BSCI410 Exam 3 + Final Stuff
1/103
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
104 Terms
Transposable Elements
DNA that has evolved the ability to move from place to place within a genome
→ “Jumping genes”
Can be highly disruptive to genes and chromosomes, yet also drive evolution
Don’t jump all the time. Movement can be activated by stressors like heat shock, radiation, tissue culturing, pathogen infection, or cancer.
They have a major impact on shaping genome structure and function
Effects of Transposable Elements
Genetic instability and disease
→ TEs can cause deleterious mutations, disrupt gene functions, and contribute to cancers or genetic disorders
Gene Regulation
→ TEs often contain regulatory sequences (promoters, enhancers) that when inserted close to genes, can rewire gene regulatory networks
Chromosome rearrangements
→ High concentrations of TEs can lead to unequal crossing over, causing deletions or duplications
Genetic Variability and Evolution
→ Through TRANSPOSITION (movement of TEs in the genome), TEs increase genetic variability, which can help populations adapt to environmental changes
Epigenetic Modification
→ Organisms use silencing mechanisms (DNA Methylation, histone modifications) to repress TEs. But this can silence the expression of genes close to the transposon.
TEs Background
Existence of TEs was inferred from genetic studies in Maize (Corn mottling was caused by movements of a TE into and out of a pigment gene)
Unrecognized for many years because DNA was believed to have fixed positions on chromosomes
TEs were thought to be “selfish” DNA. Have now been found in all organisms.
12.5% of the Drosophila genome are TEs
90% in Maize
TE Abundance in Human Genome
44% of Human Genome
Made of Retrotransposons
-LINEs: 20%
-SINEs: 13%
-HERVs: 8%
and DNA Transposons: 3%
Only a few LINEs and SINEs in the human genome are able to move due to mutations in the sequences needed for transposition
The HERVs and DNA transposons in the human genome are immobile relics.
DNA Transposon Mechanism
