Chapter 15 - Regulation
1/85
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
86 Terms
What do the mechanisms of gene regulation, or genetic control, determine?
where, when, and how much a gene is expressed
every gene is a ________ (transcribed region) of DNA
RNA-coding region
What are transcription factors?
DNA-binding proteins that recognize specific sequences within the regulatory regions near the gene to either activate or repress transcription
operon
a cluster of structural genes with related functions under the control of a common regulatory system that can respond to changes in environmental conditions
common in prokaryotes but rare in eukaryotes
The products of the structural genes of the lac operon in E. coli are involved in what?
utilization of lactose for energy
lac operon: these genes will only be expressed if _____ is available; but also only if the cell needs to use lactose for energy
lactose
When are the genes of lac operon expressed?
only if glucose is absent and lactose is available
lacZ gene product
B-galactosidase
What is B-galactosidase?
an enzyme that breaks down lactose (a disaccharide) into galactose and glucose
can also isomerize lactose into allolactase
lacY gene product
permease
What is permease?
a membrane transporter for lactose and facilitates the entry of lactose into the bacterial cell
lacA gene product
transacetylase
What is transacetylase?
not involved in lactose metabolism and involved in the removal of by-products of lactose digestion from the cell
structural genes under operon control
lacZ, lacY, and lacA
control regions
lacP (promoter or Plac)
lacO (lac operator)
lacP
binding site for RNA polymerase
lacO
binding site for the regulator protein; it overlaps lacP
regulator locus
lacI
Pi
lac(I)
the gene that codes for the regulator protein
Pi
the promoter for the lacI gene
What happens in the absence of lactose?
the regulator protein (repressor) binds to the operator, blocking RNA polymerase from binding to the promoter
repression of the structural genes, but this system is leaky
What is the operon inducer?
allolactose which binds to the repressor protein and inactivates it
this activates the expression of the structural genes
if glucose is available, the lac structural genes are ___, even if lactose is present
off
When does the binding of RNA polymerase to the promoter occur?
only if glucose is absent
adenylate cyclase
enzyme that catalyzes the hydrolysis of ATP into cAMP+PP is induced when glucose is absent
cytoplasmic levels of cAMP (catabolite) increase
What does cAMP activate?
catabolite activated protein (CAP)
the activated CAP binds to a site next to the _____ and facilitates the binding of RNA polymerase to it
lac promoter
What is cAMP?
catabolite that binds to the catabolite activated protein (CAP) to activate it
What is the catabolite activated protein (CAP) necessary for?
stable binding of RNA polymerase to the lac promoter
only active when glucose is absent
What happens in the presence of glucose?
cAMP levels decrease and CAP remains inactive
RNA polymerase does not bind the promoter efficiently
the operon is OFF (no expression of the structural genes)
What are the products of the five structural genes of trp operon?
enzymes involved in the biosynthesis of the amino acid tryptophan (Trp)
if Trp is absent
structural genes are expressed
if Trp is present
structural genes are turned off by Trp itself
promoter (trpP)
binding site for RNA polymerase
operator (trpO)
binding site for the repressor protein
repressor gene (trpR)
codes for the repressor protein (repressor gene promoter PR)
Trp absent
the repressor protein remains inactive
RNA polymerase binds to the promoter and initiates the transcription of the structural genes
Trp present
Trp binds to the repressor protein and activates it. The binding of the Trp-activated repressor to the operator prevents the binding of RNA polymerase to the promoter.
EMSA (electrophoretic mobility shift assay)
a method used for detecting specific DNA-protein interactions, such as the specific binding of transcription factors and other regulatory proteins to regulatory regions of the chromosome
example of EMSA
establishing the specific binding of the trip repressor protein to the trp operator (trpO) in the presence of tryptophan
mix in a test tube:
a fragment of DNA containing trpO
E.coli cell extract (contains all cellular proteins)
tryptophan (Trp)
specific antibodies against the trp repressor and other cellular proteins, such as the lac repressor
five experiments
DNA alone
DNA + cell extract (no Trp)
DNA + cell extract + Trp
DNA + cell extract + Trp + anti-lac repressor antibody
DNA + cell extract + Trp + anti-trp repressor antibody
in the five experiments, what to do?
slow gel electrophoresis of samples from the contents of each tube
electrophoretic mobility shifts occur due to the different sizes of the complexes that form
RNA polymerase I
the larger rRNA genes (5.8s, 18s, 28s)
What transcribes the eukaryotic genes?
