Week #8 - Membrane Structure and Transportation
1/57
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
58 Terms
What is the plasma (cell) membrane?
The plasma membrane is a semi-permeable boundary that separates the cell from its environment.
It is primarily composed of a phospholipid bilayer where the molecules are held together by weak Van der Waals forces and hydrophobic interactions—not covalent bonds—allowing for flexibility and movement.
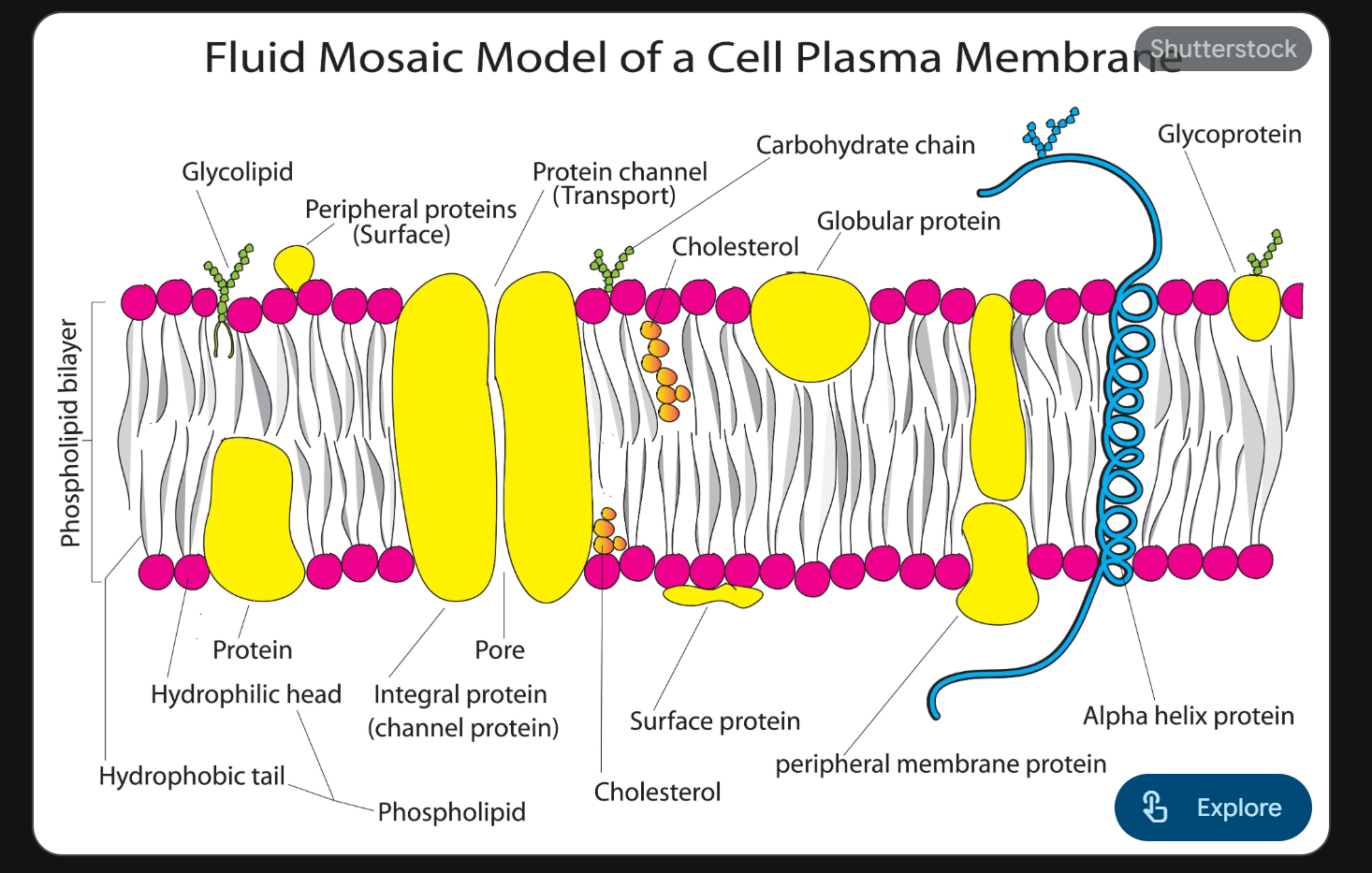
According to the Fluid Mosaic Model, the membrane is a functional "mosaic" of:
Lipids: Phospholipids (amphipathic base) and Cholesterol (fluidity buffer).
Proteins: Integral and peripheral proteins that act as transporters, receptors, and enzymes.
Carbohydrates: Surface molecules used for cell-to-cell recognition.
What are the six primary functions of biological membranes?
Define and enclose specific compartments within the cell (cell boundary)
Control the movement of materials into and out of the cell
Allow the cell to respond to external stimuli
Enable interactions and communication between cells
Provide a scaffold for biochemical activities
What is the most studied cell membrane?
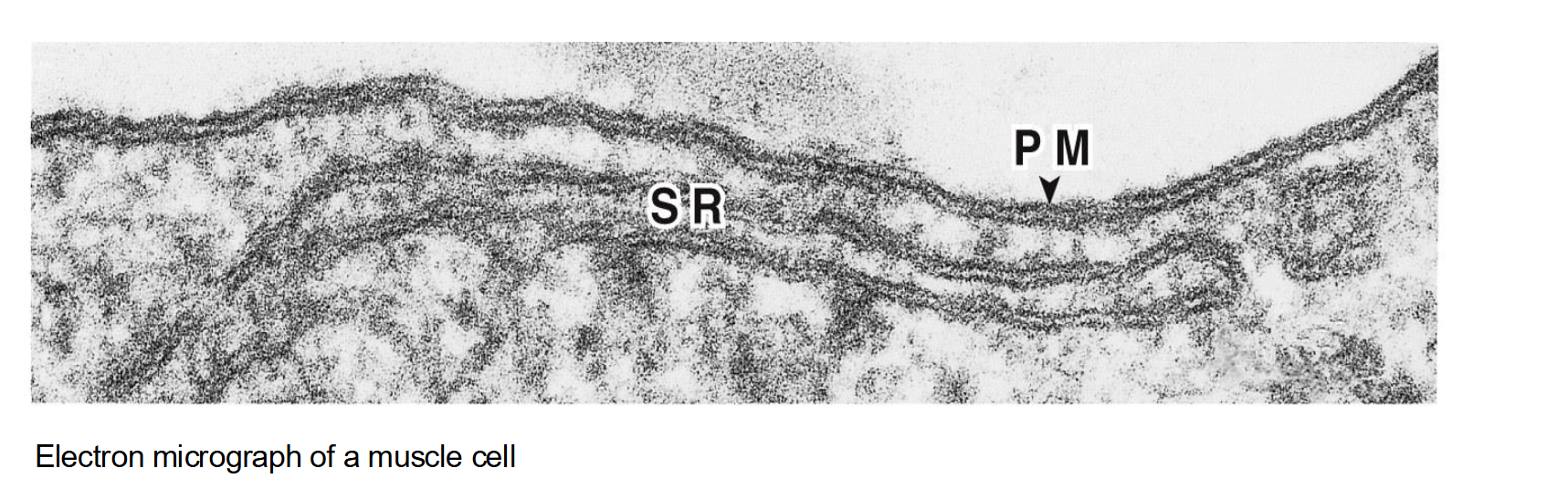
The PM is the most studied cell membrane
But there is another membrane…the sarcoplasmic reticulum (SR)!
The sarcoplasmic reticulum (SR) is a specialized type of smooth endoplasmic reticulum found in muscle cells.
Its main function is to store and regulate calcium ions (Ca²⁺), which are essential for muscle contraction
Why are red blood cells (RBCs) used as a model for studying membrane structure?
RBCs are ideal because they do not contain nuclei or internal membranes (organelles), allowing researchers to study the plasma membrane in isolation
How many layers does the plasma membrane have?
Three Layers!
Trilaminar structure made of a phospholipid bilayer
White Middle → Hydrophobic!
“Brown” Ends → Hydrophilic!
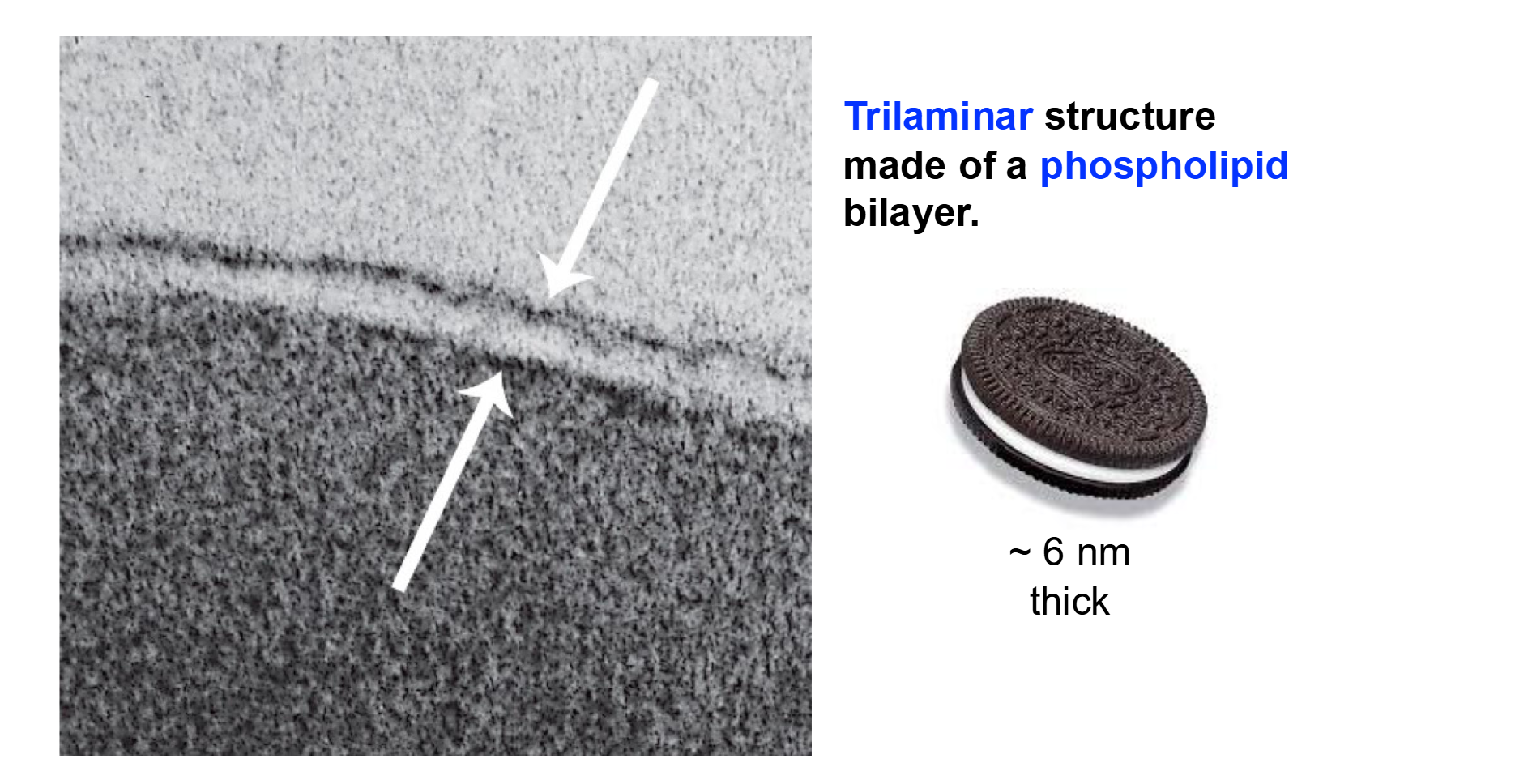

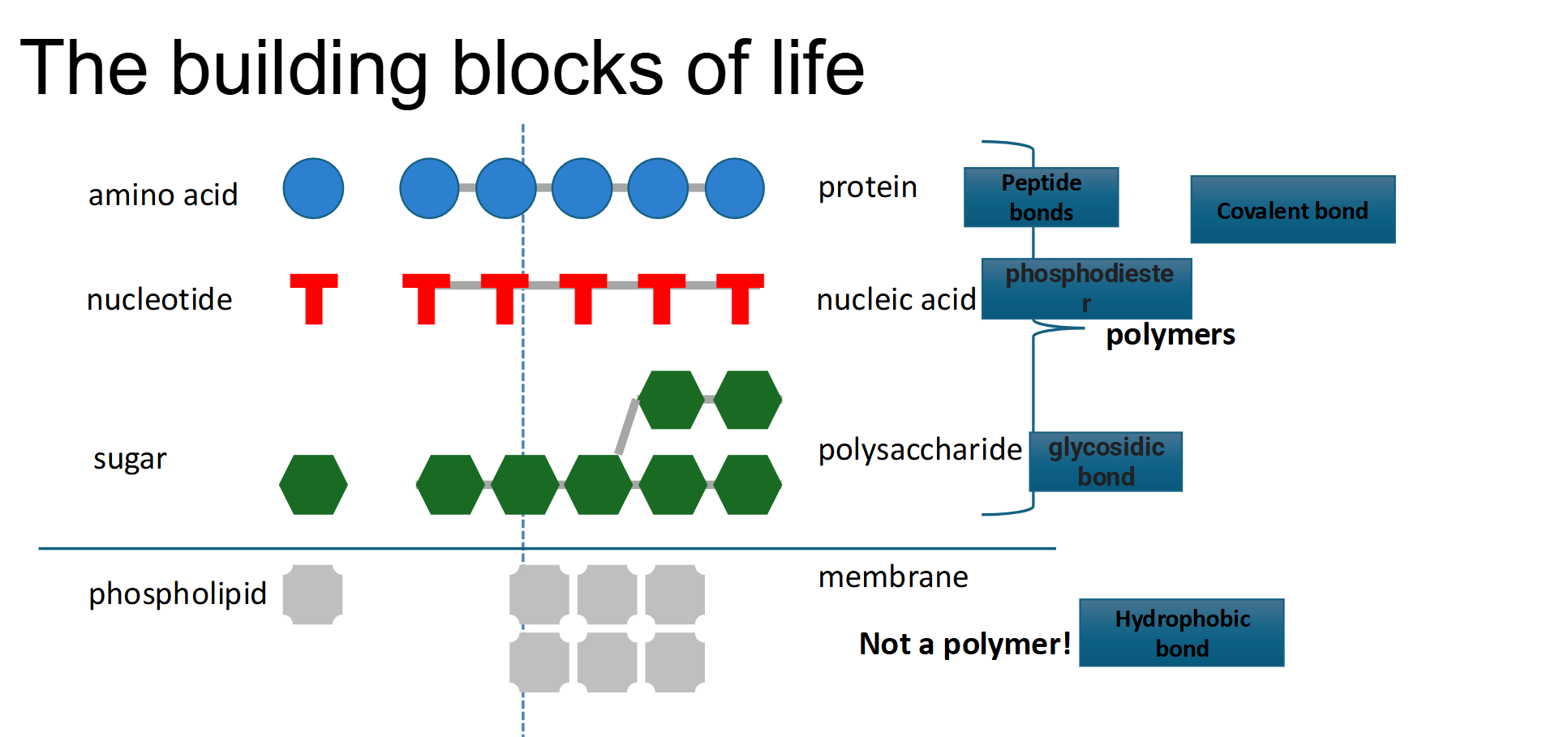
Important Distinction with Phospholipids
Unlike the other building blocks of life, they are not a polymer (Do not form any covalent bonds!)
Instead they connect by interactions → Hydrophobic interactions
What does it mean for a phospholipid to be "amphipathic"?
It means the molecule has both opposing chemical properties: a hydrophilic (water-loving) head and a hydrophobic (water-fearing) tail .
Head: Consists of a charged phosphate-containing group and glycerol.
Tail: Consists of two nonpolar hydrocarbon fatty acid chains.
Carbons of tail can be joined by single or double bonds
Saturated (no double bonds) versus unsaturated
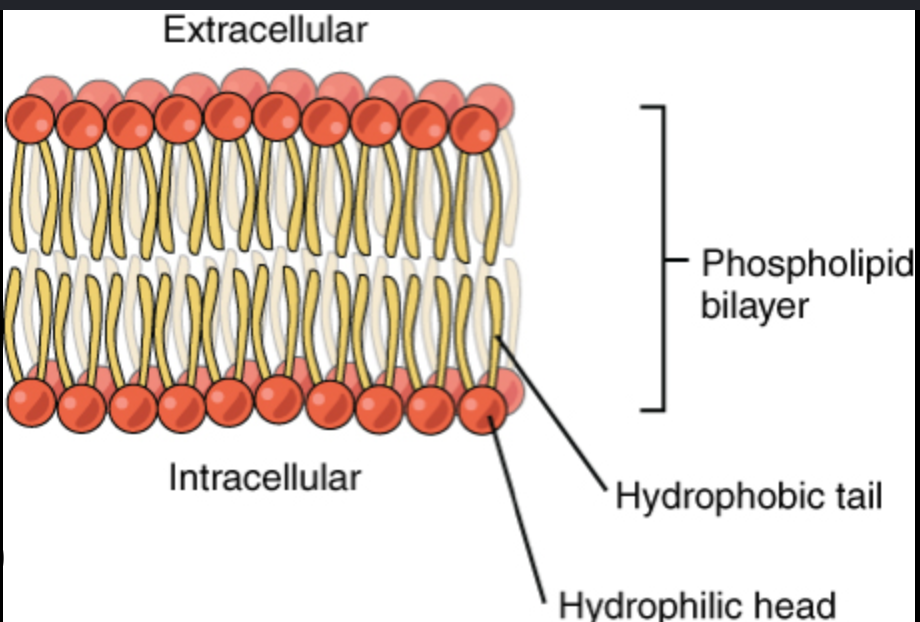
Contrast Saturated vs. Unsaturated Fatty Acids in the membrane.
Saturated: No double bonds between carbons; the chains are straight, allowing them to pack tightly together
Unsaturated: Contain one or more double bonds that create "kinks" in the chain. These kinks prevent close packing, increasing membrane fluidity
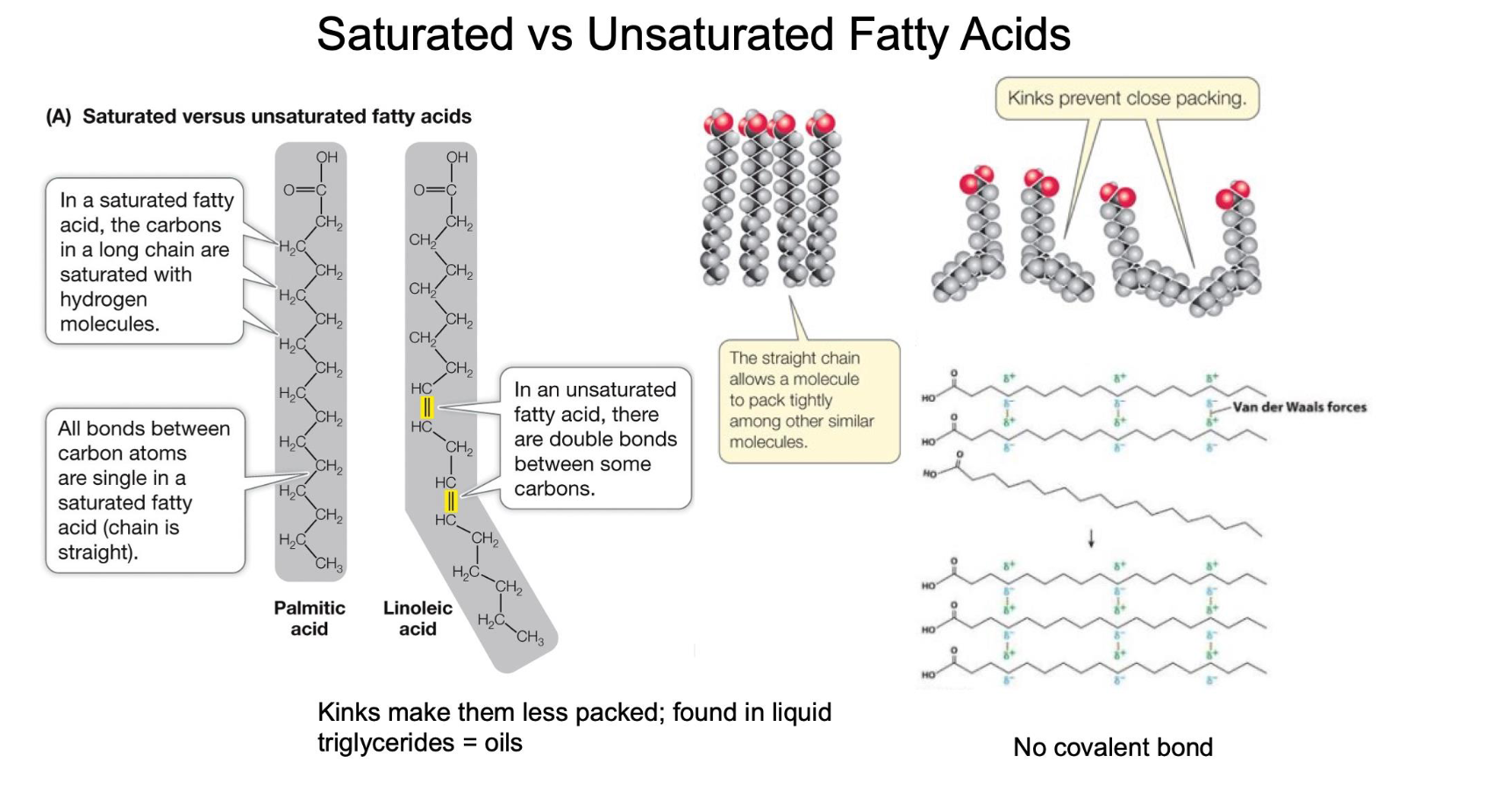
How do double bonds in fatty acid tails affect the structure and melting point of a triglyceride?