Deletion and then goes into the target site
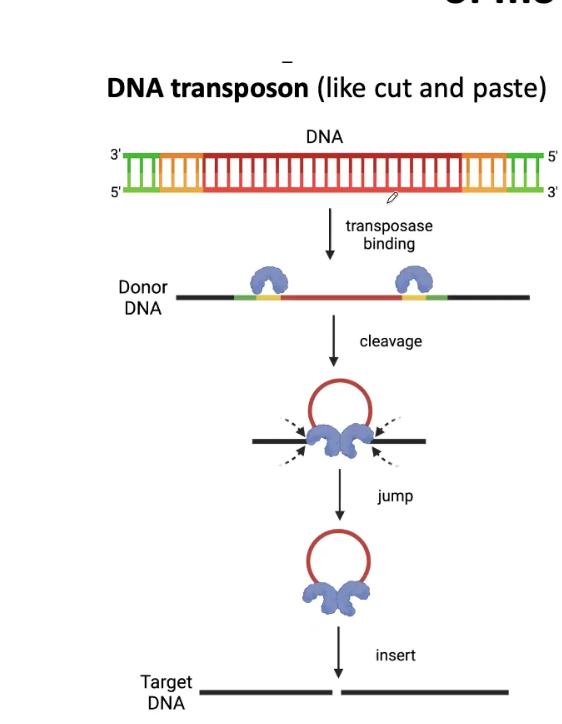
Retrotransposon Mechanism
Copies itself and then goes into target site
Can grow in quantity across the genome
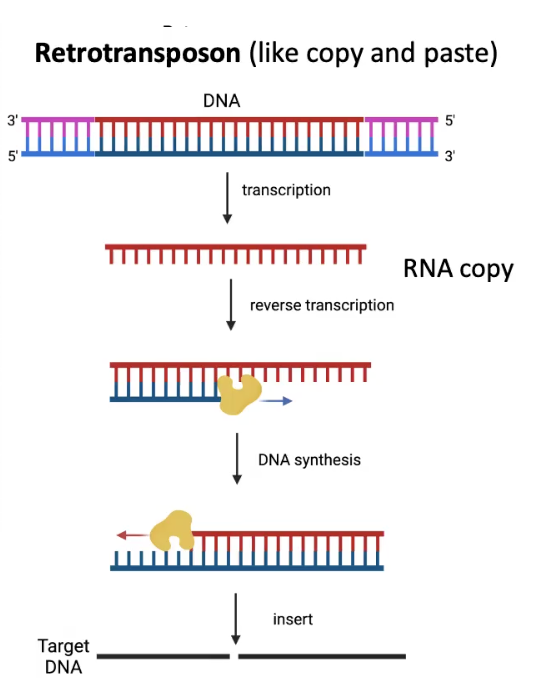
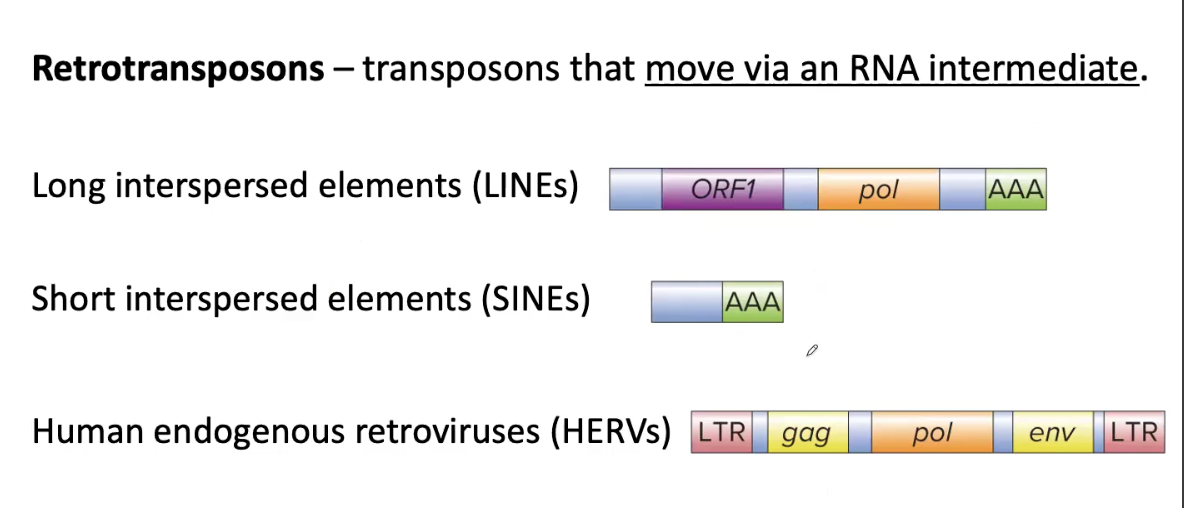
3 Types of Retrotransposons
Long Interspersed Elements (LINEs)
Short Interspersed Elements (SINEs)
Human Endogenous Retroviruses (HERVs)
LINEs and HERVs have a gene encoding REVERSE TRANSCRIPTASE called pol that transcribes mRNA into single-stranded cDNA
HERVs are similar in structure to retroviruses (RNA tumor viruses) and are flanked by LONG TERMINAL REPEATS (LTRs)
Retrotransposons: Transposons that move via an RNA INTERMEDIATE
How LTR (Long Terminal Repeat) Retrotransposons Move
Transcription of retrotransposon pol
Synthesis of cDNA by REVERSE TRANSCRIPTASE
Staggered cut is made in the genomic target site
Retrotransposon cDNA inserts into the target site
Original copy in the genome remains while new copy inserts into the other genomic location
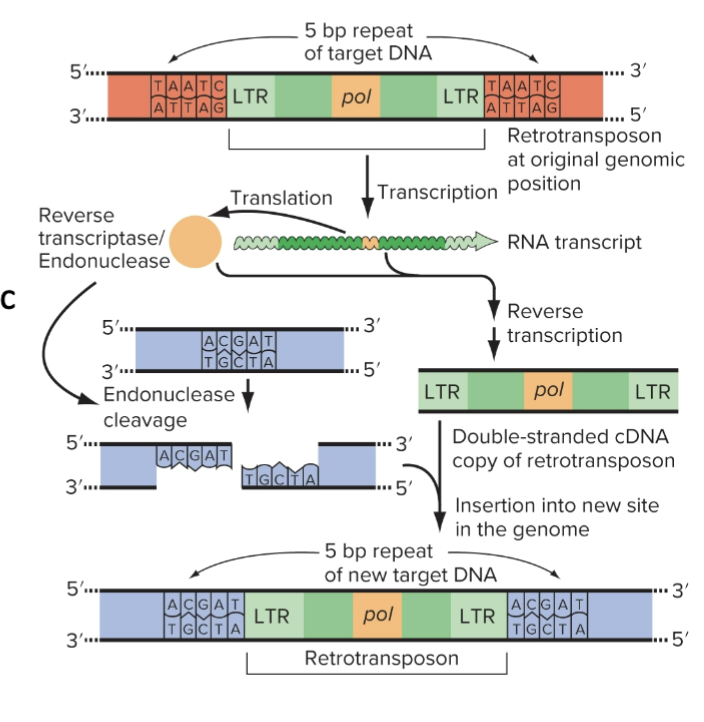
DNA Transposons
Most have INVERTED REPEATS (IRs)
→ 10-200bp long at each end
and a gene encoding TRANSPOSASE gene, which recognizes the IRs and cuts at the border between IR and genomic DNA
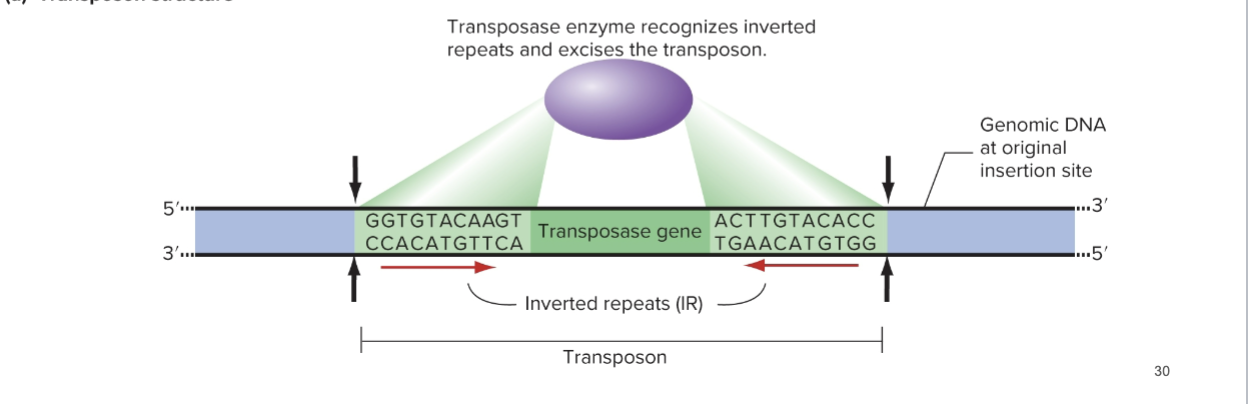
How DNA Transposons Move
Transposase cleaves the P-element (Inverted Repeats + Transposase gene in the middle) and excises it
Then adds it to a new location
Repair of the original gap using a sister chromatid or homologous chromosome occurs. The DNA Transposon can either remain or be removed from its original location depending on whether it exists on the homologous chromosome or sister chromatid.
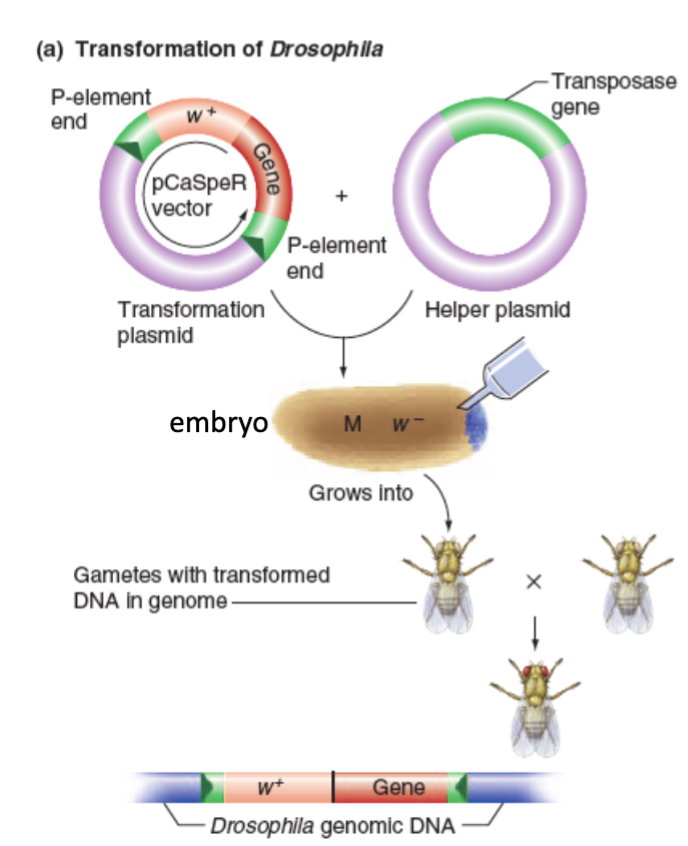
Modified P Elements to Modify Flies
Insert the desired DNA between the P element Inverted Repeats of a transformation plasmid (no transposase gene here)
Then insert a helper plasmid that encodes for the transposase gene. Thus, transposase is formed and can move the first plasmid’s segment, but not itself since it lacks IRs of its own.
