Cell S&F Lecture 34
1/31
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
32 Terms
In transcriptional regulation, what is the primary difference between cis-acting and trans-acting elements?
Cis-acting: DNA segments (physical sequences on the same molecule of DNA).
Trans-acting: Protein factors (molecules that bind to the DNA, often coming from elsewhere in the cell).
List the four cis-acting DNA segments involved in transcriptional regulation
1. Promoter
2. Enhancer
3. Silencer
4. Control elements
List the four trans-acting protein factors involved in transcriptional regulation.
1. General transcription factor
2. Activator
3. Repressor
4. Specific (Regulatory) transcription factor
True or False: Brain cells and liver cells contain different cis-acting (DNA) elements because they perform different functions.
False. All cells have the full genome and contain the same cis-acting elements. The difference lies in which elements are utilized in specific tissues.
Distinguish between the roles of General Transcription Factors and RNA Polymerase
General Transcription Factors: Assist the polymerase in getting started.
RNA Polymerase: The "executioner" that performs the actual polymerization (building the RNA strand).
What is the primary role of Specific Transcription Factors in tissue-specific activation?
They specify which genes to transcribe in a given cell. While general factors are broadly active, specific factors determine the unique "identity" of the cell's output.
According to the Combinatorial Model, if both liver and brain cells have the same Albumin gene/DNA, why is it transcribed at a high level in the liver but a low level in the brain?
Because of the regulatory transcription factors available in each cell.
The Liver has the specific combination of activators (red/orange/yellow shapes) that bind to the control elements.
The Brain lacks that specific combination, so even though the DNA is there, the gene isn't "turned on" effectively.
Based on the Albumin gene diagram, which components of the transcription machinery remain constant across different cell types, and which are variable
Constant: The DNA sequence (Control elements/Promoter), General Transcription Factors, and RNA Polymerase.
Variable: The specific Regulatory Transcription Factors (the shapes/colors available in the cytoplasm/nucleus).
How do you artificially (ectopically) express a brain-specific gene in liver
1. Deliver a copy of the brain-specific gene into liver cells
2. Deliver the liver-specific promoter and enhancer into brain cells
3. Deliver liver-specific transcription factors into brain cells
4. Link the coding sequence of the brain-specific gene to the promoter and enhancer of a liver-specific gene, and then deliver this hybrid gene into the liver cells
Response elements:
binding sites on DNA for regulatory transcriptional factors
Specialized control elements that respond to a signal
Same response element found in the control regions of all genes responding to the same signal
Allow coordinated expression of a group of genes scattered across the genome
What are nuclear hormone receptors (like the Glucocorticoid Receptor), and how are they activated?
They are ligand-activated transcription factors. They remain inactive in the cytoplasm until a specific signaling molecule (ligand), such as cortisol, binds to them.
In the glucocorticoid pathway, what is the function of Hsp (heat shock) proteins?
They act as molecular chaperones. They bind to the receptor (GR) in the cytoplasm to keep it in an inactive state until the hormone arrives to displace them
Describe the 4-step sequence of Glucocorticoid Receptor (GR) activation
1. Binding: Steroid hormone (cortisol) enters the cell and binds to the Receptor-Hsp complex.
2. Release: Hsp proteins are released, activating the receptor.
3. Translocation: The hormone-receptor complex moves from the cytoplasm into the nucleus.
4. Transcription: The complex binds to the Glucocorticoid Response Element (GRE) on the DNA to activate transcription.
DNA microarray
a chip with DNA deposited in a fixed format
– Numerous spots of known DNA sequences
– Each DNA has a known location
– DNA can hybridize with labeled “probe”
Posttranscriptional Regulation Of Gene Expression
RNA splicing
Translational control
Translation efficiency
mRNA stability (half-life)
miRNA and small Interfering RNA
What is Ferritin and what does it do
Iron Storage protein
Shield excessive iron, prevent it from damaging other molecules
Release iron when needed
Synthesis enhanced by high levels of irons in the cells
Transferrin
Iron delivery protein
Transferrin receptor
PM receptor of transferrin/iron
Describe the cellular response to Low Iron concentration regarding Ferritin and Transferrin receptor mRNA.
The Iron Response Protein (IRP) binds to the Iron Response Element (IRE) on both mRNAs:
Ferritin: IRP binds to the 5' end, blocking translation (no need to store iron if it’s scarce).
Transferrin Receptor: IRP binds to the 3' end, stabilizing the mRNA (promotes translation to bring more iron into the cell).
Describe the cellular response to High Iron concentration regarding Ferritin and Transferrin receptor mRNA.
Iron binds directly to the IRP, causing it to detach from the mRNA:
Ferritin: Translation is activated (the cell can now safely store the excess iron).
Transferrin Receptor: The mRNA is degraded (the cell stops bringing in more iron to avoid toxicity).