Saturated (Single Bonds Only): The hydrocarbon chains are straight, allowing them to pack tightly together. This results in more Van der Waals interactions and a higher melting point (typically solid at room temperature, like butter).
Unsaturated (One or More Double Bonds): Double bonds create "kinks" in the chain. These kinks prevent tight packing, resulting in fewer Van der Waals interactions and a lower melting point (typically liquid at room temperature, like vegetable oil).
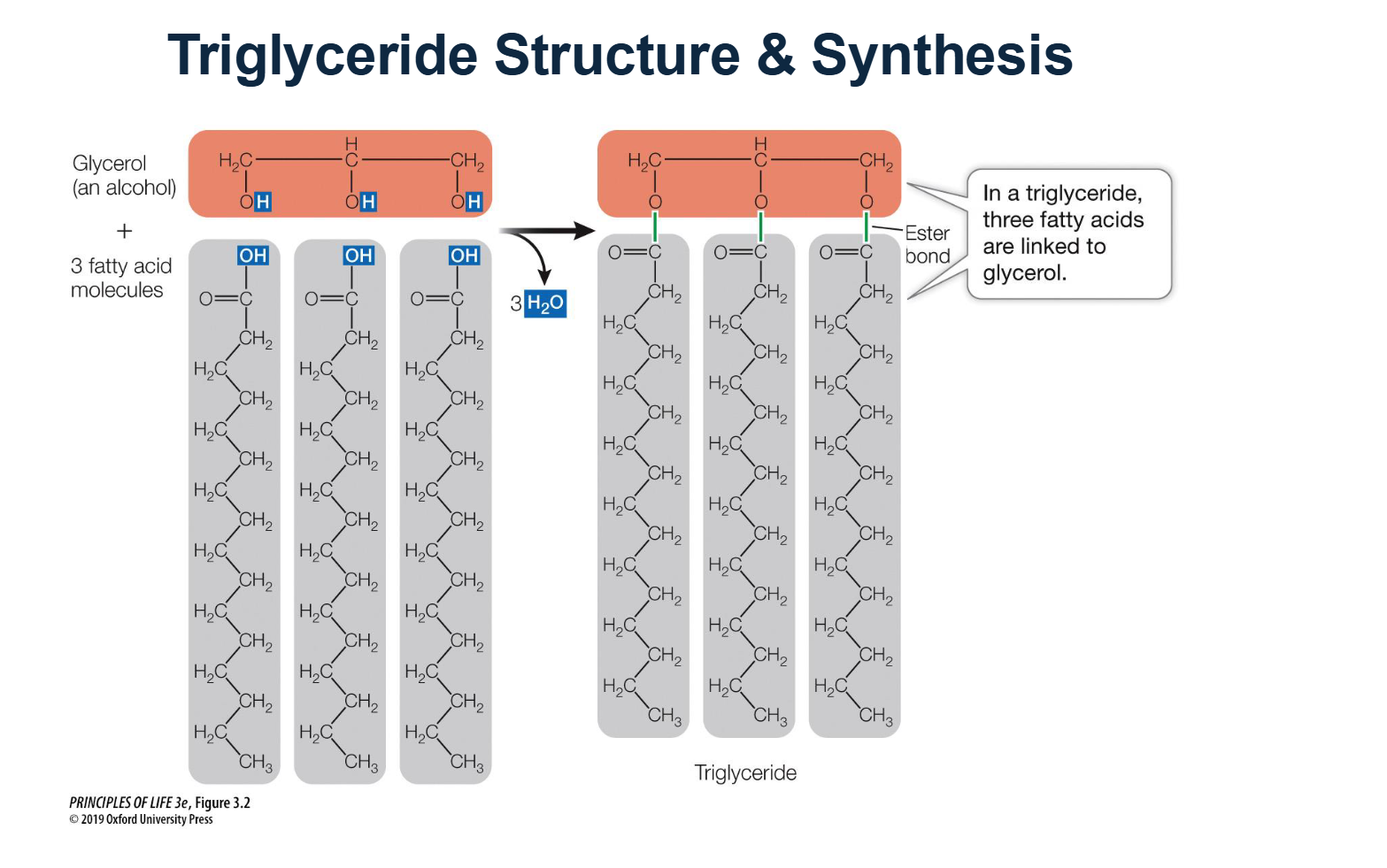
How are triglycerides synthesized, and what chemical bond is formed?
Triglycerides are synthesized through a reaction between one glycerol molecule (an alcohol) and three fatty acid molecules.
The Process: This is a dehydration synthesis reaction that results in the release of three water (H2O) molecules.
The Bond: The specific chemical bond formed between the glycerol and each fatty acid is called an ester bond.
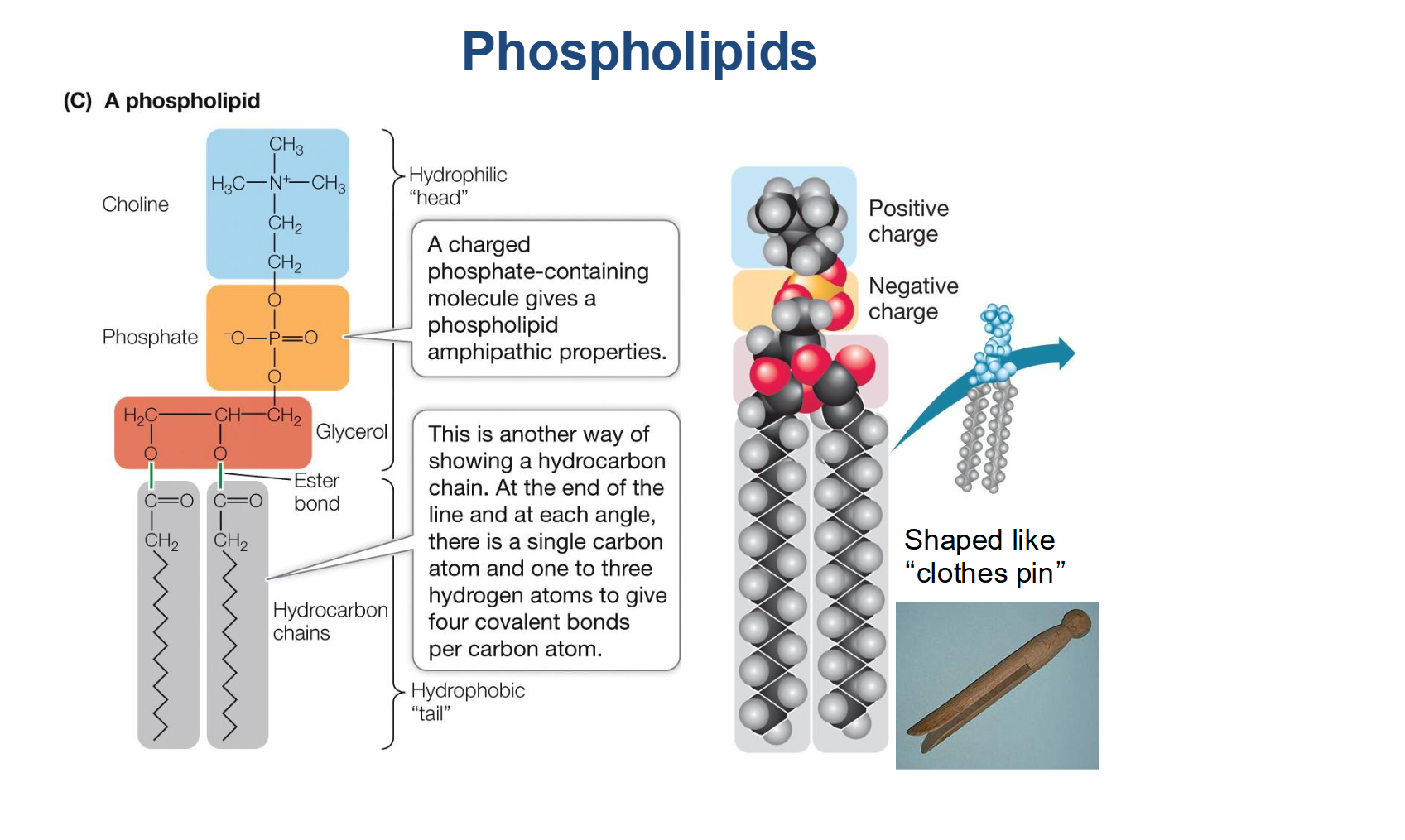
How are phospholipids held together in the bilayer?
They are not connected by covalent bonds. Instead, they are held together by weak Van der Waals forces and hydrophobic interactions.
This lack of strong bonding is critical because it allows the membrane to be flexible and for molecules to move
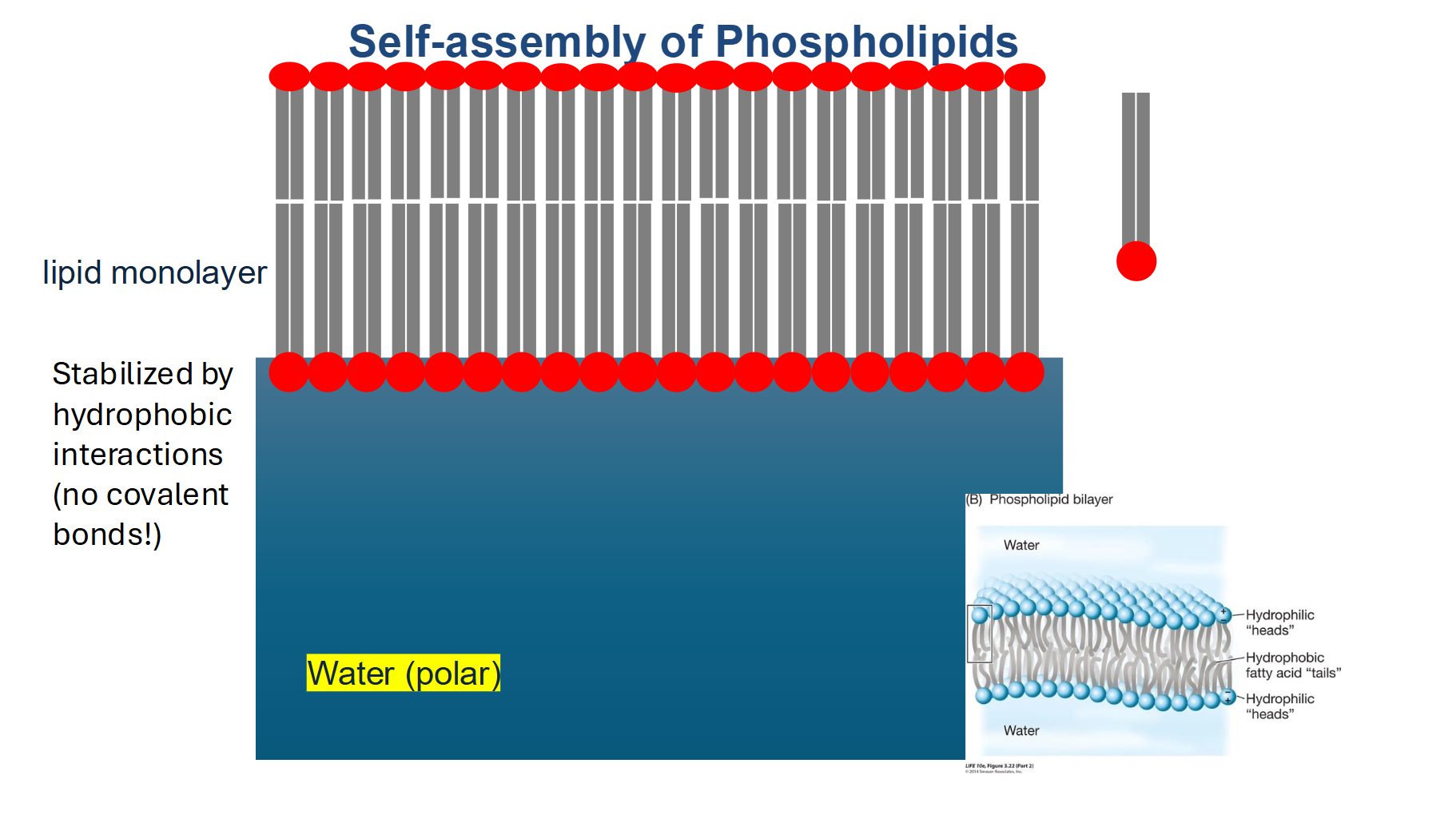
Plasma Membrane Summary
The trilaminar structure is a lipid bilayer.
The plasma membrane is about 6 nm thick—very consistent between cell types.
The lipid bilayer is made up of phospholipids
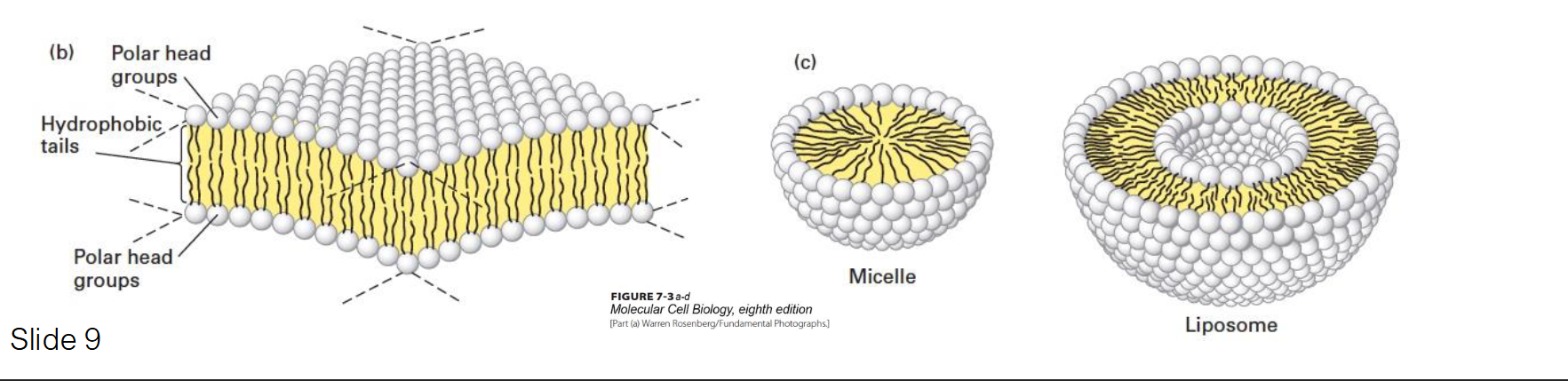
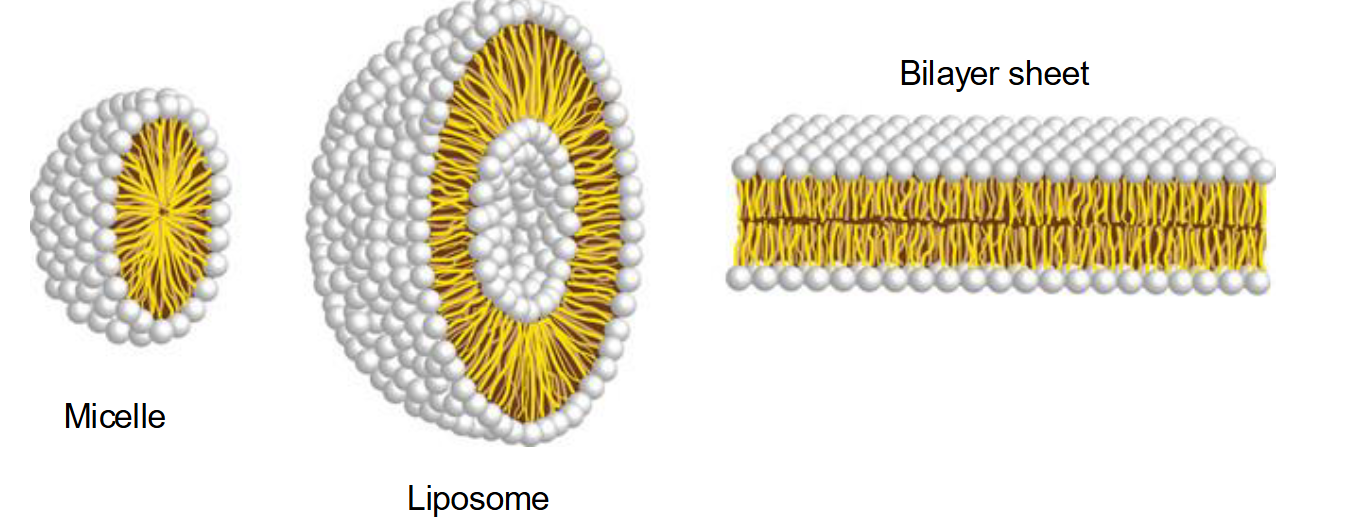
Why do phospholipids spontaneously aggregate in water, and what structures can they form?
They aggregate to bury their hydrophobic tails away from water while exposing their hydrophilic heads.
Micelles: Usually formed by fatty acids with a single hydrophobic chain.
Liposomes: Hollow spheres formed by a bilayer.
Bilayer Sheets: Large, stable sheets that form the basis of cell membranes.
Define "Selective Permeability" and how it applies to different molecules.
The membrane regulates what enters and exits based on size and charge
Easy Passage: Small, non-polar molecules like O2 and CO2
Difficult Passage: Small polar molecules (like water) have a harder time passing through the hydrophobic core.
Requires Channels: Large or charged molecules (ions) cannot pass through the lipids and must use specific protein channels.
What are the primary building blocks used to synthesize phosphatidic acid (simplest phospholipid)?
The Components: Two fatty acids are linked to a glycerol-3-phosphate molecule.
The Result: This creates Phosphatidic Acid, which is the simplest phospholipid and serves as the precursor for more complex ones.
Key Detail: Note that at this stage, the molecule already has its two hydrophobic tails and a phosphate-based "head" area, but it lacks a specific polar group (like choline).
Where specifically in the cell are phospholipids synthesized?
Synthesis occurs on the cytosolic face of the Smooth Endoplasmic Reticulum (ER) membrane, where the necessary enzymes are located.
