BICH 431 TAMU Park Exam Practice Questions
1/49
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
50 Terms
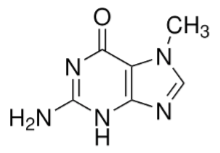
What is this structure?
a) 1-methylguanine
b) 3-methylguanine
c) 5-methyl adenosine
d) 7-methylguanine
e) 9-methylguanine
d) 7-methylguanine
What is the approximate length of the DNA fragment in a typical BAC used for human genomics?
Hint 1 μm = 1000 nm 1 mm = 1000 μ
a) 1 μM
b) 10 μM
c) 100 μm
d) 1 mmm
c) 100 μm 0.34 nm X 300,000 = 100,000 nm = 100 μm
How many possible hydrogen bonding patterns would be possible for a protein binding to the minor groove
of a two base pair region of the minor groove of B DNA?
a) 4
b) 8
c) 16
d) 32
e) 64
a) 4
Which of the following promotes a B to Z transition in the DNA of living cells?
a) Methylation of the 5-position of G
b) Overwinding
c) Osmotic stress
d) None of these
a) Methylation of the 5-position of G
Nucleic acid structures
a) The double stranded regions of RNA typically have a C 2’ endo sugar pucker
b) A-DNA contains the same Watson-Crick hydrogen bonds as B DNA, but they are more tilted relative to the
helical axis
c) “Hoogsteen” pairing can only occur in RNA because RNA is more flexible that DNA
d) The glycosidic bonds in double strand DNA are always in syn vs. rather than anti configuration
b) A-DNA contains the same Watson-Crick hydrogen bonds as B DNA, but they are more tilted relative to the
helical axis
You are doing a Southern blot with human DNA. The procedure you are following was designed to only
detect the gene most homologous to the probe. However, you also want to detect paralogous genes. Which of the
following should you increase?
a) Salt concentrations
b) Urea concentration
c) Temperature
d) pH
a) Salt concentrations
The approximate size the genome of E. coli is _____bp and human ___ bp.
a) 4 x 103 3 x 106
b) 4 x 104 3 x 107
c) 4 x 105 3 x 108
d) 4 x 10^6 3 x 10^9
d) 4 x 10^6 3 x 10^9
If a 2500 bp circular plasmid DNA in which one phosphodiester bond has been cleaved is incubated with DNA
gyrase and ATP this will result in
a. overwound supercoils
b. underwound supercoils
c. circular DNA with a writhing number of zero.
d. Decreased mobility on a nondenaturing agarose gel
c. circular DNA with a writhing number of zero.
Gibson Assembly
a) Requires the non-templated C residues that reverse transcriptase adds the end of all transcripts and special
oligonucleotides which terminate with several G residues.
b) Requires at least one restriction enzyme the forms “sticky ends”
c) Requires reverse transcriptase
d) Requires DNA ligase
d) Requires DNA ligase
Which of the following would be best general method to determine the position of genes in a physical map?
a. RNA-Seq
b. Western blotting
c. ChIP-Seq
d. quantitative PCR
a. RNA-Seq
You have optimized a real time (quantitative) PCR assay so that amplification be more than 99% efficient for
each cycle. In samples from the brain, transcripts from your favorite gene reached the critical threshold after 20
cycles. In samples from liver, transcripts of your favorite gene reached the critical threshold after 23 cycles. A
control “housekeeping” gene reached the threshold in 27 cycles in both samples. What is the ratio of
transcripts for your favorite gene in brain/liver.
a) 3
b) 4
c) 8
d) 1/3
e) 1/4
f) 1/8
c) 8
Nanopore sequencing
a) Uses fluorescent nucleotides, but does not use dideoxy-NTP
b) Typically uses ion channel proteins fused to DNA gyrase
c) Uses DNA unwinding proteins attached to precisely sized holes which are drilled in semi-conductor chips
using lasers
d) Requires ATP
d) Requires ATP
Polyacrylamide gels containing SDS (sodium dodecyl sulfate) are typically used in
a) Southern blots
b) Norther blots
c) Western blots
d) Electromobility shift assays
c) Western blots
Which of the following is NOT TRUE about nucleosome core particles?
a) The amino terminal tails can be modified by kinases, methylases and acetylases.
b) They contain two copies each of H1, HA2A, H2B, H3 and HA4
c) They contain 146-147 bp of DNA wrapped around the histone octamer.
d) They can contain alternate histones such as H2AZ and H3.3.
b) They contain two copies each of H1, HA2A, H2B, H3 and HA4
Proteins that bind to methylated lysine residues in the histone tail often contain:
a) bromodomains.
b) chromodomains.
c) histone-fold motifs
d) binding sites for transcription factors
b) chromodomains.