RNA polymerases
RNA polymerase II
all the protein-coding genes
RNA polymerase III
the small rRNA (5s) gene and all the tRNA genes
What are some small nuclear RNAS (snRNAS) and small cytoplasmic RNAs (scRNAs) transcribed by?
RNA polymerase II and others by RNA polymerase III
heterochromatin
transcriptionally inert due to the tight association of DNA with histones, which prevents RNA polymerase from binding to gene promoters
Why is the ability of the cell to alter the association of DNA with histones essential?
to allow gene regulatory proteins and RNA polymerases to bind to DNA
What does histone acetylation weaken?
the interaction between basic histones and the acidic DNA molecule, causing chromatin decondensation
What does not bind to DNA but regulate histone acetylation?
HATs and HDACs
What doe positively charged tails of nucleosomal histone proteins interact with?
negatively charged phosphate groups of DNA
Acetylation of the tails weakens their interaction with what?
DNA and may permit some transcription factors to bind to DNA
What do transcription activators recruit?
histone acetyl transferases (HATs), causing chromatin decondensation
act as coactivators but do not bind to DNA
What do transcription repressors recruit?
histone deacetylases (HDACs) causing chromatin condensation
act as corepressors but do not bind to DNA
What does the flowering locus C (FLC) code for?
a transcription factor that represses flowering but is only expressed if the histones on the locus are acetylated
What is the gene product of flowering locus D (FLD)?
a histone deacetylase that specifically inactivates the FLC to allow flowering
Arabidopsis thaliana (Thale cress)
Most widely studied plant in biology. It has contributed our understanding of the genetic, molecular, cellular, and develop-mental biology of flowering plants. It is as important to botany as Drosophila is to animal genetics
What do acetyl groups on histone proteins destabilize?
chromatin structure
What does the term transcription factor refer to?
any transcription regulator protein that binds to a specific DNA sequence
What are cis-acting elements?
DNA sequences that are necessary for the control of transcription (the regulatory regions of chromosome): promoters, enhancers, and silencers
What are trans-acting factors?
proteins that are necessary for the control of transcription: the transcription factors that bind to the cis-acting elements
promoters
the binding sites for the RNA polymerase II transcription initiation complex
a key component of many eukaryotic promoters is a TATAAA sequence within them known as the TATA box
enhancers
binding sites for transcriptional activator proteins
silencers
binding sites for transcriptional repressor proteins
What are enhancers required for?
stimulated transcription
DNA sequences for enhancers vary widely and are recognized by what?
a large variety of transcription activators
What will the activation of a gene in any particular cell depend on?
whether the cell has the right activators to bind to the gene enhancers
What happens by facilitating up-regulation?
enhancers determine where, when, and how much transcription occurs
promoter alone
only the basal transcription factors may bind to DNA, and only the basal transcription apparatus may form
only very low or undetectable transcription can occur
With the right enchancers
transcriptional activators bind to enhancers
stimulated transcription (biologically significant)
What positions can enhancers be?
distant upstream, downstream, or inverted positioin
still be effective in stimulating transcription (CANNOT MOVE PROMOTER)
Insulators (boundary elements)
DNA sequences that block the effect of enhancers in a position-dependent manner when insulator-binding proteins are bound to them
This prevents enhancers from stimulating the transcription of the wrong genes on the same chromosome
What is the purpose of the RNA polymerase II transcription initiation complex?
to direct RNA polymerase II to the correct place on the promoter behind the gene transcription initiation site (+1), because the polymerase itself does not recognize any specific DNA sequences
TFIIs
transcription factors of RNA polymerase II
TFIID
transcription factor D of RNA polymerase II
this recognizes the TATA box and binds to it
What do transcriptional activators bind to?
upstream and in some cases downstream sequence elements (enhancers) which are necessary for stimulated, biologically significant, levels of transcription
RNA polymerase II transcription initiation complex (basal transcription apparatus)
contains TFIID, other TFIIs, and RNA polymerase II. DNA loops to allow transcriptional activators, bound to enhancers, to interact with the complex
Why does DNA loops?