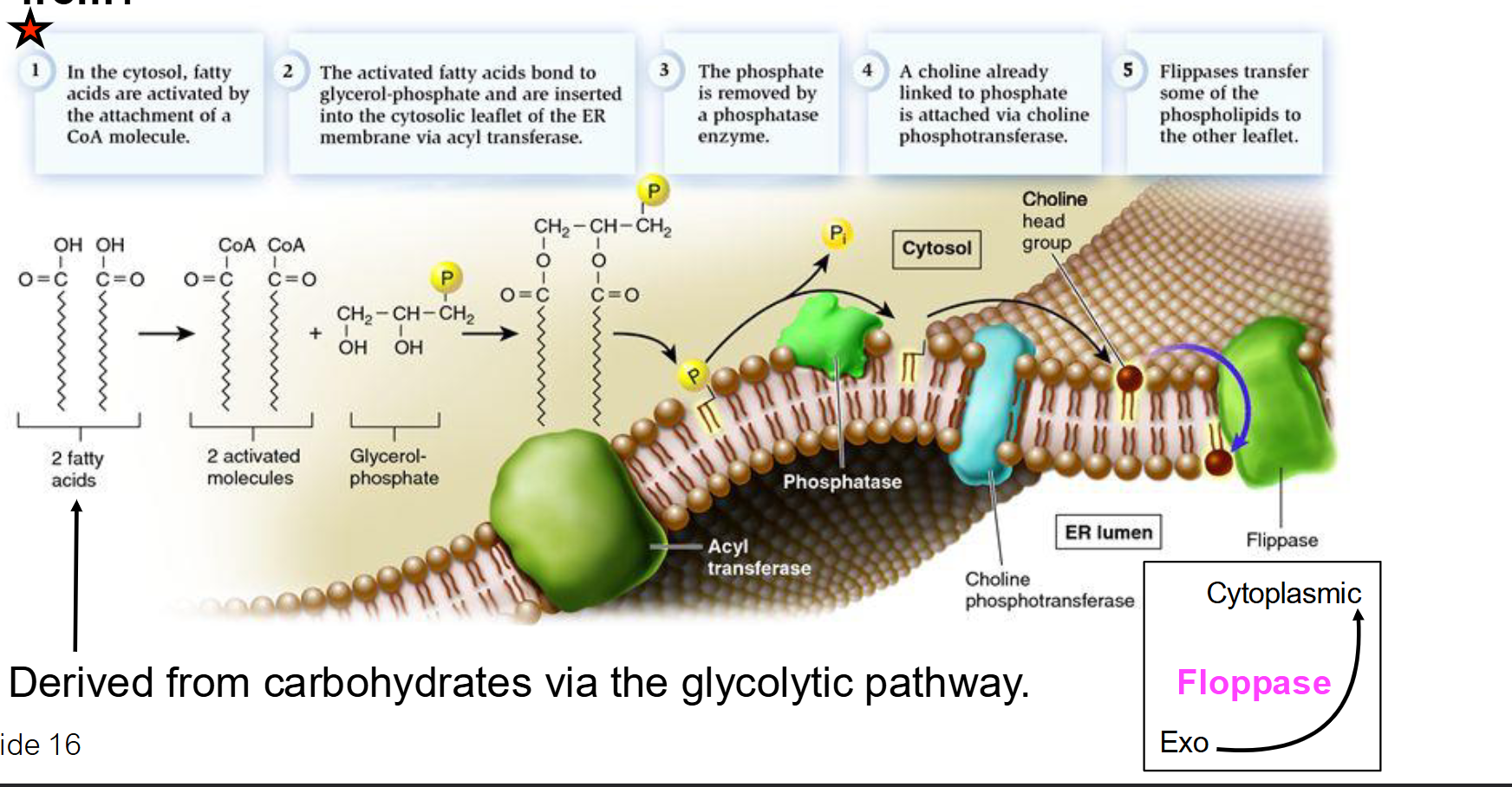
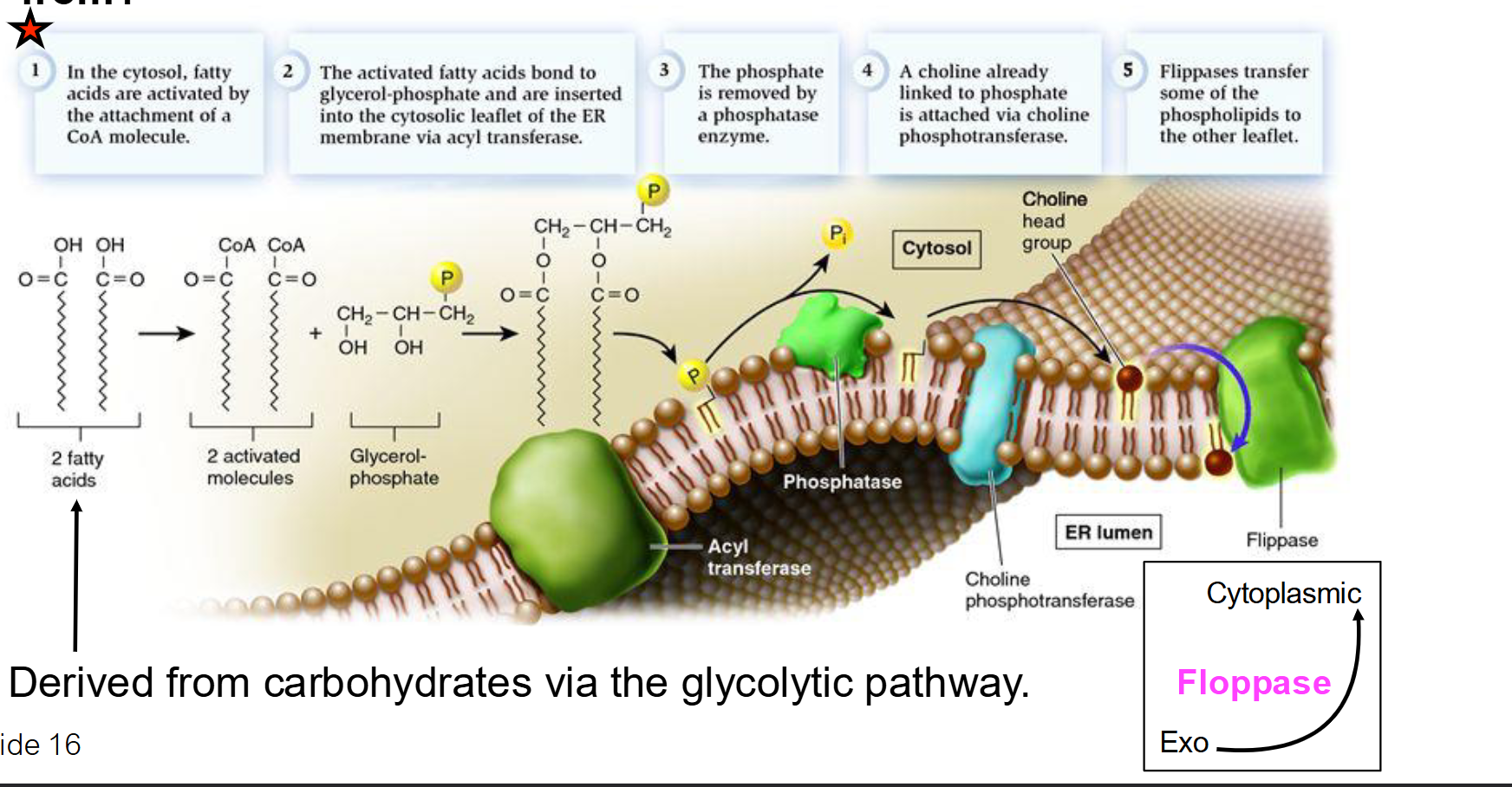
What are the 5 major steps of phospholipid synthesis in the ER membrane?
Activation: Fatty acids in the cytosol are "activated" by attaching to a CoA molecule. This provides the energy needed for the next step.
Insertion: Acyl transferase bonds fatty acids to glycerol-phosphate and inserts them into the cytosolic leaflet (the half of the bilayer facing the cytosol).
Dephosphorylation: Phosphatase removes the phosphate group, preparing it for the attachment of a specific head group.
Attachment: A polar head group (e.g., Choline) is added to complete the phospholipid.
Translocation: Flippases move phospholipids to the opposite leaflet (side) to balance membrane growth.
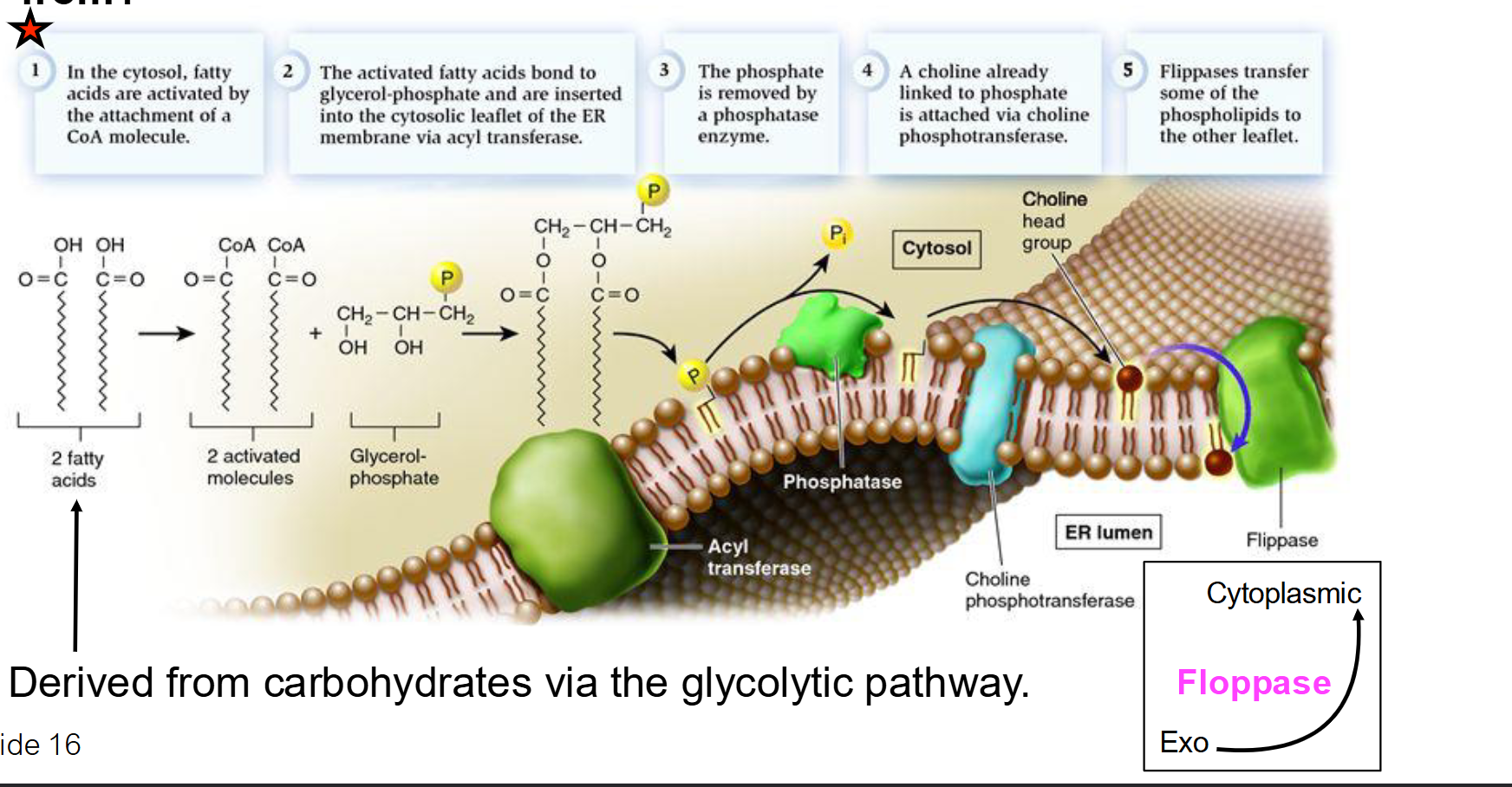
How is a specific "head group" (like Choline) added to the phospholipid?
The head group is usually "activated" by being attached to a carrier molecule like CDP.
This activated head group is then joined to the phosphatidic acid backbone by enzymes in the ER membrane.
If phospholipids are only added to the cytosolic side of the ER, how does the membrane grow evenly?
Enzymes called flippases (or scramblases) catalyze the "flip-flop" movement, transferring newly made phospholipids from the cytosolic leaflet to the exoplasmic (lumen) leaflet.
How do newly made phospholipids travel from the ER to the Plasma Membrane?
They travel via vesicular transport.
A portion of the ER membrane buds off to form a vesicle, which then travels through the cytosol and fuses with the plasma membrane, incorporating the new lipids into it.
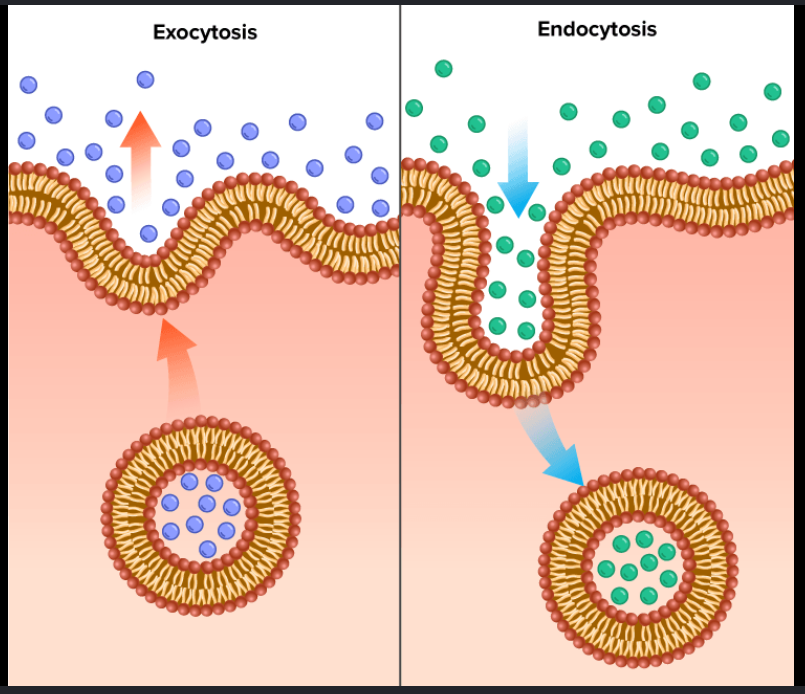
How do Exocytosis and Endocytosis affect the plasma membrane's size and composition?
Exocytosis: Increases membrane surface area by fusing ER/Golgi-derived vesicles with the plasma membrane, delivering new lipids and proteins.
Endocytosis: Decreases surface area by "pinching off" sections of the plasma membrane to bring materials into the cell for recycling or degradation.
Balance: Together, they maintain the steady-state size and specific protein/lipid makeup of the cell's exterior.
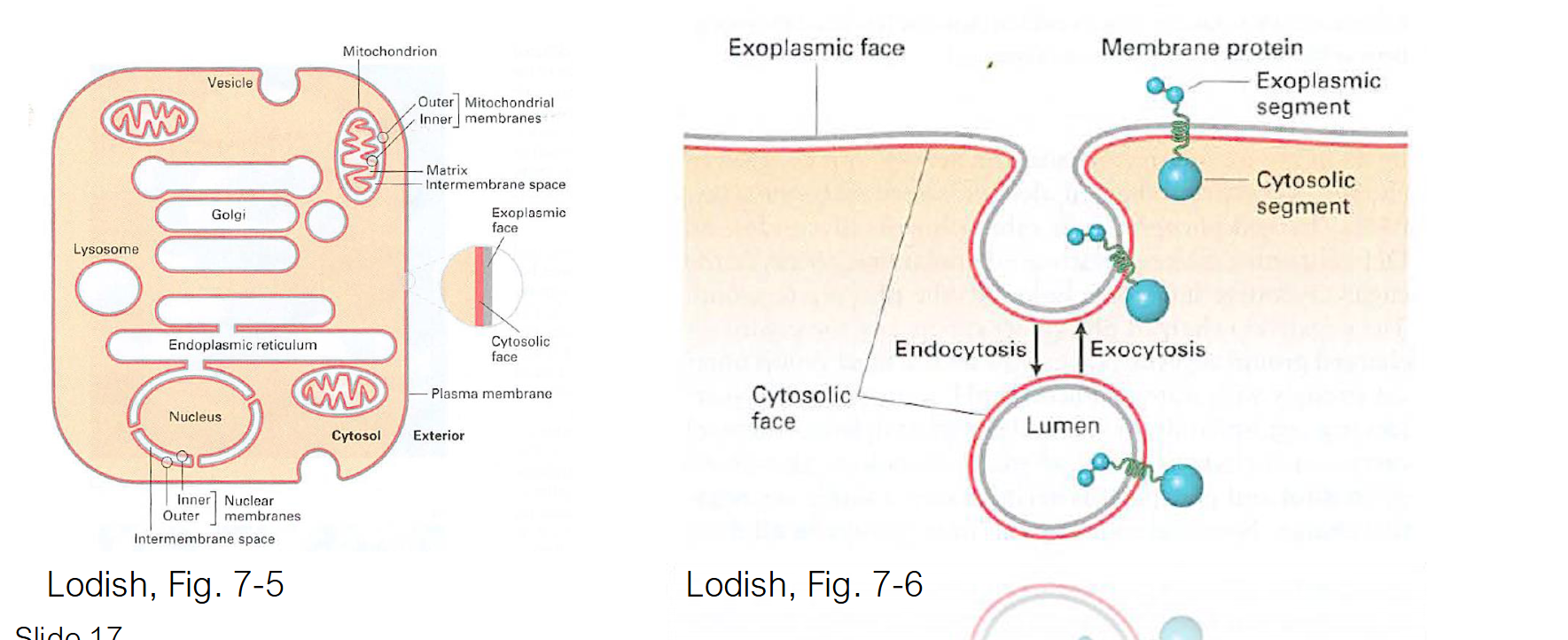
Phospholipid Orientation
Leaflets (the two halves of the bilayer) maintain their relative orientation during transport
Cytosolic Leaflet: The half of the membrane that faces the cytosol while it is part of the ER will always face the cytosol, even after it is transported to the plasma membrane.
Exoplasmic Leaflet: The half that faces the "inside" (lumen) of the ER becomes the "inner" leaflet of the transport vesicle. When that vesicle fuses with the plasma membrane, this leaflet will face the exterior of the cell.
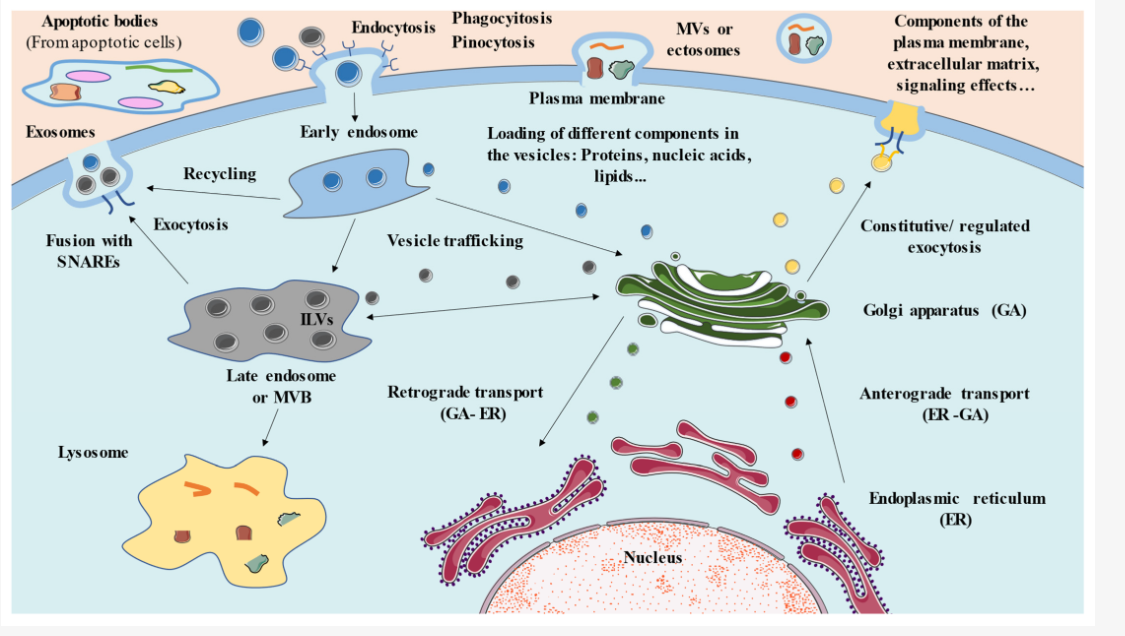
True or False: The leaflet of a vesicle that faces the cytosol will eventually face the outside of the cell after fusion.
