BIOL3060 #2 Foundations of Genetics: Structure, Replication
1/49
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
50 Terms
Why is it called DNA (Deoxyribonucleic Acid)?
Deoxy → Missing an oxygen atom at the 2′ carbon of the • deoxy- "without oxygen."
Ribo → The sugar is based on ribose.
Nucleic → Found in the cell nucleus
Acid → Because of the negatively charged phosphate groups, DNA is an acid
Why is it called RNA (Ribonucleic acid)?
Ribo- → comes from the sugar ribose, which is part of each RNA nucleotide. •
Nucleic → because it's a nucleic acid (like DNA)
Acid → due to the negatively charged phosphate groups, which make the molecule acidic.
Nucleotides
Repeating unit of DNA or RNA made up of a sugar, a phosphate, and a nitrogenous base.
Nitrogenous bases in DNA
Adenine, Thymine, Cytosine, Guanine
Nitrogenous bases in RNA
Adenine, Uracil, Cytosine, Guanine
purines
Adenine and Guanine (2 rings)
Pyrimidines
cytosine, thymine, uracil (single ring)
Nucleotides link via
phosphodiester bonds
how do phosphodiester bonds form?
Condensation reaction: these bonds form through a dehydration synthesis (water is removed)
DNA strand direction
5' to 3'
Nucleotides are always added to the ___ end of a growing chain.
3′
5′ prime end
phosphate attached to the 5th carbon of the ribose
3′ prime end
free hydroxyl group (-OH) on the 3rd carbon of the sugar.
Where do phosphodiester bonds form?
5' phosphate group of one nucleotide and 3' OH group of another
DeoxyRibonucleoside triphosphates (dNTPs)
A monomer used by DNA polymerase to polymerize DNA. Consists of the sugar deoxyribose, a base (A, T, G, or C), and three phosphate groups.
DNA vs RNA replication
DNA replicates in the nucleus, RNA replicates in the cytoplasm
Prokaryotic vs eukaryotic replication
SIMILARITIES
- both bi-directional processes
- both require primers to start the process
- both have leading and lagging
- DNA polymerase enzymes work from the direction of '5-'3 so new nucleotides are added to the '3 end of a primer.
DIFFERENCES
prokaryotes have only one site of replication
leading strand
The new continuous complementary DNA strand synthesized along the template strand in the mandatory 5' to 3' direction towards the fork
lagging strand
A discontinuously synthesized DNA strand that elongates by means of Okazaki fragments, each synthesized in a 5' to 3' direction away from the replication fork.
parent strand
original strand of DNA
daughter strand
the newly made stand in DNA replication
DNA polymerase III
adding bases to the new DNA chain; proofreading the chain for mistakes
Helicase
An enzyme that untwists the double helix of DNA at the replication forks.
Topoisomerase
releases torsional strains due to unwinding of DNA
dna ligase
enzyme that chemically links DNA fragments together
when does replication happen?
S phase Interphase
Primease
creates rna primer so polymerase knows where to start
DNA polymerase I
removes the RNA primer and replaces it with DNA
A-T have how many bonds
2 hydrogen bonds
C-G have how many bonds
3 hydrogen bonds
Denaturation
- Breaks hydrogen bonds
- Heat provides energy to overcome the hydrogen bonds between base pairs
- creates single stranded dna
Gregor Mendel
- Father of genetics
- Principles of Heredity
- 1866
Rosalind Franklin (1920-1958)
- British chemist and X-ray crystallographer
- She took Photo 51, an X-ray diffraction image of DNA that clearly showed the helical structure. (1953)
- She died young (age 37) from ovarian cancer
- Franklin's work was critical to discovering the double helix
James Watson & Francis Crick
The scientists credited with building the first correct model of the structure of DNA
- noble prize awarded in 1962
Human Genome Project
An international collaborative effort to map and sequence the DNA of the entire human genome.
1990 - 2003
Gene Expression
the process by which the information encoded in a gene is turned into a function
genome
complete set of genetic instructions for any organism
Even though every cell has the same DNA, only a _____ of genes is active in each cell type.
subset, This allows cells to perform specialized functions for the organism.
Human Genome
20,000 protein-coding genes.
A typical human cell expresses about 2,000 of those genes (~10%).
housekeeping genes
genes expressed in almost all cells
(i.e genes involved in glycolysis, because all cells need energy from glucose)
Locus
a position on a chromosome where a specific gene is located
beta globin locus
a stretch of DNA on chromosome 11 that contains the genes for several beta-like globin proteins, which are part of hemoglobin
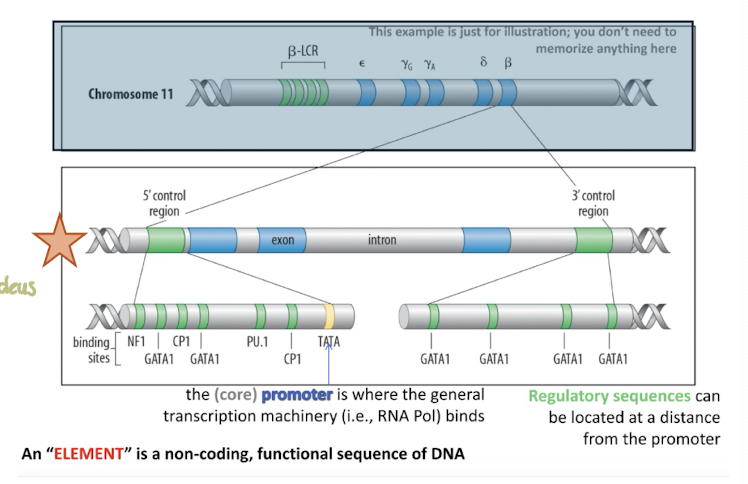

Beta Globin Locus: Locus Control Region (LCR)
powerful regulatory element located far upstream of the gene cluster
- these sequences determine whether the protein-coding parts of the gene are turned on or off
Beta Globin Locus: Exons
Coding regions that make the protein
Beta Globin Locus: Intron
Non-coding sequences between exons
Beta Globin Locus: 3′ control region
Downstream regulatory sequences that can also influence gene expression.
- increase or decrease the production of that protein
Beta Globin Locus: 5′ control region
At the Upstream "start" end of the gene, contains regulatory sequences.
Beta Globin Locus: Core promoter
TATA box, where the general transcription machinery (like RNA polymerase) binds to start transcription
Beta Globin Locus: transcription factors
Collection of proteins that mediate the binding of RNA polymerase and the initiation of transcription. proteins that bind DNA to turn gene on or off
Beta Globin Locus: Regulatory promoter
DNA sequence located immediately upstream of the eukaryotic core promoter; contains consensus sequences to which transcription factors bind.