MB Lab - Quiz 5
1/63
There's no tags or description
Looks like no tags are added yet.
Name | Mastery | Learn | Test | Matching | Spaced | Call with Kai |
|---|
No analytics yet
Send a link to your students to track their progress
64 Terms
Bacterial Unknown Day #4
START hurray
All slants should be: ???? (for inoculation)
Streaked AND stabbed
SIM Deep (how to inoculate)
SINGLE stab into center of media
Make sure to enter and exit at same (because M = motility)
Simons Citrate Test (reagents + what test)
Tests for: use citrate as SOLE carbon source
NO REAGENTS TO ADD
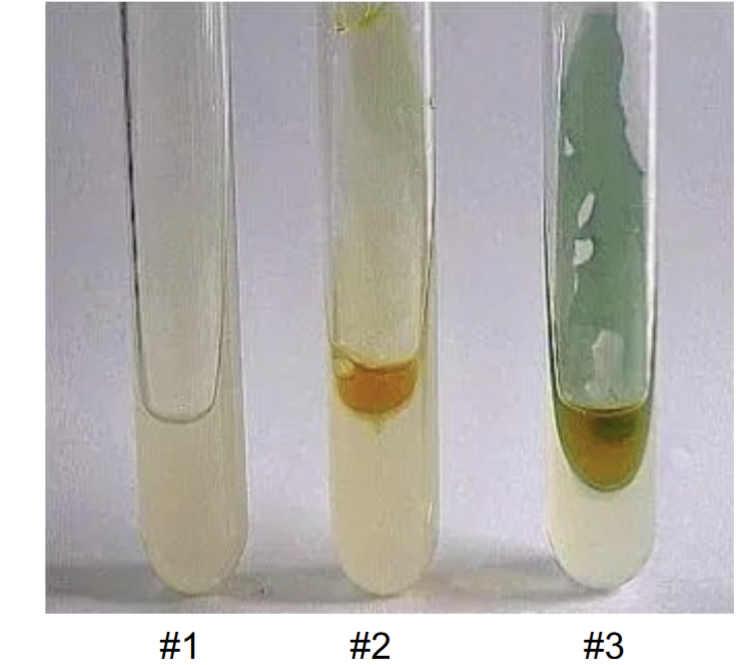
Simmons Citrate (results)
Positive = prussian blue
Ngative = green color
Phenylalanine Deamination Test (Test for + reagent)
Tests for
Presence of PHENYLALANASE (breaks down phenylalanine into phenylpyruvic acid (PPA))
Reagent
10% Ferric chloride
Why? Detects PPA with color change
Phenylalanine Deamination Test Results
Results
Positive = DEEP GREEN
Negative = YELLOW (color of reagent aka NO color change)
#1 in picture = or reagent or inoculation

SIM Deep (test for + Reagents) (tests for 3 things) know reagents for each
Tests for (and reagent that does it)
S = tests for H2S production
Reagent = Iron salt in media
I = detect INDOLE = presence of tryptophanase
Reagents: Chloroform and KOVAC’s reagent
Chloroform solubilizes the semi solid media
Makes permeable to Kovac’s
M = Motility = semi solid media
TTC (2,3,5- triphenyl tetrazolium chloride)
Colorless + soluble when oxidized; red when reduced
Makes it easy to see the results because the red color APPEARS along growth pattern
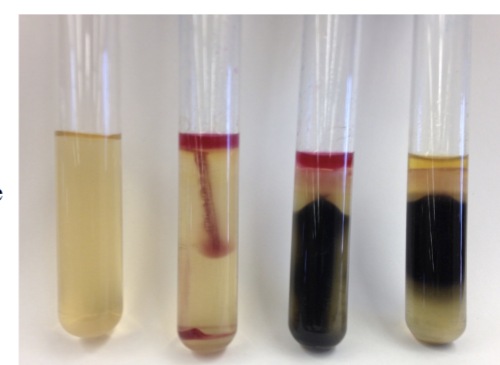
SIM Results + What reagents do you ACTUALLY ADD
ACTUALLY ADD = chloroform + Kovac’s (chlorfomr to solubilize media)
Positive
H2S = dark precipitate of iron sulfide
Tryptophanase/ Indole = red ring at top of media
Motility = growth radiating from red line
Negative = none of those/ opposite of those
Day #5 Hydrolysis tests start
Tubes vs agar plates incubate temp and time
Both incubate at 37
Tube = 48 hours
Agar plate = 24 hours
Urea Hydrolysis (test for + reagent)
Tests for: urease presence
Hydrolyzes urea into ammonia + Co2
SO detects ammonia
Reagent = Phenol red (already in media)
Detects ammonia, ammonia changes raise pH
Red = pH over 7
Yellow = pH under 7
Urea hydrolysis results
Positive = red (fuchsia)
Negative= yellow (orange)
Tryptone Hydrolysis (tests for + reagent)
Tests for: Tryptophanase presence
Hydrolyzes tryotphan into INDOLE + PYRUVIC acid
Reagent
Kovac’s reagent
Detects INDOLE (like the other test, SIM)
Tryptone Hydrolysis
Positive= red ring at top
Negative = no red ring
Fat Hydrolysis/ Spirit blue agar (test for + reagent)
Tests for: lipase presence
Which can HYDROLYZE fats into glycerol + fatty acids
Reagent (present in media)
Spirit blue agar = Contains LIPIDS