False. The leaflet that faces the cytosol in the ER and the transport vesicle will always face the cytosol, even after it fuses with the plasma membrane.
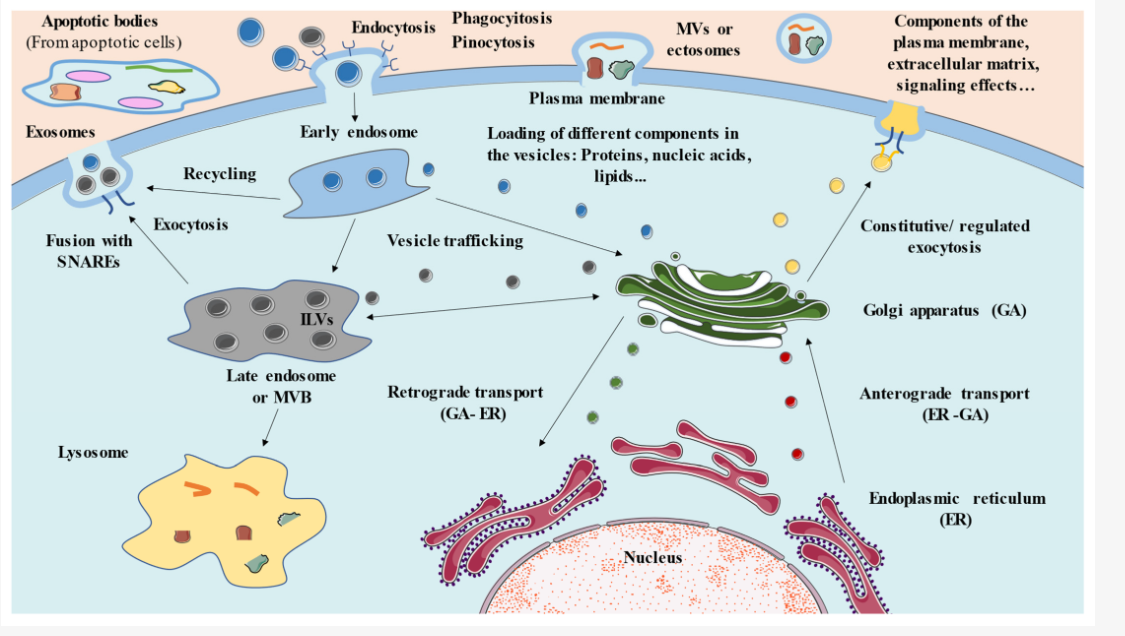
Why is the plasma membrane described as a "Mosaic"?
The membrane is a "mosaic" because it is a complex mixture of different molecules.
Phospholipids: The "lake" in which everything else floats. Form a fluid and flexible bilayer.
Proteins: These are the functional "tiles" of the mosaic. They move laterally (side-to-side) within the fluid lipid layer.
Often serve as gates and pumps for substances
Carry out biochemical reactions
Cholesterol: Found in animal cells, it sits between phospholipids to maintain the right level of stiffness.
Carbohydrates: Located on the outer surface only, attached to proteins (glycoproteins) or lipids (glycolidids). They act as recognition sites (like cellular "ID tags").
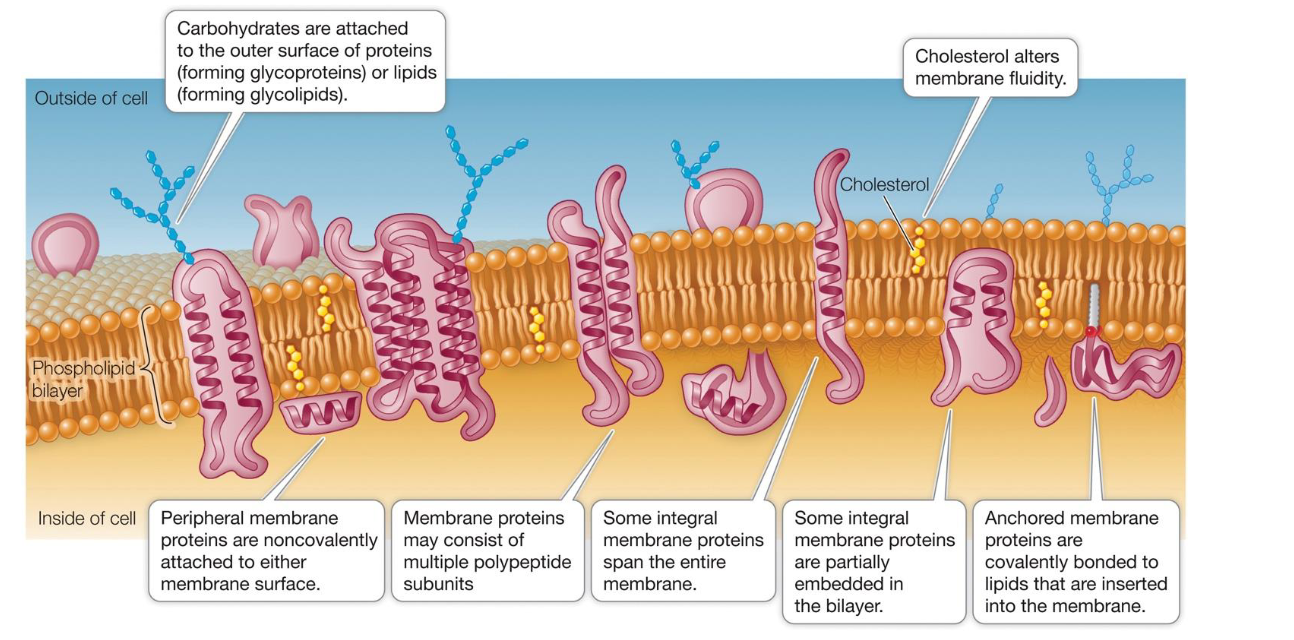
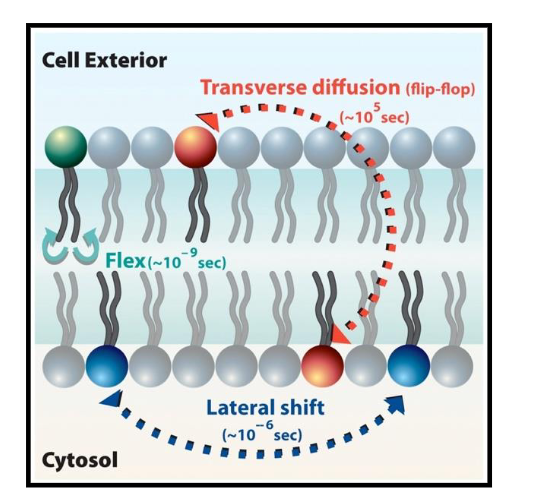
Contrast Lateral Diffusion vs. Transverse Diffusion (Flip-Flop).
Lateral Diffusion: Movement side-to-side within the same layer; happens rapidly and spontaneously.
Most lipids and proteins have “weak” hydrophobic interactions; proteins can move laterally in bilayer too
A few proteins are covalently attached (anchored) to membrane lipids (or to cytoskeleton proteins)
Transverse Diffusion: Movement from one layer to the opposite layer; happens very slowly/rarely because the polar head must cross the hydrophobic core.
This only happens efficiently when aided by enzymes like flippases.
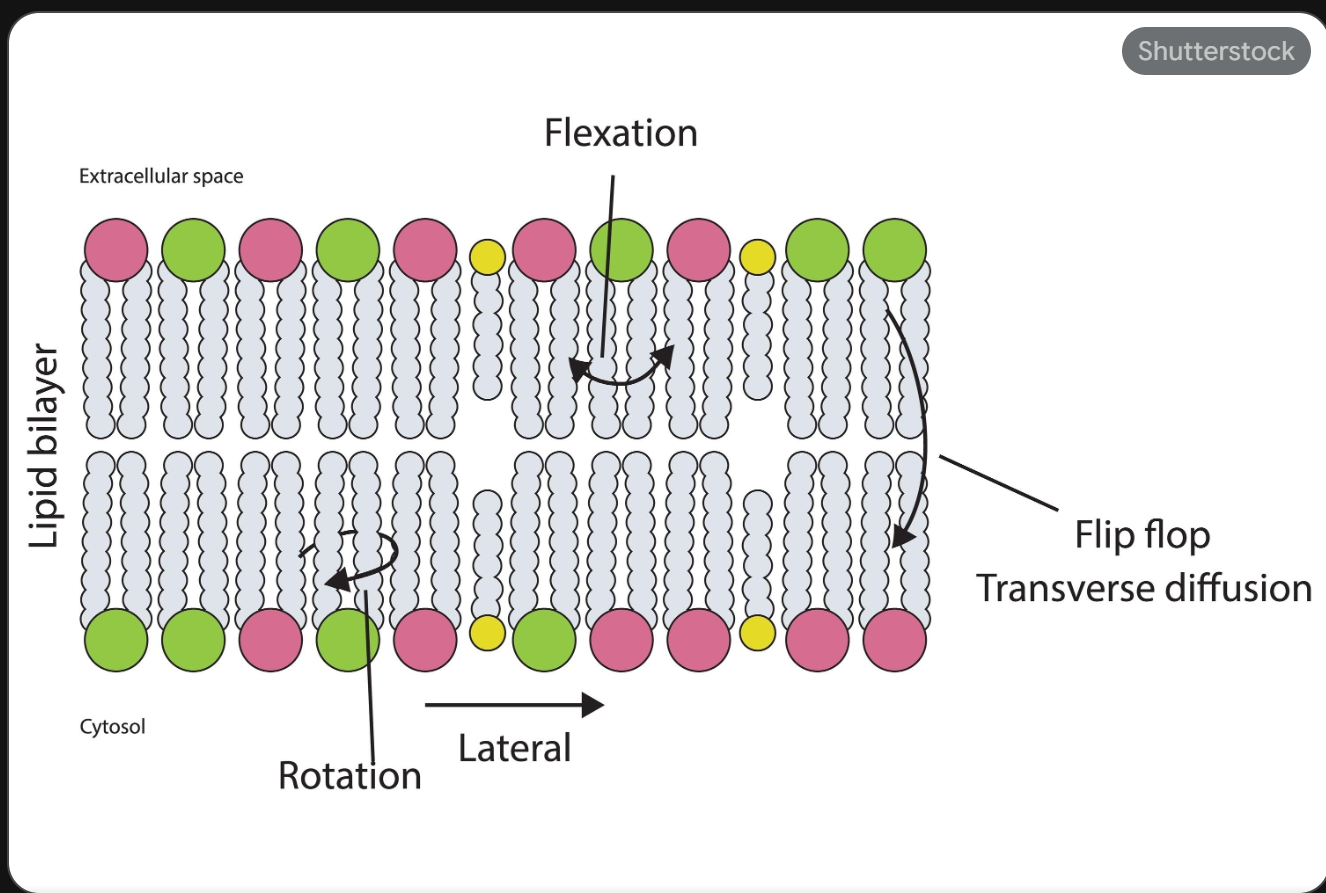
Summary of Fluid Mosaic Model
Proposed by Seymour Jonathan Singer and Garth Nicolson in 1972 → The plasma membrane is viewed as a two-dimensional liquid that restricts the diffusion of membrane components
Fluid — individual lipid molecules move
Mosaic — diverse ‘particles’ like proteins, carbohydrates, and cholesterol penetrate the lipid layer
Components are mobile.
Components can interact
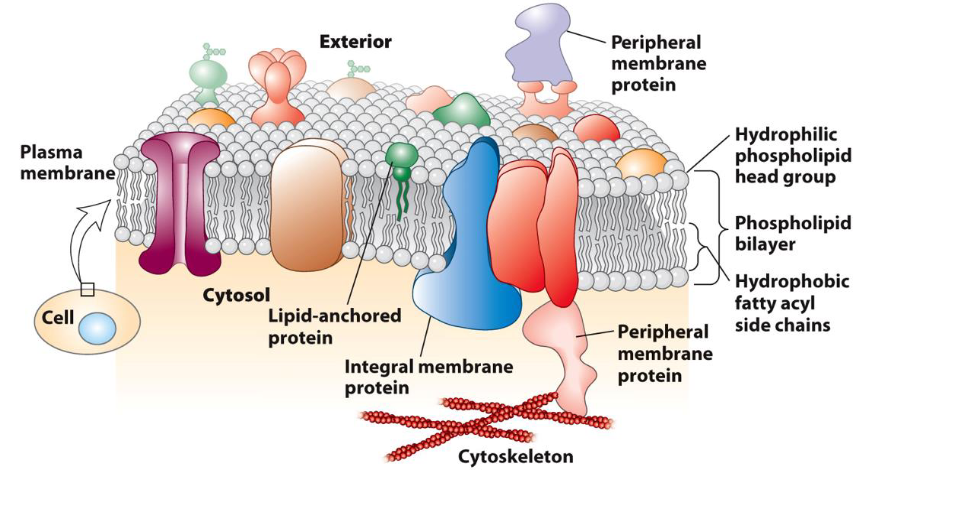
What are the defining characteristics and roles of Integral Proteins?
Placement: They are permanently embedded in the bilayer; most are "transmembrane," spanning from one side to the other.
Chemistry: They have hydrophobic regions that sit among the lipid tails and hydrophilic regions that face the water.
Movement: They can move laterally within the fluid membrane (proven by the Frye-Edidin experiment).
Removal: Because they are deeply embedded, they can only be removed by disrupting the entire bilayer (usually with detergents).
Functions: Act as channels/transporters, signal receptors, and cell-recognition markers (glycoproteins).
Facilitate the movement of polar molecules, ions, and large nutrients across the hydrophobic lipid bilayer, which these substances cannot cross on their own
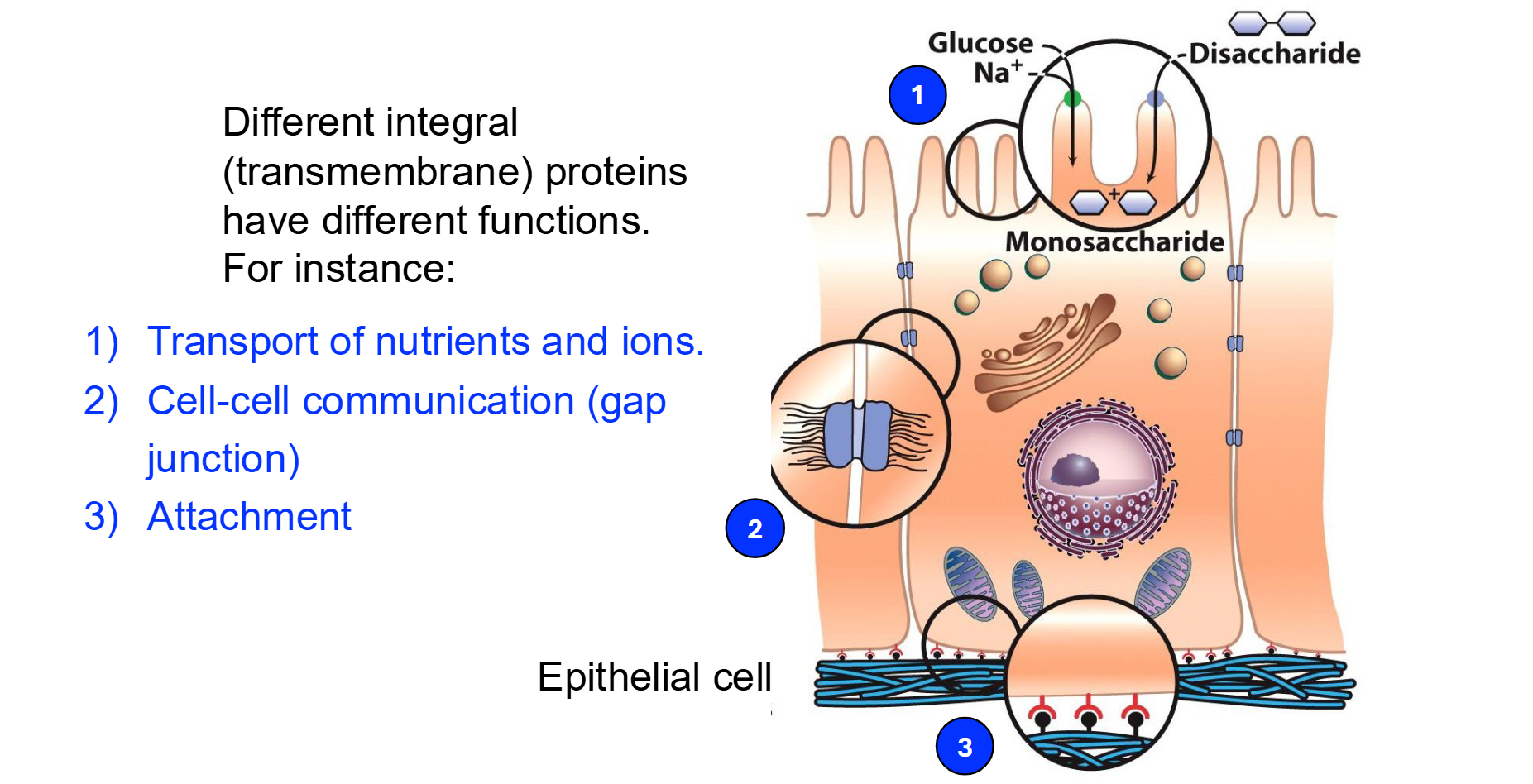
What are the defining characteristics and roles of Peripheral Membrane Proteins?
Placement: They are located entirely outside the lipid bilayer, positioned on either the cytoplasmic or the extracellular side.
Bonding: They do not penetrate the hydrophobic core. Instead, they stay attached via weak non-covalent interactions (like ionic or hydrogen bonds) with integral proteins or the polar heads of phospholipids.
Chemistry: They are composed of hydrophilic amino acids, making them soluble in aqueous buffers.
Removal: They can be removed from the membrane relatively easily using changes in pH or salt concentration, without disrupting the bilayer itself.
Functions: They often act as enzymes, signaling molecules, or anchors that connect the membrane to the cell's internal cytoskeleton.
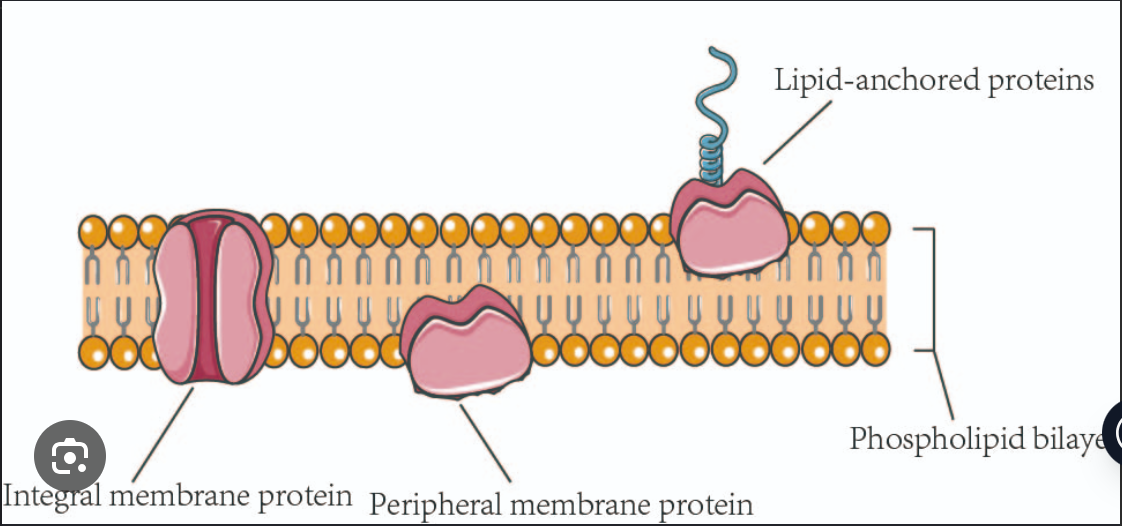
Dynamics of Plasma Membrane
Lipids move easily, laterally, within leaflet (Lateral, Rotation, Flexing).
