ap bio fuzzy areas
1/53
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
54 Terms
why does oxygen consumption show metabolism
Measure how much O₂ an organism uses per unit time
Higher O₂ use = higher cellular respiration = higher metabolic rate
👉 Why it works:
Aerobic respiration requires oxygen to make ATP
why does CO2 production show metabolism
Measure how much CO₂ is released per unit time
More CO₂ = more respiration happening
Why does measuring body heat output show metabolism
Metabolism = chemical reactions → releases heat
Why does heat increase metabolism
enzyme activity increases (to a point)
respiration increases
metabolic rate increases
Metabolism
all chemical reactions in the body
It includes:
building molecules (anabolism)
breaking molecules down (catabolism)
making ATP (cellular respiration is a big part of this)
DNA/protein synthesis
active transport, etc.
Metabolic rate
how fast those reactions are happening
So it’s:
the rate (speed) at which energy is being used / reactions are occurring
Think of ATP like electricity in metabolism:
More respiration = more electricity generated
Not every device runs faster
But more devices can run at once
What is surface area to volume ratio?
The amount of surface available for exchange compared to the amount of internal volume that needs resources.
What happens to SA:V ratio as organisms get larger?
It decreases because volume increases faster than surface area.
Why does a higher SA:V ratio increase exchange with the environment?
Because more surface is available per unit of volume, allowing more diffusion per unit time and shorter diffusion distances.
Why do small organisms have higher metabolic rates per unit mass?
a greater proportion of their body is exposed to the environment relative to their heat-storing volume.
small organisms must produce heat faster
which requires:
higher cellular respiration rate
higher metabolic rate per unit mass
cyanobacteria with thylakoids are now _____________ in eukaryotes, so there was compartments even in prokaryotes
chloroplasts
what is the endomembrane system made of
nuclear membrane, rough er, smooth er, vesicles, lysosomes, golgi complex
The endomembrane system is dynamic because its components are connected through continuous or vesicle-mediated membrane flow, allowing membranes and their associated proteins to be modified, transported, and recycled between the nuclear envelope, endoplasmic reticulum, Golgi apparatus, and plasma membrane.
transcription factors enter the nucleus through
nuclear pores
sites or rRNA from left to right
E, P, A
when do ribosomes go to rough er
when protein is destined for membrane, golgi, lysosome, other organelles, or export
chloroplast and mitochondria both have
own circular DNA (chromosome), bacterial ribosomes to make own proteins, double membrane (outside was the vesicle from the eukaryote’s plasma membrane)
smooth er has
enzymes
smooth er
detoxifying toxins into soluble form, lipid synthesis, carb breakdown and synthesis
golgi
receives vesicles from rough and smooth er and chemically modifies contents
packs into vesicles
lysosomes are only in
animal cells
cytoskeleton
dynamic network of protein fibers
enables cell to move membrane
enable cells to move materials and organelles
cellulose is a key component in
water conducting tubes in plant stems (xylem)
the phospholipid bilayer is stabilized by weak bonds between tails called
vander wells
what can go through membrane easily
oxygen, nitrogen, carbon dioxide
facilitated diffusion
facilitated down gradient, using kinetic energy from diffusing molecules
diffusion occurs because kinetic energy in higher concentration
random motion
statistical imbalance
net movement emerges automatically
does exocytosis need input of energy to occur
Yes, exocytosis requires energy because ATP is used to transport vesicles along the cytoskeleton and to facilitate membrane docking and fusion with the plasma membrane.
Neurotransmitter binds receptor
Ion channels open
Small voltage change (EPSP/IPSP)
These sum together
If threshold reached → voltage-gated channels open
Action potential starts
hypotonic
more water, less solute
hypertonic
less water, more solute
water diffuses from
hypotonic (more water, less solute) to hypertonic (less water, more solute)
osmoregulate
regulate osmotic balance
how does more solute in a solution effect water potential
decreases it
water moves from high water potential to low water potential. explain
molarity in solution decreases water potential. water moves from hypo to hyper, so it moves to more solute which makes water potential negative. pressure in a solution increases water potential. water moves to lower water potential, away from this pressure.
Describe what would happen if you placed a plant cell in a 0.0 M solution.
a) The cell would lyse
b) The cell would be turgid
c) The cell would plasmolyze
d) The cell would be flaccid
plasmolyze and pull away from cell wall
In liquid water, hydrogen bonds are constantly breaking and reforming
In ice, they become more fixed and organized
in evaporation, what happens to bonds
hydrogen bonds break and the water molecules have kinetic energy and separate far apart like gas
When KE increases (heating):
molecules move faster
collisions become stronger
eventually they break intermolecular hydrogen bonds
As kinetic energy increases, water molecules move faster and eventually overcome hydrogen bonds between molecules, allowing them to separate into the gas phase while remaining H₂O.
