Microbio Lec 13 EXAM 3
1/10
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
11 Terms
Origin of Earth + Cellular Life
Origin of Earth
is ~4.5 billion yrs old
water appeared ~4.3billion
cellular life MAY have originated @ hydrothermal systems on ocean floor
abundant supply of energy (H2 and HsS) prob found around these sites
geochemistry support abiotic prod of molecules req. for life (amino acids, lipids, sugars, + nucleotides)
Origin of Cellular Life
life may have begun w/ RNA
RNA essential cofactors +molecules
can bind to small molecules
has catalytic activity → may have catalyzed its own synthesis
earliest virus may have evolved from RNA genome cell-like structures
PROTEINS EVENTUALLY REPLACED RNA as catalysts
DNA became genome + template
earliest cells prob had DNA, RNA, protein, + membrane sys. for energy conservation
LUCA existed ~3.8-3.7 billion years ago then bacteria + archaea diverged
Metabolic Diversification: Consequences of Earth’s biosphere
Metabolism of primitive cells was EXCLUSIVIELY anaerobic
obtained carbon from CO2 (autotrophy) → eventually evolved ability to use N2
obtained energy from H2; early chemolithotrophic metabolism would’ve supported prod. of large amnts of organic compounds
phototrophs used energy from SUN to oxidize H2S, H, + H2O to syn. complex organic molecules
1st phototroph were anoxygenic (EARTH was prev. anoxic)
Cyanobacteria → O2 prod → oxygenic phototrophs evolved
phototrophic bacteria (cyanobacteria + chloroflexus) from modern stromatolites (ancient stromatolites contain fossils similar to modern phototrophic bacteria)
Photosynthesis + Oxidation of Earth
~2.5-3.3 billion years ago cyanobacteria evolved a photosystem that could use H2O instead of H2S making O2
allowed evolution life to exploit energy from O2 respiration
respiring O2 was energetically advantageous
O2 reacted spontaneously w/ reducing iron → formed iron oxide instead of accumulating
GREAT OXIDATION EVENT → ~2.4 billion yrs ago
laminated sedimentary rocks → banded iron formations from precipitated iron oxide
atmosphere gradually became OXIC
made ozone shield/ layer(O3)
earth’s surface previously inhospitable
shield allowed organisms to expand range over surface
Living Fossils: DNA Records the History of Life
Phylogeny → evolutionary history of related DNA sequences
CARL WOESE → tree of life
CREATED universal tree of life (PHYLOGENETIC TREE) based on nucleotide seq. similarity in ribosomal RNA (rRNA)
established 3 domain of life (BAE)
root rep. when all life shared LUCA (was likely prokaryotic w/ DNA genome + ability to transcribe + translate proteins)
EX: 60+ genes shared by nearly all cells → must’ve been present in LUCA
Divergence of Eukarya from archaea resulted in membrane enclosed nucleus → organelles gave rise to eukaryotic cell structures
Endosymbiosis Hypothesis
hypothesis for origins of Eukaryotic cells
proposed a series of engulfment events where host cells engulf bacteria
Focus on sequential organelle acquisition (metabolic dependency)
Contend that mitochondria arose from stable incorporation of an aerobic respiring bacterium in the cytoplasm Of early eukaryotic cells
Chloroplasts arose from incorporation of Cyanobacterium-like Cell into cytoplasm of Eukaryotic cell leading to Eukaryotic photosynthesis
Oxygen spurred evolution of organelle containing eukaryotic organisms
Genome structure of mitochondria and chloroplast supports endosymbiosis hypothesis
Formation of Eukaryotic Cell