Lipids movement to other leaflet is difficult and slow
Membrane proteins diffuse within the bilayer:
Movement of proteins is restricted.
Rapid movement is spatially limited.
Long range diffusion is slow.
Biochemical modification can alter protein mobility in the membrane—an important feature for signal transduction
What factors influence membrane fluidity, and how do they work?
Temperature: Heat increases fluidity (movement increases); cold increases viscosity (solidification).
Saturation: Unsaturated tails (with kinks) increase fluidity by preventing tight packing. Saturated tails (straight) decrease fluidity by packing tightly.
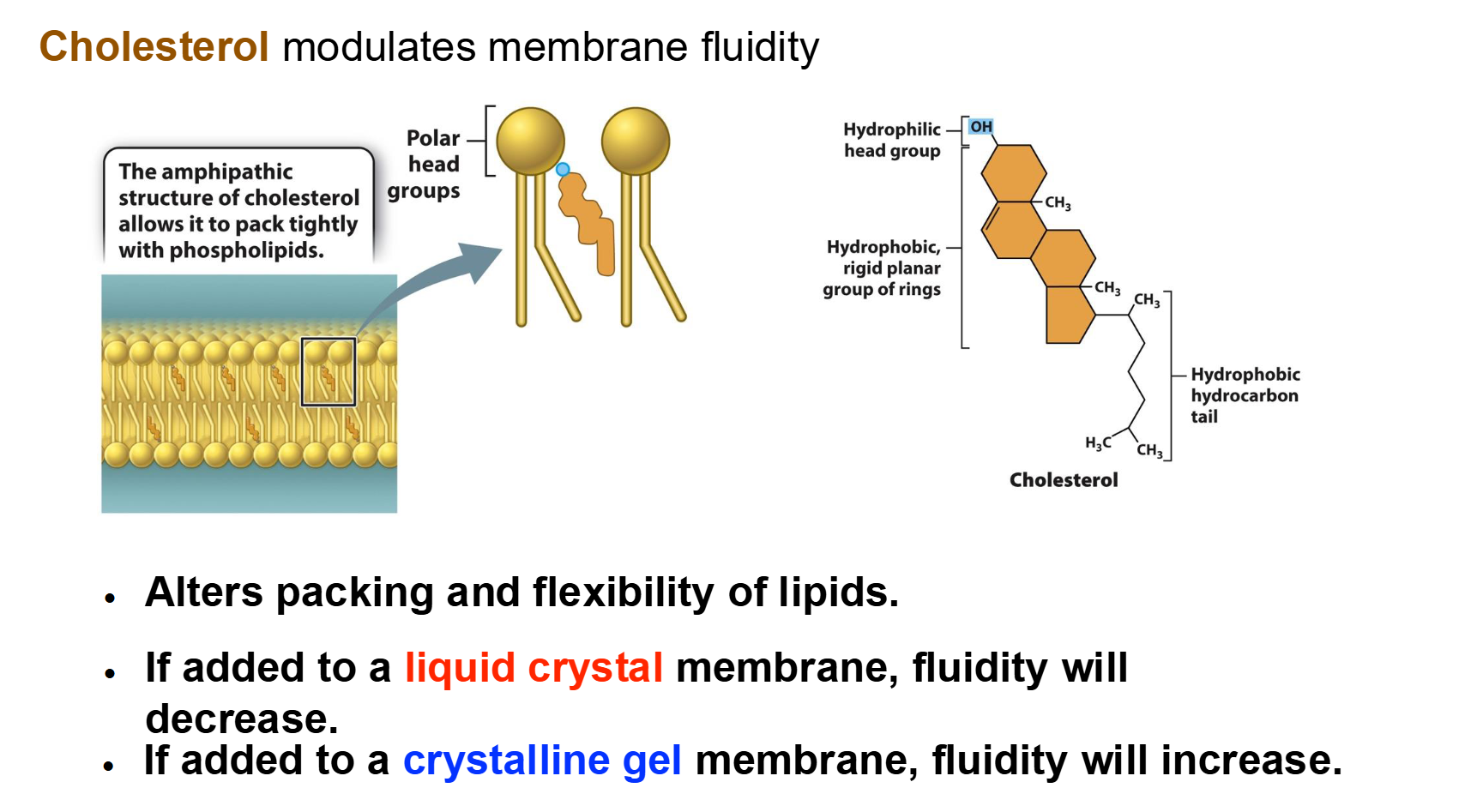
Cholesterol: Acts as a buffer.
At warm temperatures, it restrains phospholipid movement so the membrane doesn't become too liquid.
At cool temperatures, it gets in the way of tight packing, preventing the membrane from solidifying.
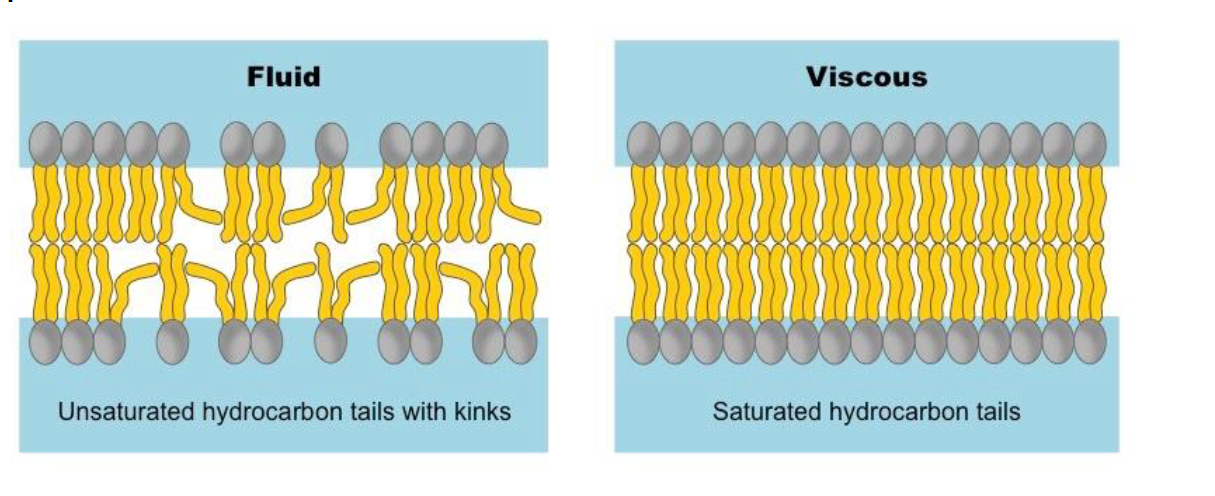
Fluidity of The Biological Membrane
Warming increases fluidity ⟶ liquid crystal
Cooling decreases fluidity ⟶ crystalline gel
Membrane fluidity is determined by the nature of lipids in membrane:
Unsaturated lipids increase fluidity
Saturated lipids reduce fluidity
Cholesterol acts as a bidirectional regulator of membrane fluidity because at high temperatures, it stabilizes the membrane and raises its melting point
At low temperatures it intercalates between the phospholipids and prevents them from clustering together and stiffening.
What was the primary conclusion of the Frye-Edidin experiment?
It proved that the plasma membrane is fluid and that membrane proteins are capable of lateral diffusion rather than being locked in fixed positions.
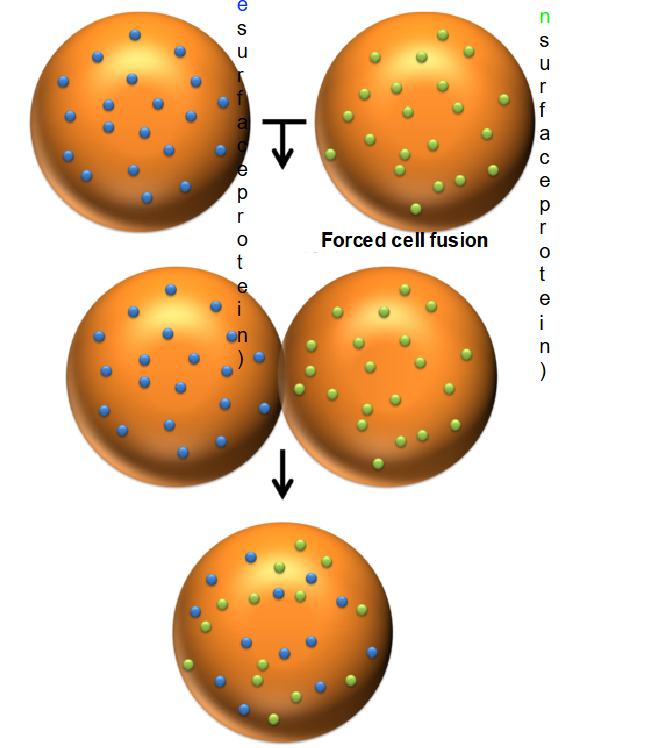
Cell Fusion: A mouse cell and a human cell are fused together to create a single "hybrid" cell.
Labeling: The surface proteins of each cell are tagged with different fluorescent markers (e.g., green for mouse and blue for human).
Initial Observation: Immediately after fusion, the proteins remain segregated—the mouse proteins stay on one half of the hybrid cell, and the human proteins stay on the other.
The Result: After 40 minutes at 37^C, the proteins have fully intermixed across the entire surface of the hybrid cell.
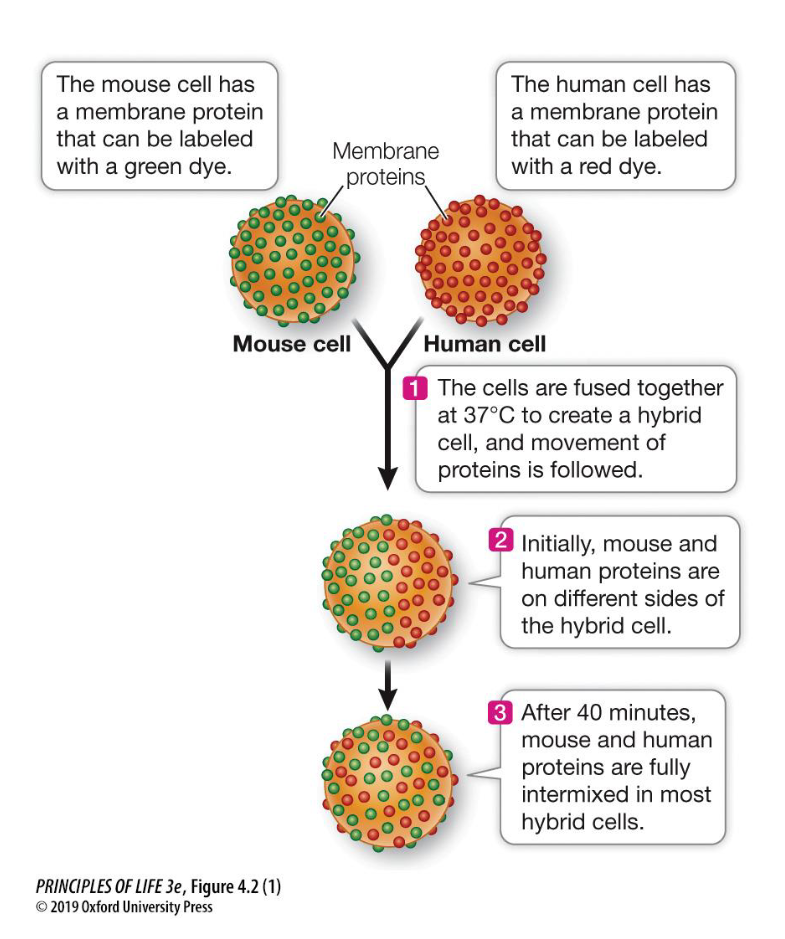
Describe the 40-minute result of the Frye-Edidin experiment at 37^C
After 40 minutes, the mouse and human surface proteins, which were initially on separate halves of the hybrid cell, became fully intermixed across the entire membrane.
Higher temperatures increase membrane fluidity, leading to faster and better intermixing.
Lower temperatures make the membrane more viscous, inhibiting the movement of proteins.
Part 1 Summary

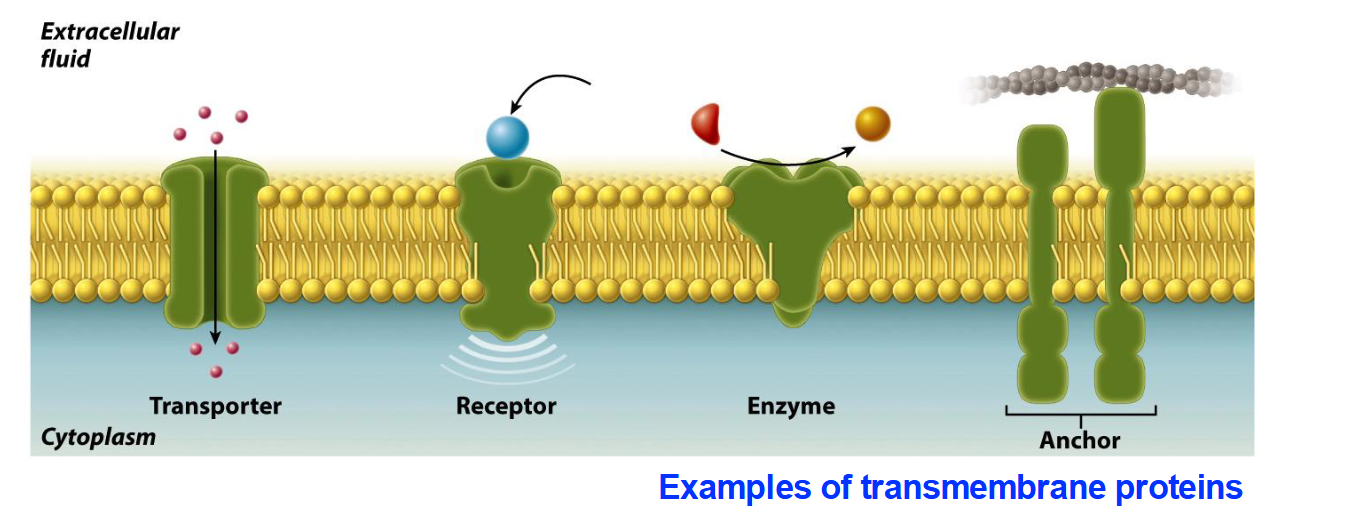
Identify the four functional types of transmembrane proteins mentioned in the slides.
Transporters: Move ions or molecules across the membrane.
Receptors: Bind to specific ligands (like substrates to enzymes) to trigger cellular responses.
Enzymes: Catalyze chemical reactions at the membrane surface.
Anchors: Attach the membrane to other structures like the cytoskeleton.
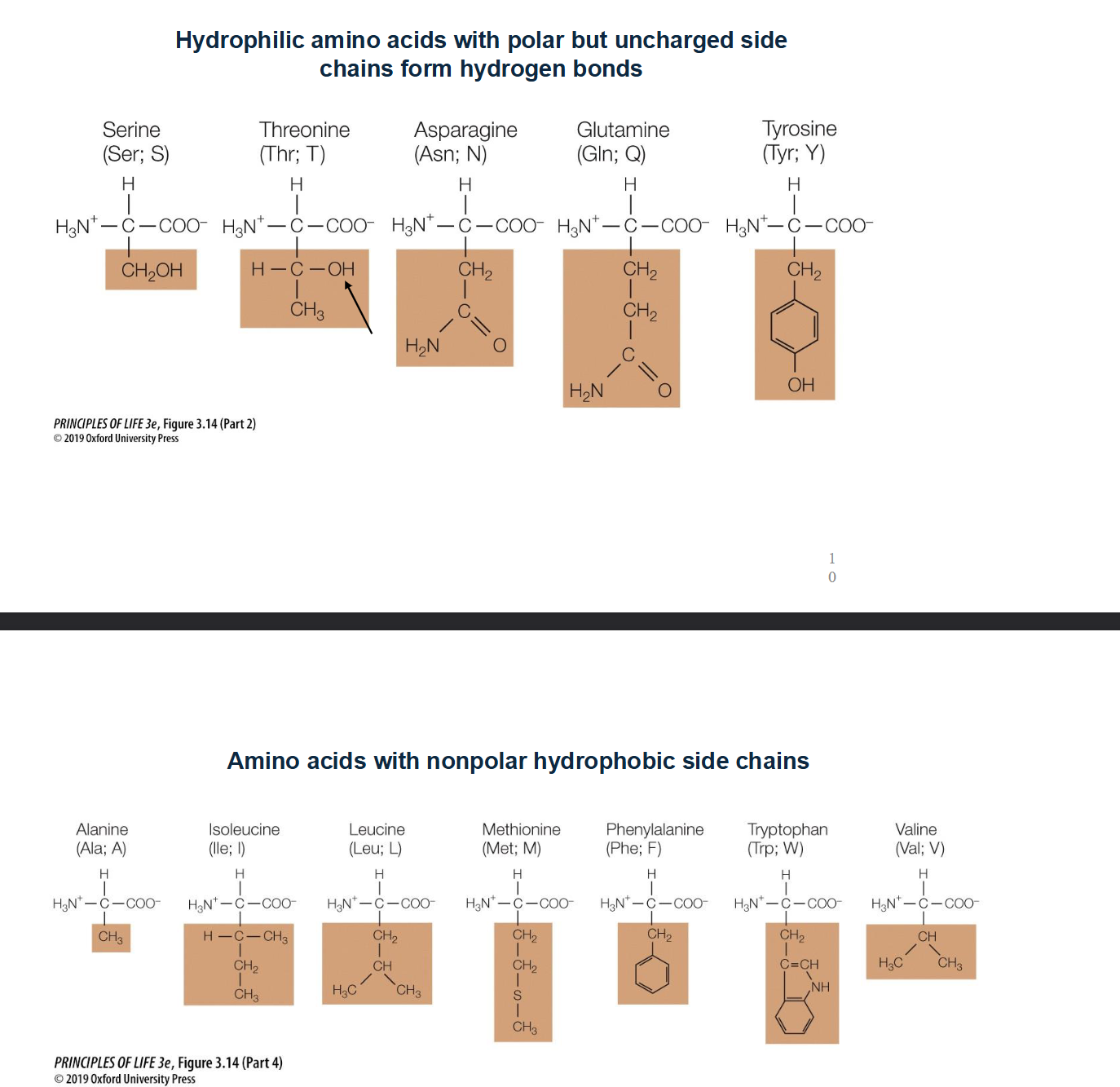
Which Amino-Acids can exist within the lipid membrane?
Non-Polar or uncharged amino acids!
Look for chains that just contain methane (CH3)
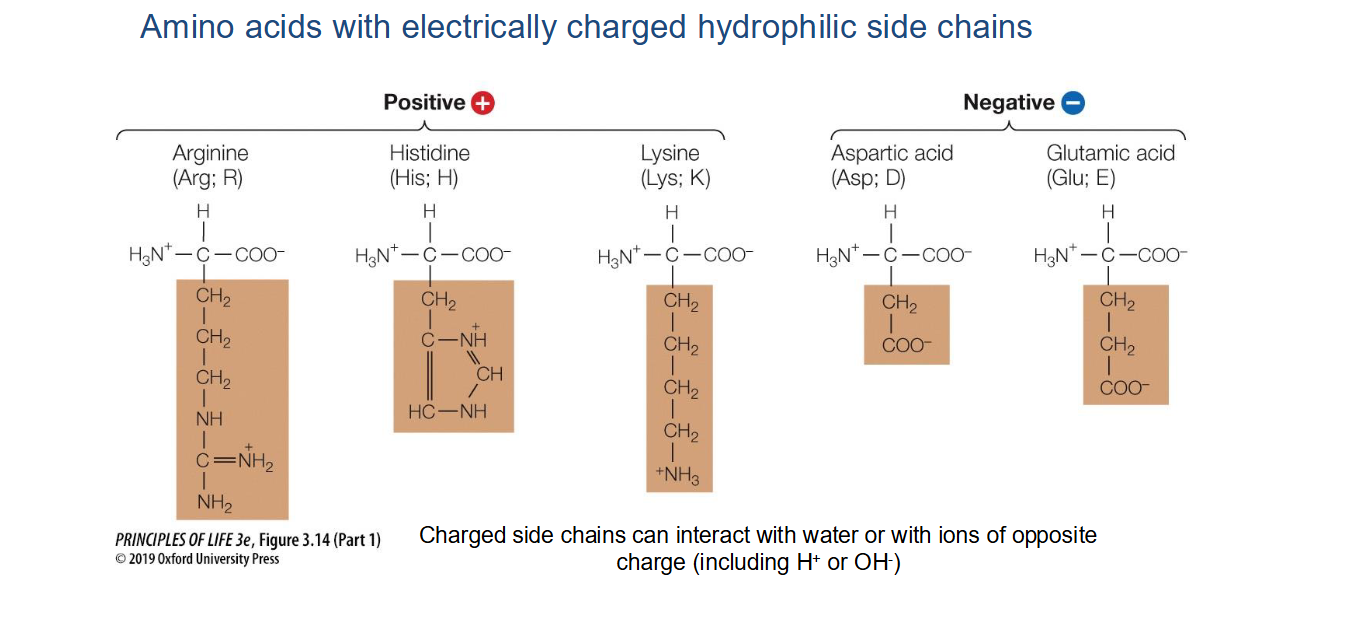
Which amino acids have electrically charged hydrophilic side chains?