Inject both into syncytial embryos.
Defective TE Copies
Many TEs end up with deletions during the transposition process
→ Deletion of PROMOTER for retrotransposon transcription
→ Deletion of REVERSE TRANSCRIPTASE/TRANSPOSASE GENE
→ Deletion of INVERTED REPEATS needed for DNA Transposons
Creates disparity between Autonomous TEs and Nonautonomous TEs
Autonomous TEs
Non-deleted TEs that can transpose on their own
Nonautonomous TEs
Defective TEs that require the activity of the autonomous enzymes in order to move (i.e. the sequence of interest in the drosophila transformation plasmid, which needs help from transposase in another plasmid)
How TEs disrupt genes and alter genomes
TE can insert within the coding region of a gene, inactivating it
TE can insert near a gene and affect its expression pattern
Alleles associated with TEs can be unstable
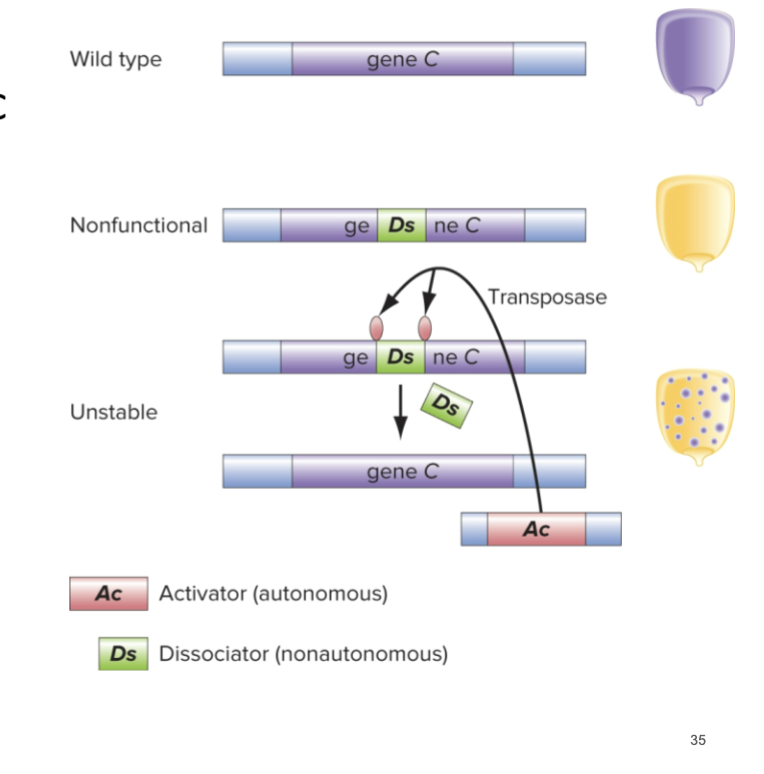
TE-Associated Mutant Alleles
Can often be unstable
A non-autonomous TE (Ds in picture) can hop into gene C and disrupt function
AC is an autonomous TE that cleaves out Ds and restores function
TEs altering genomes
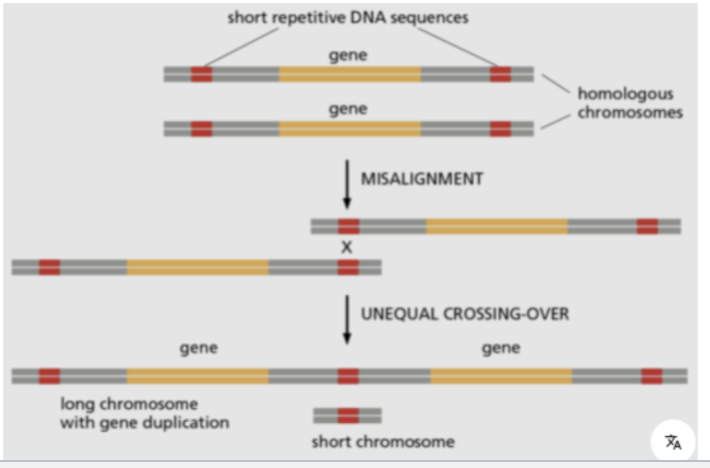
TEs can also trigger spontaneous chromosomal rearrangements by unequal crossing over between TEs
→ Misalignment of homologous chromosomes or sister chromatids during meiosis/mitosis
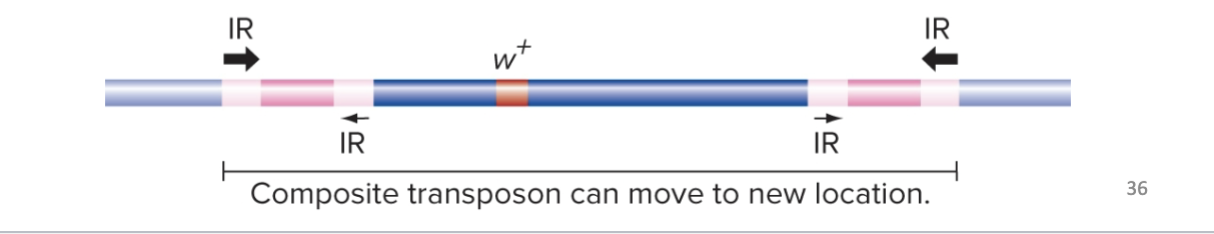
Transposition can also cause relocation of a gene in the genome
→ Can occur when two copies of a TE integrate in nearby locations on the same chromosome
→ Then the transposase recognizes the outermost IR sequences and moves the intervening sequences to a different location
Gene Duplication caused by TEs
RETROTRANSPOSONS can cause gene duplication
By mistakenly reverse transcribing cellular mRNAs and inserting them somewhere into the genome
OR by unequal recombination between repeated sequences due to misalignment (unequal crossing over) during meiosis
New Functions acquired by Duplicated Genes
