BSC 1010 Exam 4 (Chapter 17)
1/63
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
64 Terms
From GENE to PROTEIN
genes specify proteins via transcription & translation.
transcription: dna-directed synthesis of rna
translation: rna-directed synthesis of polypetide
mutation: 1/+ nucleotides can affect protein struc & function.
DNA
inherited by org leads to specific traits by dictating synthesis of proteins
proteins →
links betw genotype & phenotype
gene expression:
2 stages: transcription & translation
In 1902, British physician Archibald Garrod…
…first suggested that genes dictate phenotypes through enzymes that catalyze specific chem rxns.
metabolic pathway:
cells synthesize & degrade molecules in a series of steps.
George Beadle & Edward Tatum →
exposed bread mold to X-rays, creating mutants: unable to survive on minimal media.
Adrian Srb & N. Horowitz →
identified 3 classes of arginine-deficient mutants- Each lacked a diff enzyme necessary for synthesizing arginine.
1 gene- 1 enzyme hypothesis:
function of a gene is to dictate production of a specific enzyme.
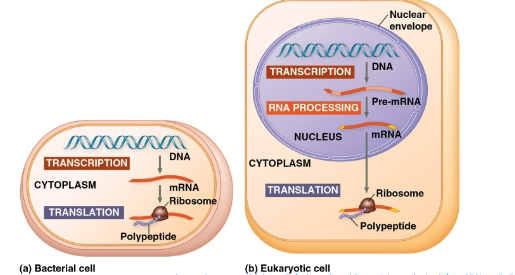
transcription:
synthesis of RNA using info in DNA- produce messenger RNA (mRNA)
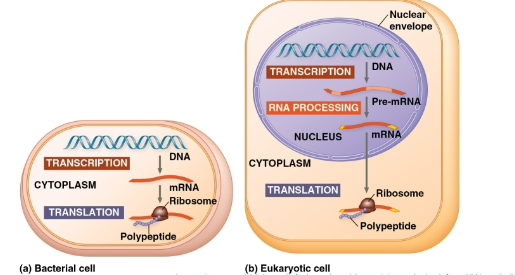
translation →
synthesis of polypeptide, using information in mRNA- Ribosomes: sites of translation.
The central dogma:
concept that cells are governed by a cell.
chain of command: DNA → mRNA → Proton
DNA sequence →
will determine aa-sequence & therefore, will determine protein function.
CODON:
every 3 nucleotides on mRNA codes for an aa
The genetic code is universal-
from bacteria to us- 4 nucleotides (A,T,C,G) code for 20 aa (the sequence of those is what make a bacteria DIFFERENT from us).
Prokaryotes: transcription..
no separation of the transcription and translation process.
Initiation (Prokaryotes)
promotor→ piece of DNA upstream that indicates where RNA polymerase should bind & start.
Elongation (Prokaryotes)
RNA polymerase → adds complimentary nucleotides (A,U,C,G) to make mRNA
Initiation (Eukaryotes: transcription..)
Promotors → much more complex than prokaryotic prom.
Eukaryotes have 3 RNA polymerases (RNA polymerase I, II, III)
transcription factors needed
(Eukaryotes: transciption..2)
Elongation
Termination (Eukaryotes: transcription..)
mRNA is released- modified before leaving nucleus for translation.
5’ Cap-
cap is added to help mRNA from being immediately degraded.
3’ Poly A Tail-
adenines added to end, helps mRNA pass through nuclear pore.
Splicing of introns-
gene has exons and introns (exons are expressed, introns are cut out.
Translation: 3 Steps
1) Initiation
2) Elongation
3) Termination
Initiation; mRNA
attaches to smaller subunit of the ribosome
Initiation; AUG
start codon- a tRNA w appropriate anti codon attaches
the larger subunit of the ribosome then comes in
Elongation
tRNAs move in w appropriate amino acid, aa chain grows using peptidyl transferase.
Termination
stop codon is reached
the amino acid chain then is processed
in eukaryotes, aa chain move into ER to be further processed.
The genetic code
20 amino acids, but there are only 4 nucleotide bases in DNA
flow of information from gene to protein →
based on a triplet code: a series of nonoverlapping, three-nucleotide words.
