Genetic Code & Transcription
1/60
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
61 Terms
Genetic Code and the Directional Flow of Genetic Info
what does DNA serve as? what does this direct?
what is the final product sometimes?
what is the central dogma of molecular biology?
DNA serves as a template for the synthesis of an RNA molecule, which then directs the synthesis of a protein product
sometimes, the RNA itself is the final product
the principle of directional information flow from DNA to RNA to protein is the central dogma of molecular biology
Transcription vs Translation
Transcription: RNA synthesis using DNA as a template
Translation: synthesis of protein using the information in the RNA
Mutations that alter sequences near the 5′ end of mRNA result in alterations near the corresponding protein’s N-terminal end, whereas mutations that alter the 3′ sequences of mRNA result in alterations in the protein’s C-terminal end. What do these findings imply?
A.The N- and C-terminal ends of proteins are susceptible to alteration.
B.The genetic code is a series of amino acids (the component parts of proteins).
C.The order of nucleotides from 5′ to 3′ in mRNA determines the order of amino acids from N- to
C-termini.
D.The genetic code is universal.
C.The order of nucleotides from 5′ to 3′ in mRNA determines the order of amino acids from N- to
C-termini.
Transcription and Translation Involve Many of the Same Components in Bacteria and Eukaryotes
what is mRNA?
rRNA?
tRNA?
Messenger RNA, mRNA, is RNA that is translated into protein
Ribosomal RNA, rRNA, is an integral component of the ribosome
Transfer RNA, tRNA, molecules serve as intermediaries, bringing amino acids to the ribosome
The latter two function in translation
Transcription & Translation image
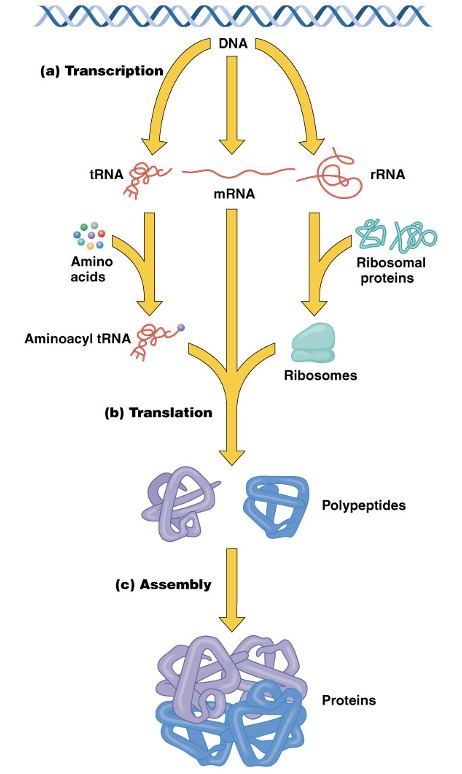
The Genetic Code
what is it?
what is a doublet and triplet code?
The relationship between the DNA base sequence and the linear order of amino acids in the protein products is based on a set of rules known as the genetic code
There are four DNA bases and 20 amino acids
A doublet code, in which two bases specify a single amino acid, is inadequate because only 16 combinations are possible
A triplet code, in which combinations of three bases specify amino acids, would have 64 possible combinations, more than enough for all 20 amino acids
Supporting a Triplet Code using Frameshift Mutations
what does inserting or deleting a nucleotide do?
what are mutations that called this called?
ex?
Inserting or deleting a nucleotide (indel mutations) causes the rest of the sequence to be read out of phase—this is a shift in the reading frame
Mutations that cause insertion or deletion of a nucleotide are thus called frameshift mutations
a) Proflavin is a mutagen that induces insertion or deletion of single nucleotides
b) experiments using Proflavin supported the idea of a triplet code
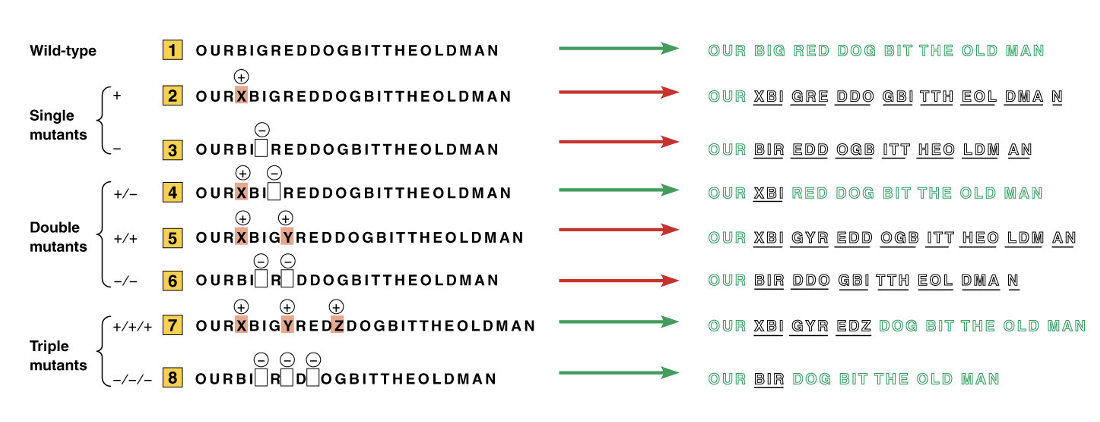
The interpretation of Revertible Mutations
single, double, triple mutants?
• No revertant single mutants
• Revertant double mutants if the two mutations are opposite
• Revertant triple mutants only if all mutations are the same
The Genetic Code Is Degenerate and Nonoverlapping
how many combinations of nucleotides are there, how many amino acids?
what does this mean?
There are 64 combinations of nucleotide triplets and only 20 amino acids
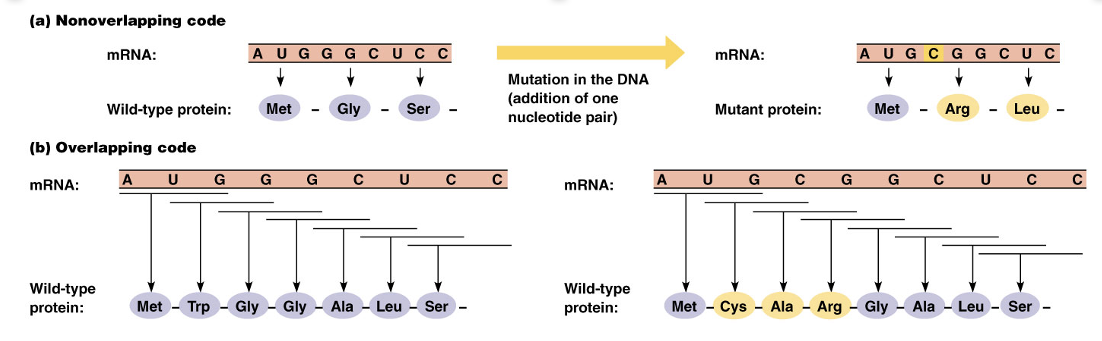
This means the genetic code is a degenerate code, meaning that a particular amino acid can be specified by more than one triplet
It is also nonoverlapping; the reading frame advances three nucleotides at a time
Nonoverlapping vs Overlapping Code
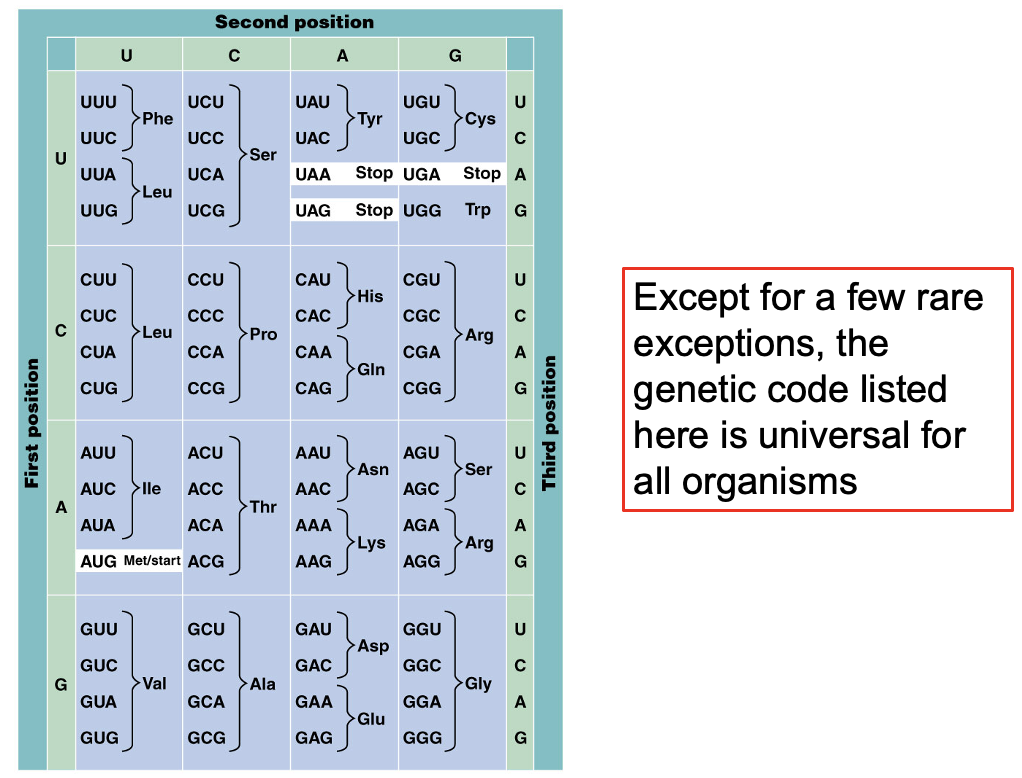
The Codon Dictionary Was Established Using Synthetic RNA Polymers and Triplets
what are codons?