Gene Families: Sets of related genes with slightly different functions; most likely arose from gene duplications
→ The two copies of a duplicated gene are related, but they can diverge in sequence and function
→ Some gene families in vertebrates have HUNDREDS of members thought to be driven by TEs
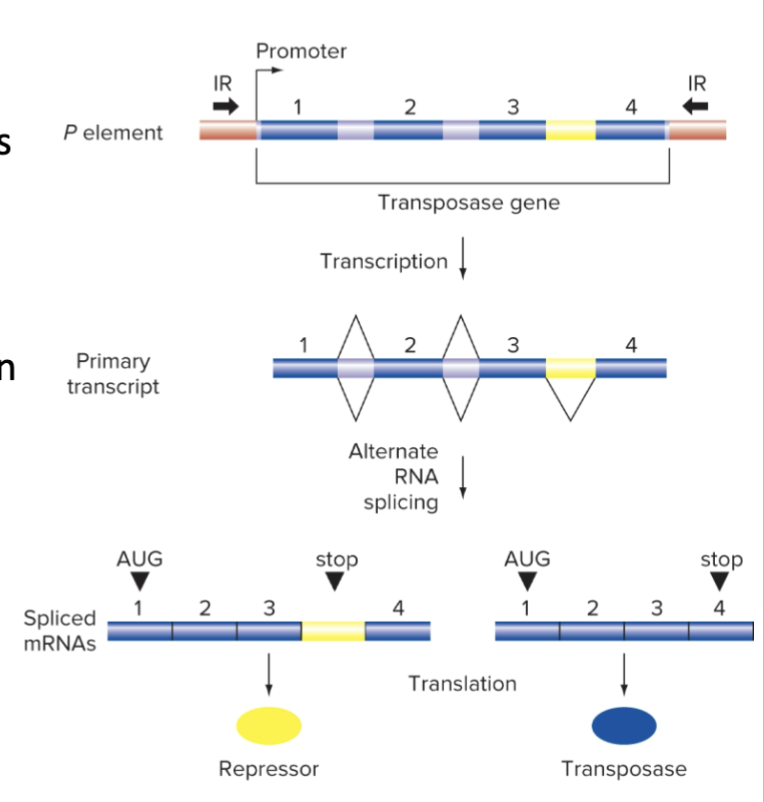
Alternative Splicing by Transposase Gene limits TE Movement
One splice form produces TRANSPOSASE
The other produces a transposition REPRESSOR
The repressor COMPETES with transposase for binding to the Inverted Repeats (IRs)
The Sequence Hypothesis
DNA, RNA, and Proteins are COLINEAR
→ They all run only in one direction. If we know the sequence of DNA, the RNA and Amino Acid sequence can be identified.
Different Types of RNA Polymerases
RNA Pol 1 is typically for Ribosomal RNA
RNA Pol 2 is for mRNAs, lnc(Long NonCoding)RNAs, and miRNAs
RNA Pol 3 is for tRNAs, U6 sn(Small Nuclear)RNA, and SS rRNA
Plants have two additional RNA pols (IV and V) which Synthesize siRNA
RNA Polymerases do not require a primer, unlike DNA Polymerases.
→ Can do “de novo” synthesis from a DNA template.
→ Adds new ribonucleotides to the 3’-OH of the growing RNA chain
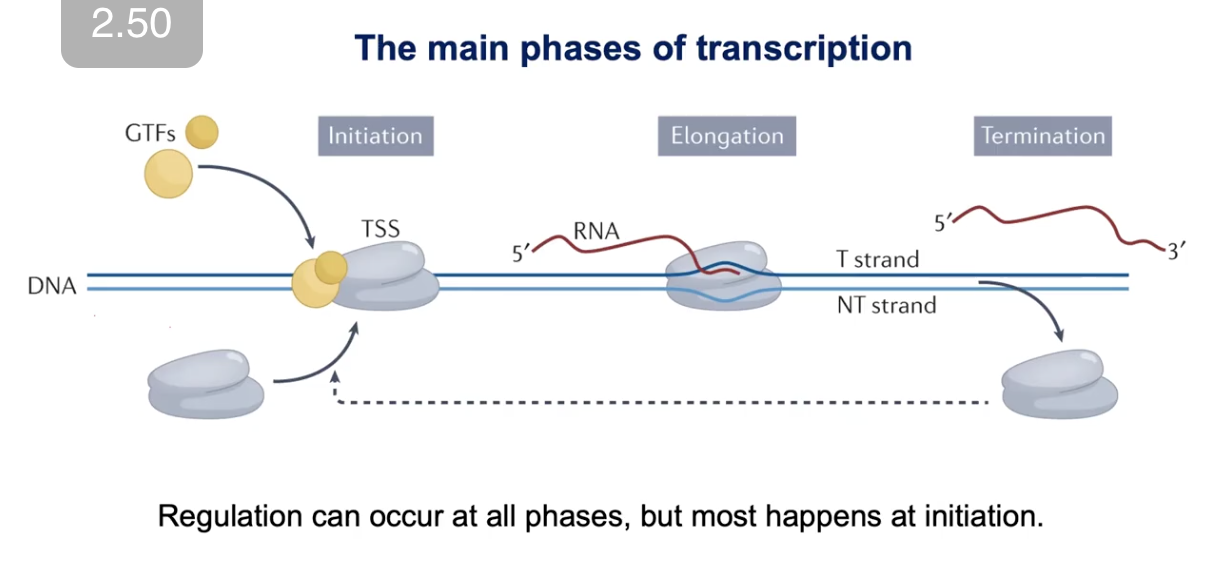
3 Main Phases of Transcription
Initiation
When RNA polymerase enzyme gets loaded onto the core promoter
→ Involves GERMINAL TRANSCRIPTION FACTORS (GTFs)
Elongation
Termination
*Regulation can happen at all phases, but mostly happens at Initiation
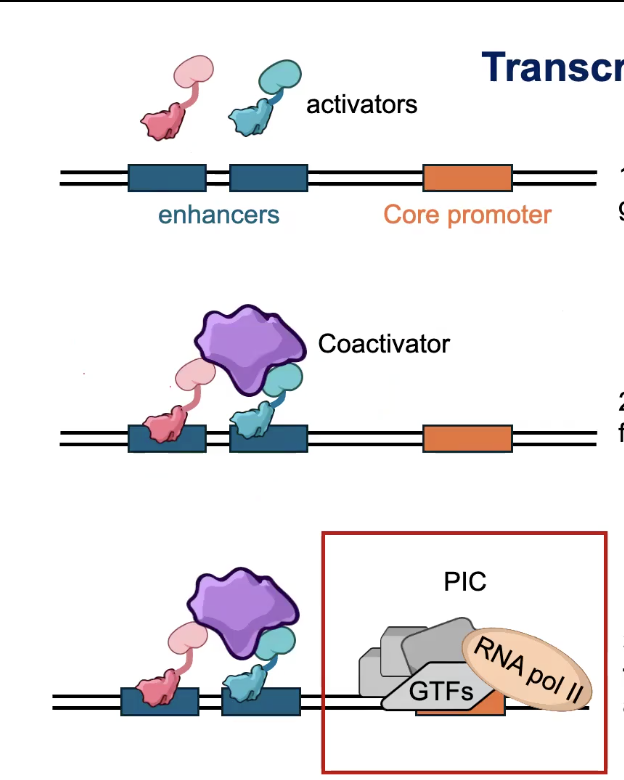
Steps of Initiation
Activator Binding
→ Transcription factor proteins bind to gene-specific cis-regulatory regions (like enhancers)
Coactivator Recruitment
→ Activators (are TFs, and don’t work alone) and recruit intermediary factors known as CO-ACTIVATORS
Pre-initiation Complex (PIC) Assembly
→ General Transcription Factors (GTFs) and RNA Polymerase II assemble into the pre-initiation complex at the CORE PROMOTER
Core Promoter
Contains specific sequences known as PROMOTER ELEMENTS that bind General Transcription Factors (GTFs).