Positively charged: Arginine (Arg), Histidine (His), and Lysine (Lys).
Negatively charged: Aspartic acid (Asp) and Glutamic acid (Glu).
Note: These side chains can interact with water or ions of opposite charge
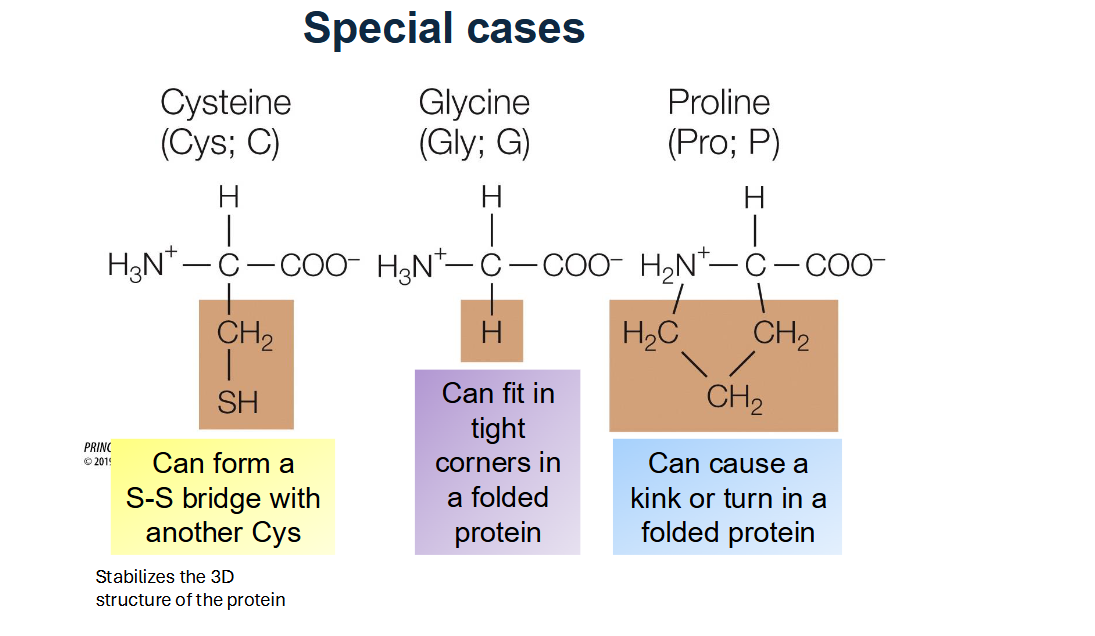
List the three "Special Case" amino acids and their unique properties.
Cysteine (Cys): Can form disulfide bridges (S-S bonds) with other cysteines to stabilize protein 3D structure.
Glycine (Gly): The smallest amino acid (R group is just H), allowing it to fit into tight corners of folded proteins.
Proline (Pro): Has a ring-shaped R group that limits rotation and can cause "kinks" or turns in a protein chain.
What is a transmembrane protein domain and its most common structural element?
A transmembrane domain is a peptide sequence that is largely hydrophobic (uncharged) and spans the plasma membrane, permanently attaching the protein to it.
The alpha helix is the most common structural element used to cross biological membranes.
The spiral shape allows the protein to "hide" its polar backbone on the inside while exposing its "greasy" side chains to the membrane lipids.
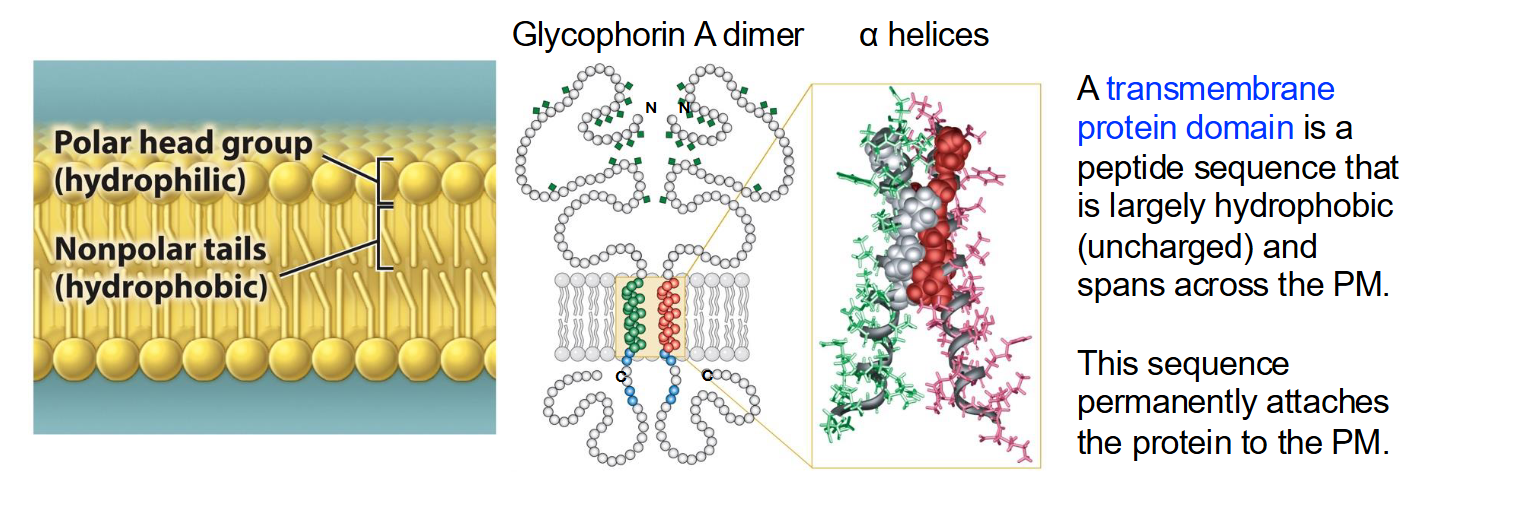
List the 9 amino acids that have hydrophobic side chains
Glycine (Gly), Alanine (Ala), Valine (Val), Leucine (Leu), Isoleucine (Ile), Proline (Pro), Phenylalanine (Phe), Methionine (Met), and Tryptophan (Trp).
These are frequently found in transmembrane domains.
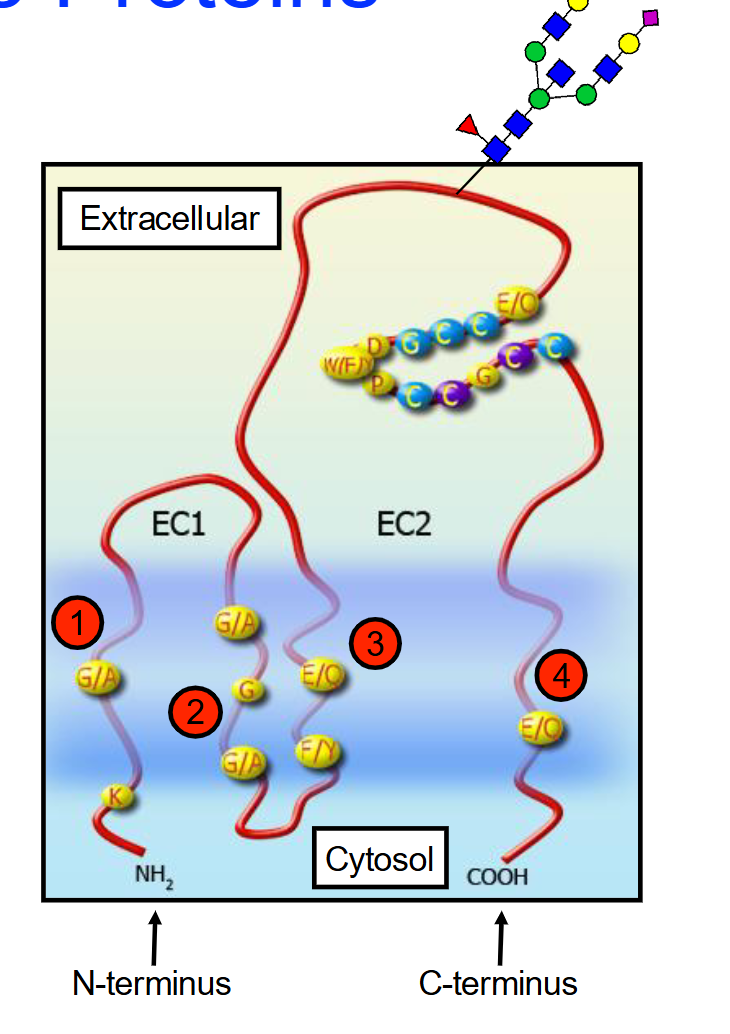
What are Tetraspanins (TM4SFs)?
A family of membrane proteins found in multicellular eukaryotes characterized by having four transmembrane alpha-helices and two extracellular domains (EC1 (short) and EC2 (longer) ).
Some tetraspanins can be glycosylated (attachment of a carbohydrate molecule) on the long extracellular loop
Function: These aren't just anchors; they act as "scaffolds" that help cells stick together (adhesion), move around (motility), and signal each other to grow (proliferation).
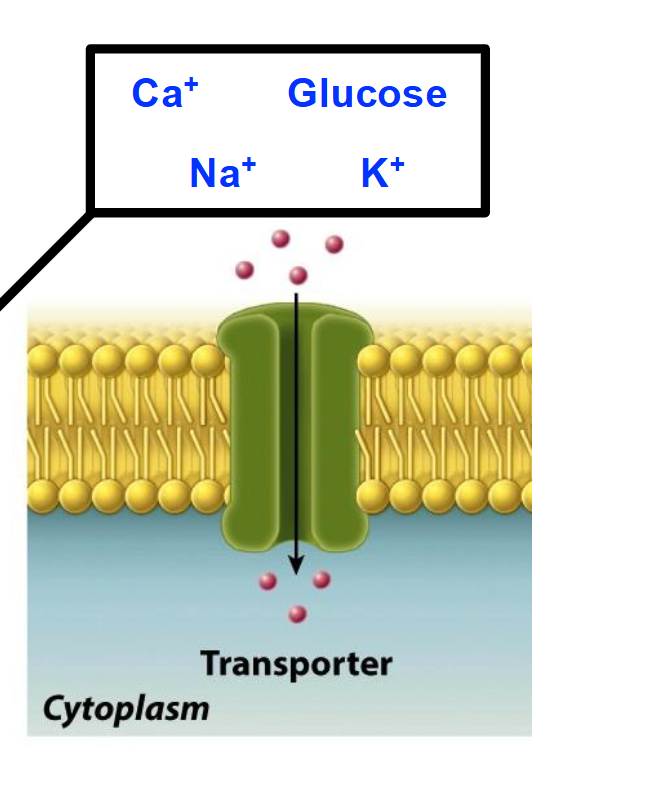
Movement of Substances Across Cell Membranes
Lipid bilayers do not allow many compounds or molecules to pass through them freely.
Small, uncharged molecules cross membranes relatively easily (O2, CO2, NO).
Large/polar/charged compounds cannot easily cross lipid bilayers.
Specific mechanisms are required for the controlled transport of many substances across membranes
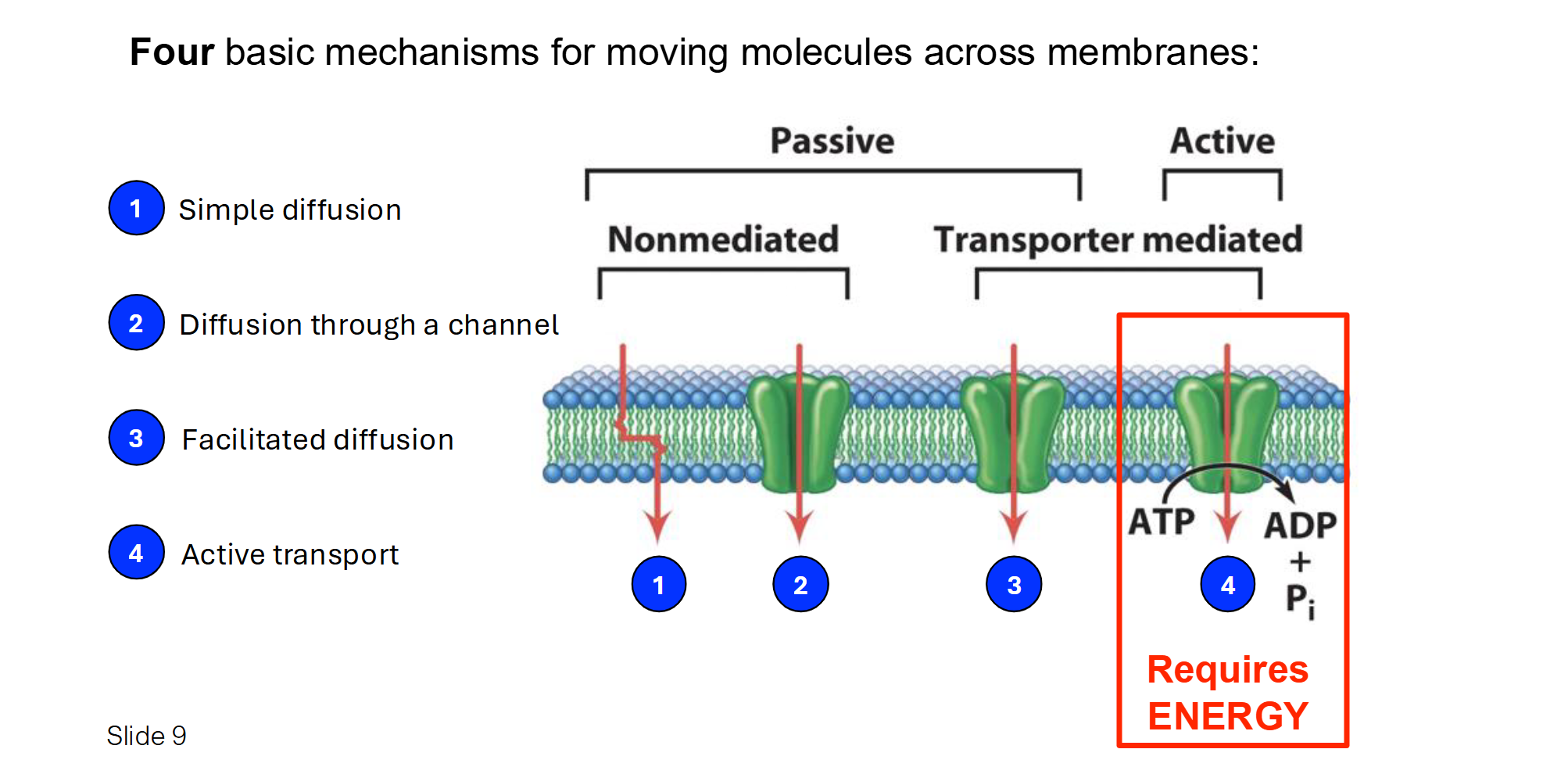
Summary of the 4 Methods of Membrane Transport
Simple Diffusion: Molecules move directly through the lipid bilayer from High to Low concentration. No energy or protein needed. (e.g., O2, CO2).
Facilitated Diffusion (Channel): Molecules move through a protein "tunnel" or pore from High to Low concentration. No energy needed. (e.g., Aquaporins for water).
Facilitated Diffusion (Carrier): Molecules bind to a protein that changes shape to move them from High to Low concentration. No energy needed. (e.g., Glucose transport).
Active Transport: Molecules are pumped against the gradient from Low to High concentration. Requires ATP (Energy). (e.g., Na+/K+ pump).
Passive Diffusion
Passive movement of substances across cell membranes relies on molecular concentrations.
DOES NOT require energy.
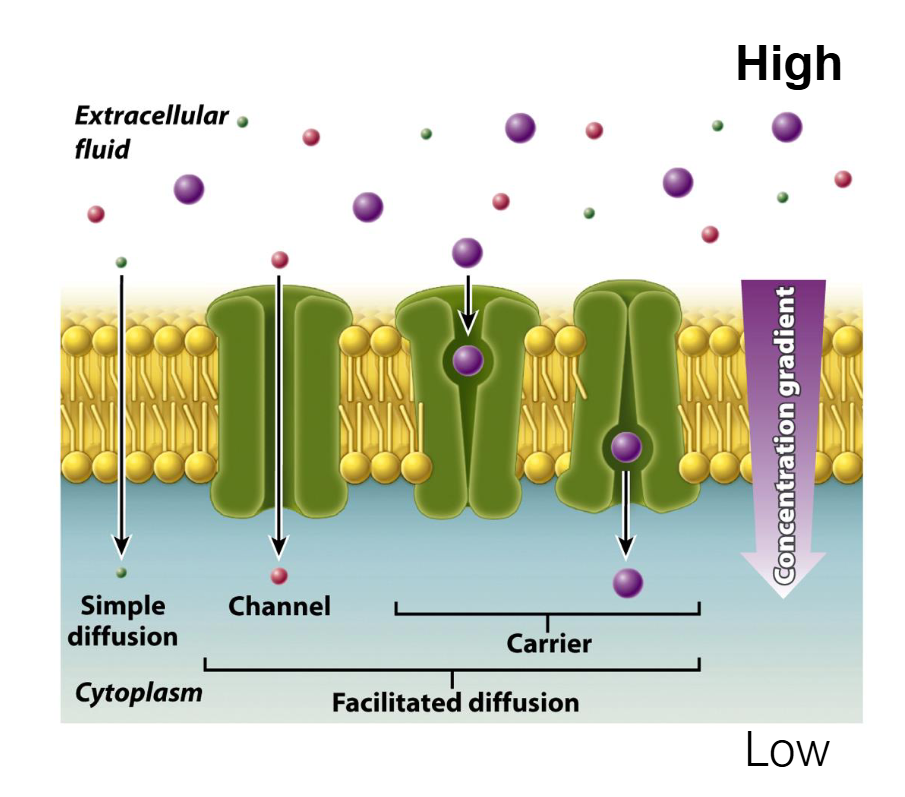
Simple Diffusion
Direction: Down the concentration gradient (High to Low).
Energy: No ATP required (Passive).
Mechanism: Molecules pass directly through the phospholipid bilayer.
Molecules: Only small, nonpolar molecules (like O2 and CO2) can do this because they can dissolve in the hydrophobic "greasy" center of the membrane.
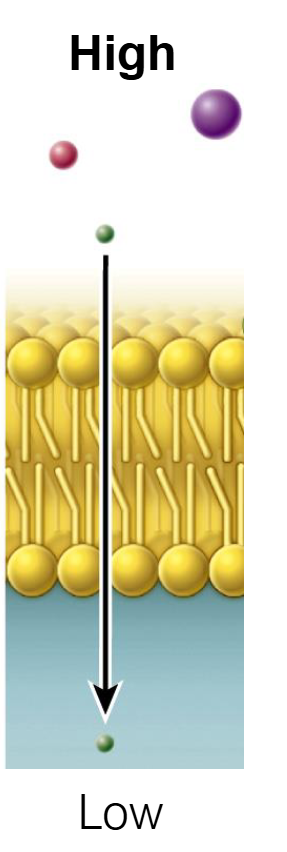
Diffusion Through A Channel (Facilitated Diffusion)
Protein channels provide another form of passive transport.