what did homopolymer experiments show?
RNA triplets, called codons, are read by the transcriptional machinery
Further homopolymer experiments showed AAA codes for lysine, and CCC codes for proline
As synthetic polymer technology progressed, production of all different codons independently led to the elucidation of the entire codon dictionary
Genetic code is universal for all organisms
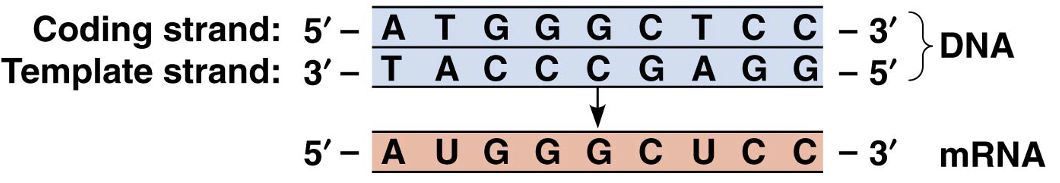
Messenger RNA Guides the Synthesis of Polypeptide Chains
how is mRNA transcribed?
mRNA is transcribed from DNA similarly to how DNA is replicated, but with two differences
In mRNA synthesis, only one DNA strand is copied, called the template strand; the other strand is called the coding strand because it is similar to the mRNA sequence
In mRNA synthesis, a uracil base (U) is used instead of thymine
Transcription Involves Four Stages: Binding, Initiation, Elongation, and Termination
what is the transcription unit?
when does transcription begin? explain the 4 steps
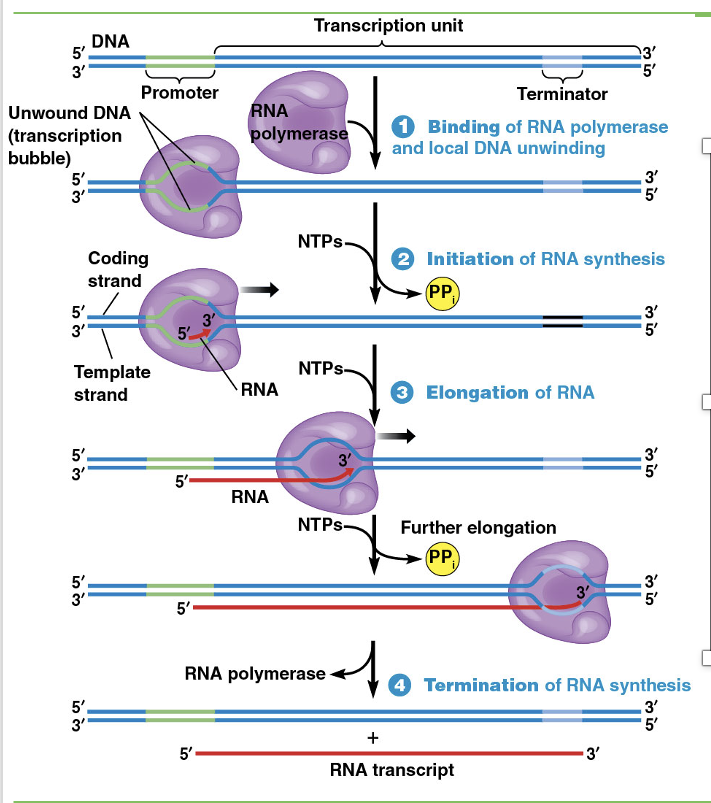
The DNA that gives rise to one RNA molecule is called the transcription unit
Transcription begins when RNA polymerase binds to a promoter sequence (1), triggering local unwinding of the double helix
RNA polymerase then initiates synthesis of RNA using one DNA strand as a template (2)
After initiation, the RNA polymerase moves along the DNA template, unwinding the helix and elongating the RNA (3)
Eventually, RNA polymerase dissociates from the DNA template, leading to termination of synthesis and release of the RNA molecule (4)
Transcription Image
what determines where RNA synthesis will start?
what does upstream and downstream mean?
•RNA polymerase binds to a DNA promoter site, a sequence of several dozen base pairs that determines where RNA synthesis will start
•The terms upstream and downstream refer to sequences located toward the 5′ or 3′ end of the transcription unit, respectively
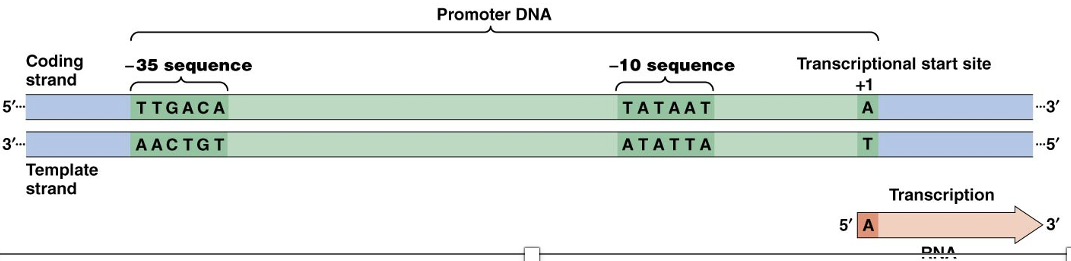
Essential Sequences in a Typical Bacterial Promoter
what is the transcription start site usually? AUTGC?
what is about 10 bp upstream of the start site?
10 sequence? -35 sequence?