No universal element. For example, only 24% of human promoters have a TATA box.
Often have multiple elements cooperate to control a single gene.
At active genes, the core promoter is in a NUCLEOSOME-FREE REGION.
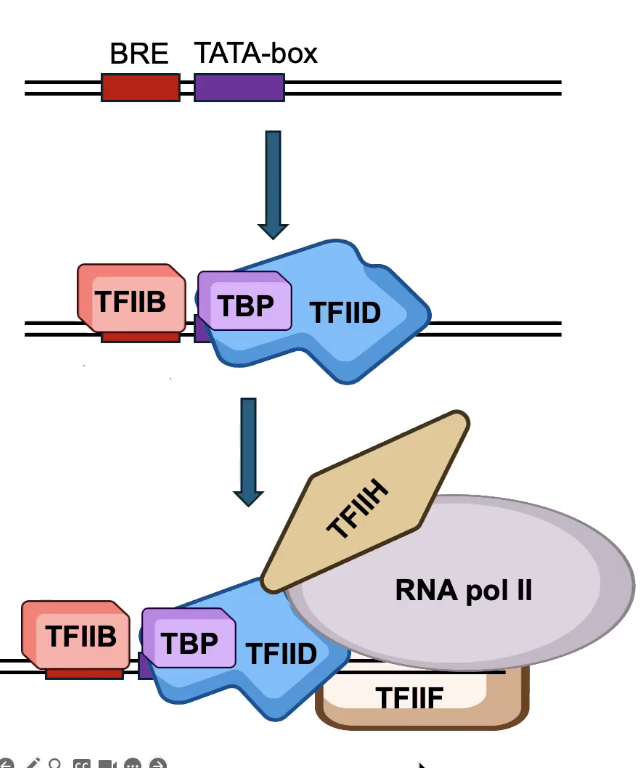
General Transcription Factors (GTFs)
Bind to the core promoter and load RNA polymerase.
TBP (TATA Box-binding Protein), a subunit of TFIID, binds the TATA box.
TFIIB binds the BRE element.
TFIIB and TFIID recruit the rest of the Pre-Initiation Complex (PIC)
→ Includes TFIIF, which loads RNA Pol II (TFIIF is a stabilizing factor for RNA Pol II)
→ Also includes TFIIH. The HELICASE subunit of TFIIH melts the promoter DNA and unwinds it to allow RNA Pol II to function.
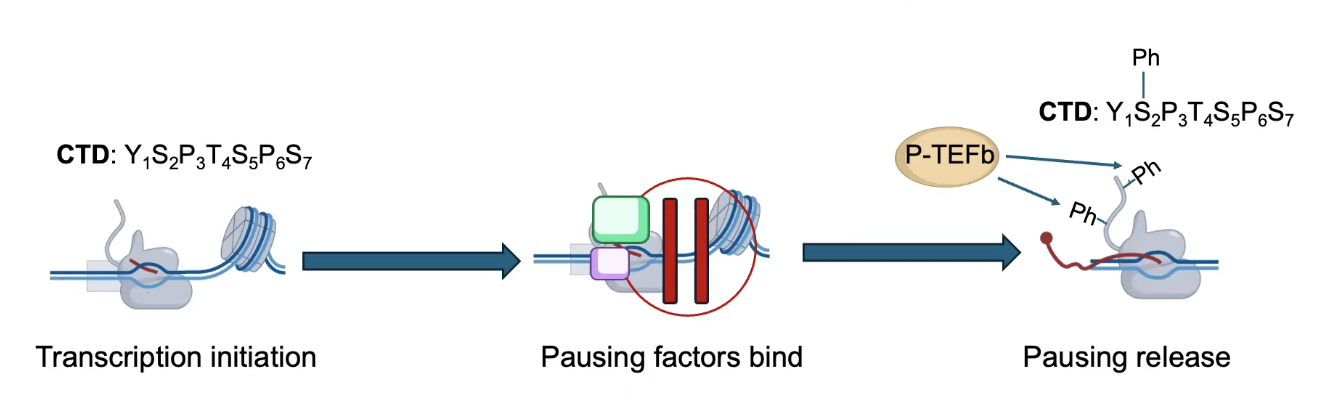
Elongation can be regulated by RNA Pol II Pausing
Elongation factor P-TEFb phosphorylates the CTD of RNA Pol II to PROMOTE elongation. Releases the pausing of RNA Pol II, starts it again.
→ RNA Pol II pausing is used to load RNA Pol molecules on genes for RAPID INDUCTION.
→ Used for loading RNA Pol II before the RNA might be needed, thus allowing transcription to start quickly again once the mRNAs are actually needed.
