L3: Taking the Measure of Microbial Ecosystems
1/29
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
30 Terms
What is a microbial community?
A group of microbial populations co‑existing and interacting in the same environment.
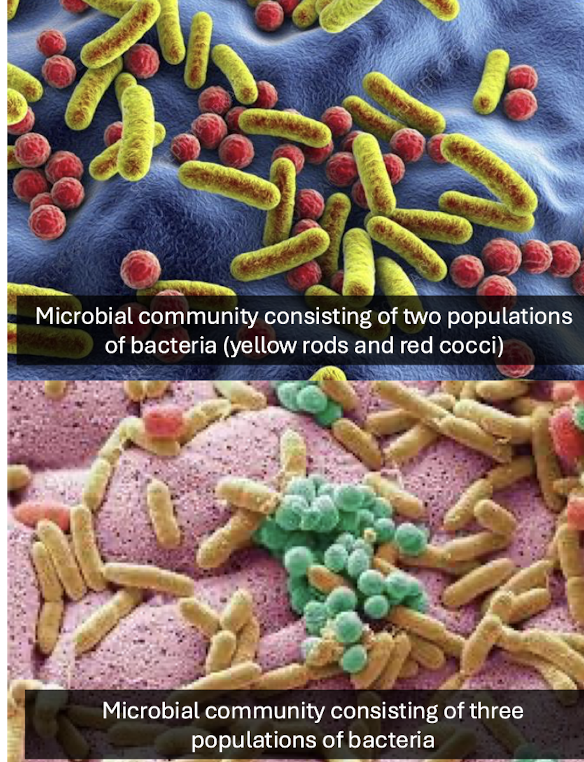
What are the two major components of microbial ecology?
Biodiversity (“who is there”) and microbial activity (“what they are doing”).
Why is microbial ecology important?
It explains how populations assemble, interact, and influence environmental processes.
What is enrichment culture?
Use of selective media/conditions to favour growth of a desired organism.