The transcription start site is almost always a purine and usually an adenine
About 10 bp upstream of the start site is the sequence TATAAT, called the –10 sequence or the Pribnow box
At or near the –35 position is the sequence TTGACA, called the –35 sequence
Bacterial RNA Polymerase Structure and Initiation of Transcription IMAGE
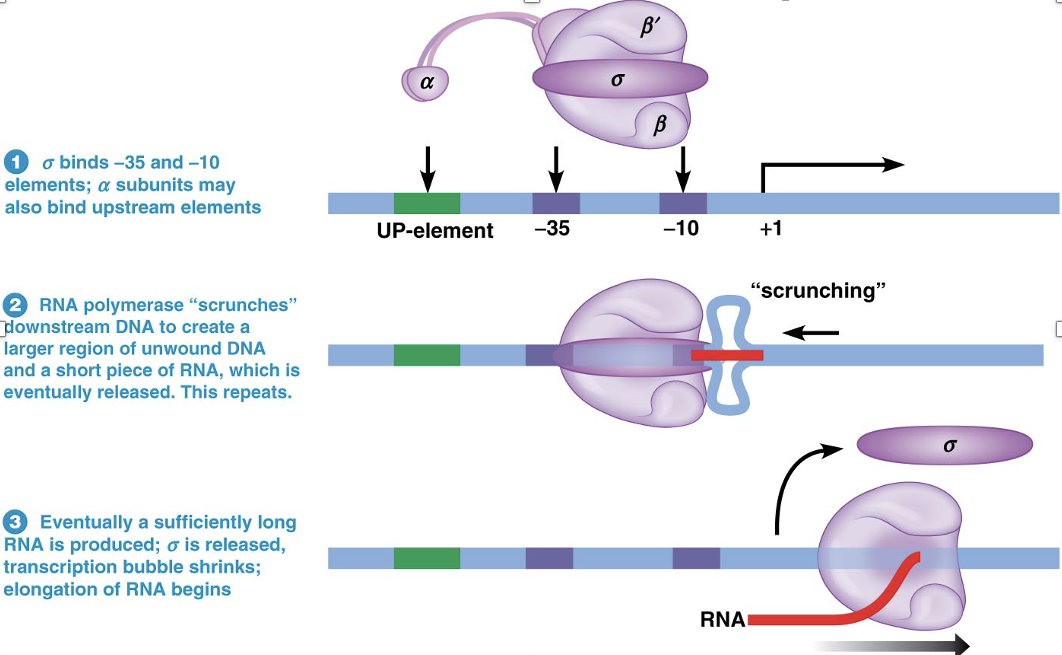
Elongation of the RNA Chain
which end is each new nucleotide added to?
what is unwound and rewound?
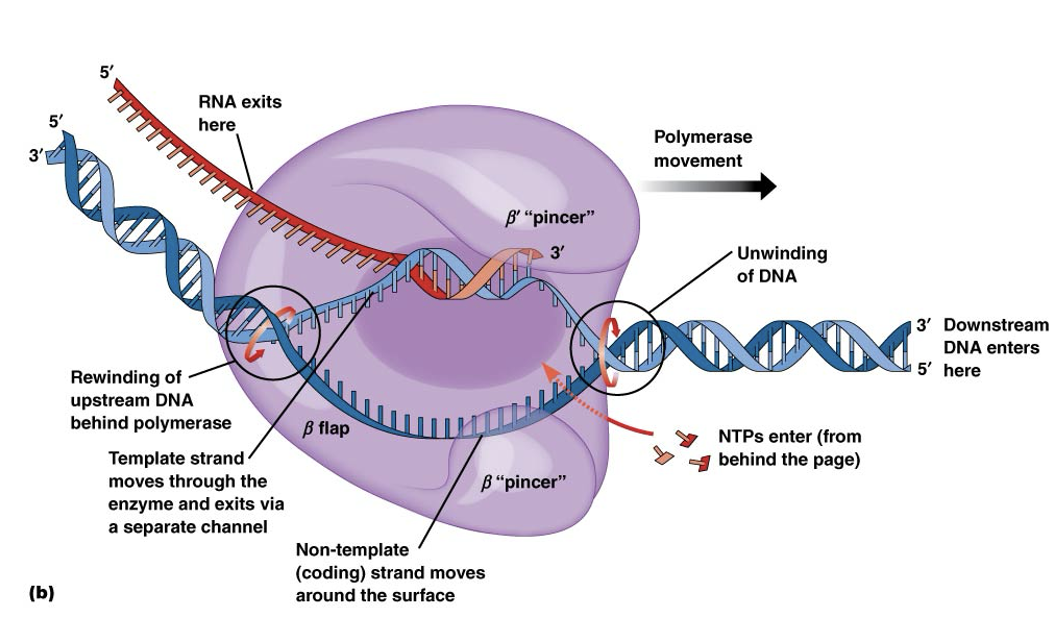
Chain elongation continues as RNA polymerase moves along the DNA molecule
The RNA is elongated in the 5′ to 3′ direction, with each new nucleotide added to the 3′ end
As the polymerase moves along the DNA strand, the double helix ahead of the polymerase is unwound, and the DNA behind it is rewound into a double helix
Elongation of RNA Chain image
Termination of RNA Synthesis
what is the termination signal?
what do many termination sequences contain?
what do they form?
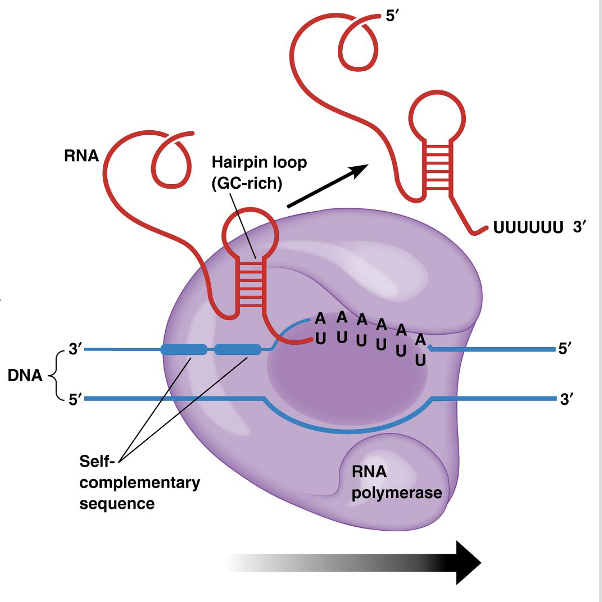
Elongation of the RNA chain proceeds until the RNA polymerase copies a sequence called the termination signal
Many termination sequences contain a short GC-rich sequence followed by several U’s
The GC region in the RNA forms a hairpin loop pulling the RNA molecule away from the DNA
Then the bonds between the U’s and the A’s of the template strand break, releasing the RNA
Transcription in Eukaryotic Cells Has Additional Complexity Compared with Prokaryotes
what does eukaryotic transcription involve?
what are the differences?
Eukaryotic transcription involves the same four stages as prokaryotic, but there are several important differences
Three different RNA polymerases transcribe one or more different classes of RNA
Eukaryotic promoters are more varied than bacterial ones; some are even located downstream of the gene
Eukaryotic Transcription
what are transcription factors?
what type of interaction is in eukaryotic transcription?
Eukaryotic transcription differs from that of prokaryotes
RNA polymerases in eukaryotes require additional proteins called transcription factors, some of which must bind before the RNA polymerase can bind
Protein-protein interactions play a prominent role in eukaryotic transcription
Eukaryotic Transcription (continued)
what is more important than termination of transcription?
what do newly forming RNA molecules undergo?
Eukaryotic transcription differs from that of prokaryotes
RNA cleavage is more important than termination of transcription in determining the 3′ end of the transcript
Newly forming RNA molecules undergo RNA processing, chemical modification during and after transcription
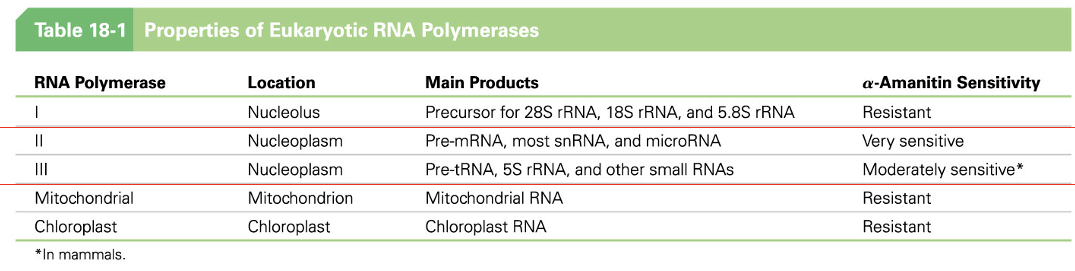
RNA Polymerase I, II, and III Carry Out Transcription in the Eukaryotic Nucleus
location?
main products?
a-Amanitin Sensitivity
(Only for RNA Polymerase II and III)
There are three RNA polymerases in the nucleus, designated RNA polymerases I, II, and III
We will focus on just II and III

Three Classes of Promoters Are Found in Eukaryotic Nuclear Genes, One for Each Type of RNA Polymerase
what is the core promoter?