This mode is particularly effective for small, charged molecules (ions) like Na+, K+, Ca2+, Cl-.
Ions move down concentration gradients — flow is downhill.
Channels are selective, allowing only particular types of ions to pass.
Ion channels are formed by integral membrane proteins—typically multiple subunits—that line an aqueous pore.
Uniporter → Transport happens in one direction
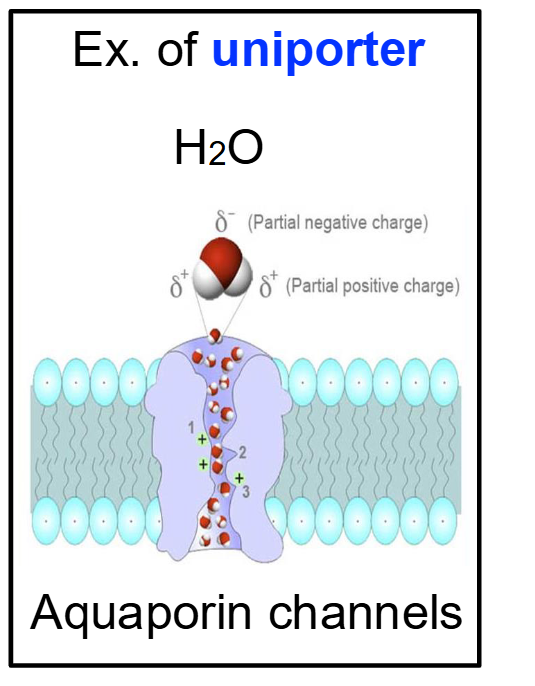
What are Ion Channels?
Channels often are gated — can be open or closed.
This is an important feature because it provides channels the ability of responding to different stimuli (e.g., neurotransmitters).
In other words, they can be turned ON/OFF in response to different signals
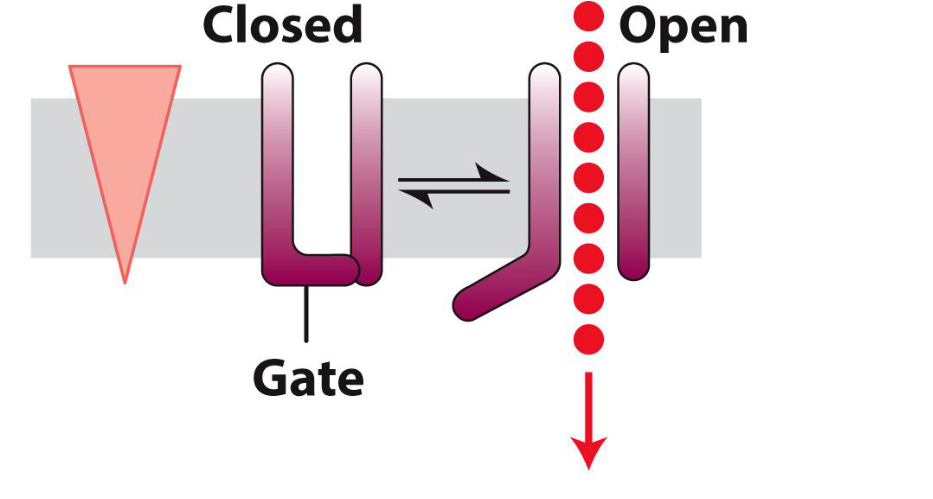
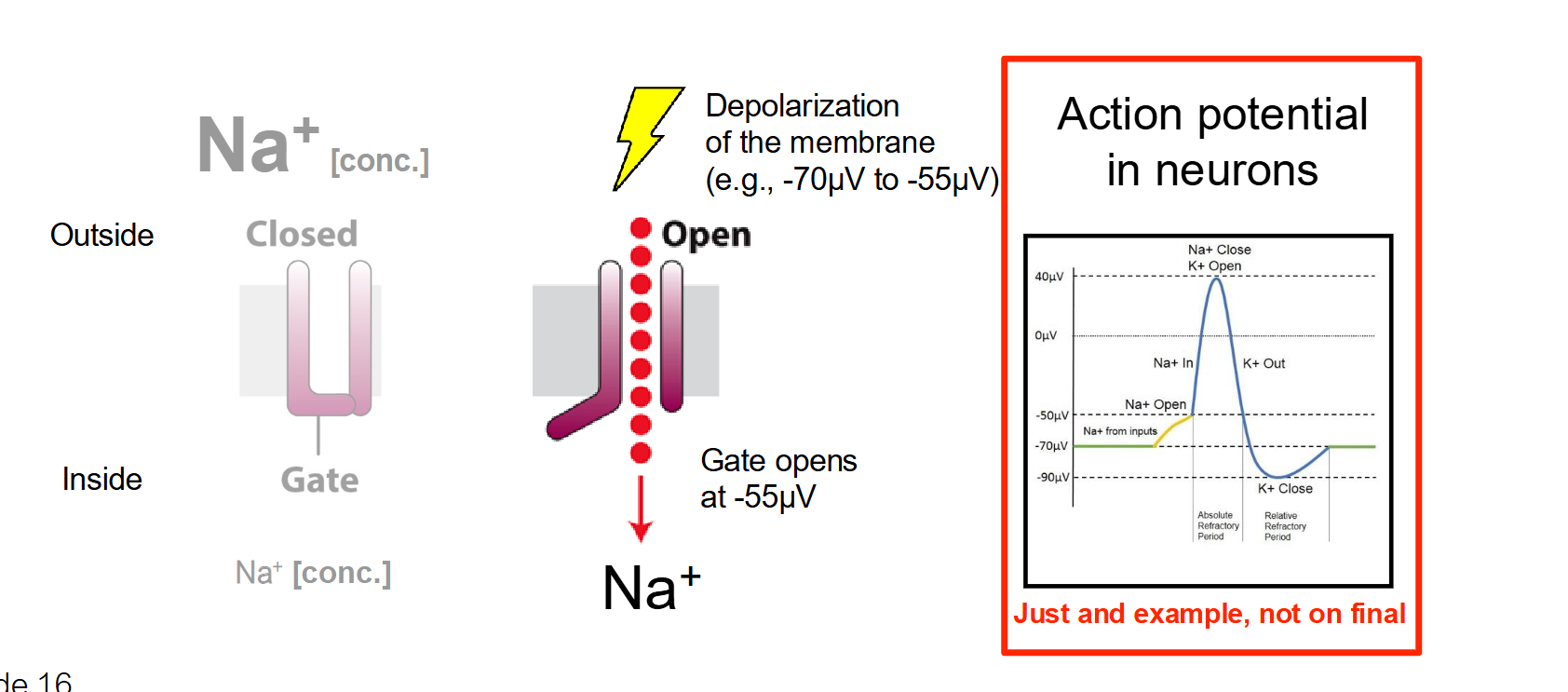
Voltage-Gated Channels
The Trigger: A change in the electrical membrane potential (voltage).
How it works: These channels have sensors that detect electrical charges. When the voltage across the membrane reaches a certain threshold (like during a nerve impulse), the channel changes shape to open its gate.
Location: Found extensively in neurons and muscle cells to facilitate action potentials.
Ex. Na+ → Under non-depolarized conditions (when they are quiet), neurons have little amount of Na+ inside them…but during the gradeint changes and they take in Na+
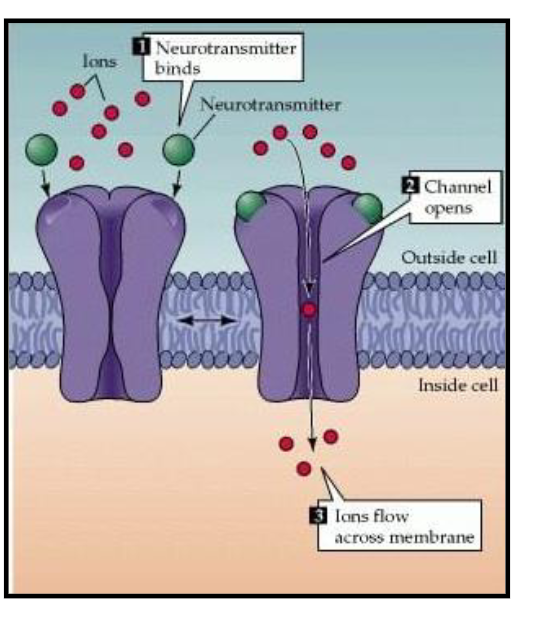
Ligand-Gated Channels
A specific molecule (ligand) binding to the outside of the channel.
Binding of a ligand produce conformational change in the structure of the receptor/channel
Think of this like a lock and key. A signaling molecule (like a neurotransmitter) binds to a receptor site on the exterior part of the protein, causing the gate to pop open.
The protein acts like a selective tunnel. When it opens, it forms a continuous pore through the membrane. Ions simply "flow" through the open gate by the thousands.
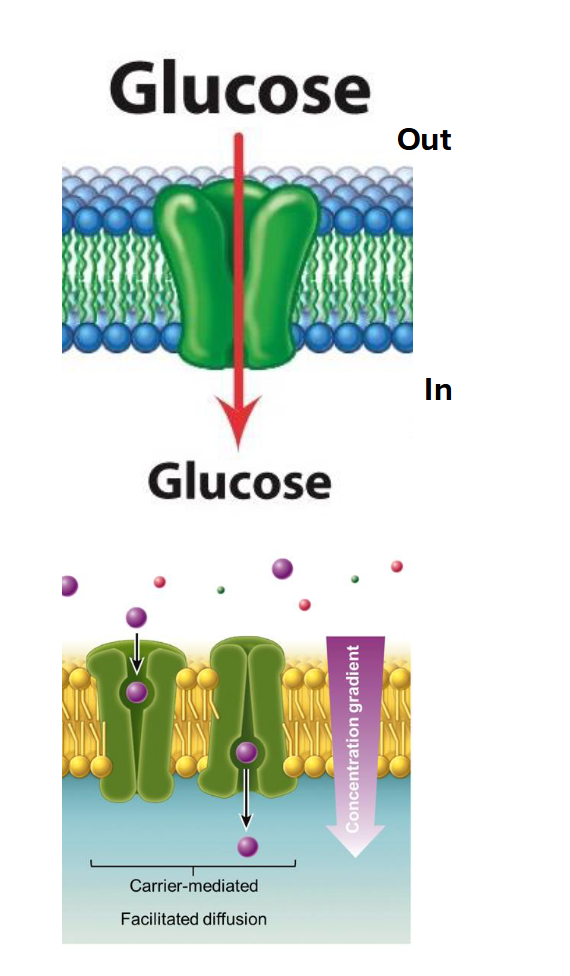
Facilitated Diffusion (Carrier)
Direction: Down the concentration gradient (High to Low).
Energy: No ATP required (Passive).
Mechanism: Uses a transporter (carrier). Unlike a simple tunnel, the carrier protein must bind the molecule, change its shape (conformational change), and then release it on the other side.
Molecules: Larger polar molecules, such as glucose or specific amino acids.
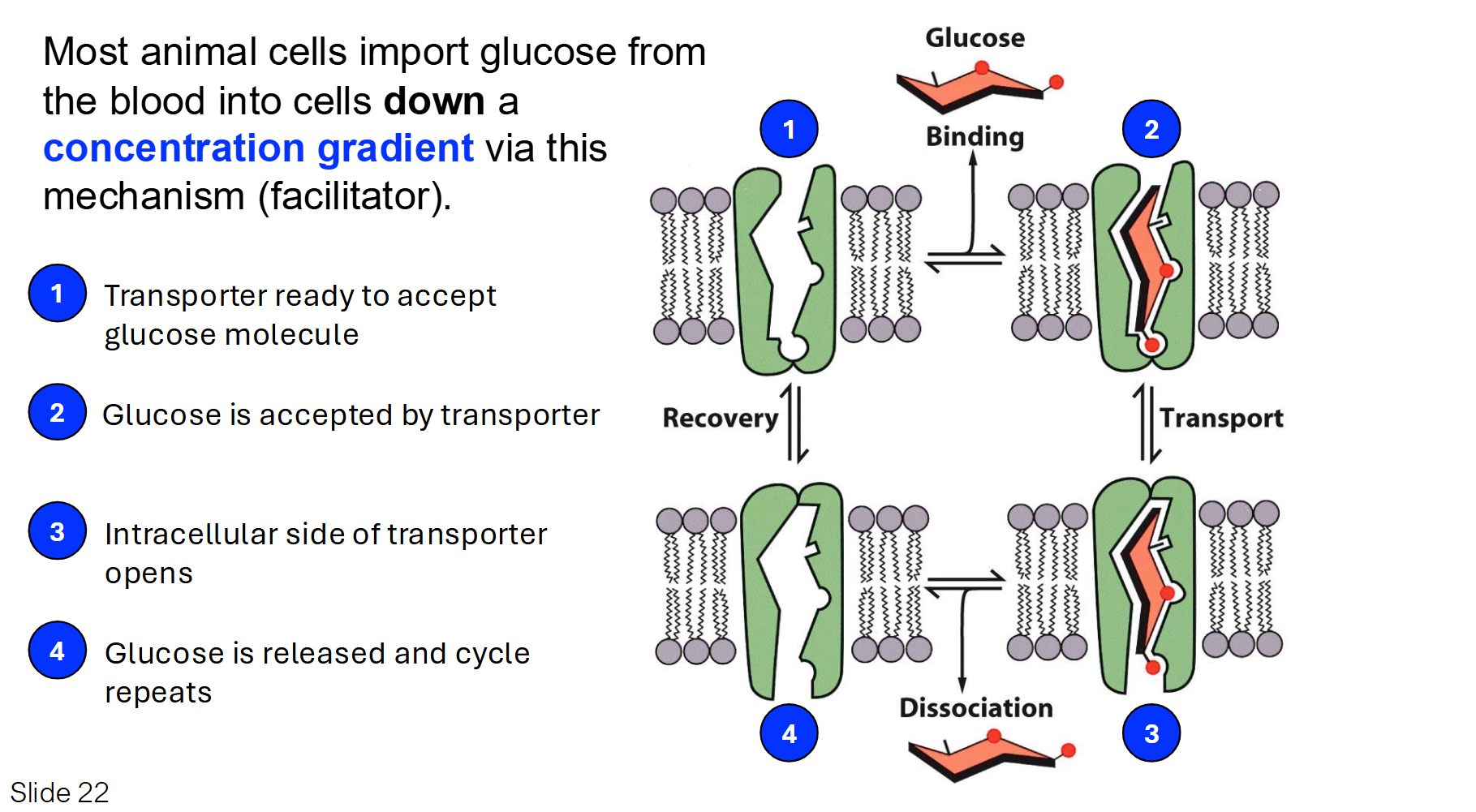
Carriers: Glucose Transporter
The glucose transporter is a classic example of a Uniport. It moves only one type of molecule at a time down its concentration gradient.
Mechanism: Glucose binds to a specific site on the transporter, which causes the protein to flip its shape and release the glucose on the other side.
Key Point: This is facilitated diffusion (passive), meaning it doesn't require ATP.
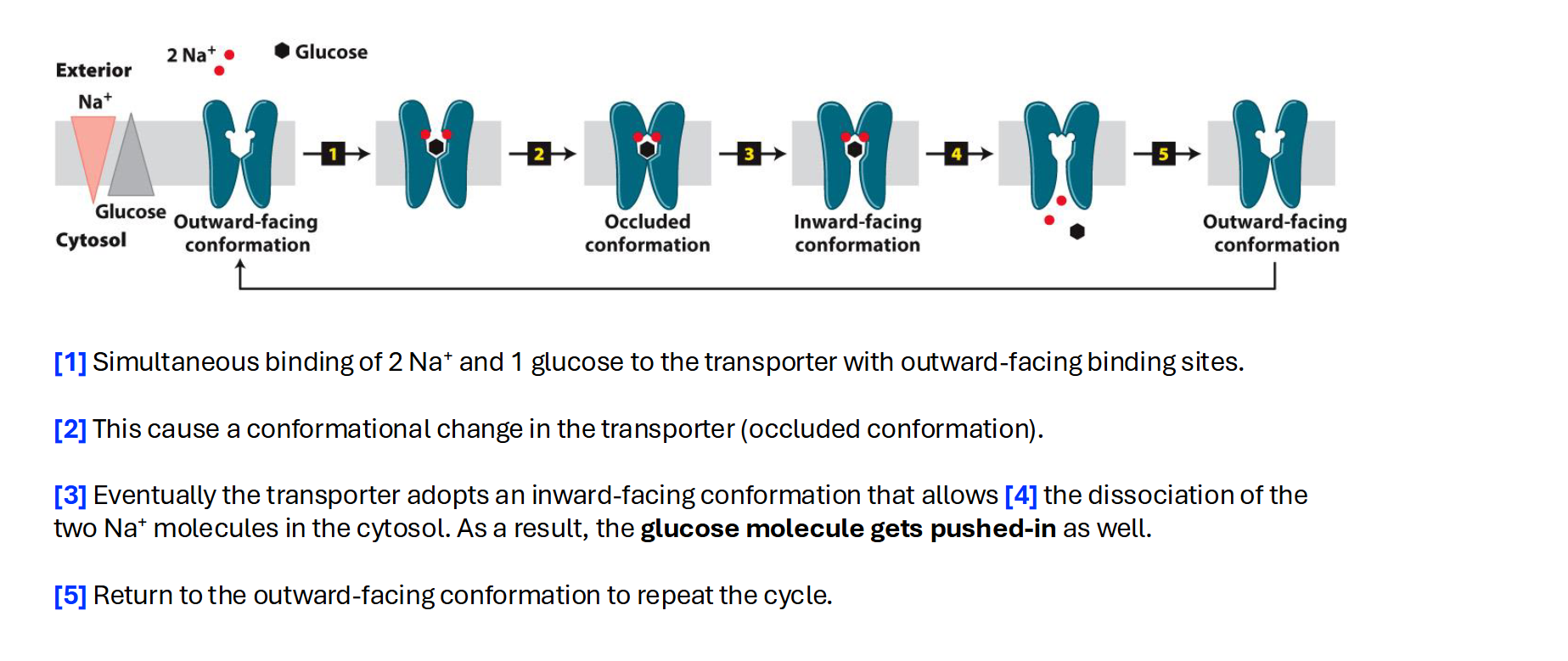
Carrier: Symporter
A symport moves two different substances in the same direction at the same time.
Mechanism: Usually, one substance (like Sodium, Na+) moves down its concentration gradient, and the energy from that movement is used to "drag" a second substance (like Glucose) against its concentration gradient.
Example from your slides: The Na+-glucose symporter.
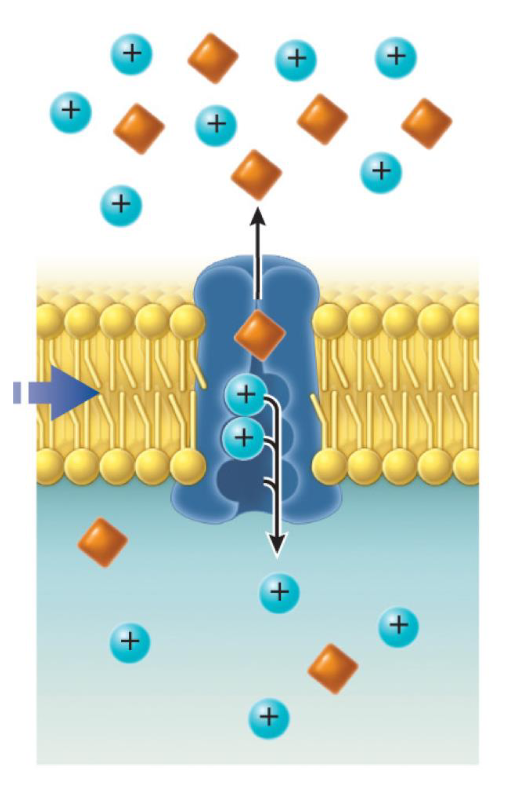
Carrier: Antiporter
An antiport moves two different substances in opposite directions across the membrane.
Mechanism: As one substance enters the cell, another substance is ejected out of the cell simultaneously.