RNA interference (RNAi):
the phenomenon of RNA-mediated inhibition of gene expression
siRNA:
small interfering RNA
Dicer:
an endonuclease
RISC:
RNA-induced silencing complex
Ubiquitin:
small proteins that modifies other proteins
E1, E2, E3:
enzymes involved in ubiquitylation of proteins
Proteosome:
Protease complex that can recognize ubiquitin
List the 5 levels of gene expression regulation summarized
1. Genome (e.g., DNA accessibility)
2. Transcription (e.g., Enhancers/Promoters)
3. RNA processing (e.g., Splicing)
4. Translation (including mRNA stability)
5. Post-translational (e.g., Protein degradation)
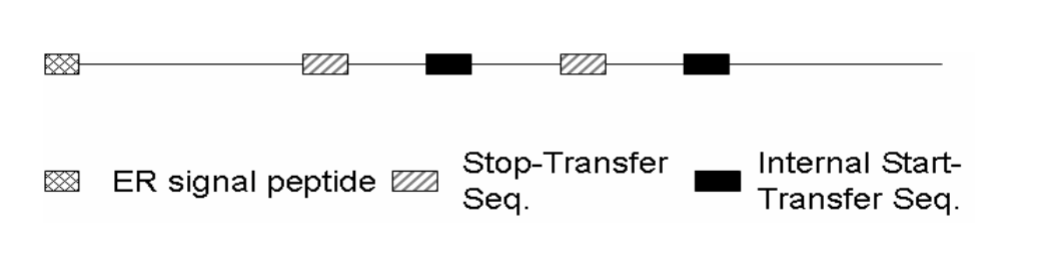
Where will the N-terminus be
The N-terminus is in the ER lumen (the initial ER signal sequence is cleaved off inside)
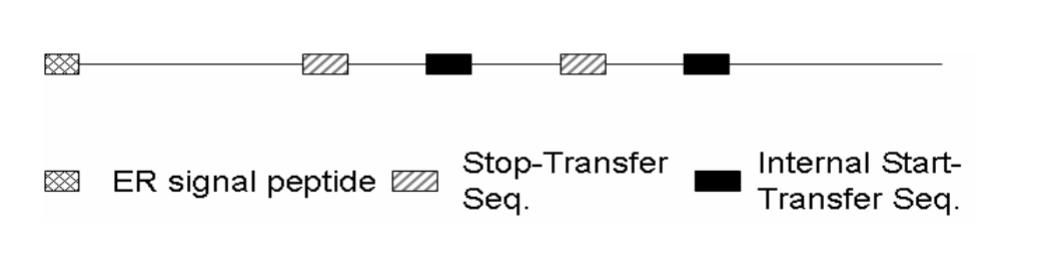
Where will the C-terminus be?
The C-terminus is located in the cytosol
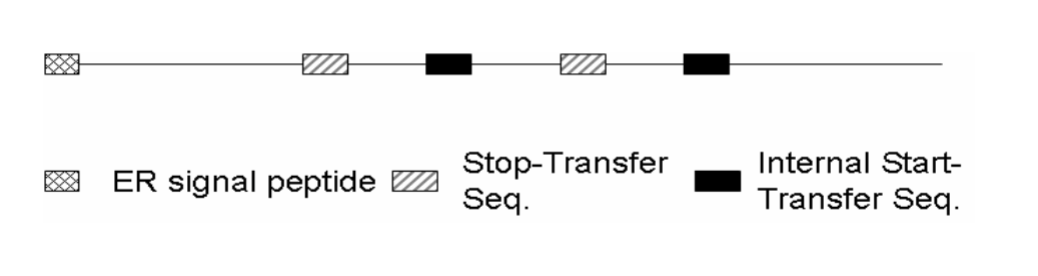
How many transmembrane domains?
There are 4 transmembrane segments. Each "Stop-transfer" and each "Internal start-transfer" sequence remains embedded in the phospholipid bilayer.
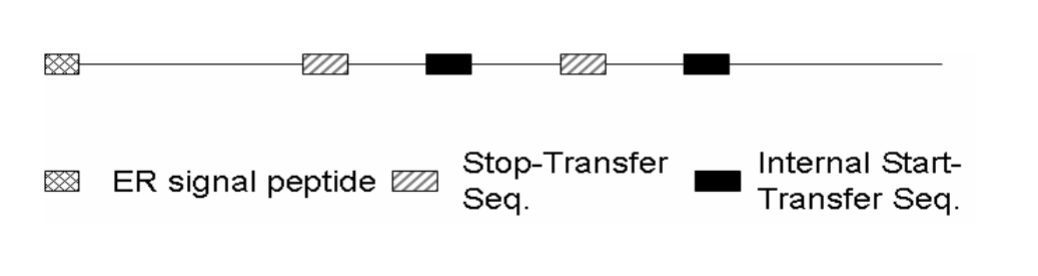
What is the name of this type of proteins?
This is a multi-pass transmembrane protein.