These words are then translated into a chain of amino acids →
forming a polypeptide
RNA synthesis-
catalyzed by RNA polymerase, which pries the DNA strands apart and joins together the RNA nucleotides
the RNA →
complementary to the DNA template strand
RNA polymerase →
does not need any primer
RNA synthesis follows same base-pairing rules as DNA…
…except that uracil substitutes for thymine.
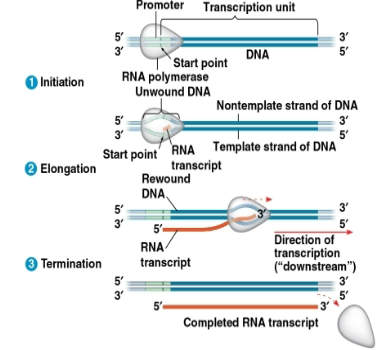
Promoter:
DNA sequence where RNA polymerase attaches
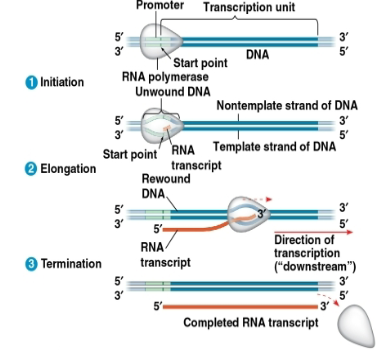
Terminator:
in bacteria, the seq signaling the end of transcription
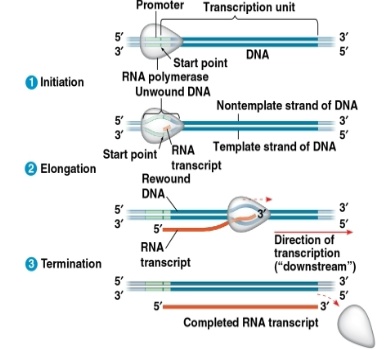
transcription unit:
the stretch of DNA that is transcribed
Promoters signal the transcription start point and…
…usually extend several dozen nucleotide pairs upstream of start point.
transcription factors →
help guide binding of RNA polymerase and the initiation of transcription.
completed assembly of transcription factors and RNA polymerase II bound to a promoter is called =
transcription initiation complex
A promoter called a TATA box …
…is crucial in forming initiation complex in eukaryotes.
As RNA polymerase moves along the DNA…
…it untwists double helix, 10-20 nucleotides at a time.
nucleotides-
added to 3’ end of the growing RNA molec.
Transcription progresses at a rate of…
…40 nucleotides per second in eukaryotes
a gene can be transcribed simultaneously by…
…serval RNA polymerases.
Termination of Transcription
mechanisms of termination- diff. in bacteria & eukaryotes
In bacteria →
polymerase stops transcription at the end of terminator and mRNA can be translated w/o further modification.
In eukaryotes →
RNA polymerase II transcribes polyadenylation signal sequence; RNA transcript is released 10-35 nucleotides past this polyadenylation sequence.
Enzymes in eukaryotic nucleus modify pre-mRNA (RNA processing) before…
…the genetic messages are dispatched to cytoplasm
During RNA processing…
…both ends of the primary transcript are altered.
in most cases…
…certain interior sections of the molecule are cut out and remaining parts spliced together.
5’ end receives a…
…modified nucleotide 5’ cap
3’ end gets a…
…poly-A tail
these modifications share several functions
facilitate export of mRNA to cytoplasm
protect mRNA from hydrolytic enzymes
help ribosomes attach to 5’ end
Most eukaryotic genes and their RNA transcripts have…
…long noncoding stretches of nucleotides that lie betw coding regions
These are removed through RNA splicing…
noncoding segments in a gene are called intervening sequences[introns]
The other regions are called exons because…
…they are eventually expressed, usually translated into aa seq’s
removal of introns is accomplished by…
…spliceosomes
spliceosomes consist of a variety of proteins and…
…several small RNAs that recognize the splice sites
the RNAs of the spliceosome also…
…catalyze the splicing rxn
A cell translates an mRNA message into protein w help of…
…transfer RNA (tRNA)