Chromosomes
a) Two or more short AT rich regions with 10 bp spacing will prevent nucleosome binding.
b) SWI/SNF can move nucleosomes from one position to another using ATP
c) Wrapping DNA around nucleosomes causes positive supercoils
d) To avoid problems with mis-segregation during mitosis and meiosis, all chromosomes only bind one
microtubule (two for the pair of chromosomes to be separated)
b) SWI/SNF can move nucleosomes from one position to another using ATP
Topologically associated domains (TADs)
a) Are a primitive type of nucleosome found only in bacteria
b) Always contain the long noncoding RNA encoded by the XIST gene.
c) Active genes in TADs are often present in phase-separated molecular condensates
d) Only contain active genes
c) Active genes in TADs are often present in phase-separated molecular condensates
Epigenetic marks
a) Require Hoogsteen base pairing
b) All remain with one parental DNA strands, which is why terminally differentiated cells usually do not
divide
c) Can be created and erased by histone acetylases and deacetylases
d) Require SW/SNF or ISWI
c) Can be created and erased by histone acetylases and deacetylases
Dosage compensation in mammals
a. One X-chromosome in females is permanently inactivated in all cells
b. The presence of Barr bodies is an unambiguous way to determine biological sex since they are present in
all females, but absent in all males
c. All genes on one of the X chromosomes in females are inactivated by the action of a long noncoding RNA
encoded by the XIST locus.
d. None of these
d. None of these
Approximately how long is a typical RNA primer used for DNA replication
a) 5
b) 7
c) 11
d) 46 or 47
e) 147
c) 11
DNA replication
a. SeqA inhibits DnaA from reinitiating by blocking Dam methylase sites.
b. All DNA and RNA polymerases require a primer with a free 3’ hydroxyl.
c. Activation of oriC requires simultaneous binding of Seq A and Dam methylase.
d. HU blocks binding of DnaA-ATP to oriC until HU is fully methylated by the DNA adenine methylase
a. SeqA inhibits DnaA from reinitiating by blocking Dam methylase sites.
Which answer is the correct order of events to initiate DNA replication at E. coli oriC?
1. Binding of DNA Pol III holoenzyme
2. Binding of DnaA
3. Binding of helicase
4. Unwinding DNA at the origin
5. Binding of primase
6. Binding of SeqA
a) 1, 2, 3, 6, 4, 5
b) 2, 4, 3, 5, 1, 6
c) 4, 3, 2, 5, 1, 6
d) 3, 4, 6, 2, 1, 5
e) 2, 6, 4, 3, 1, 5
b) 2, 4, 3, 5, 1, 6
DNA polymerases
a) E. coli pol III has a 5’ to 3’ exonuclease activity
b) E. coli pol III is more abundant than pol I
c) DNA must be released from pol I for mismatched bases to be removed,
d) The Klenow fragment of E. coli DNA pol I lacks the 5’ to 3’ exonuclease
d) The Klenow fragment of E. coli DNA pol I lacks the 5’ to 3’ exonuclease
What is responsible for increasing processivity of DNA Pol III?
a. Presence of the dNTP loader complex.
b. The hand-shaped structure of the polymerase core which is unique to Pol III.
c. The sliding clamp that allows it to remain associated with the DNA template.
d. The “thumb” structure of the core that holds the DNA template tightly.
c. The sliding clamp that allows it to remain associated with the DNA template.