The core promoter is the smallest set of DNA sequences that initiates transcription
We will compare the core promoter elements for the promoters used by RNA Polymerases II and III
The Promoter for RNA Polymerase II
what are the 4 types of DNA sequences that are involved in the core promoter function?
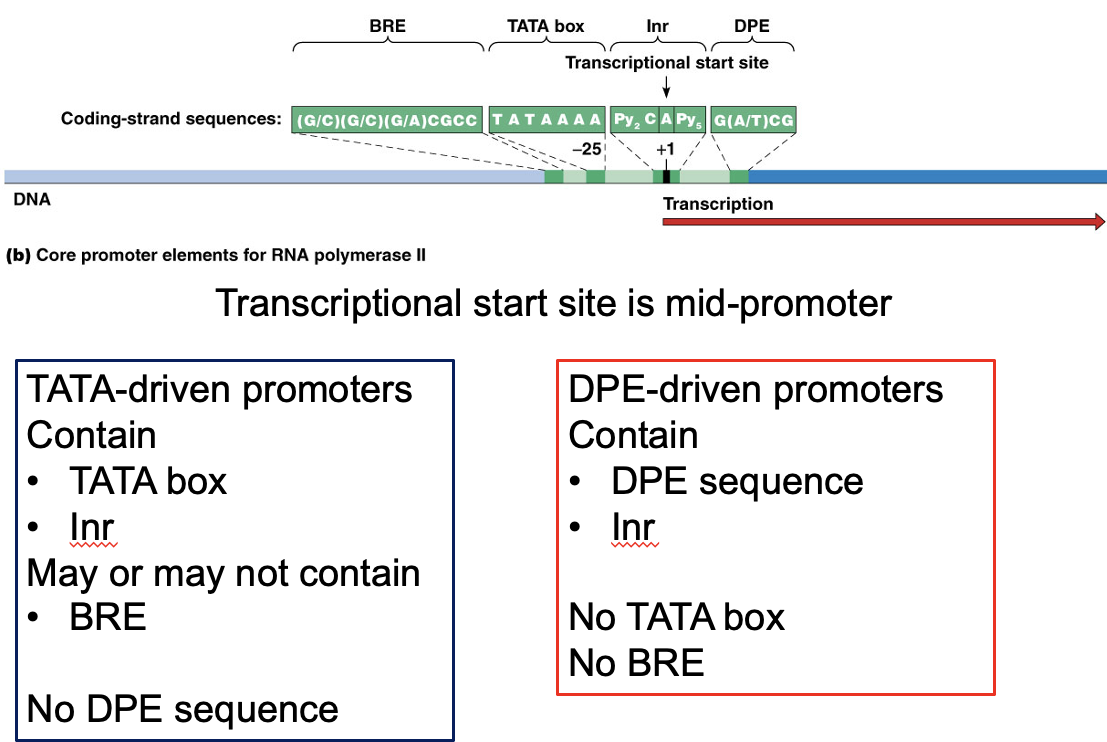
At least four types of DNA sequences are involved in core promoter function
A short initiator sequence surrounds the transcription start point
2.The TATA box, a consensus sequence of TATA followed by two to three A’s, is located about 25 nucleotides upstream of the start point
3.The TFIIB recognition element (BRE) is located slightly upstream of the TATA box
4.The downstream promoter element (DPE) is located about 30 nucleotides downstream from the start point
TATA-driven Promoters
TATA-driven Promoters:
Contain:
TATA bos
Inr
May or may not contain:
BRE
No DPE Sequence
DPE-driven promoters
DPE-driven promoters
Contain:
DPE sequence
lnr
NO TATAbox
NO BRE
Promoters for RNA Polymerase III
what location are the promoters at that are used by RNA polymerase III
what boxes does tRNA have rRNA have?
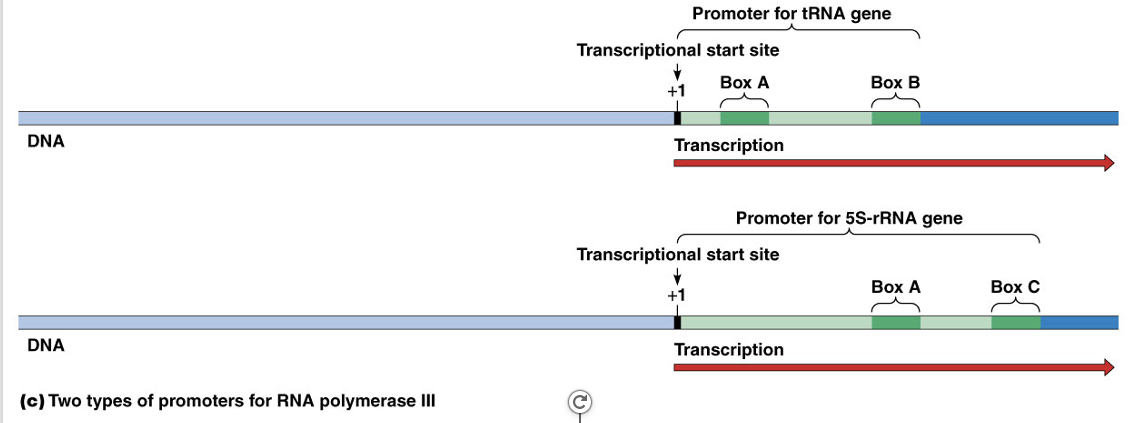
RNA polymerase III uses promoters that are entirely downstream of the start point
In both 5S RNA and tRNA, the promoters are different, but both consensus sequences fall into two blocks of about 10 bp each
tRNA has box A and box B; rRNA has box A and box C
where does transcription start?
what are consensus sequences transcribed into?
Transcriptional start site is at upstream end of promoter
Consensus sequences are all transcribed into RNA
Additional Control Elements
how much transcription are core promoters able to drive?
what improves promoters efficiency?
what are proximal control elements?
what are those farther away called?
Core promoters are capable of driving only a basal (low) level of transcription
Additional short sequences upstream (upstream control elements) improve the promoter’s efficiency
Those within 100–200 nucleotides of the start point are called proximal control elements
Those farther away are called enhancer elements or distal control elements
General Transcription Factors Are Involved in the Transcription of All Nuclear Genes
what are general transcription factors required for?
what factors do eukaryotes have?
what does a large complex of proteins form?
A general transcription factor is always required for RNA polymerase binding to promoters
Eukaryotes have many such factors, called TFs, that bind the promoter in a defined order starting with TFIID
Eventually, a large complex of proteins forms a preinitiation complex on the promoter
Initiation at a RNA Polymerase II Promoter
what is essential for beginning the process?
what does it do?
what lead to the recruitment of RNA polymerase II?
