Bio chem Chat review
1/101
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
102 Terms
Factors that INCREASE fluidity
More unsaturated fatty acids, Shorter fatty acid chains, Higher temperature
Factors that DECREASE fluidity
More saturated fatty acids, Longer fatty acid chains, Lower temperature
Cholesterol in the membrane
Stabilizes membrane fluidity, Prevents membranes from becoming too rigid or too fluid
Liquid Ordered (Lo)
More tightly packed, More cholesterol, longer saturated fatty acid chains, low temperatures
Liquid Disordered (Ld)
More fluid, More unsaturated fatty acids, Higher temperature
Triacylglycerol
Energy storage, Cannot form bilayers
Phospholipids
Amphipathic, Form bilayers
Sphingomyelin
Common in myelin, Found in lipid rafts
Cholesterol
Membrane fluidity buffer, Present in lipid rafts
Arachidonic acid
Precursor to inflammatory signaling molecules
Phospholipases
PLA1, PLA2, C, D
PLA1
Cuts fatty acid at carbon 1
PLA2
Cuts fatty acid at carbon 2, Often releases arachidonic acid
PLC
Cuts before phosphate group, Produces DAG + phosphorylated head group
PLD
Cuts after phosphate group, Produces phosphatidic acid
Channels
Very fast, Usually not saturable in same way as enzymes, Allow diffusion down gradient
Transporters