Ter sites
a) help load helicase onto the oriC after it is melted by DnaA
b) are only present in linear chromosomes
c) are involved in blocking movement of replication forks
d) are only active when fully methylated.
c) are involved in blocking movement of replication forks
DNA damage/repair
a) Exposure to X-rays often causes formation of 50-100 thymidine dimers per second per cell
b) 5-bromouracil increases the mutation rate in E. coli by inhibiting DNA scanning by MutS
c) Xeroderma Pigmentosa in humans is usually due to a defect in the photolyase DNA repair system
d) None of these
d) None of these
Introns
a) Typically have an AG on their 5’ end and a GU on the 3’ end
b) U1 binds both the 5’ end of the intron and the lariat site
c) U5 binds to the ends of both exon 1 and exon2
d) U4 binds to both the 5’ end of the intron and U2
c) U5 binds to the ends of both exon 1 and exon2
Recombination
a) Holliday junctions are usually resolved by the action of AP endonuclease
b) Rec BCD makes double strand cuts near chi sequences that simulate homologous recombination
c) RecA can only catalyze assimilation of a strand of DNA into a homologous fragment of linear duplex DNA
d) none of these
d) none of these
DNA repair
a) Translesion DNA synthases always cause mistakes, but prevents lethality
b) RuvC nuclease is always required to repair double strand breaks in DNA
c) Spo II is only attached to the ends of double strand breaks during meiosis
d) None of these
c) Spo II is only attached to the ends of double strand breaks during meiosis
DNA repair
a) Fork regression occurs most often after double strand breaks in the DNA
b) Fork regression usually occurs before DNA damage is repaired
c) Fork regression requires Spo II
d) If E. coli DNA pol III encounters an AP or abasic site, the replication fork will collapse
b) Fork regression usually occurs before DNA damage is repaired
DNA repair and Recombination
a) Recombination between sister chromatids can cause the progeny of heterozygous cells to become homozygous
b) RuvC nuclease is only required for recombination between non-sister chromatids
c) Ku70 and Artimis are required for recombination during meiosis
d) None of these
d) None of these
What bacterial DNA template strand would produce a mRNA having the sequence 5’ AUGCUACCGUUA 3’.
Note that all answers are arranged 5’ to 3’
a) 5’ ATGCTACCGTTA
b) 5’ ATTGCCATCGTA
c) 5’ TAACGGTAGCAT
d) 5’ TACGATGGCAAT
c) 5’ TAACGGTAGCAT
RNA processing
a) Approximately half of human genes contain introns
b) Addition of the 7mG cap occurs after synthesis of 8-9 bases of mRNA
c) Introns and exons can be included or removed from precursor mRNA, but their order is never changed
d) None of these
c) Introns and exons can be included or removed from precursor mRNA, but their order is never changed
The correct order of transcriptional events occurring after RNAP binds to the promoter is
a) Start of RNA synthesis, open complex formation, closed complex formation, promoter clearance
b) Closed complex formation, open complex formation, promoter clearance, start of RNA synthesis
c) Closed complex formation, open complex formation, start of RNA synthesis, promoter clearance
d) Open complex formation, closed complex formation, start of RNA synthesis, promoter clearance
c) Closed complex formation, open complex formation, start of RNA synthesis, promoter clearance
NAP and sigma factors
a) RNAP can produce mRNA from constitutive “housekeeping” genes without a sigma factor but requires them
for inducible gene expression and to deal with special circumstances such as heat shock
b) Most sigma factors besides sigma 70 recognize promoters which contain -10 sequences enriched in G & C
c) Sigma factors contain a separate exonuclease site for proofreading.
d) Sigma factor 70 remains associated with E. coli RNAP during abortive transcription.
d) Sigma factor 70 remains associated with E. coli RNAP during abortive transcription.
Transcription in bacteria
a) Closed complex formation is irreversible and requires interaction of sigma with the -10 and -35 sequences.
b) During closed complex formation, the RNA exit channel of RNA polymerase is blocked by NusA
c) RNA synthesis is less processive during initiation than during elongation
d) E. coli RNAP typically produces 500-1000 nucleotides of mRNA per second
c) RNA synthesis is less processive during initiation than during elongation
The average RNAP error rate is approximately
a) 10-2 - 10-3
b) 10^-4 - 10^-5
c) 10-6 - 10-7
d) 10-8 - 10-9
b) 10^-4 - 10^-5
Proofreading reactions of bacterial RNAP
a) Hydrolytic editing produces rNTP
b) Pyrophosphorolysis requires pyrophosphatase.
c) Pyrophosphorolysis removes bases from the 5’ end of mRNA
d) E. coli RNAP has an intrinsic nuclease activity that uses water as nucleophile.
d) E. coli RNAP has an intrinsic nuclease activity that uses water as nucleophile.