What do you need for enrichment?
Appropriate inoculum
Selective medium
Specific incubation conditions
Example:
Selecting aerobic N₂‑fixing bacteria like Azotobacter by removing NH₄⁺/NO₃⁻ from medium.
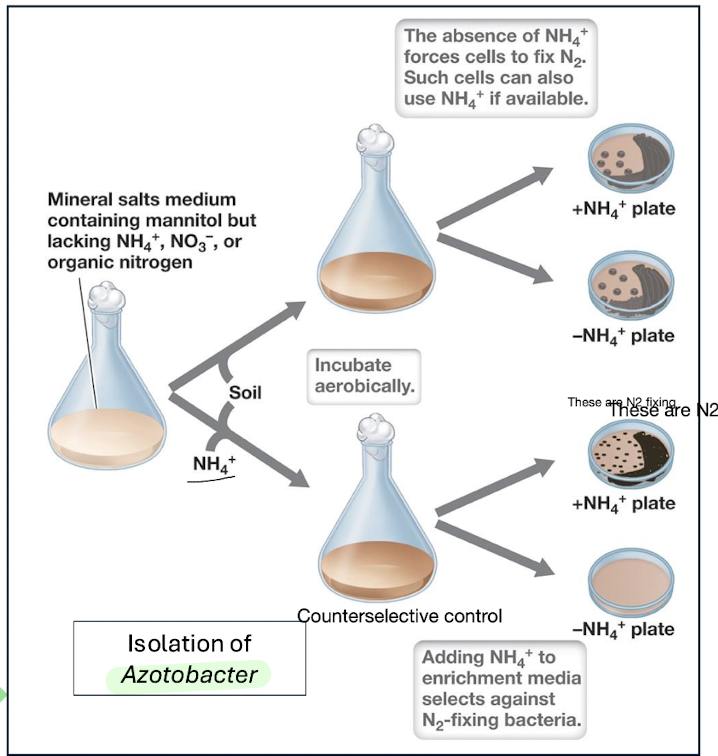
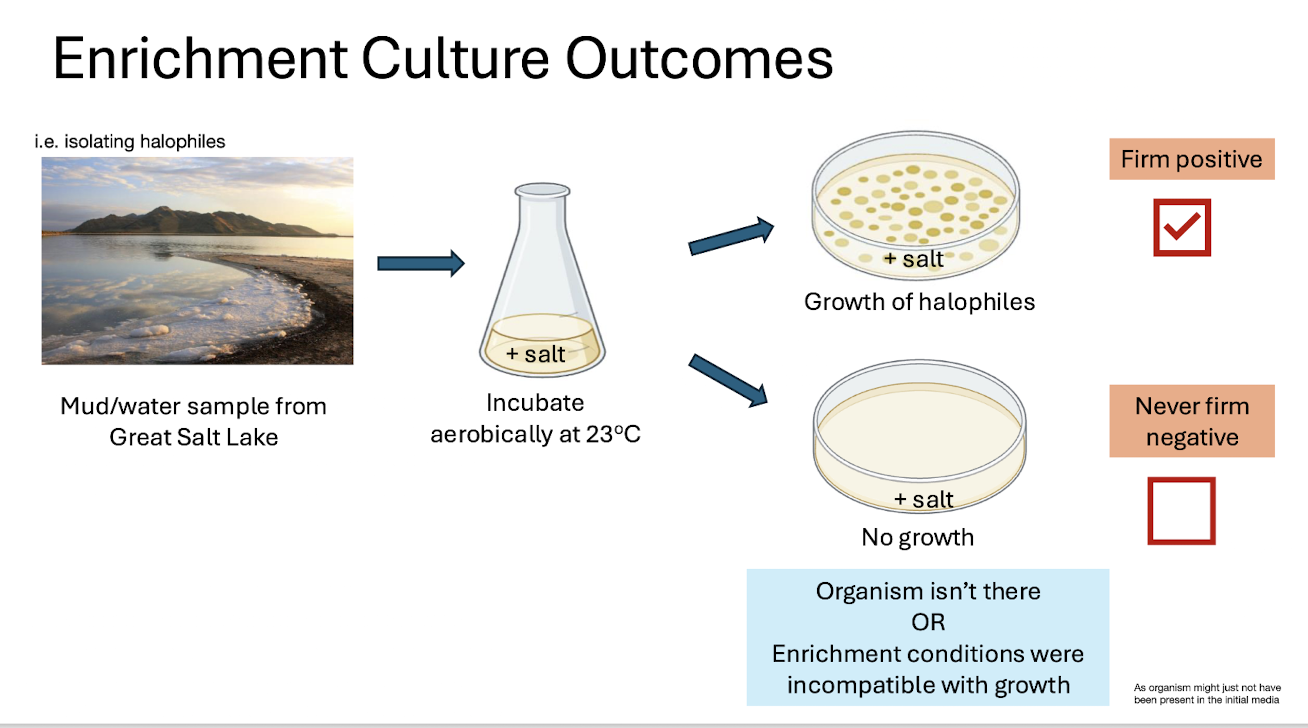
Enrichment Outcomes
Firm positive: Growth under selective conditions.
Never firm negative: Absence of growth may mean organism absent OR conditions unsuitable.
What is enrichment bias? and how can it be avoided
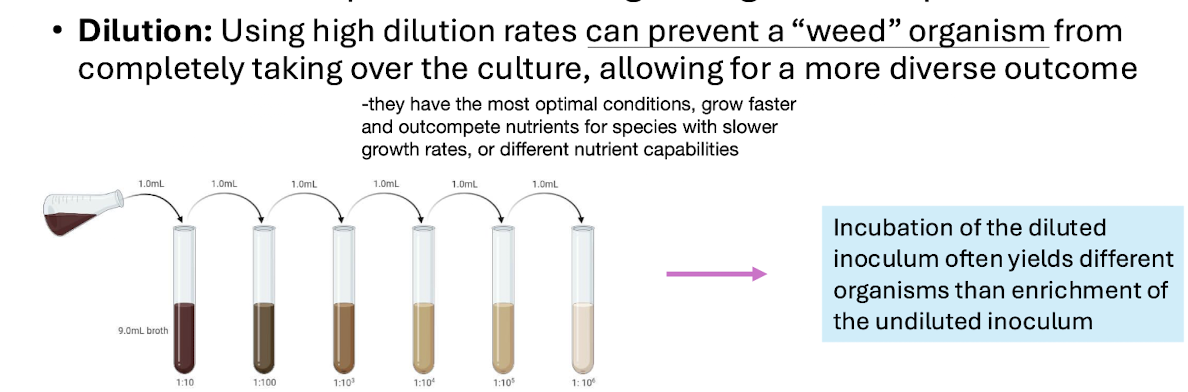
Favouring fast‑growing “weed” species; does not reflect natural abundance.
Dilution helps reduce dominance of weed species.