Slower, Saturable kinetics, Often undergo conformational changes
P-type ATPases
Become phosphorylated during transport cycle, Example: SERCA
SERCA Mechanism
photo
F-type ATPase
Produces ATP using proton gradient
V-type ATPase
Acidifies intracellular compartments
ABC transporter
ATP-powered transporter, Multidrug resistance
Aquaporin
Water transport
SNARE proteins
facilitate membrane fusion
Caveolin
Caveolae formation
Transmembrane α-helices are rich in
Ile, Val, Leu, Phe
Charged residues are unfavorable in membrane core
Lys, Arg, Asp, Glu
Free Energy Transport Equation
ΔG = RT ln(C2/C1) + zFΔΨ
Key idea of ΔG
Negative ΔG = favorable, Positive ΔG = unfavorable
RTK Key sequence
RTK → IRS-1 gets P → P-IRS-1 binds to Grb2 → Grb2 binds to Sos → Sos bind to Ras → Ras stimulate GDP/GTP exchange →Ras binds to Raf → Raf p-lates MEK → MEK p-lates ERK
Ras
small GTPase
Sos
GEF
Raf
MAPKKK
MEK
MAPKK
ERK
MAPK
Amplification (Signal Transduction)
One signal activates many downstream molecules
Modularity (Signal Transduction)
Scaffold proteins organize pathways
Divergence (Signal Transduction)
One signal activates multiple outputs
Integration (Signal Transduction)
Multiple pathways combine signals
Localized response (Signal Transduction)
Signal confined to specific region
Steps of cell division
G1 → S → G2 → M → G0