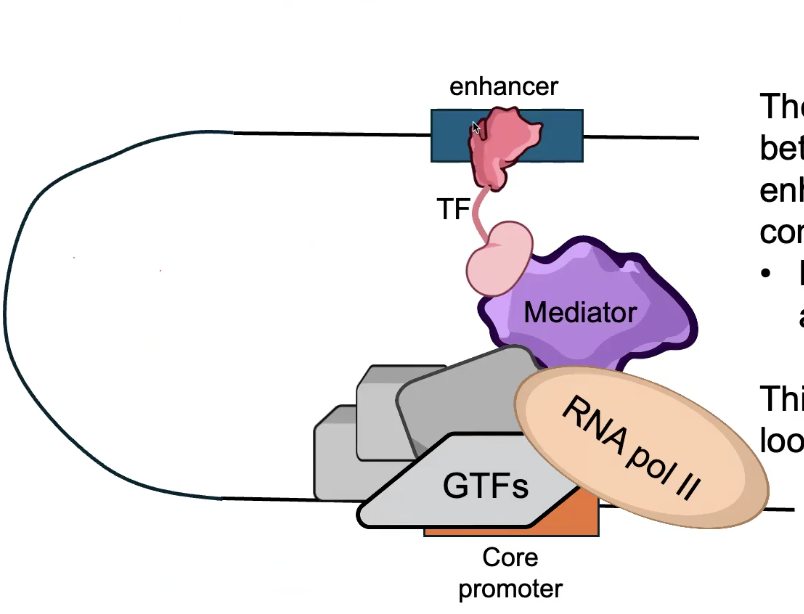
DNA Looping
Can Mediate LONG-RANGE interactions between CORE PROMOTER and distal CIS-REGULATORY SITES
Mediator complex forms a bridge between TFs bound at enhancers, and the Pre-Initiation Complex (PIC) bound at the CORE PROMOTER.
→ Mediator has 30 subunits in animals.
This interaction occurs through looping of intermediate DNA.
Transcription Factors
Are DNA binding proteins that control the rate of RNA Synthesis
Are SEQUENCE SPECIFIC
Are often MODULAR Proteins:
→ DNA BINDING DOMAIN directly interacts with a specific DNA sequence
→ DNA binding domains contain highly conserved regions
→ TFs can be grouped into FAMILIES by the type of DNA binding domain they contain
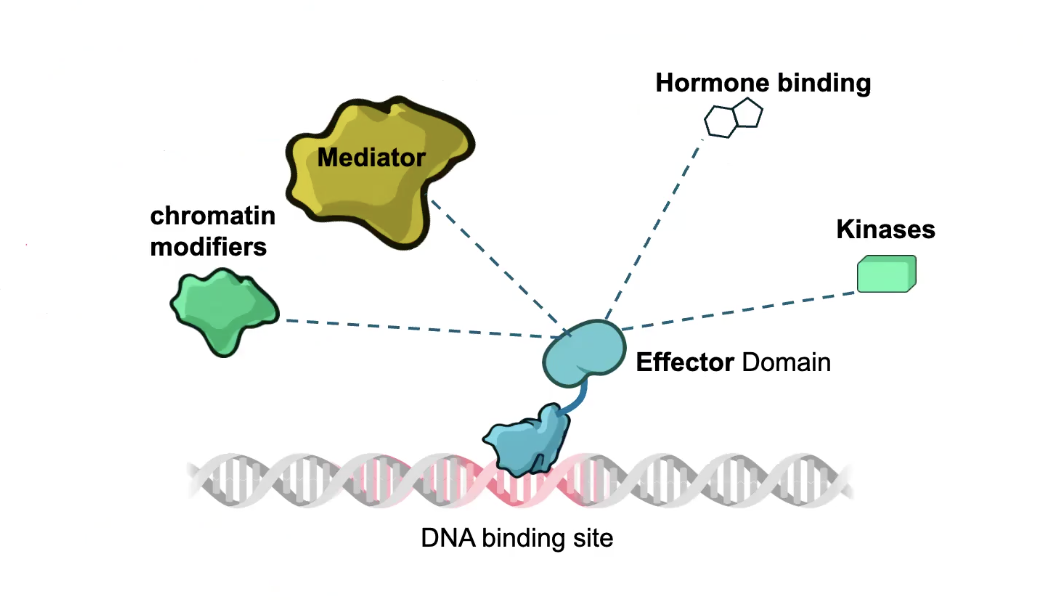
→ EFFECTOR DOMAIN alters the rate or probability of transcription (ACTIVATOR domain or REPRESSOR domain)
→ Are less well conserved than DNA binding domains
*The more complex an organism is, the more TFs they tend to have
Helix-Loop-Helix Transcription Factor Example
The HOMEODOMAIN is a highly conserved DNA binding domain found across eukaryotes
→ Many major TFs that regulate development contain Homeodomains (like HOX proteins)
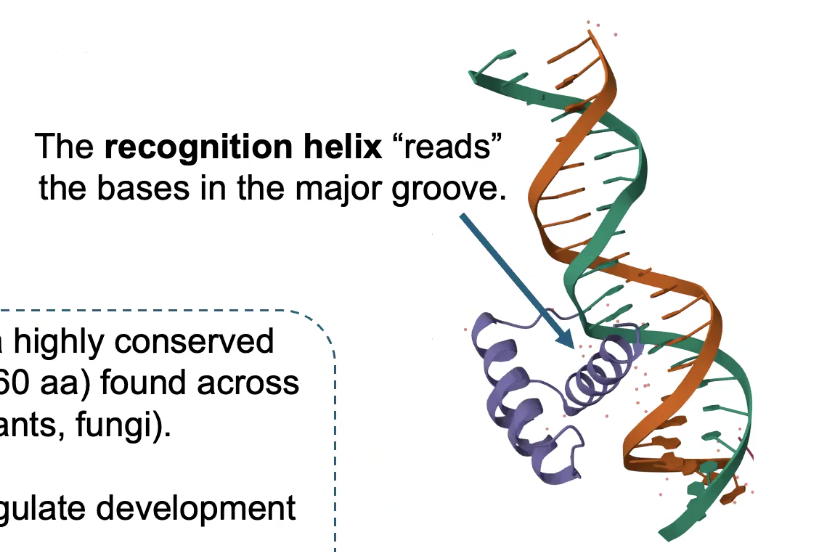
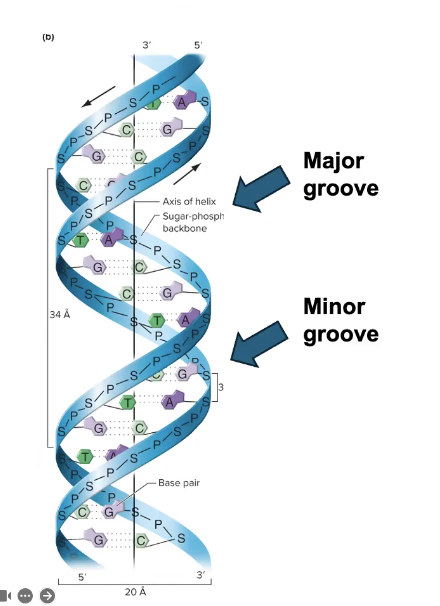
How TFs “read” DNA Sequences
TFs make SPECIFIC interactions w/DNA
→ Direct interactions between specific amino acids and bases in the MAJOR GROOVE
Major groove is larger and has more opportunities for interactions and bonds to occur
→ Different minor groove interactions are not unique between some nucleobase combinations, thus not good for reading DNA
TFs as Multimers
Some TFs bind DNA as dimers, trimers, tetramers, etc.
i.e. p53 is a tumor suppressor protein
→ Is the most commonly mutated gene in human cancers
→ and is a TETRAMERIC transcription factor
Chromatin
In the eukaryotic nucleus, DNA is packaged together with proteins to make CHROMATIN.