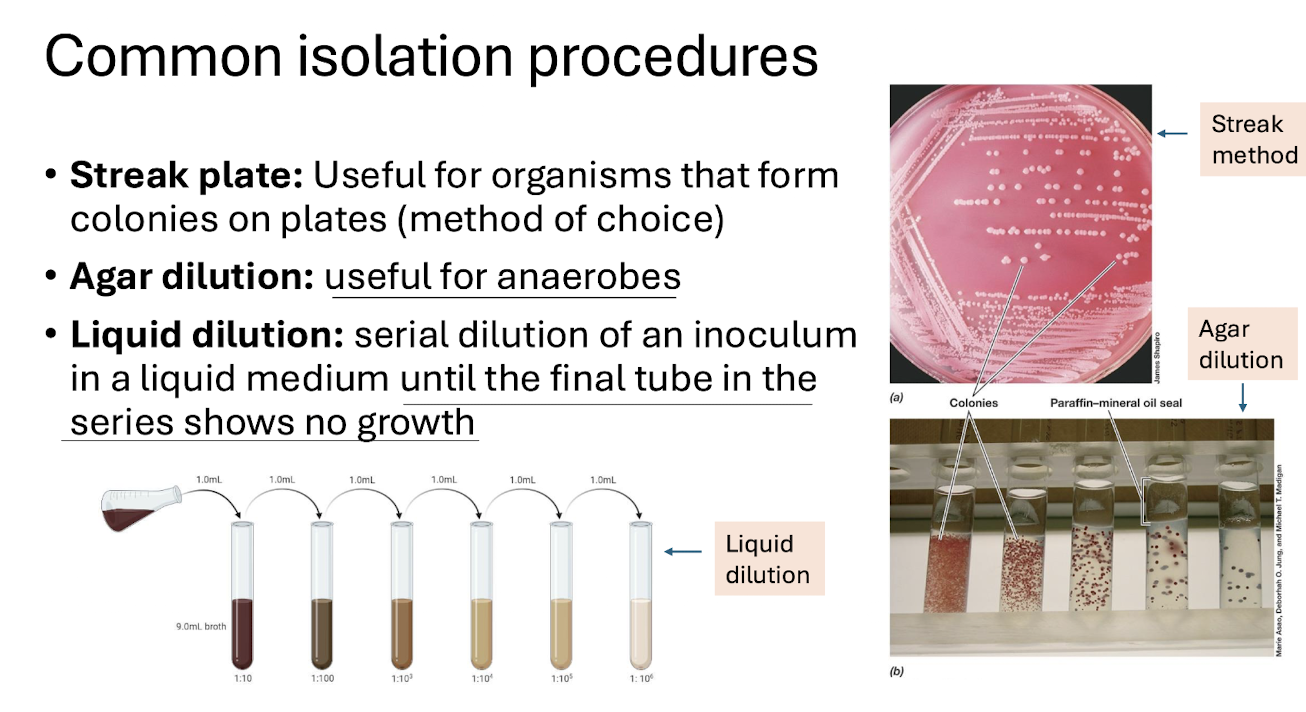
What are the 3 main classical isolation techniques?
Streak plate — isolates colonies on solid media.
Agar dilution — useful for anaerobes.
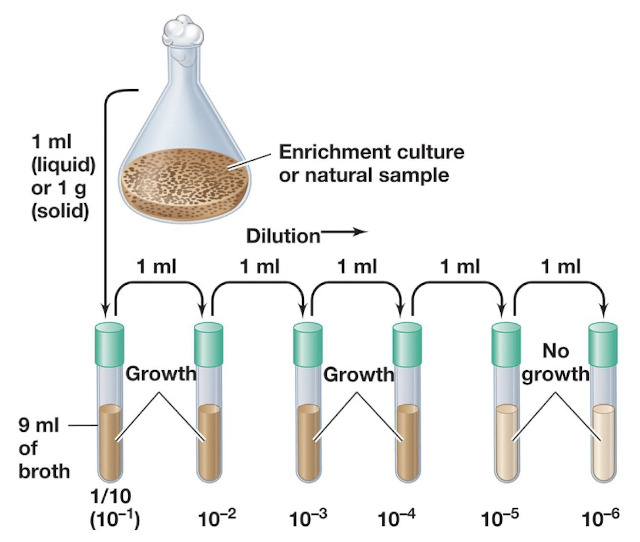
Liquid dilution — serial dilution until no growth.

What is MPN? (Most Probable Number)
A statistical method estimating microbial numbers via serial dilution in liquid media.
Key idea: Last tube showing growth likely originated from ≤10 cells
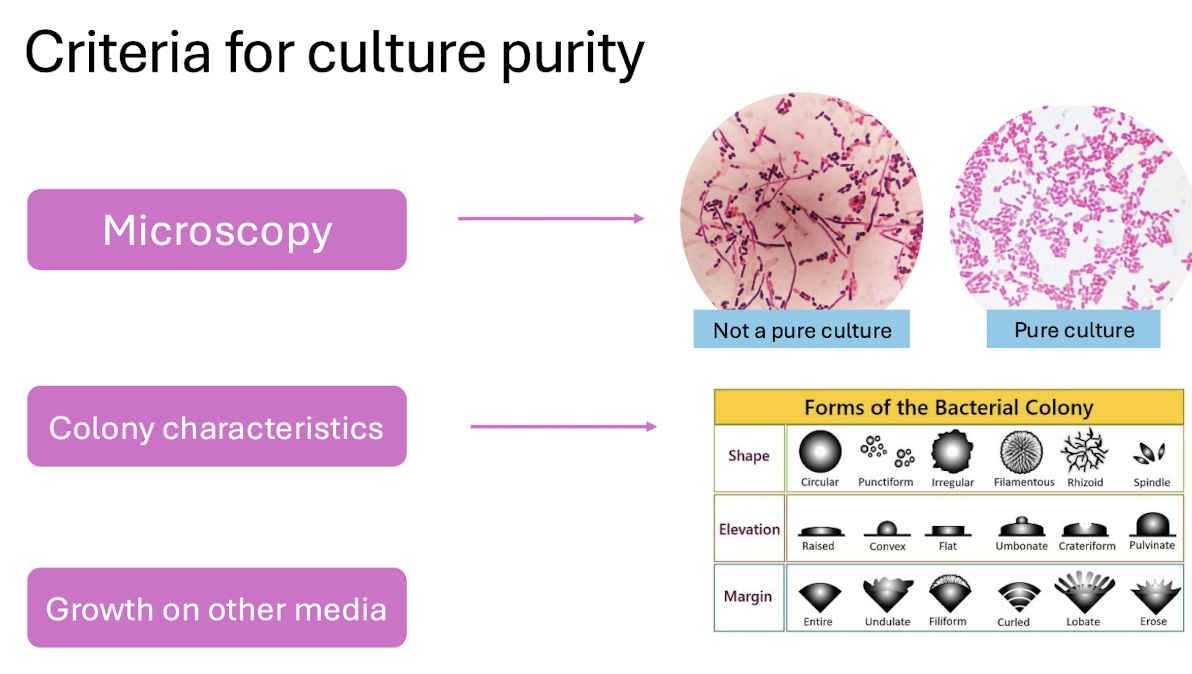
Criteria for Culture Purity
Microscopy (single morphology).