TFIID is essential for beginning the process
TFIID recognizes and binds DNA because of its TATA-binding protein (TBP) subunit
Other TFs follow TFIID, many binding to each other and not directly to DNA
Leads to recruitment of RNA Polymerase II
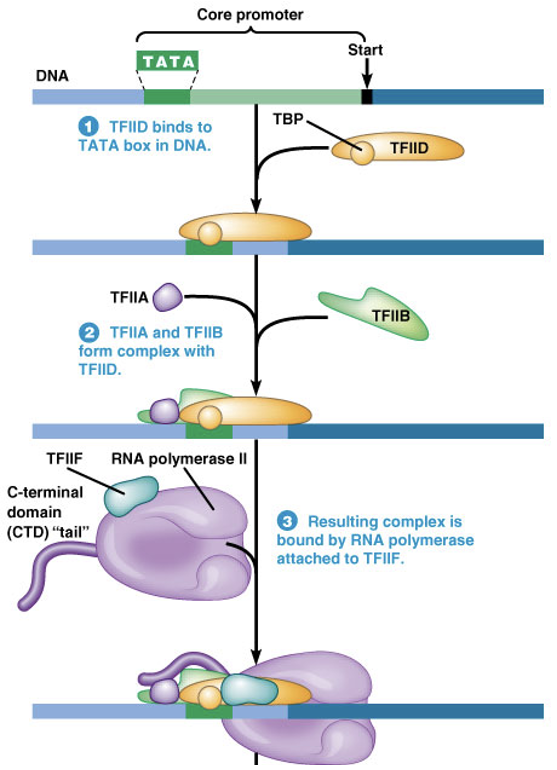
Initiation at a RNA Polymerase II Promoter
what activity does TFIIH have?
what does this promote?
TFIIH has both helicase and kinase activity
•Local unwinding of DNA
•Phosphorylation of the C-terminal domain (CTD) of RNA Polymerase II
Both promote the release of RNA Polymerase II from the initiation complex and the beginning of transcription
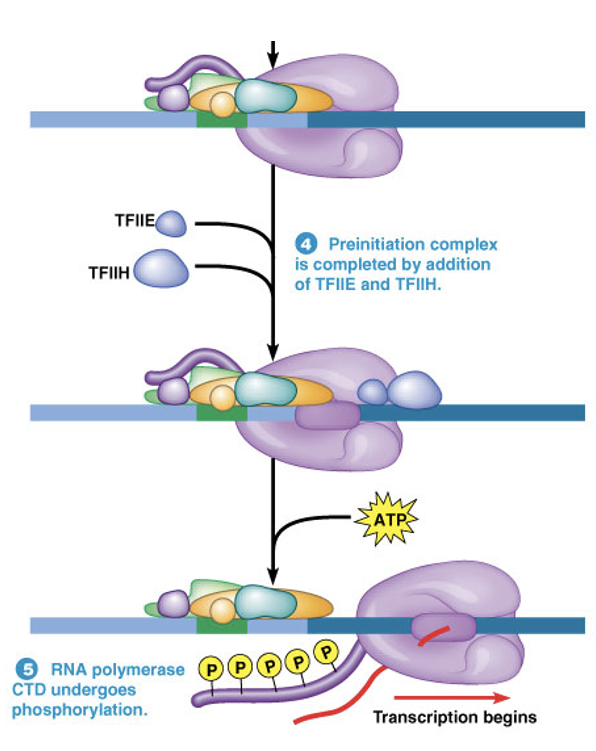
Termination of Eukaryotic RNA Synthesis
what is termination governed by?
for RNA polymerase III what do the termination signals include?
what happens to the RNA polymerase II transcripts?
where is the cleavage sit?
Termination is governed by signals that differ for each type of RNA polymerase
For RNA polymerase III, termination signals include a short run of U’s, and no protein factors are required for their recognition
For RNA polymerase II, transcripts are cleaved at a specific site before transcription ceases
The cleavage site is 10–35 nucleotides downstream of a AAUAAA sequence in the RNA
Termination of Transcription
For RNA polymerase III, termination signals include a short run of U’s, and no protein factors are required for their recognition
For RNA polymerase II, transcripts are cleaved at a specific site before transcription ceases
The cleavage site is 10–35 nucleotides downstream of a AAUAAA sequence in the RNA
Cleavage location image
RNA Polymerase II continues through the AAUAAA sequence
The pre-mRNA is cleaved 10-35 nt after the AAUAAA and released
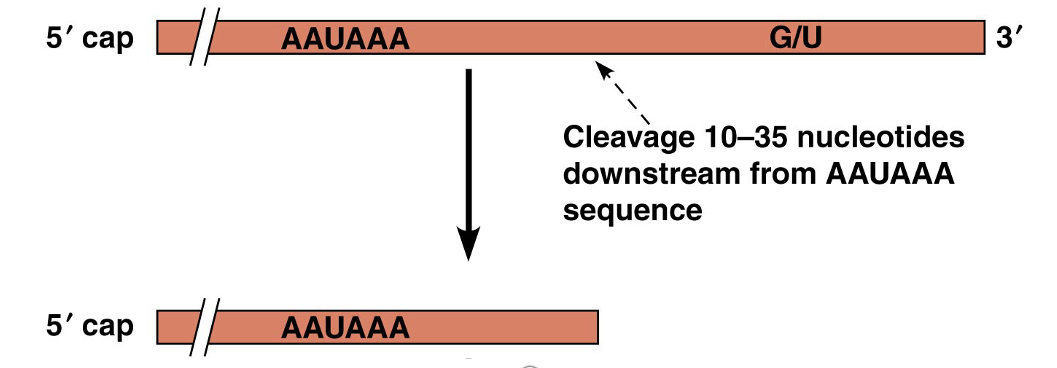
RNA Processing and Turnover
what is the primary transcript?
what must is undergo?
A newly produced RNA molecule is called the primary transcript
It must undergo RNA processing (chemical modification) before it can function in the cell
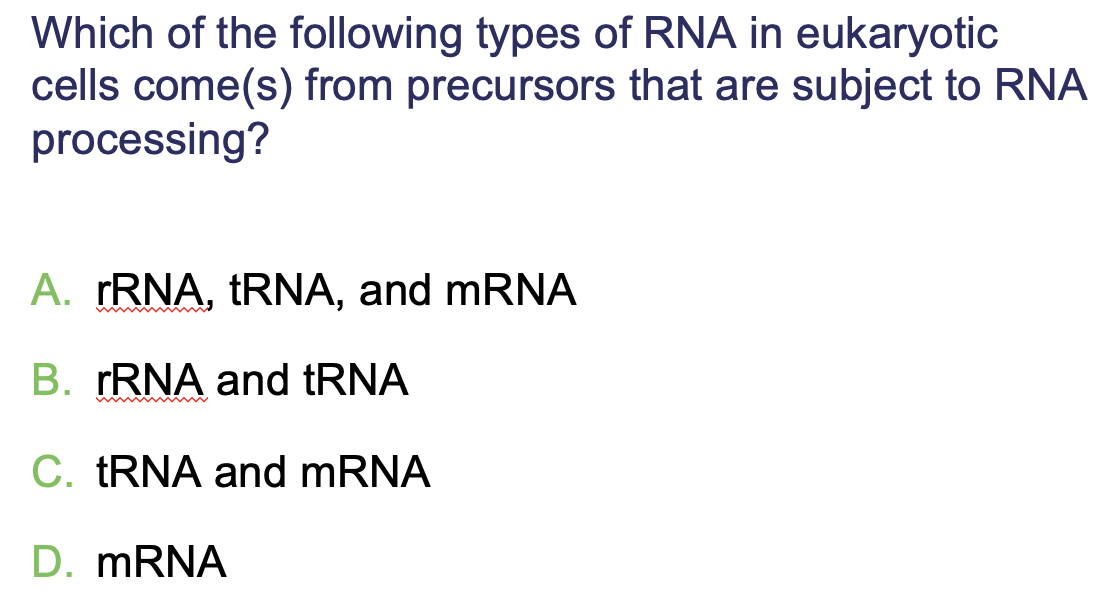

A. rRNA, tRNA, and mRNA
Transfer RNA Processing Involves Removal, Addition, and Chemical Modification of Nucleotides
what do cells do?
what does their secondary structure contain?
what structure to tRNAs have?