The concentration gradient of one molecule is used to transfer a second molecule in opposite directions.
Example: The sodium-proton exchanger (Na+/H+ exchanger) in the nephron of the kidney. This antiporter transport Na+ into the cell and protons H+ out of the cell.
This carrier is specifically responsible to maintain pH and sodium levels in specific kidney cells
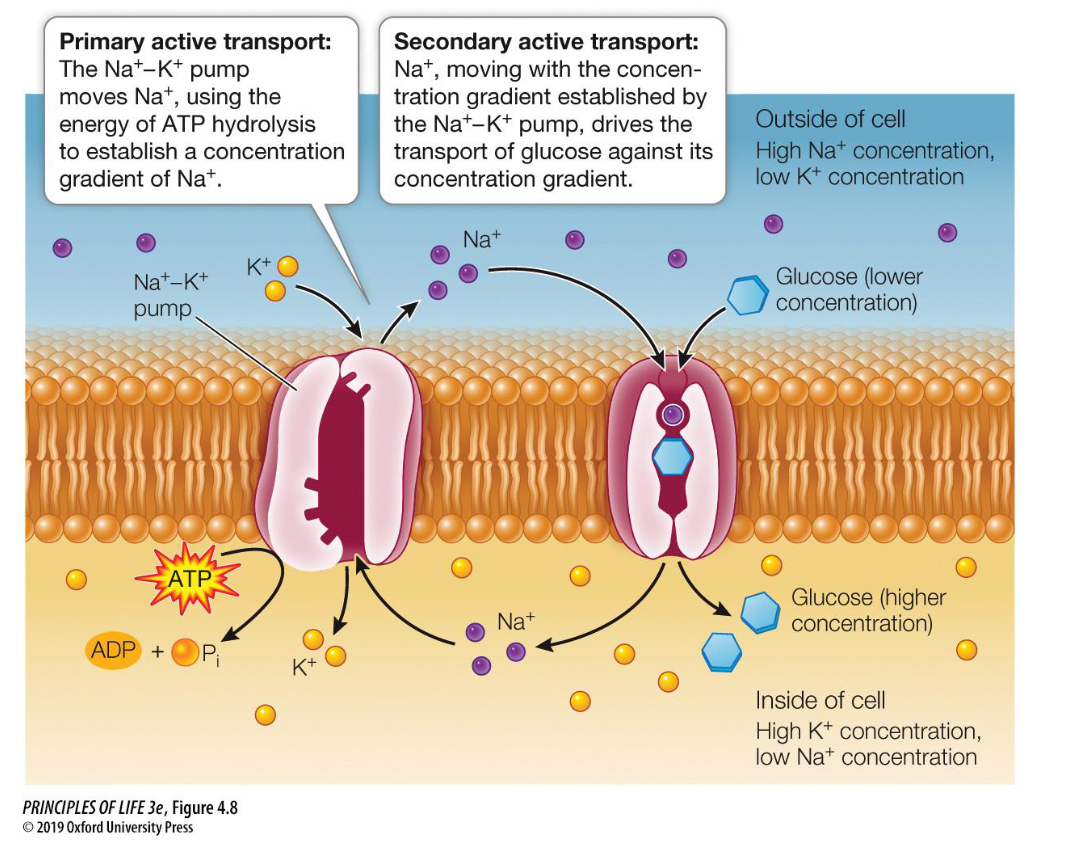
Primary Active Transport
Active transport is like swimming upstream; it requires "effort" in the form of ATP (chemical energy).
Definition: The movement of molecules or ions against their concentration gradient (from Low to High concentration) by directly using chemical energy.
Energy Source: It relies on the hydrolysis of ATP.
Key Example: The Na+/K+ Pump. It uses 1 ATP to pump 3 Na+ out and 2 K+ in.
The Na+/K+ pump is a primary active transporter (an antiport) that maintains a high concentration of Sodium (Na+) outside the cell and a high concentration of Potassium (K+) inside the cell.
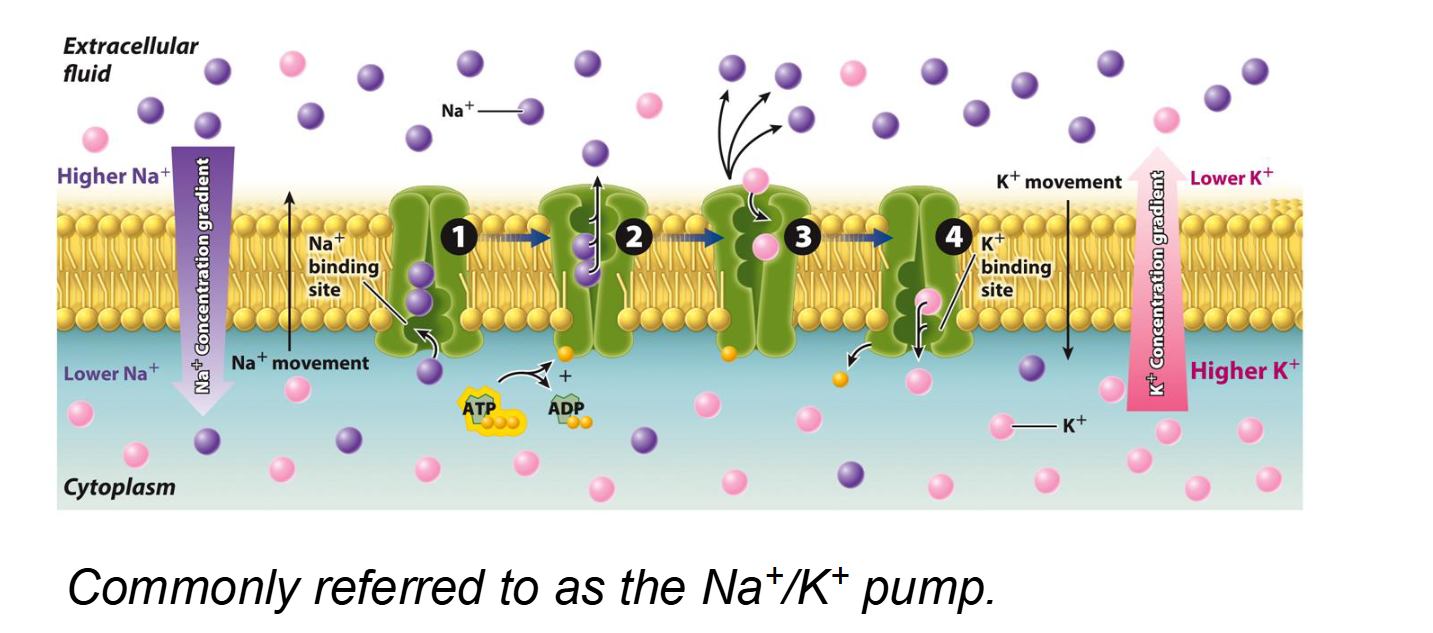
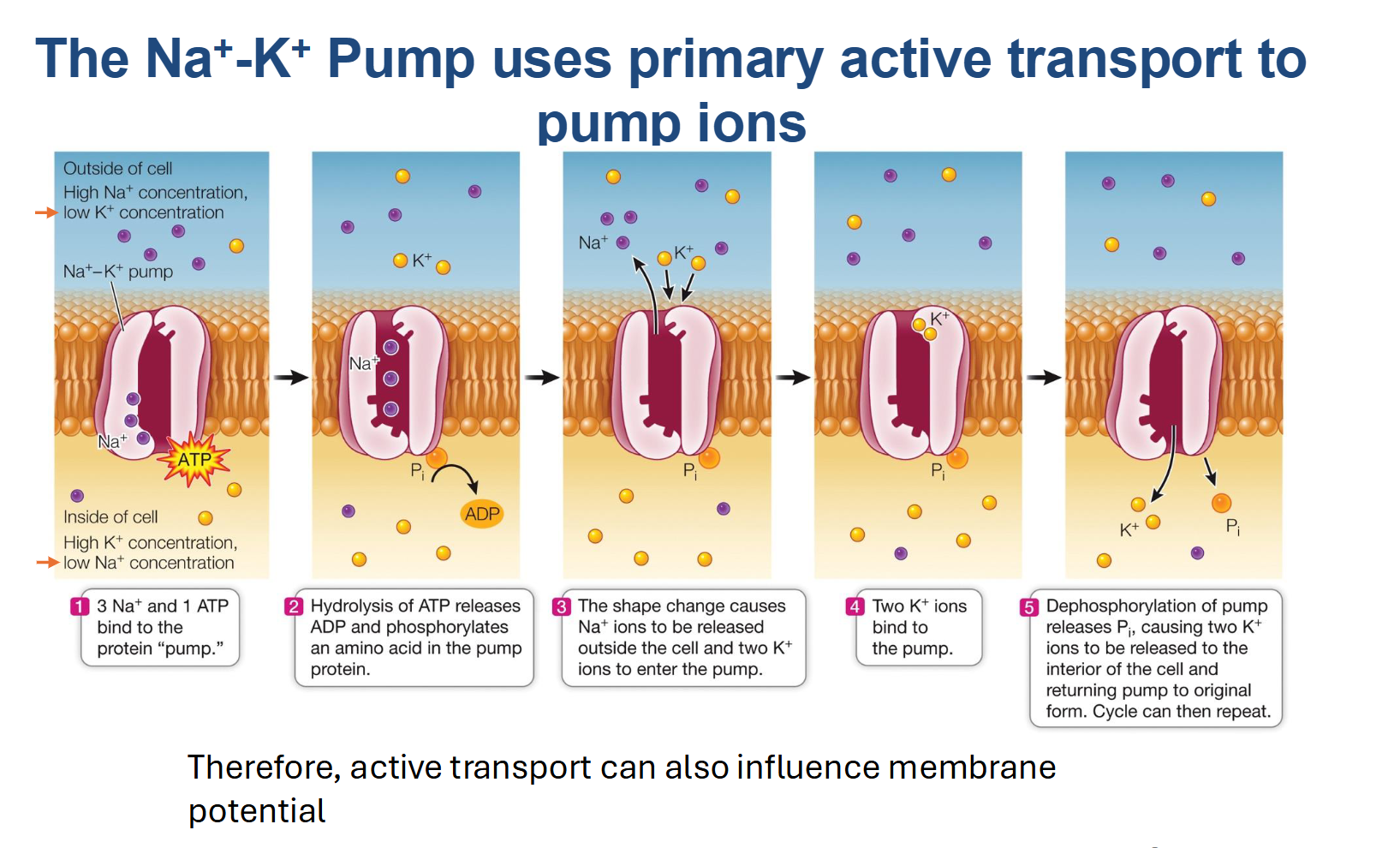
The Na+/K+ Pump
Type of Transport: Primary Active Transport (specifically an Antiport).
Energy Requirement: Requires 1 ATP molecule per cycle.
Net Movement: Pumps 3 Sodium (Na+) ions OUT against gradient and 2 Potassium (K+) ions IN against gradient
Importance: Maintains electrochemical gradients, regulates cell volume, and provides the "energy reservoir" for Secondary Active Transport (like the Na+-glucose symporter)
By actively pumping ions like Na+ out, the cell prevents itself from becoming too "salty" inside (Remember osmosis!).
Very important to maintain higher concentration of Na+ outside than inside the cell
What is Secondary Active Transport and how does it differ from Primary Active Transport?
Definition: The movement of a substance against its concentration gradient by using the energy stored in an electrochemical gradient (usually Na+) rather than using ATP directly.
The "Coupled" Mechanism: It relies on a Primary active transporter (like the Na+/K+ pump) to first create a steep concentration gradient. The "downhill" movement of one ion then powers the "uphill" movement of another molecule.
Found in intestinal and kidney cells
Key Example: The Na+-glucose symporter.
Na+ moves down its gradient into the cell.
This provides the energy to pull Glucose against its gradient into the cell simultaneously.
Distinction: * Primary: Uses ATP directly.
Secondary: Uses an existing ion gradient (indirect energy).
What is Osmosis and which protein facilitates its speed?
Definition: The passive diffusion of water across a selectively permeable membrane.
Direction: Water moves from an area of high water concentration (low solute) to an area of low water concentration (high solute).
Helper Protein: While some water can cross the bilayer slowly, Aquaporins are specialized channel proteins that allow for the rapid facilitated diffusion of water.
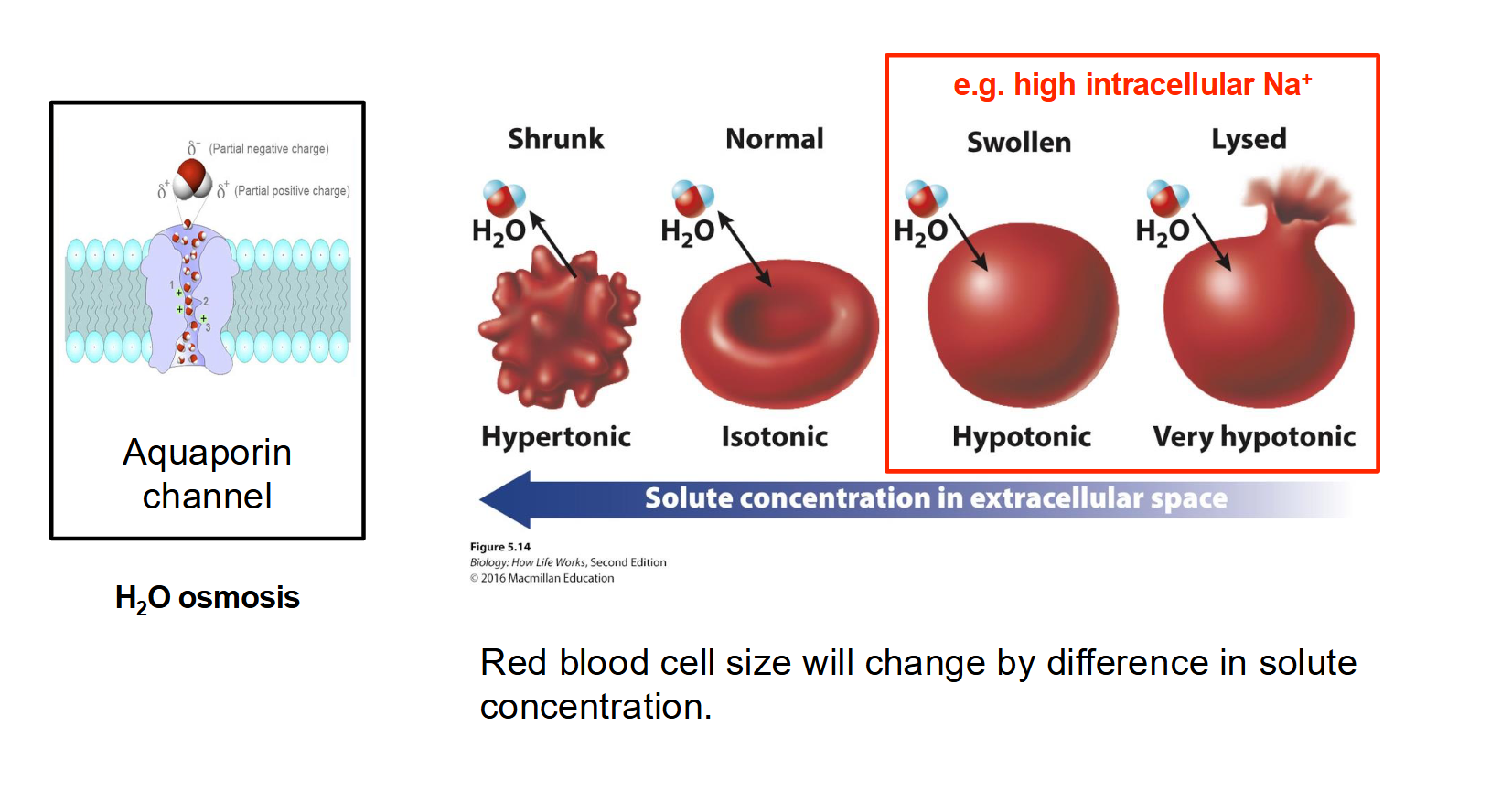
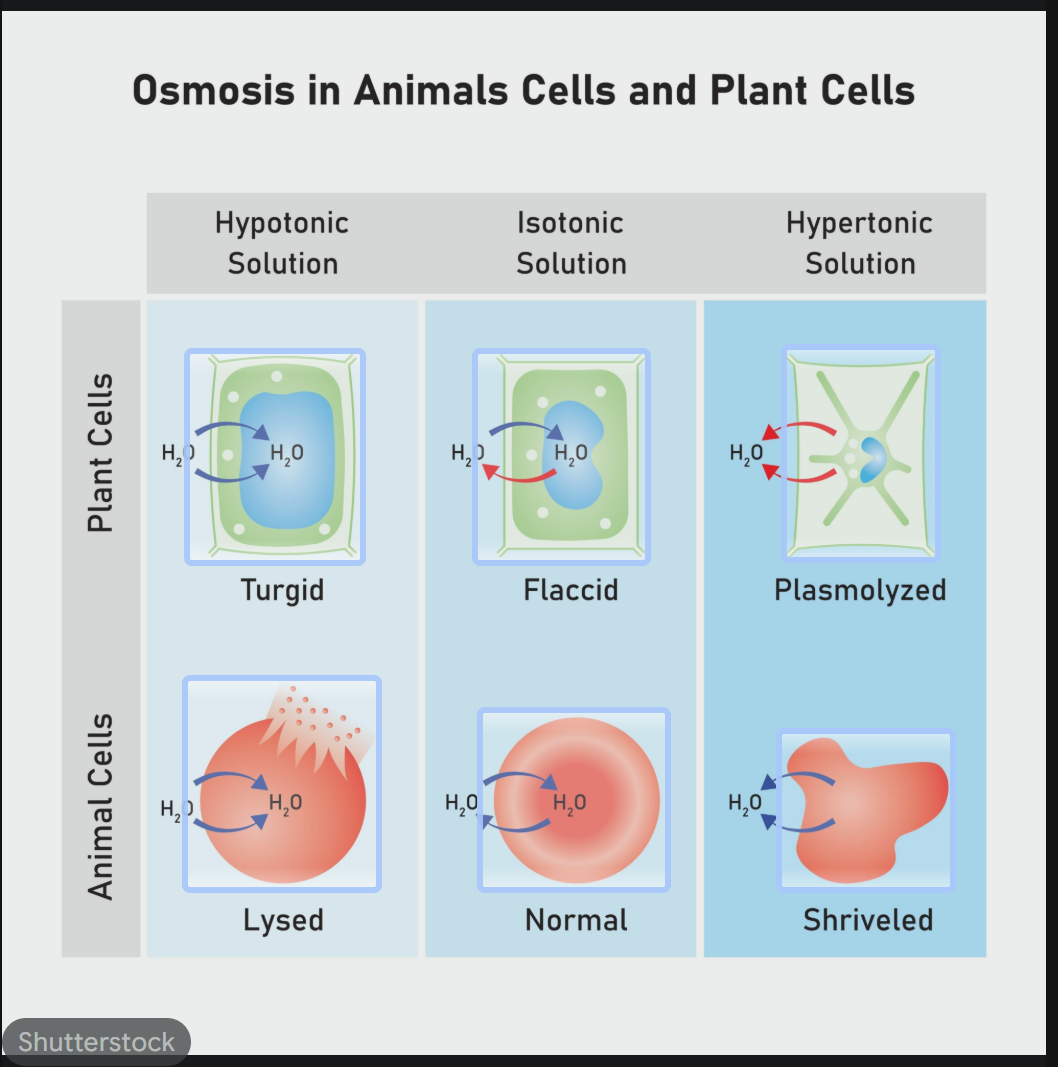
How does tonicity affect the physical state of a cell (Osmosis in action)?
Hypertonic Environment: Higher solute concentration outside the cell. Water leaves the cell, causing it to shrink/shrivel.
Isotonic Environment: Equal solute concentration inside and out. There is no net movement of water; the cell remains normal.
Hypotonic Environment: Lower solute concentration outside the cell. Water rushes into the cell, causing it to swell.
In Animal Cells: The cell may burst (Lysis).
In Plant Cells: The cell becomes firm due to Turgor Pressure (the plasma membrane pushes against the cell wall, but the wall prevents bursting).