Colony characteristics (shape, margin, elevation).
Growth on other media.
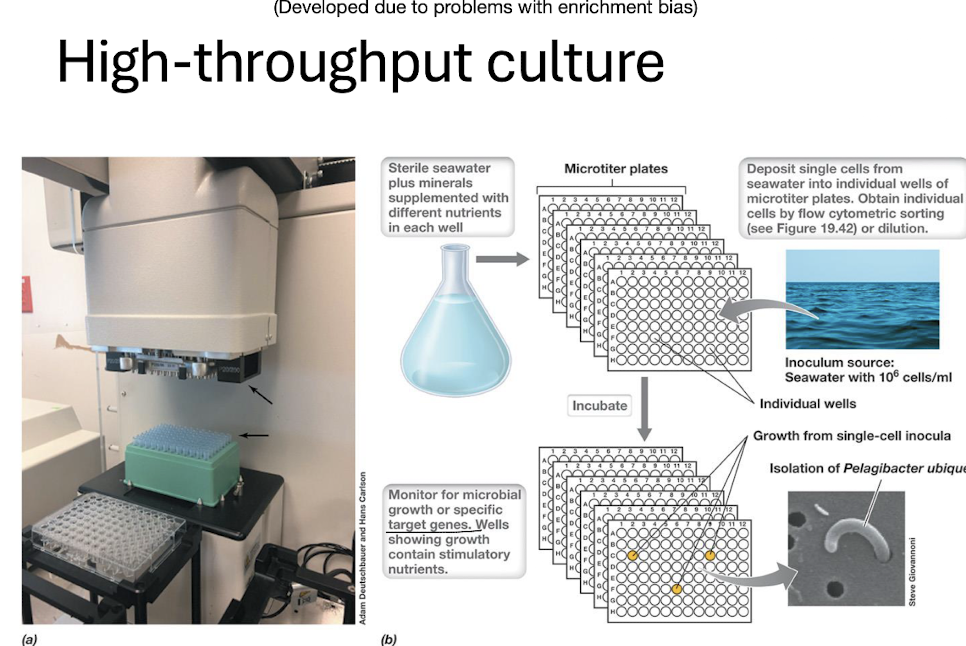
What is high‑throughput culturing?
= Robotic or microtiter‑based isolation of single cells under many nutrient conditions.
Example: Isolation of Pelagibacter ubique, the most abundant marine bacterium.
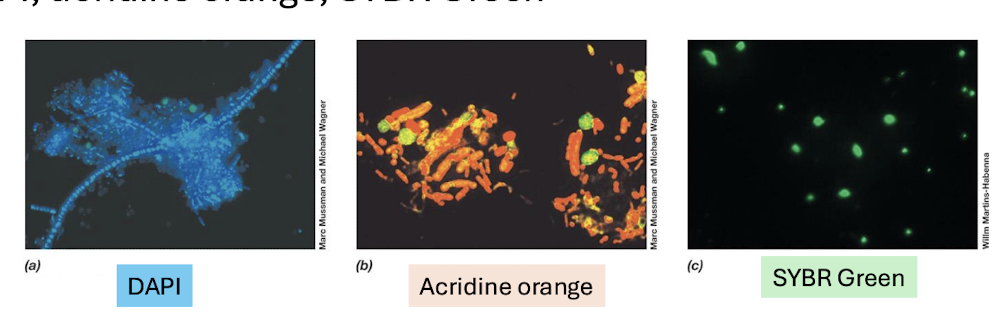
What are the 3 main general fluorescent stains and their functions?
DAPI, Acridine Orange, SYBR Green
→ Bind nucleic acids; stain all cells (live + dead)
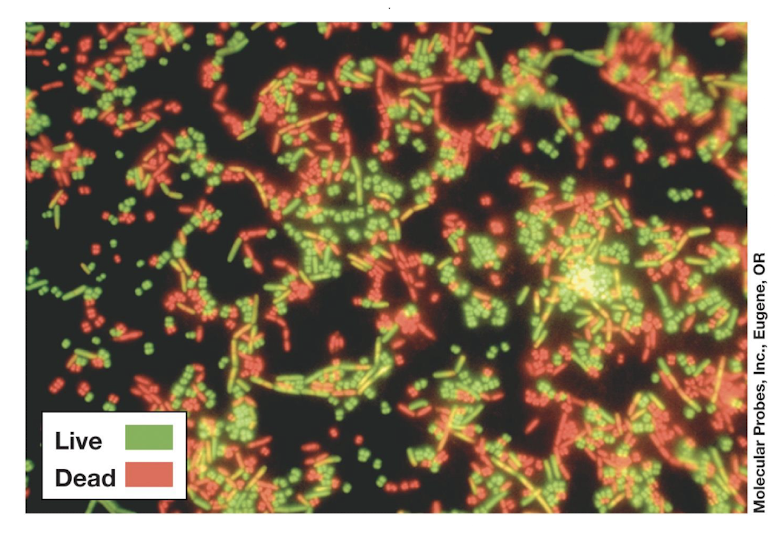
What is Viability Staining (LIVE/DEAD)? + limitations
Green dye → all cells.