Cells synthesize several dozen kinds of tRNA molecules
They fold into a secondary structure, most containing four hairpin loops; but some have a fifth region called a variable loop
tRNAs have a cloverleaf structure and are synthesized as pre-tRNAs, followed by processing
Primary Transcript for yeast tyrosine tRNA vs Mature tRNA, secondary structure
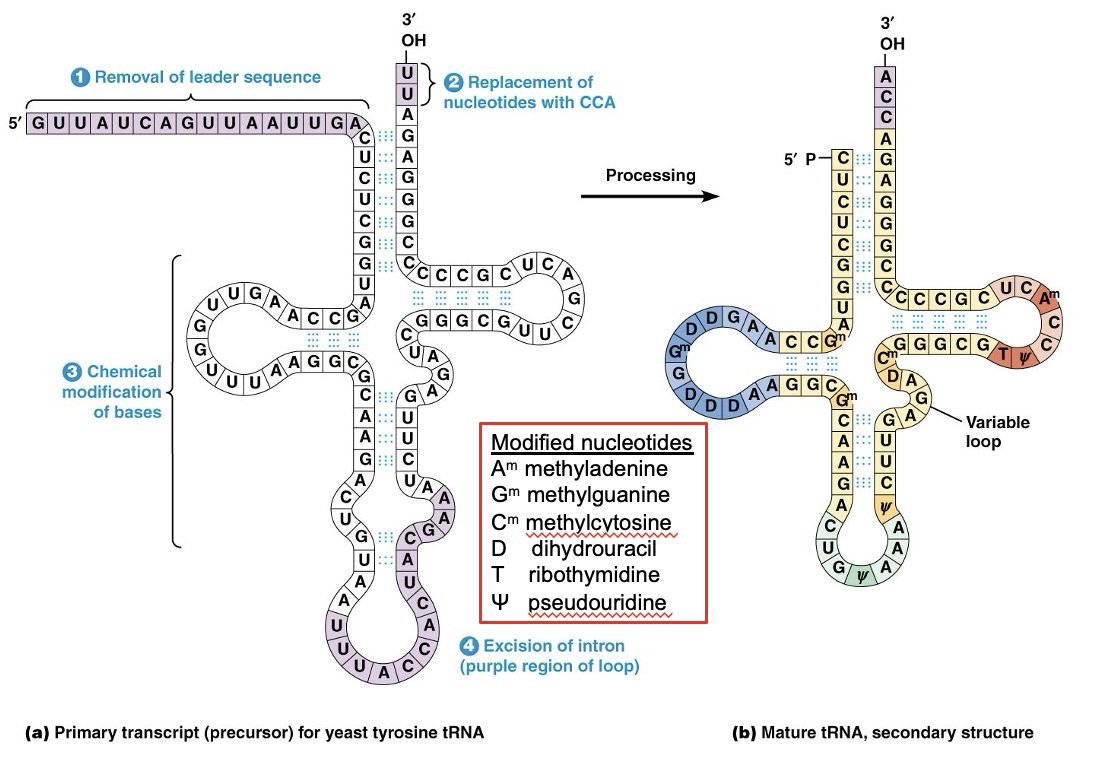
Eukaryotic Transcripts
what is hnRNA?
how are Pre-mRNAs processed?
what does the C- terminal domain of one of the subunits of RNA polymerase II act as?
hnRNA is a mixture of mRNA molecules and their precursors, pre-mRNA
Pre-mRNAs are processed by removal of sequences and addition of 5′ caps and 3′ tails
The C-terminal domain of one of the subunits of RNA polymerase II acts as a platform for protein complexes involved in processing
5′ Caps and 3′ Poly(A) Tails
what is the 5’ cap?
how is it bound by the RNA molecule?
Eukaryotic mRNAs have a modified nucleotide called the 5′ cap, and the 3′ ends have a long stretch of adenines called the poly(A) tail
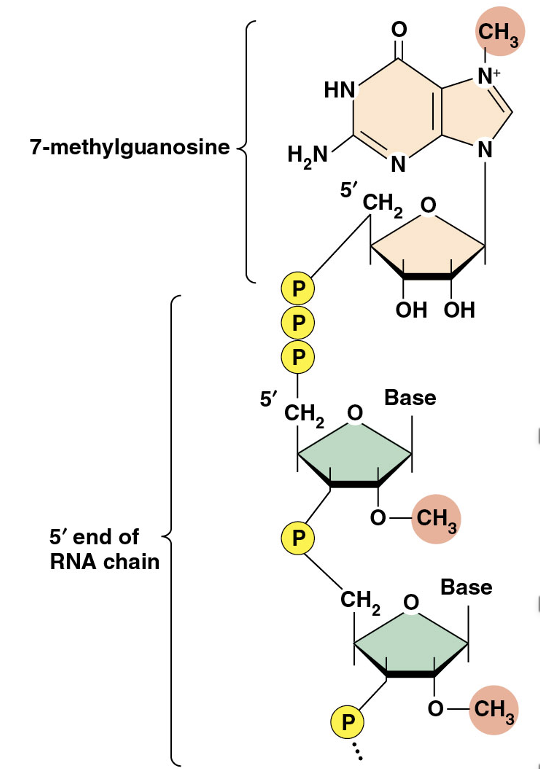
The 5′ cap is a guanosine that is methylated at position 7 of the purine ring
It is bound to the RNA molecule by a 5′→5′ linkage rather than the usual 3′→5′ bond
5’ Cap
what does it contribute to?
what role does it play?
•The 5′ cap is a guanosine that is methylated at position 7 of the purine ring
•It is bound to the RNA molecule by a 5′→5′ linkage rather than the usual 3′→5′ bond
•The cap contributes to mRNA stability by protecting the RNA from nucleases
•The cap also plays a role in positioning the RNA on the ribosome for initiation of translation
5’ Cap Photo
The Poly(A) Tail
how many nucleotides long is it?
what is it added by?
where is the signal for the addition of the poly(A) tail located? (sequence)
function?
what is it required for?
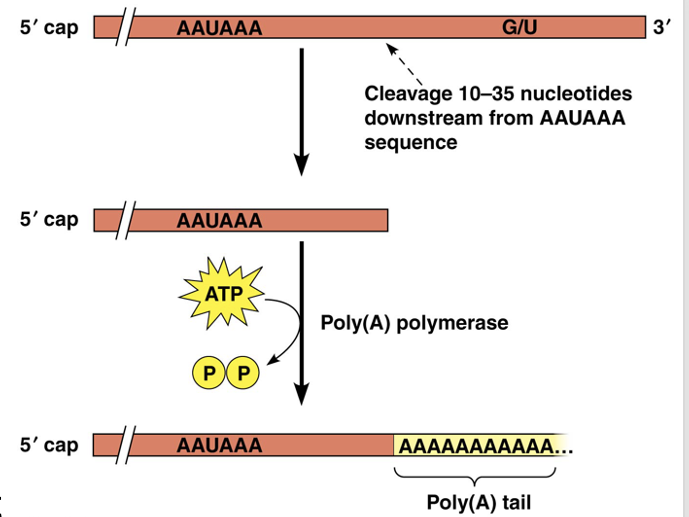
The poly(A) tail ranges from 50 to 250 nucleotides long and is added by the enzyme poly(A) polymerase
A signal for addition of the poly(A) tail, AAUAAA, is located just upstream of the polyadenylation site, and a GU- or U-rich element is located downstream of it
•The poly(A) tail protects the mRNA from nuclease attack; the length of the tail influences stability
•It is also required for export of the transcript to the cytoplasm
•It may also help ribosomes recognize and bind mRNAs
The Poly(A) Tail Image
Spliceosomes Remove Introns from Pre-mRNA
what is RNA splicing?
Sequences commonly found at the intron-exon boundaries likely determine by what?
what does the 5’ end and start with and terminate with at the 3’ end?
