DNA Replication
1/24
Earn XP
Description and Tags
phase 1 of protein synthesis
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
25 Terms
where does DNA replication occur
nucleus
what is the main goal of this process
to replicate genetic information before the splitting of the cytoplasm
stage 1
separating the DNA strand
what enzymes are used in stage 1
DNA helicase, DNA gyrase and single stranded binding proteins
DNA helicase
unwinds and splits double helix by snipping H bonds between nitrogenous base pairs
what is formed after DNA helicase splits nitrogenous bases apart
replication bubble
at the end of each replication bubble is
a replication fork (y-shaped region where DNA is actively opening)
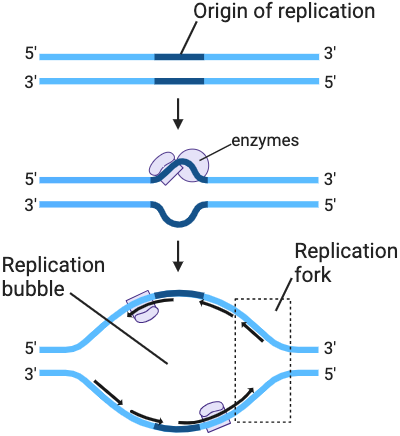
DNA gyrase
relives tension caused during unwinding
single stranded binding proteins (SSB’s)
keep the 2 strands separated by binding to the exposed DNA sites to block H-bonding from re-occuring
stage 2
building the complementary strands
what enzymes are used in stage 2
primase, RNA primer and DNA polymerase III
primase
synthesizes RNA primer and lays it down on different sites of the single strands of DNA
RNA primer
acts as a marker that tells DNA polymerase III where to bind (must be activated and layed down by primase in order to work)
DNA polymerase III
binds to the RNA primer sites and lays down nucleotides along the daughter stands in the 5’— 3’ direction, using the template strand as a guide
what direction is the template strand read vs the direction in which DNA polymerase III builds
template strand reads 3’—5
DNA polymerase III builds in 5’ — 3’
template strand vs daughter strand
template strand runs antiparallel and reads in the 3’ — 5’
daughter strand is the new DNA being formed and built in the 5’—3’
types of daughter strands
leading and lagging
leading strand
DNA polymerase III continuously builds strand towards the replication fork
lagging strand
DNA polymerase III builds in okazaki fragments (short snippets of DNA) away from the replication fork
stage 3
proof reading
what enzymes are used in stage 3
DNA polymerase I and DNA ligase
DNA polymerase I
removes RNA primers and proof reads okazaki fragments
DNA ligase
joins the okazaki fragments together
quality control of new DNA strands
DNA polymerase I and III can also act as exonucleases
exonucleases/exonucleic enzymes
act as molecular scissors that cut and remove any incorrect nucleotide pairs and replaces it with the correct complementary base pair