Red dye (propidium iodide) → only cells with damaged membranes.
Limitations
Small cells may be missed.
No phylogenetic information
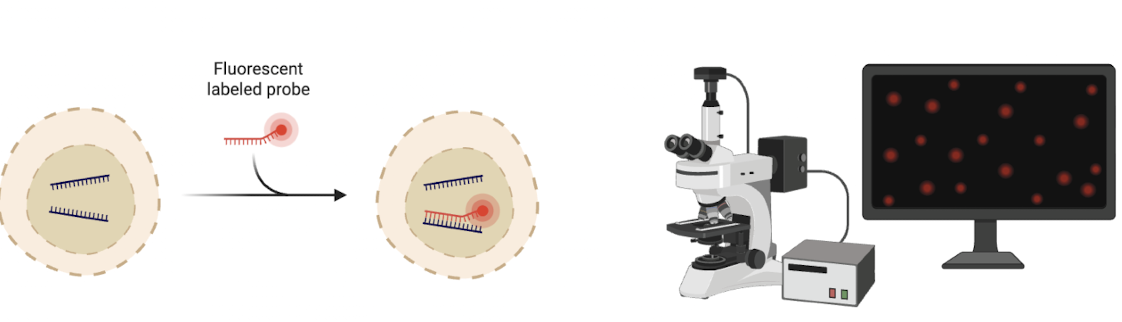
What is FISH?
(Fluorescence In Situ Hybridisation)
Fluorescent probes hybridise to complementary sequences inside cells.
Uses:
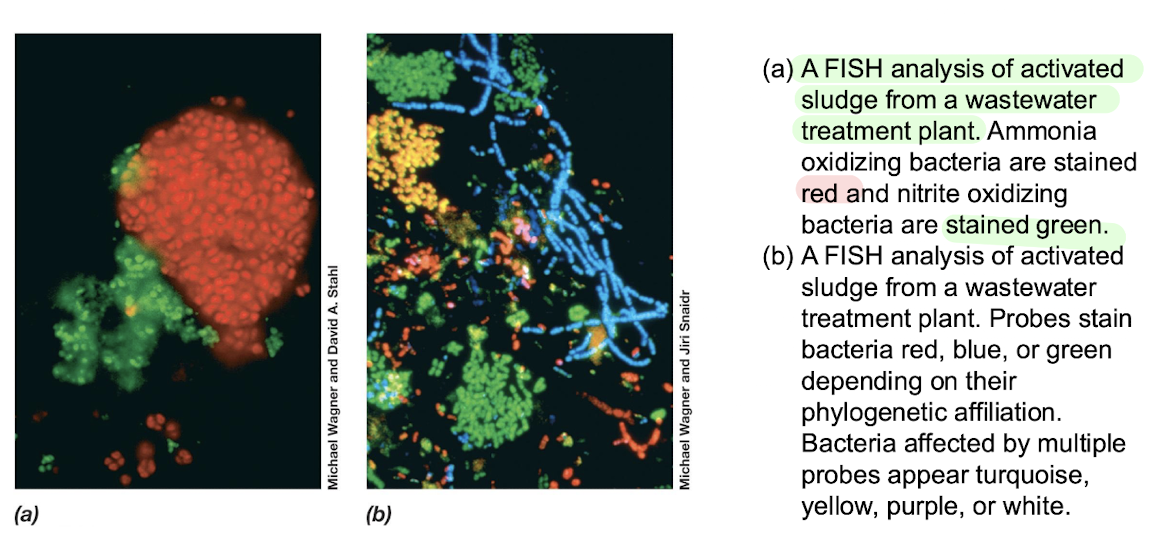
Identify taxa, quantify specific groups, visualise spatial organisation.Example:
Activated sludge: ammonia oxidisers (red), nitrite oxidisers (green).
PCR in Community Analysis: Why use bacterial/eukaryotic rRNA genes?
Universal, conserved + variable regions, good phylogenetic marker.