RNA polymerase
a) Alpha amanitin preferentially blocks RNA pol II
b) Actinomycin D preferentially blocks bacterial, but not eukaryotic RNAP
c) RNA pol I produces all of the structural RNAs in ribosomes, but not tRNA or mRNA
d) DNA in front of RNA pol which is producing mRNA becomes underwound
a) Alpha amanitin preferentially blocks RNA pol II
Polymerases
a) The error rate of reverse transcriptase is approximately 1 in a million
b) All RNA polymerases consist of 4 subunits which are homologous from bacteria to animals
c) MicroRNA is produced by RNA pol II
d) RNA pol 1 is the only RNA polymerase in eukaryotic cells that does not require TATA binding protein for
activation
c) MicroRNA is produced by RNA pol II
How long is the RNA/DNA hybrid in E. coli RNAP to which NusA is bound
a) It does not contain any RNA/DNA hybrid
b) 4-5
c) 8-9
d) 12-15
c) 8-9
Termination
a) Removal of all Rut sites would block rho independent termination, but not rho dependent termination.
b) Rho independent terminators contain a hairpin/stem-loop followed by a string of 3-8 G residues.
c) Animals rely on the 3’ to 5’ exonuclease activity of Xrm2 to terminate RNA pol II transcription.
d) none of these
d) none of these
Promoters and polymerases
a) TFIID binding, RNA Pol II binding, TFIIH binding, promoter clearance, phosphorylation of CTD Ser2
b) TFIID binding, RNA Pol II binding, phosphorylation of CTD Ser2, promoter clearance,
c) RNA Pol II binding, TFIID binding, TFIIH binding, promoter clearance, phosphorylation of CTD Ser2
d) TFIIH binding, RNA Pol II binding, phosphorylation of CTD Ser2, promoter clearance
a) TFIID binding, RNA Pol II binding, TFIIH binding, promoter clearance, phosphorylation of CTD Ser2
Which of the following sequences is NOT generally found in the core promoter of RNA Polymerase II?
A. BRE
B. DPE
C. Enhancer
D. Inr
C. Enhancer
Promoters and polymerases
A. Pol I makes tRNA
B. Pol III promoter elements are typically downstream of the start site.
C. TBP creates an 80-degree bend in DNA, but only in promoters that contain a TATA.
D. TPB is often found as a component of TFIIE
B. Pol III promoter elements are typically downstream of the start site.
CTD of Pol II
A. Is attached to pol II by TFIID during formation of the pre-initiation complex
B. is first phosphorylated during the transition between open complex formation and promoter clearance
C. Phosphorylation of Ser5 increases as RNAP transcribes genes and helps recruit termination factors
D. is cleaved into 7 amino acid segments during termination
B. is first phosphorylated during the transition between open complex formation and promoter clearance
TBP
a) Directly binds the TATA box in the promoters for the precursor for rRNA
b) Bends promoter DNA after TFIIF and RNA pol II bind
c) Directly binds to the downstream promoter elements found in most RNA pol II promoters
d) Binds before TFIIE
d) Binds before TFIIE
Poly A
a) is added to the mRNA of all organisms except for a few extremophiles
b) is added to the 3’ end of mRNA with the aid of Xrn2 after RNA pol II terminates transcription.
c) is present on all human mRNAs
d) is added by a special form of RNA polymerase that does not require a template.
d) is added by a special form of RNA polymerase that does not require a template.
What is the structure of a typical mRNA cap?
a) 7 methyl-GMP in a 5’-3’ linkage to the first base of the mRNA
b) 7 methyl-GMP in a 5’-5’ linkage to the first base of the mRNA
c) 7 methyl-GTP in a 5’-3’ linkage to the first base of the mRNA
d) 7 methyl-GTP in a 5’-5’ linkage to the first base of the mRNA
d) 7 methyl-GTP in a 5’-5’ linkage to the first base of the mRNA
The nucleophile for the first step of spliceosome pre-mRNA splicing is
a) hydroxyl of the branch point adenosine
b) hydroxyl of a guanine nucleotide non-covalently bound to SNURP U5
c) hydroxyl of the 5’ splice site
d) hydroxyl of the 3’ splice site
a) hydroxyl of the branch point adenosine