Packaging is dynamically controlled, and chromatin compaction is therefore not homogenous in the nucleus.
→ Regulated by specific enzymes.
→ CHROMATIN REMODELING ENZYMES and HISTONE MODIFYING ENZYMES
Nucleosome
A basic unit of chromatin, consisting of ~145bp of DNA wrapped around an octamer of histone proteins
Histones: Highly conserved BASIC (positively charged) proteins that bind to DNA
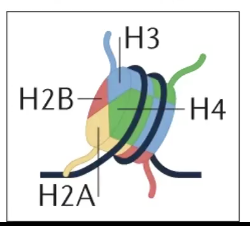
→ Core histones include H2A, H2B, H3, and H4
→ Made of 2x H2AH2B dimers, and 1x H3H4 tetramer
Flexible N-Terminal tails extend out of the core octamer
→ Are subject to EXTENSIVE PTM
Histone proteins block the initiation of transcription (if at the promoter), but NOT elongation.
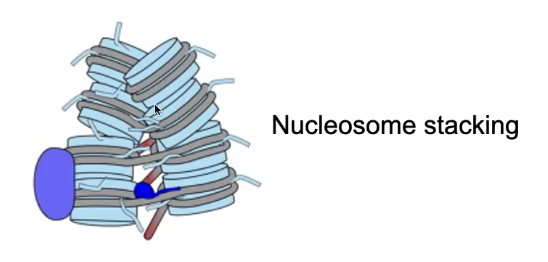
Nucleosome Stacking
Is favored because of interactions between the negatively charged DNA and the positively charged N-Terminal tails of the Histones (Lysine-rich)
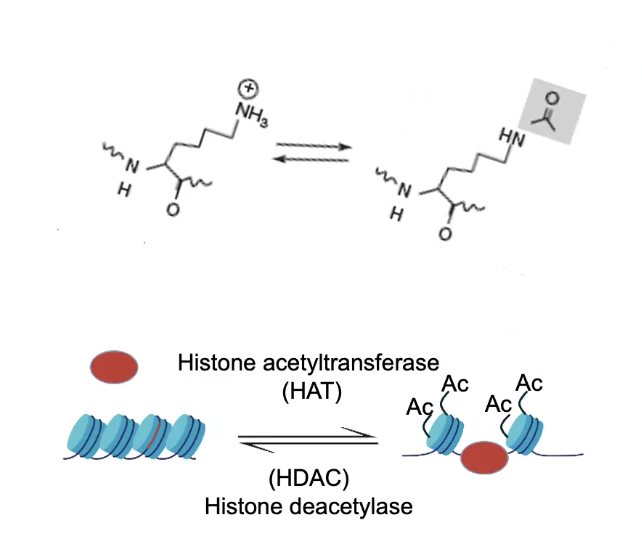
Histone Acetylation
Associated with Transcriptional Activity
Is controlled by HISTONE ACETYLTRANSFERASE (HAT) and HISTONE DEACETYLASE (HDAC) enzymes
Compaction occurs when the Positive Lysine residues on the N-Terminal tails of the Histones interact with the negatively-charged DNA
→ Addition of an acetyl group neutralizes the positive charge on lysine, uncompacts DNA
→ Thus opens up DNA and allows transcription to occur
Acetylation can recruit additional proteins
→ Some protein domains are “readers” of histone modifications. I.e. BROMODOMAINS (BD) which bind acetylated lysine residues
→ CBP is a TF and a HAT (therefore a writer) that also has a BD (a reader)
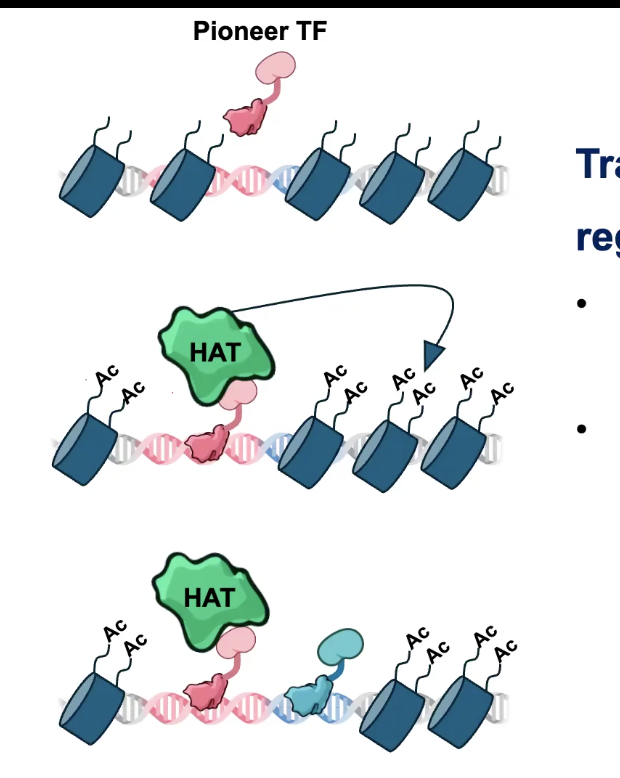
TFs and Histone Acetylation
CBP being both a reader and writer allows it to spread and maintain acetylation on chromatin beyond where the TF initially binds
However, CBP is a COACTIVATOR and has no sequence specificity. Thus, needs to be recruited by an ACTIVATOR, another TF called DORSAL (Drl)
Drl recruits CBP to drive transcription of genes.
Dorsal (Drl) can also act as a repressor
Alone, Drl acts as an activator.
When Drl binding occurs in conjunction with Dead Ringer (Dri) binding, the two proteins recruit GROUCHO, a transcriptional REPRESSOR protein
Thus, Drl can act as both an activator and a repressor TF.