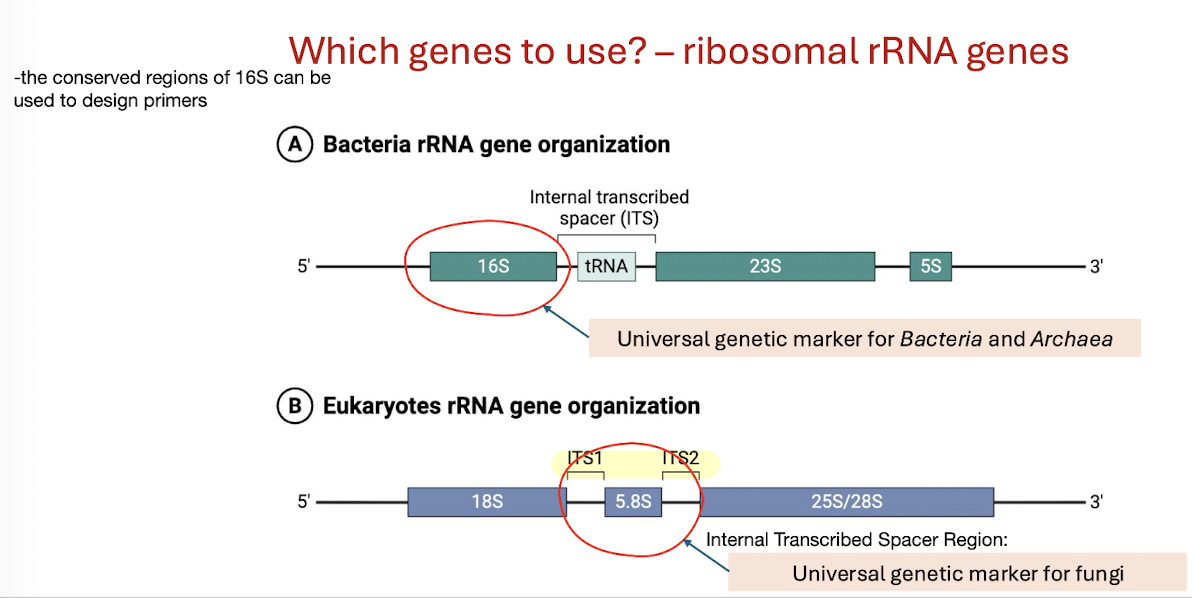
Which part of bacterial/archaeal and eukaryotic rRNA genes is considered a universal genetic marker?
Bacteria/Archaea → 16S rRNA gene;
Eukaryotes → 18S rRNA gene.
Fungi → 5.8S rRNA gene found between the ITS1 and ITS2 regions
What are phylotypes /OTUs?
(Operational Taxonomic Units) are groups of organisms with ≥ 97% similar marker gene sequences e.g. 16S rRNA
Used as species approximations in microbial ecology.