Is chimeric and made of genes from both bacteria and archaea
Hypothesized Explaination:
1) Serial Endosymbiosis hypothesis → 1st began as nucleus-bearing lining that split from archaea then later acquired mitochondria and chloroplast from endosymbiosis
occurred when line engulfed a bacterial cell that survived and replicated
2) Symbiogenisis Hypothesis → arose from symbiotic relationship between archaea and bacteria; bacteria partner was engulfed to form mitochondria
Hydrogen hypothesis → arose from H2 prod. Bacterium and H2 consuming Achaea
Genes for lipid biosynthesis were transferred from bacterial symbiont to archaeal host
Phylogenetic Tree
Composition and construction that depicts evolutionary history and resembled family tree
composed of nodes and branches
Branch tips → species that exist today
Nodes → ancestor diverged into 2 lineages
Branch length → # of changes that have occurred along the branch
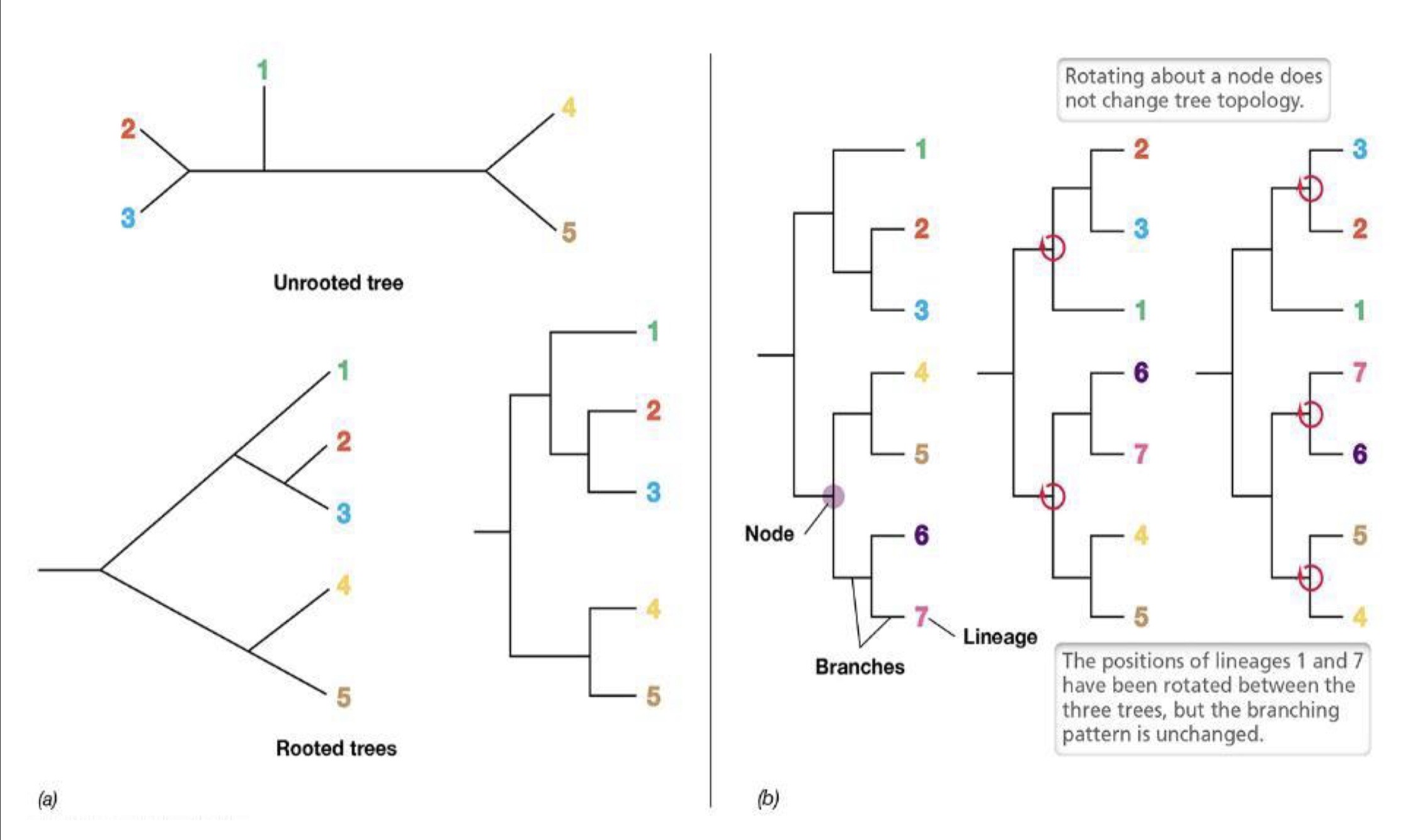
Taxonomic Methods in Systematics
Gene Sequence Analyses
*****SSU rRNA is universal****
functionally constant
Highly conserve (meaning slow evolution)
Adequate length (not too long and short, making it good for study)
SSU rRNA not always useful for distinguishing closely related species
Other highly conservative genes (EX: recA and gyrB) useful for distinguishing at species level
MLST (multilocus Sequence typing)
Another way to differentiate different organism that have similar genes
Method in which several different "housekeeping genes" from an organism are sequenced
housekeeping genes = essential functioning genes
Genome Fingerprinting
Ribotyping → method of identifying microbes from analyzing DNA fragments generated from restriction enzyme digestion of gene encoding SSU rRNA generating a pattern called a ribotype
used in bacterial identification in clinical diagnostics and microbial analyses of food, water, and beverages
Phenotypic Analysis
EX: fatty acid analysis
Types and proportions of fatty acids in cytoplasmic membrane lipids
FAME (fatty acid methyl ester)
Widespread use in clinical, public health, and food/water inspection laboratories to identify pathogens
Relies on variation in composition of fatty acids in membrane lipids for specific prokaryotic groups
Classification and Nomenclature
Taxonomy → how organisms are classified and named
Classification → organization of organism into progressive more inclusive groups on the basis of either phenotypic similarity or evolutionary relationship
Species → 1 to several strains
Genus (genera) → group of several species
How Are Species Named
Prokaryotes given descriptive genus using binomial system of nomenclature used throughout biology
Names for new species and higher groups of prokaryotes regulated by International Code of Nomenclature of Bacteria
New isolates are examined to see if it’s sufficiently different to be a new taxon.
Requires detailed description of characteristics/traits and proposed name
At least 2 international culture collections
IF organism is well-characterized BUT NOT yet cultured, provisional taxonomic name with Candidatus can be used
Bergey’s Manual of Systematic Bacteriology most widely accepted classification of prokayotes
no official classification released yet
Culture Collections
National microbial culture collections are an important foundation.
• catalog and store
• protect diversity
• store viable cultures (frozen or free-dried)
• act as repositories for type strains