to allow transcriptional activators, bound to enhancers, to interact with the complex
What are the products of the GAL structural genes in yeast?
proteins that metabolize the sugar galactose
What does the transcription of GAL structural genes requires?
enhancer UASG (upstream activating sequence of GAL genes) which is permanently occupied by the transcription activator GAL4
What happens in the absence of galactose?
the GAL80 protein binds to GAL4 and prevents it from activating the transcription of the GAL structural genes
What happens when galactose is available?
it activates the GAL3 protein, which interacts with GAL80 to displace it and expose the GAL4 trans-activating domain
What is required that interacts with GAL4 and the transcription initiation complex?
mediator protein
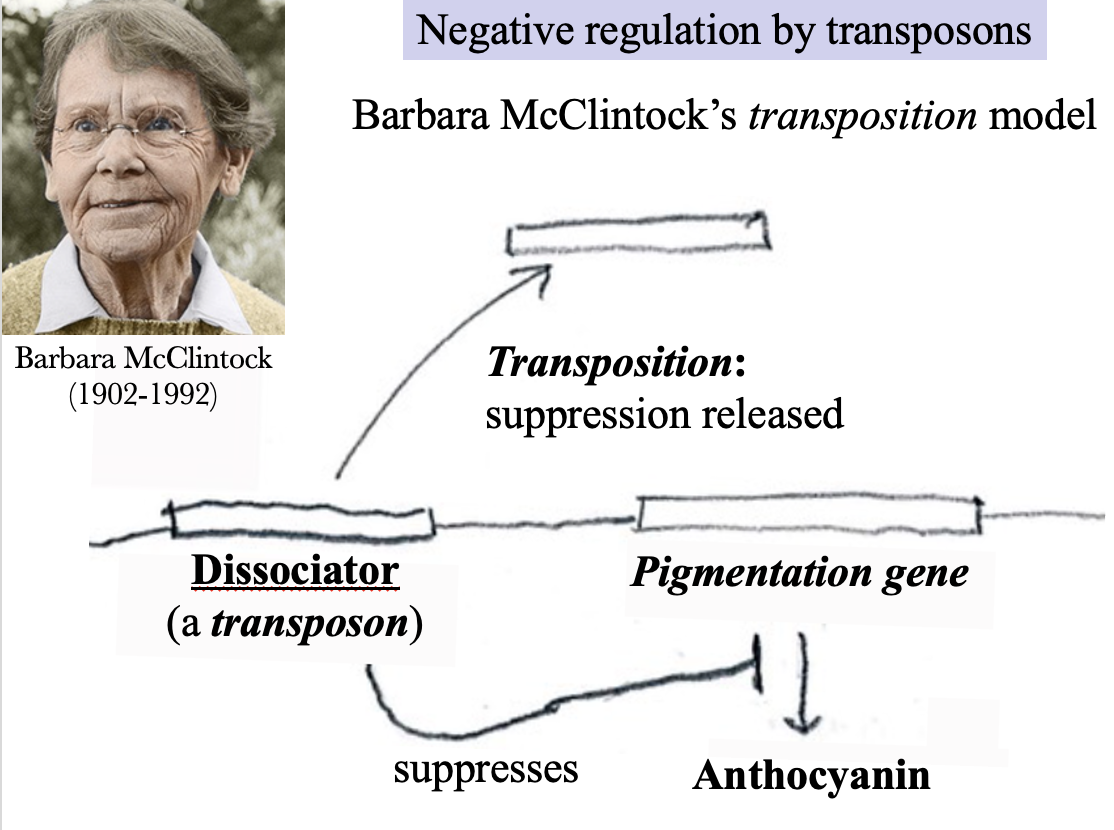
Barbara McClintock’s transposition model (1950)
The expression of a gene that is necessary for the synthesis of the pigment anthocyanin can be suppressed by a silencer that she named Dissociator. It is a transposon: a segment of DNA that can move from one location on a chromosome to another. If Dissociator is transpositioned, the suppression of the pigmentation gene is released, and anthocyanin can be made.
How long the Dissociator stays next to the pigmentation gene before “jumping” affects the degree of pigmentation and mottling in a maize kernel. The reddish-purple patterns caused by the transposon may be dots, blotches, or streaks. If the transposon stays next to the pigmentation gene long enough, the grain will be completely unpigmented.