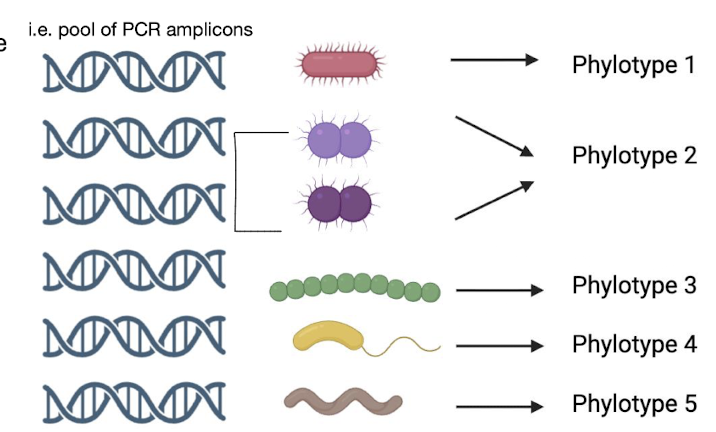
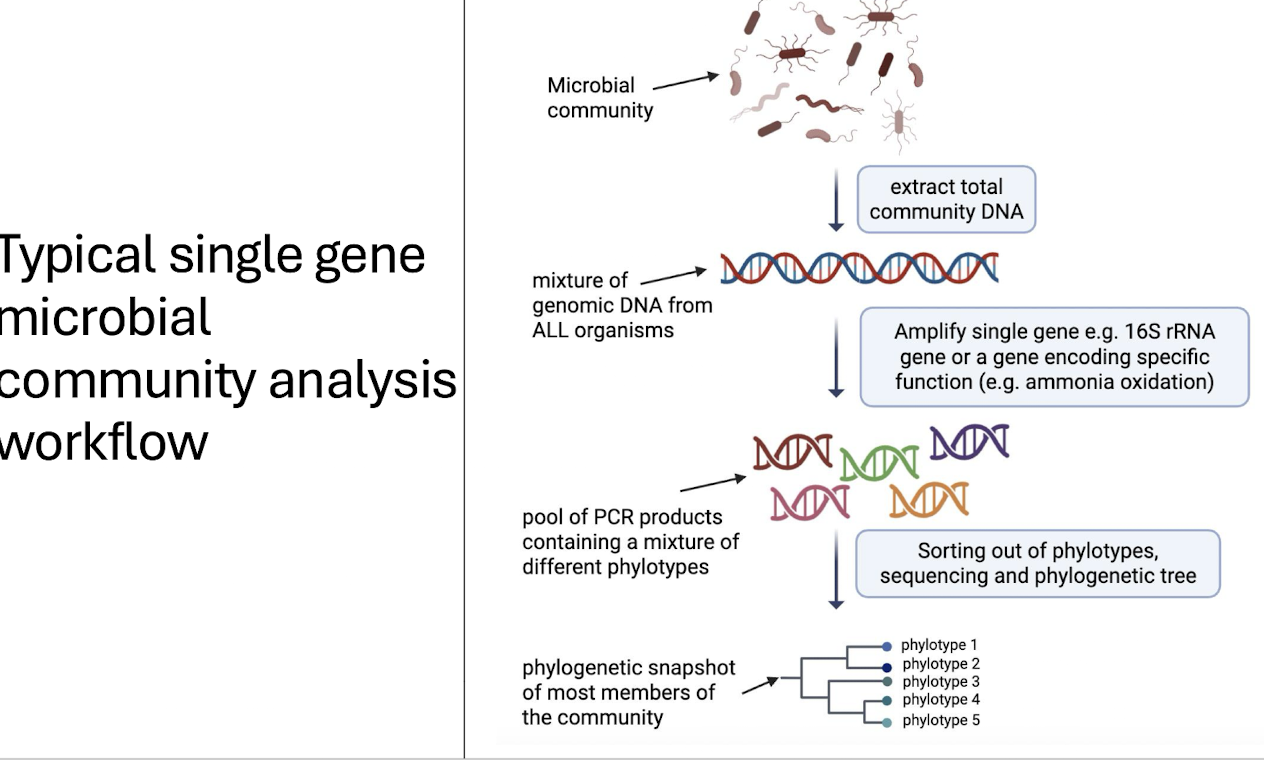
How do we separate phylotypes?
DGGE (Denaturing Gradient Gel Electrophoresis)
NGS (Next Generation Sequencing)
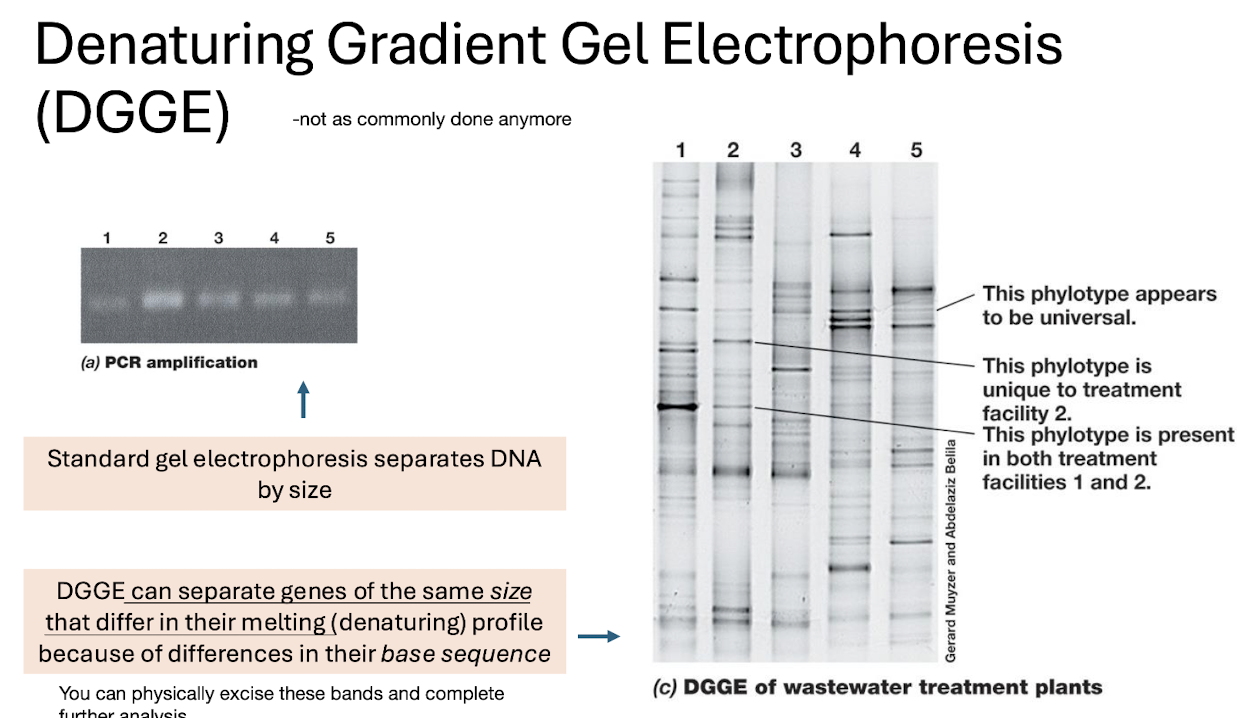
How does DGGE work?
Separates DNA of same size but different melting behaviour, using a gradient of denaturants
Produces banding patterns representing phylotypes.
How does NGS work?