secondary structure of protein
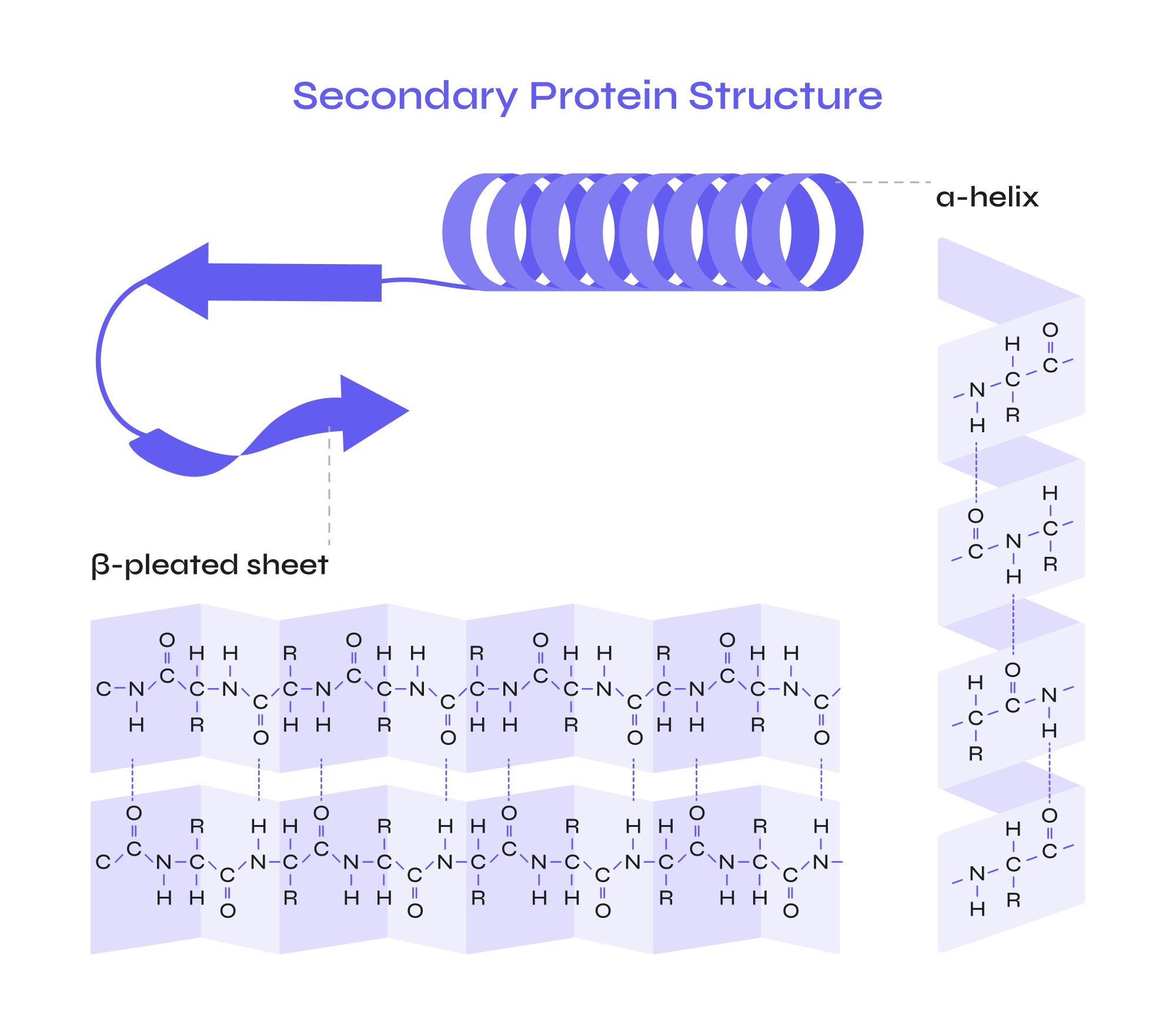
percent change is
( final - initial ) / initial x 100
Why do metabolic pathways have many steps?
Energy Efficiency: Releases energy in small, manageable amounts to capture it as ATP rather than losing it all as heat.
Regulation: Creates multiple "checkpoints" where enzymes can speed up or slow down the process based on the cell's needs.
Versatility: Produces intermediate molecules that can be siphoned off and used in other pathways (like building blocks for amino acids).
Compare intracellular vs membrane receptors (speed + regulation)
Intracellular: slower response; directly regulate gene expression
Membrane: faster response; indirectly regulate cell activity through signaling cascades (can eventually affect gene expression)
phosphotase
remove phosphate group
How is a protein kinase typically "turned on"?
Phosphorylation: An upstream enzyme (another kinase) attaches a phosphate group to a specific site on the kinase.
Once activated, what does a protein kinase do "over and over"?
The Relay: It grabs fresh ATP from the cell, snips off the phosphate, and glues it onto a different target protein (often another kinase).
Signal Amplification: A single active kinase can catalyze this reaction hundreds of times, activating a massive "army" of downstream proteins from just one initial signal.
What are the 3 primary ways a phosphatase is activated?
Negative Feedback: A downstream kinase phosphorylates the phosphatase to trigger a shutdown.
Ion Binding: Increased levels of Calcium (\(Ca^{2+}\)) bind to the enzyme to "toggle" it on.
Recruitment: The phosphatase is physically moved (localized) to the area where the active kinases are.
Intracellular signaling is slower because it involves changes in gene expression, requiring transcription and translation, whereas membrane signaling modifies existing proteins and produces a faster response.
meiosis 1 nondisjunction result
50% of the gametes are normal, 25% have an extra chromosome, and 25% are missing one.
meiosis 2 nondisjunction result
Sister Chromatids | 2 normal gametes, 2 abnormal gametes. |
why do a testcross
determining if a parent is homozygous recessive or heterozygous. if one of the kids is recessive, then they are hetero, if all of the kids are dominant, they are homozygous dominant.
where is the tata box
In eukaryotes, many promoters contain a famous sequence called the TATA box (literally a string of Ts and As), which is easy for proteins to recognize and pull apart because A-T bonds are weaker than G-C bonds.
describe the steps of rna transcription
Recognition: Transcription factors (helper proteins) bind to the promoter on the DNA.
Binding: RNA Polymerase attaches to the promoter/transcription factor complex.
Initiation: The DNA double helix is unzipped at the promoter site.
Elongation: RNA Polymerase moves away from the promoter and down the gene to start building the RNA strand.
what happens in rna processing in eukaryotes
introns removed (splicing)
exons kept
5’ cap + poly-A tail