G1 (cell division)
cell making RNA and proteins
S (cell division)
DNA synthesis
G2 (cell division)
RNA and protein synthesis
M (cell division)
Nucleus and cell division
G0 (cell division)
terminally differentiated cells
Cyclins
Levels fluctuate, Synthesized and degraded each cell cycle
CDKs
Kinases activated by cyclins
CDK activation Requires
cyclin binding, activating phosphorylation, removal of inhibitory phosphate
Apoptosis Characteristics
Programmed cell death, Caspase activation, DNA fragmentation, Cell shrinkage, Minimal inflammation, can occur during development
Apoptosis can be triggered by
DNA damage, developmental signals, death receptor signaling
Oncogenes
Gain-of-function mutations, Promote cell division
Examples of oncogenes
Ras, erbB2
Tumor suppressors
Loss-of-function mutations, Normally inhibit cell division
Examples of tumor suppressors
p53, Rb, BRCA1
SERCA
P-type ATPase, pumps Ca2+ from cytosol to ER lumen
Lipid rafts
cholesterol + sphingolipids
Unsaturated FA
increase fluidity
Saturated FA
decrease fluidity
Caspases
drive apoptosis
tracrRNA
naturally occurring bacterial immune locus (RNA involved in guide RNA processing)
Cas9
DNA cutting enzyme (breaks double bonds)
PAM
recognized DNA sequence adjacent to target DNA
Homology-directed repair