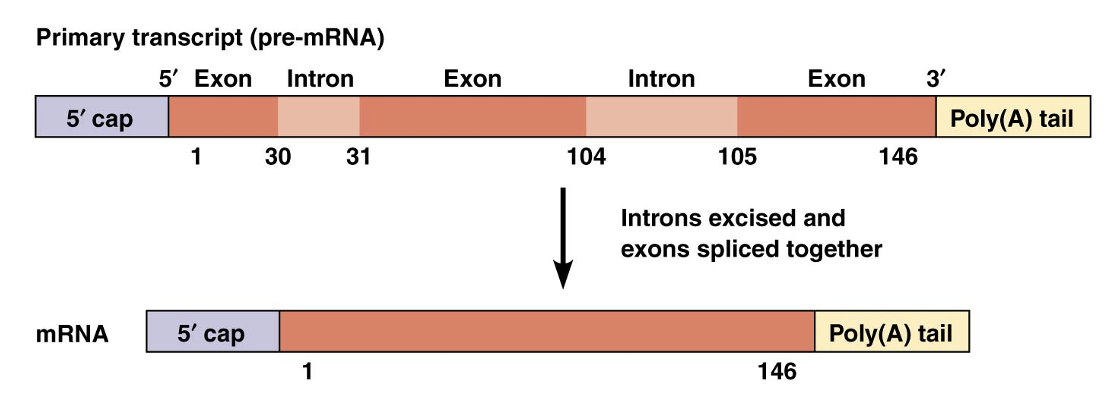
The process of removing introns and joining the exons is RNA splicing
Sequences commonly found at the intron-exon boundaries likely determine the 5′ and 3′ splice sites
Analysis of base sequences of hundreds of different introns revealed that the 5′ end of an intron typically starts with GU and terminates with AG at the 3′ end
Primary Transcript (pre-mRNA)
The Splicesome
what are spliceosomes?
where do they assemble?
what are snRNPs?
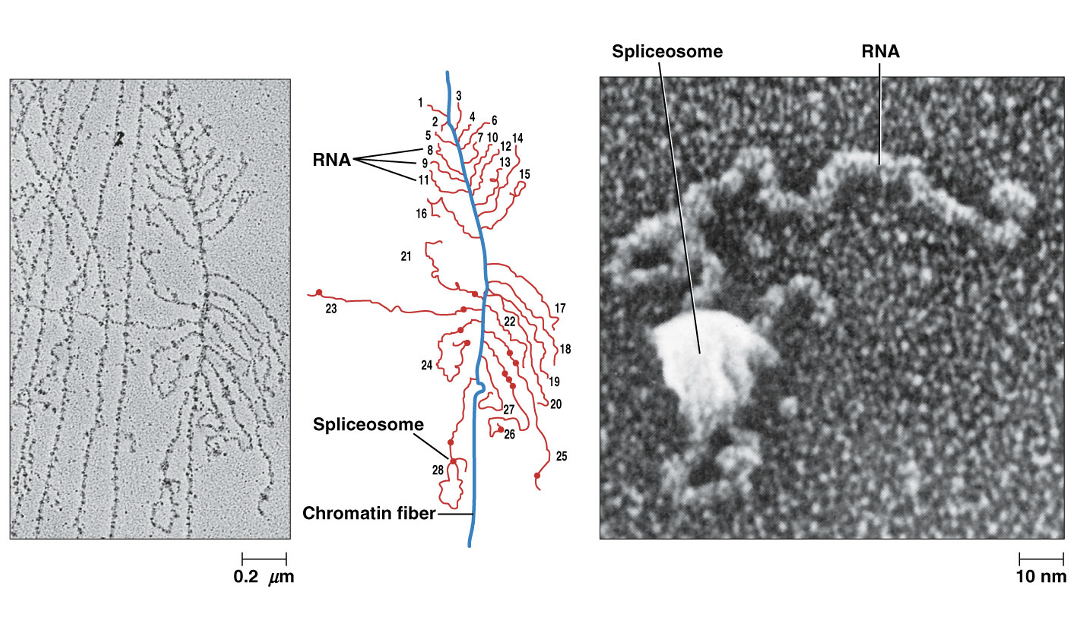
Intron removal is catalyzed by large molecular complexes called spliceosomes, consisting of five types of RNA and many proteins
Spliceosomes assemble on transcripts from a group of smaller RNA-protein complexes called snRNPs (small nuclear ribonuclearprotein complexes), each containing one or two snRNA molecules (small nuclear RNA)
Spliceosome Image
Splicesome Assembly
what are they assembled by?
what are the steps?
Spliceosomes are assembled by sequential binding of snRNPs to pre-mRNA
The first step is the binding of a snRNP called U1, which contains an RNA that can base-pair with the 5′ splice site
A second snRNP called U2 binds to the branch-point sequence
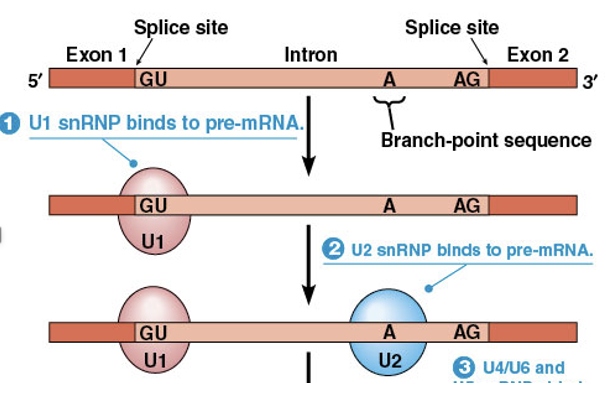
Splicesome Assembly (continued)
4,5,6 step
resulting structure?
Finally a group of snRNPs (U4/U6 and U5) brings the ends of the intron together to form a mature spliceosome
The pre-mRNA is cleaved at the 5′ splice site, which is joined to an adenine residue located at the branch-point sequence
The resulting structure is called a lariat
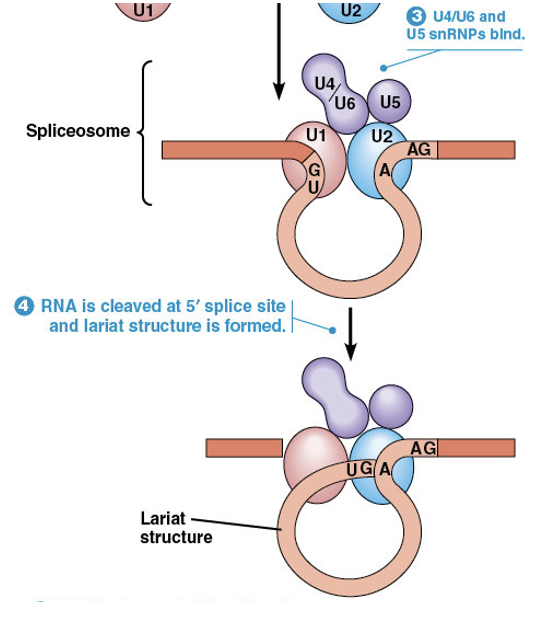
Spliceosome Assembly (continued)
what happens after the lariat is formed?
what is an exon junction complex (EJC)? where is it deposited?
