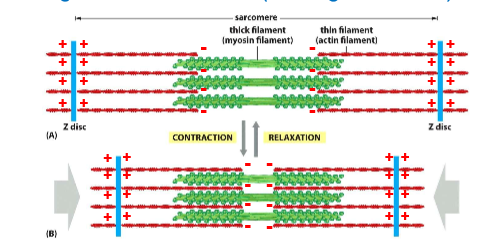
Sarcomere
Thin actin filaments on the outside, thich myosin filaments in the middle
Formin-nucleated linear filaments
Formins bind the plus end
Promote rapid monomer addition
Produce long, unbranched filaments
Bundled by fascin in filopodia
Contractile bundles
Actin filaments assemble spontaneously
Myosin II forms bipolar filaments
Crosslinkers (α‑actinin, etc.) organise anti‑parallel arrays
Myosin sliding → contraction
Muscle contraction
No nucleotide (rigor state): Myosin head tightly bound to actin.
ATP binding: Reduces affinity → myosin releases actin.
ATP hydrolysis: Myosin head “cocks” into a high‑energy conformation.
Pi release: Triggers the power stroke—myosin head pivots, pulling actin filament.
ADP release: Returns to rigor state, ready for another cycle.
Cell migration
Myosin I:
Anchored to the plasma membrane.
Walks along actin filaments using its motor domain.
Generates local tension and helps pull the membrane forward.
Myosin II:
Forms small bipolar “minifilaments”.
Uses its motor domains to contract actin meshwork at the rear of the cell.
Drives retraction of the trailing edge and forward movement of the cell body.
Microtubule nucleation
The centrosome is the primary site
It contains centrioles surrounded by pericentriolar material rish in γ-tubulin ring complexes (γ‑TuRC) which act as templates for MT assembly
γ‑TuRC binds αβ‑tubulin dimers.
The first ring of tubulin forms around the γ‑TuRC.
The minus end of the MT remains anchored at the centrosome.
The plus end extends outward into the cytoplasm.
Dynamic Instability
Each tubulin dimer binds GTP, but only β‑tubulin’s GTP is hydrolysed after incorporation.
Growing MTs have a GTP‑tubulin cap at the plus end.
When the cap is lost (due to hydrolysis outpacing addition), the lattice becomes unstable → catastrophe (rapid depolymerisation).
Regaining a GTP cap → rescue (growth resumes).
Dynamic instability allows MTs to “search and capture” cellular targets:
MTs grow and shrink randomly until they encounter specific structures (e.g., kinetochores, cell cortex).
Stabilisation occurs upon binding → defines cell polarity, spindle formation, or organelle positioning.
Microtubules in axons
relativley stable
long parallel bundles
Unifrom polarity allows directional transport: Kinesin (+ ended) → anterograde transport, Dynein (- ended) → retrograde transport
Microtubules in cilia/flagella
9+2 arrangment: Nine outer doublet MTs, Two central singlet MTs
All MTs have + ends at the top and - ends at basal body
Extremely stable
Dynein arms on A‑tubules “walk” along adjacent B‑tubules.
Nexin links convert sliding into bending → ciliary beating.
Microtubules on mitotic spindle
Kinetochore MTs
Attach to chromosomes
Plus ends at kinetochores
Minus ends at centrosomes
Interpolar MTs
Overlap in the spindle midzone
Stabilised by MAPs
Astral MTs
Radiate outward to the cortex
- ends at centrosomes, + ends extend outwards
Dynamic instability is essential
MT dynamics + motor proteins generate:
Chromosome alignment
Chromosome segregation
Spindle elongation