precise DNA repair mechanism
non-homologous end joining
error prone repair mechanism
sgRNA
synthetic RNA combining two RNA components
Aquaporins
channel proteins that facilitate the passage of water
flippase
requires ATP to transport lipids from one leaflet to another
scramblase
randomized phospholipid distribution across leaflets
caveolin
important in caveolae formation
t-snare
facilitates membrane fusion
voltage-gated channel
open and close in response to changes in membrane potential
ligand-gated channel
binding of ligand causes allosteric transition that opens and closes channel
1 mM
10^-3 M
eicosanoids
lipid derived signaling molecule involved in inflammation
cholesterol
sterol abundant in mammalian plasma membranes
SERCA Pathway
ATP binds to Ca2+ → Asp is P-lated → Ca2+ released into lumen → ADP is released → deP-lation of p domain →original domain conformation is reset
B-adrenergic receptor desensitization pathway
cAMP activate PKA → PKA P-lates BARK → BARK binds to GbGy receptor → receptor is p-lated → Barr binds to receptor → GbGy released → Barr receptor complex is endocytosed (turned off)
cAMP activates
PKA
RTK has
intrinsic tyrosine kinase activity
MAP kinase cascade
signaling pathway that uses RTK, IRS-1, Grb2, Sos, Ras, Raf, MEK, ERK
cyclin