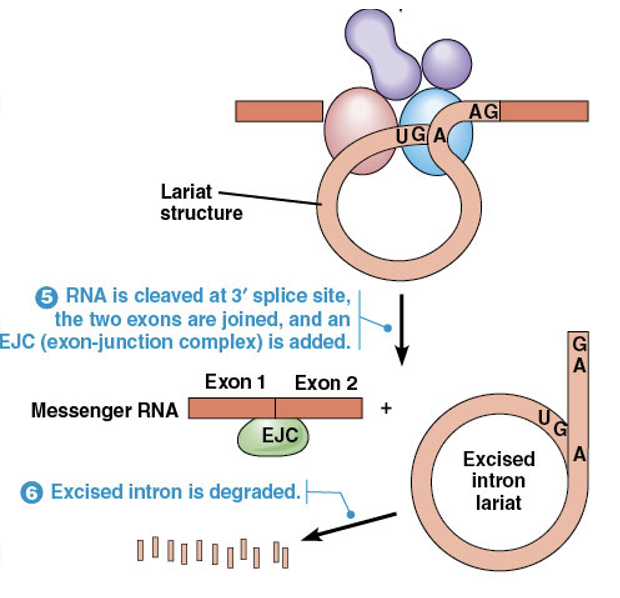
After the lariat forms, the 3′ splice site is cleaved, and the two ends of the exon are joined together
A multiprotein complex called an exon junction complex (EJC) is deposited near the boundary of each newly formed exon-exon junction
Alternative Splicing
how is it possible? via what mechanisms?
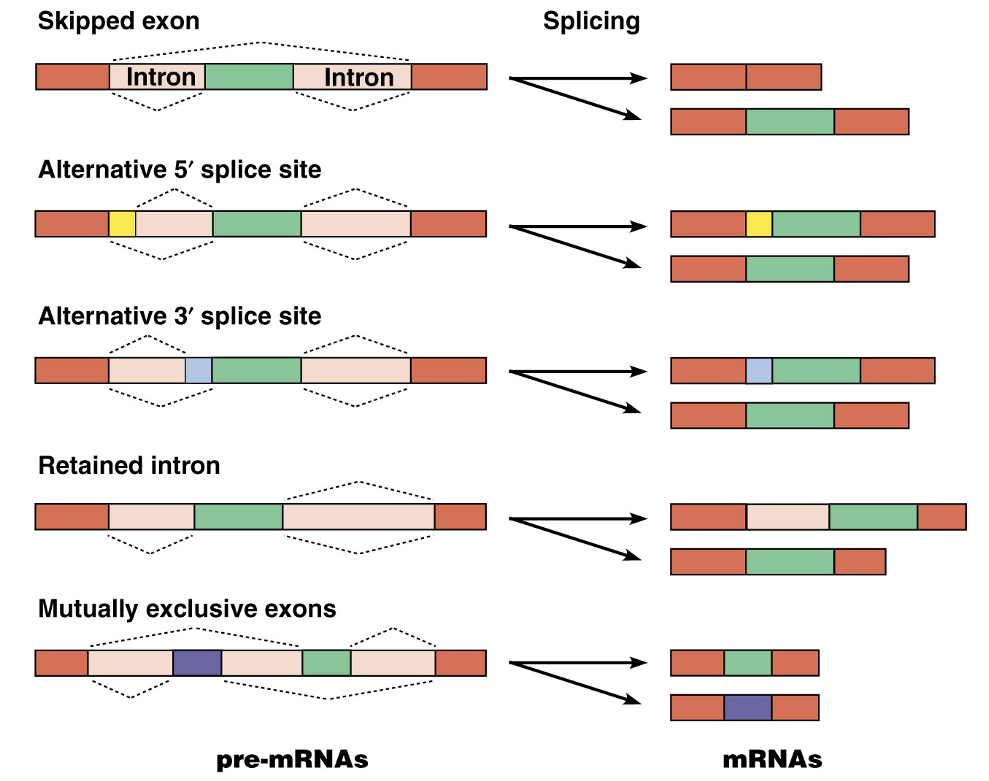
The presence of introns allows each gene’s pre-mRNA molecule to be spliced in multiple ways, leading to production of multiple protein products
This alternative splicing is possible via mechanisms allowing certain splice sites to be activated or skipped
Alternative Splicing Image
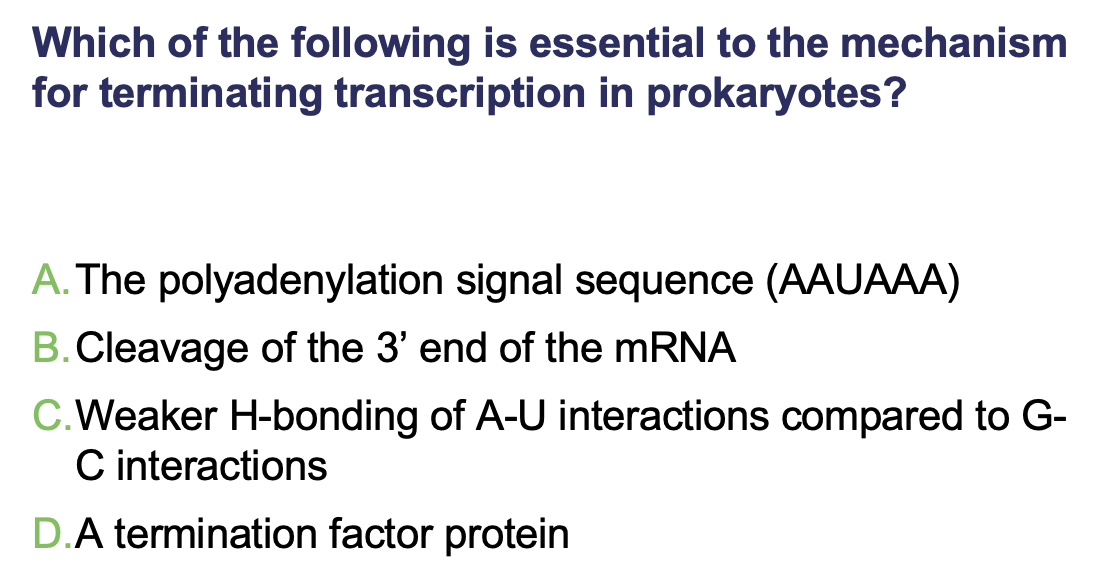

C.Weaker H-bonding of A-U interactions compared to G-C interactions
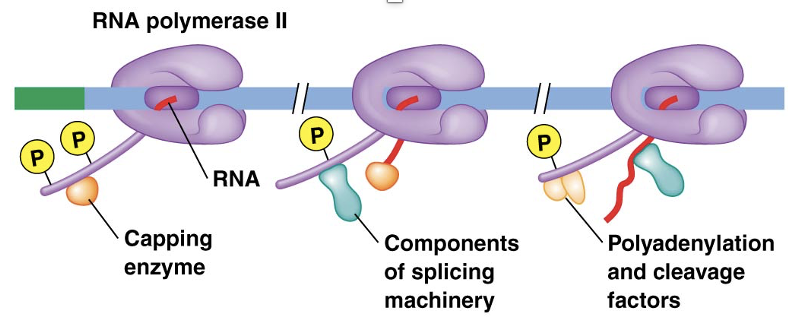
The C-Terminal Domain of RNA Polymerase II Coordinates RNA Processing
what occurs cotranscriptionally?
what domain is responsible for this processing?
what do repeats of a seven amino acid sequence of the CTD do?
•Many RNA processing events occur cotranscriptionally
•The long C-terminal domain (CTD) of RNA polymerase is responsible for this processing
•Many repeats of a seven-amino-acid sequence on the CTD bind enzymes needed for capping, splicing, and cleavage/polyadenlylation
Most mRNA Molecules Have a Relatively Short Life Span
what do most mRNA molecules have?
how are turnover rates measured?
how long are the half-lives of mRNA molecules of eukaryotes?
what about bacteria?
Most mRNA molecules have a high turnover rate (rate at which molecules are degraded and replaced)
It is measured in terms of half-life, the time required for 50% of the molecules to degrade
mRNA molecules of eukaryotes have half-lives of several hours to a few days; in bacteria, the half-lives are usually only a few minutes
The Abundance of mRNA Allows Amplification of Genetic Information
what provides the opportunity for amplification of genetic info?
ex?
mRNA can be synthesized again and again from a piece of template DNA, providing an opportunity for amplification of genetic information
For example, the haploid genome of the silkworm has only one copy of the fibroin gene, but about 104 copies of the mRNA are present in the cell at any given time