Next Generation Sequencing (NGS) is a high-throughput method that allows for the direct rapid sequencing of large amounts of DNA (PCR products) by simultaneously analysing millions of fragments, enabling comprehensive profiling of microbial communities → reveals rare biospheres
Done by devices like Illumina sequencers, Pacbio, Nanopore (4th gen)
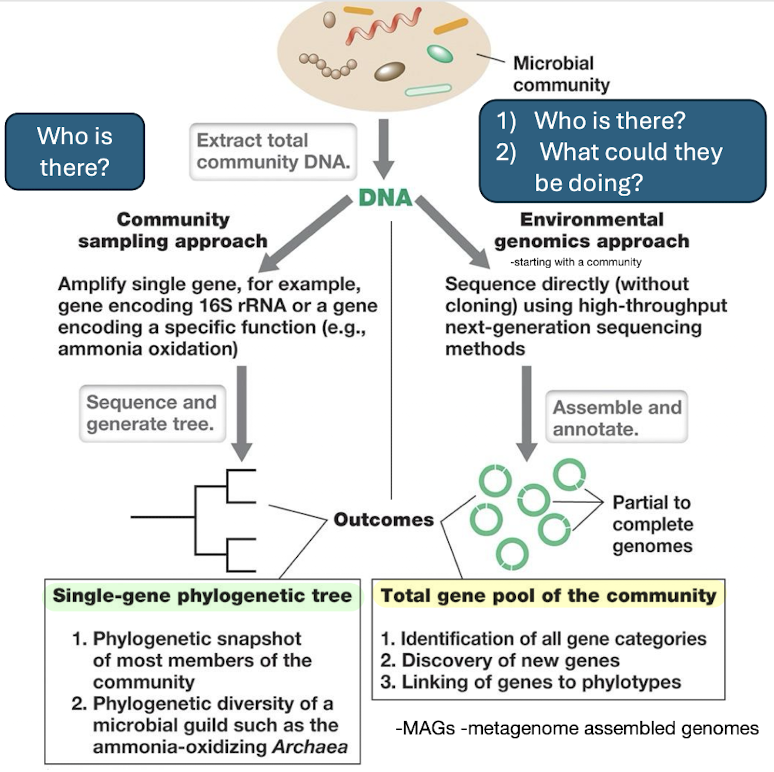
What is metagenomics and what are the main outputs produced as a result of it?
Sequencing all genes in an environmental sample.
Outputs:
Who is there
What they can do
New genes
Link genes to phylotypes
What is the difference between metatranscriptomics and metaproteomics ?
Metatranscriptomics: Which genes are being expressed.
Metaproteomics: Which proteins are present (direct functional insight).
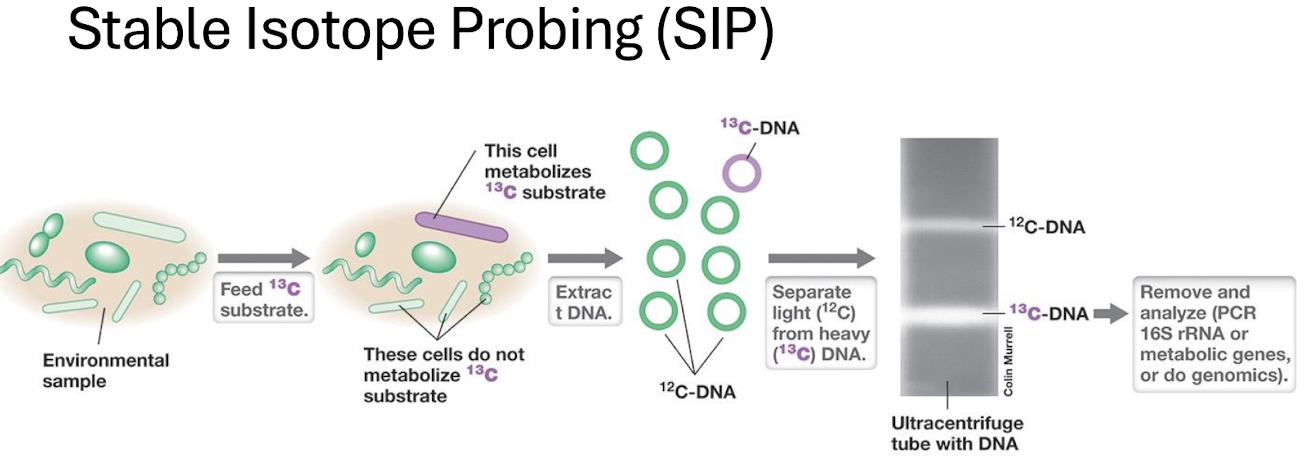
Stable Isotope Probing (SIP)
Feed community a ¹³C‑labelled substrate → organisms metabolising it incorporate ¹³C into DNA.
Heavy (¹³C) DNA separated from light (¹²C) DNA by density gradient centrifugation.
Links activity to identity.
Which statement about enrichment culture is TRUE?
B. It selects for organisms able to grow under chosen conditions
Which stain differentiates live vs dead cells?
C. Propidium iodide + green dye
FISH identifies microorganisms based on:
C. Hybridisation of fluorescent probes to target sequences
A phylotype is defined as:
B. A group of organisms sharing ≥97% sequence similarity
Metagenomics provides:
C. Total gene content of a community
SIP allows researchers to:
B. Link metabolic activity to specific organisms
Which method separates DNA fragments of identical size but different sequences?
B. DGGE