protein synthesized during every cell cycle
ubiquitinated cyclin
tags proteins for degradation
phosphotase
removes inhibitory phosphate from CDK
destruction box recognition protein
triggers cyclin ubiquitination
CDK
cyclin dependent kinase
mutant RTK active even without ligand binding...
promotes cell division, oncogene
loss of p53 =
increase cancer risk
erbB2
oncogene
crRNA
RNA containing sequence complementary to target DNA
phosphatidylcholine
2 fatty acid chains + phosphate, CH2, CH2, N+, 3CH3
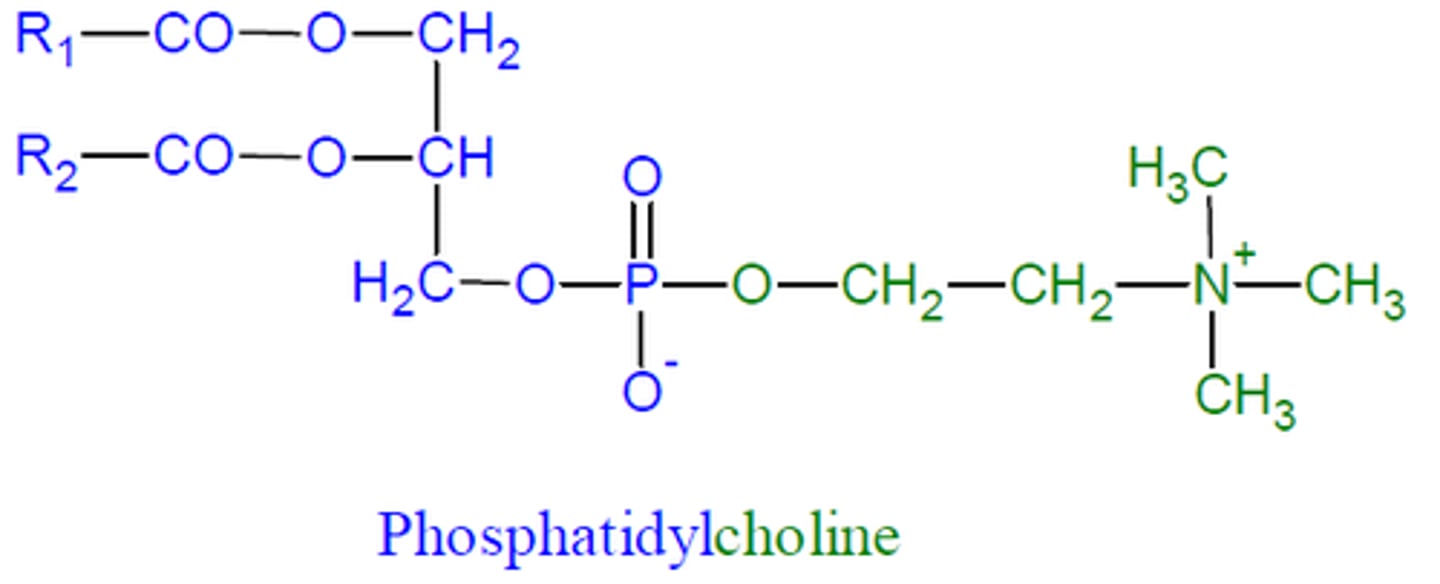
phosphatidylethanolamine
2 fatty acid chains + phosphate, CH2, CH2, NH3+
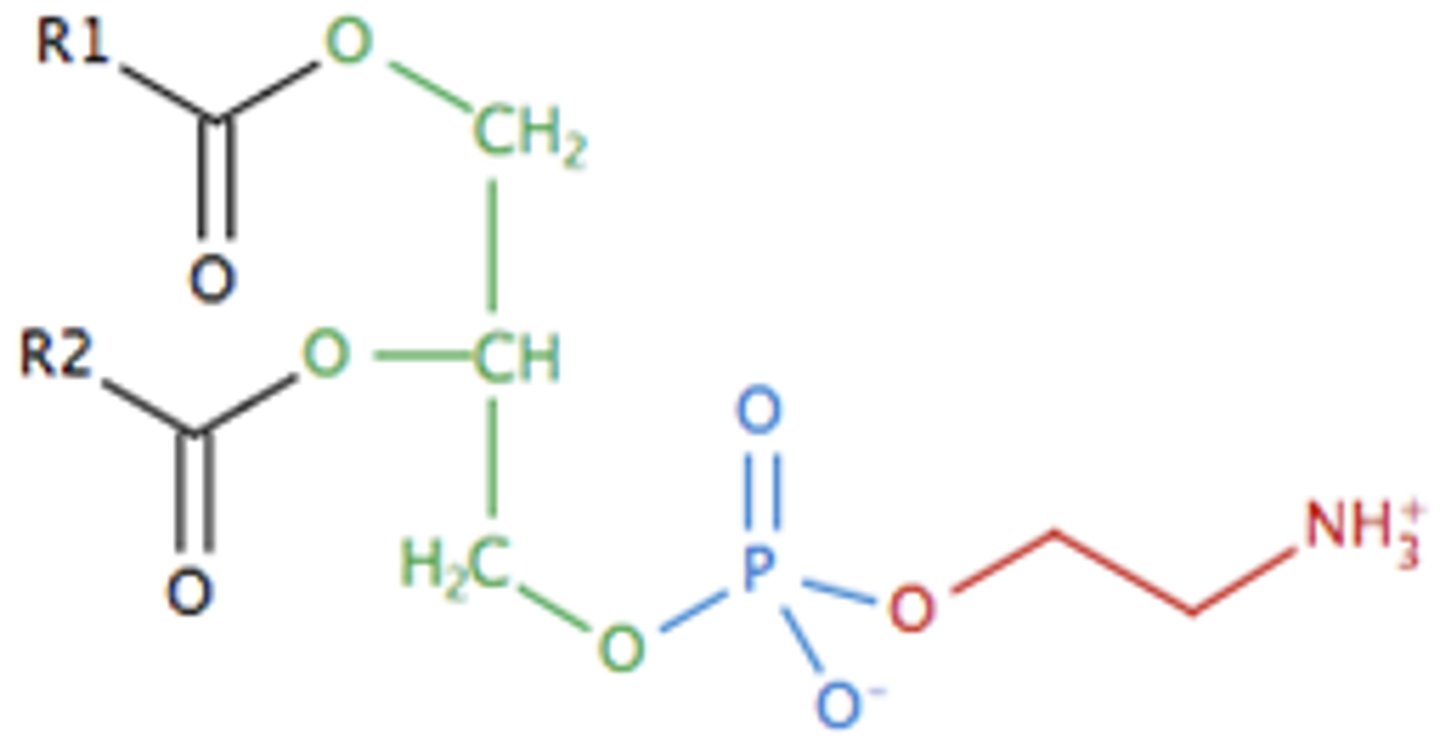
phosphatidic acid
2 fatty acid chains + phosphate, H
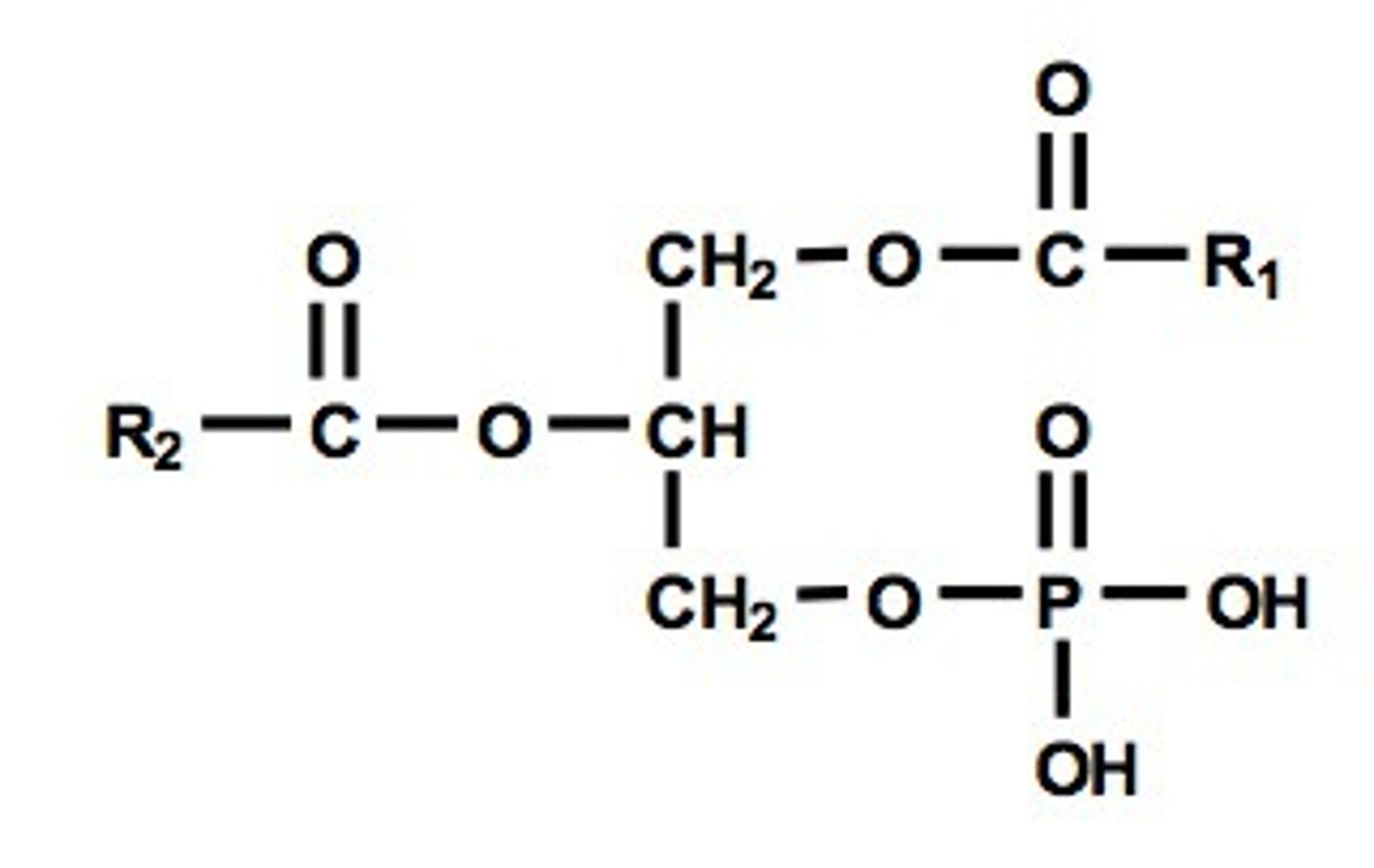
phosphatidylserine
2 fatty acid chains + phosphate, CH2, C, NH3+, COO-, H
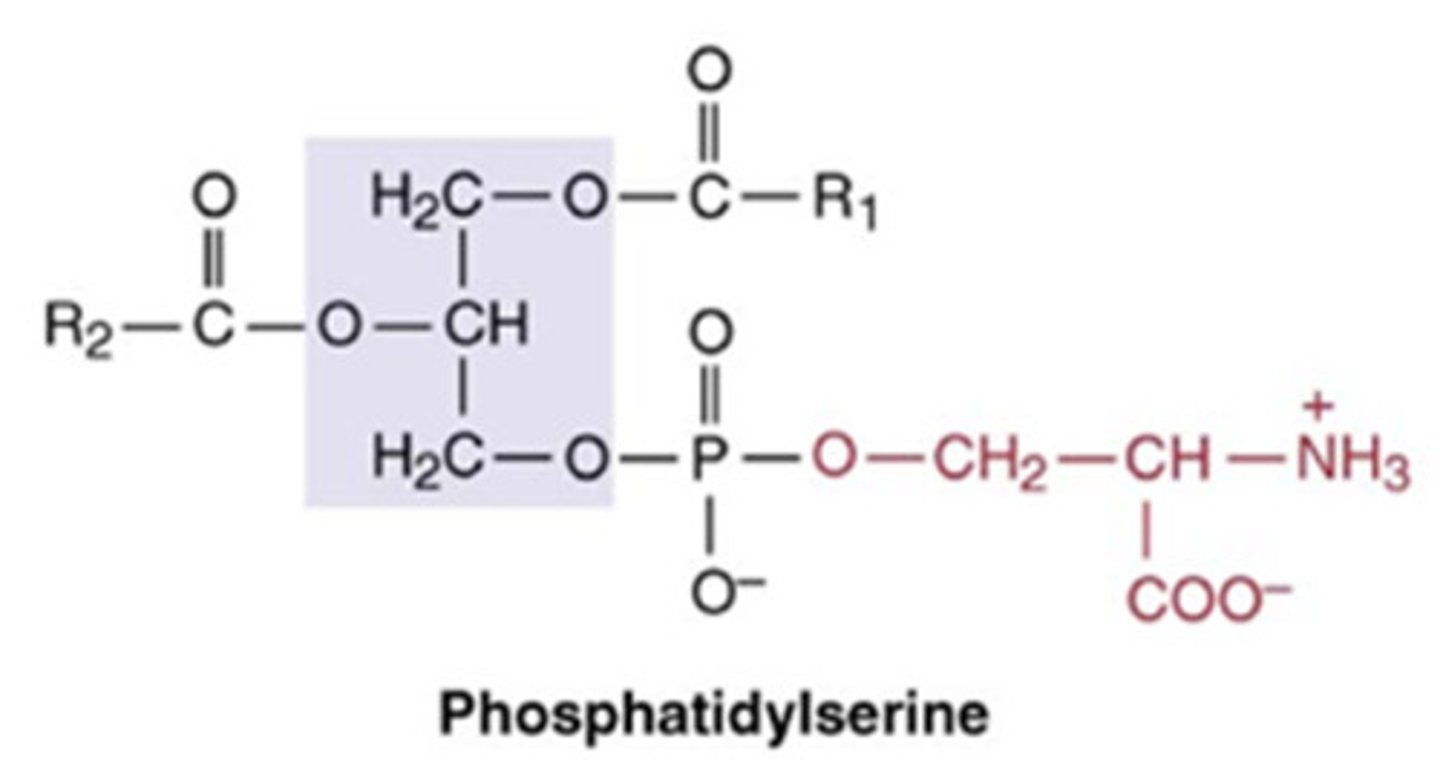
Phosphatidylinositol 4,5-bisphosphate
2 fatty acid chains + phosphate, 6 carbon ring, -OH on C 2 3 and 6, OP on C 4 and 5
floppase
moves phospholipids from cytosol to outer leaflet
sphingomyelin
membrane lipid abundant in myelin
lipid derived signal molecules involved in inflammation
prostaglandins, leukotrienes, thromboxanes
lipids that can form bilayers
phospholipids and sphingolipids
signaling pathway that activates Akt
PI3K