TF Cooperativity
TFs can act cooperatively to regulate genes
→ Can assist the binding of additional TFs by remodeling the surrounding chromatin
PIONEER TFs are a special class of TF that can “open” Chromatin and pave the way for other TFs
Effector Domain receives signals to control TF Action
These signals can be HORMONES, which are perceived by NUCLEAR RECEPTOR proteins
→ Nuclear Receptor proteins are both hormone receptors AND TFs
i.e. Cortisol, which interacts with the Ligand Binding Domain of GR
→ GR gets freed and can then enter the nucleus
→ The GR DNA binding domain (DBD) interacts with cis-regulatory elements known as GLUCOCORTICOID RESPONSE ELEMENTS (GREs)
Chromatin Immunoprecipitation with sequencing (ChiP-Seq)
An affinity purification based method allowing us to map the genomic binding sites of chromatin proteins.
→ To find out where TFs bind in the genome
Protocol:
1) Cross-link/fix samples with Formaldehyde
2) Lyse cells and isolate Chromatin
3) Fragment Chromatin into smaller pieces (100-600bp) (using Sonication or Enzymatic Digestion)
4) Affinity purify target using an antibody (Immunoprecipitation)
5) Isolate and sequence DNA
ChiP-seq can also be used to map the genomic locations of any chromatin-bound protein like Histone modifications. You just need an antibody.
ChiP-Seq revealed that TFs can have hundreds-thousands of binding sites in the genome
→ Identifying DNA patterns of TF binding motifs is called MOTIF ANALYSIS
Cell Differentiation
TF cascades generate patterns of gene expression
TF proteins can act hierarchically. A MASTER/INITIATOR TF can activate or repress downstream TFs.
The assortment of TF proteins in a cell determines the expression state of genes, and therefore cell structure and function.
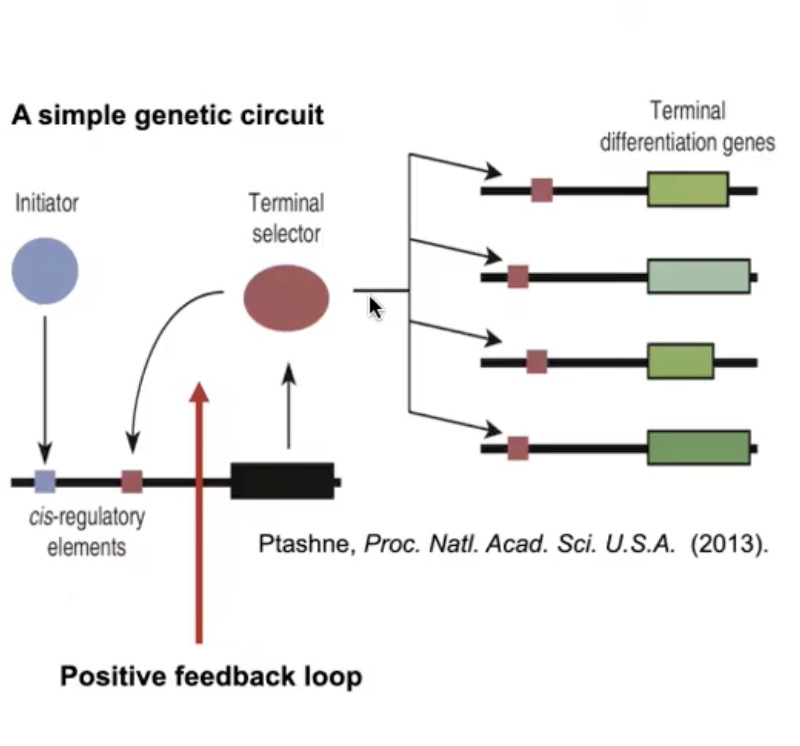
Positive Feedback in Transcriptional Regulation
An initiator TF activates expression of a terminal selector TF
The terminal selector TF activates downstream differential genes, but ALSO binds to cis-regulatory cites upstream of ITS OWN gene.
This simple genetic circuit maintains a pattern of gene expression that PRESERVES CELL FATE
Gene Regulatory Networks
Small genetic circuits are combined to make Gene Regulatory Networks
A network of regulatory actions make a Program of Gene Expression
RNA Processing
The nascent pre-mRNA must be processed before it’s ready for translation
→ Occurs CO-TRANSCRIPTIONALLY (occurs simultaneously with transcription)
→ ONLY mature mRNA is exported to the cytoplasm for translation
The coding sequence in Eukaryotes is interrupted by non-coding INTRONS. RNA splicing removes introns from the message.
→ Whereas in bacteria it’s typically ready for translation immediately
The now-MATURE mRNA contains an uninterrupted coding sequence sandwiched between two non-coding UnTranslated Regions (UTRs).
→ The UTR’s function is in translation, mRNA localization, and mRNA Stability (like mRNA metadata)
3 Key Steps of RNA Processing
5’ Capping
RNA splicing
3’ cleavage and poly-adenylation
RNA Pol II C-Terminal Domain (CTD)
The CTD is a platform for recruiting RNA processing enzymes
→ Consists of tandem repeats of the hepta-peptide YSPTSPS
→ The CTD is phosphorylated at different residues over the course of transcription. Called the CTD CYCLE
5’ Capping Enzyme
One enzyme recruited by the CTD is the 5’ capping enzyme
→ adds a 7-METHYLGUANOSINE (m7G) cap to the 5’ end of the pre-mRNA
The 5’ cap is crucial for:
mRNA Stability
mRNA Exporting
mRNA Translation
CAP BINDING PROTEINS interact with the 5’ cap
DECAPPING ENZYMES can remove the 5’ cap to trigger mRNA degradation
RNA Splicing Details
There are conserved sequences at the 5’ and 3’ ends of introns that are recognized as SPLICE SITES
→ By enzyme called the SPLICEOSOME
The 5’ end of the intron is cleaved and then reacts with the -OH of the BRANCHPOINT ADENINE (occurs within the intron itself)
The free 5’ splice site is then joined with the 3’ splice site to LINK the two exons.
The circularized intron is released as an INTRON LARIAT
→ Can be degraded by the cell
Spliceosome
Catalyzes RNA Splicing (removal of introns)
Has a core of 5 Small Nuclear RiboNucleoProteins (snRNPs)
Each snRNP is a complex of proteins and a small nuclear RNA (snRNA)
→ snRNAs form the catalytic core of the spliceosome.
*The spliceosome is an example of a RIBOZYME (an RNA enzyme)