Need lipids to be hydrolyzed
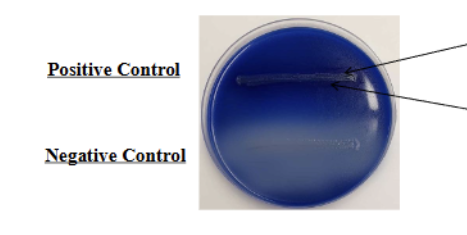
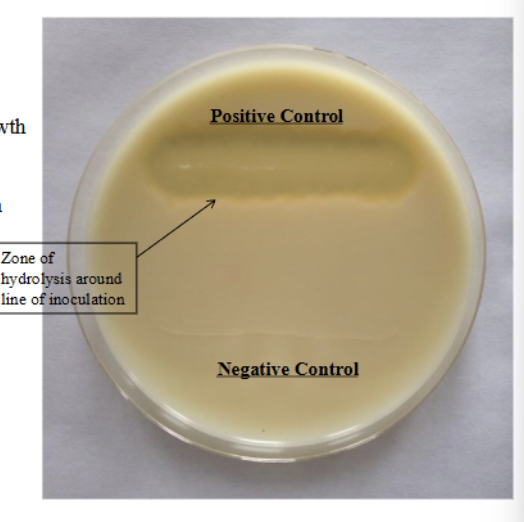
Fat hydrolysis/spirit blue test
Positive = Dark precipitate around inoculation (due to LOWERING of PH)
Lipid depletion around inoculation aka ZONE of hydrolysis
Negative = NO dark precipitate

Casein Hydrolysis (Milk Agar) ( tests for + reagent)
Tests for: Caseinase = hydrolyzes casein
Casein = predominant protein in milk (gives milk a white color)
Resagent = NONE to add
Media = milk agar aka has casein/milk
Casein hydrolysis- results
Positive = clear zone of hydrolysis around inoculation
Negative = NO zone around growth, remains white
Starch Hydrolysis (tests and reagent)
Tests for: Presence of amylase
(which can hydrolyze starch)
Reagent
Media = Starch
Iodine = add GRAMS IODINE
Make thin layer across surface of agar
Gorms complex/ binds to INTACT STARCh
turning NEGATIVE REULTS to blue/black
If it did, will stay clear beacue it wont bind
Starch Hydrolysis (results)
Positive = clear zone around growth (zone of hydrolysis)
No color change
Negative = NO clear area around growth
Media is STAINED brownish blue by starch iodine complex
IMViC test (what are the 4, and what is pos/neg for each category (2 categories)
4 physiological tests used to differentiate between ENTEROBACTERIAEASE or ENTERIC BACTERIA
4
Indole
Methyl red
Voges Proskaeur
Citrate
E.coli = ++—
Indole + Methyl red = POSITIVE
E. aerogenes = —++
VP + Citrate = POSITIVE
What does second I stand for in IMViC
Just for pronunciation
API 20E Identification for Enterobacteriaeae and other Gram negative
Start
What is the API 20E system
Minaturized version of convential l TUBE TESTS for bacterial unknowns
Gram negative rod that is member of enterobacteriaceae
Inoculation of test strip (important steps)
Fill EACH DEPRESSION with a drop of so of whater
DO NOT OVERFLOW
Used to create humid environment
Create suspension of bacterial unknown
MAKE SURE TO MAKE TURBID
Use sterile Pasteur pipet w/ rubber bulb
Tube vs Cupule
Tube = lower part
Cupule = part open to the air
Which tests to NOT fill suspenion up to cupule (aka which nee dot be anaerobic) - 5 add MINERAL oil
ADH
LDC
ODC
H2S
URE
Think —- HALO U
TDA Reagent + result
Reagent = add ferric chloride
Positive = brown-red
Negative = Yellow
VP Reagent + result
Reagent = Barritt’s (a-naphthol + KOH)
Positive = pink or red (appears within 10 minutes)
IND (reagent and tests)
Kovac’s reagent, NO CHLOROFORM NEEDED??
Positive= red ring within 2 minutes
NIT
Reagents = sulfanilic acid + NN, dimethyl 1 napthylamine
Positive = red color
Negative = ADD ZINC to test
UV Light Lethal effects start
START ya
How to swab plate
TRYINT TO MAKE a fully inoculated plate
Swab with cotton swab in 3 diff directiosn in zig zag
What is UV light + total range
electromagnetic energy
1nm to 400nm
3 groups of UV light
UV A, B, C
UV -A
315-400 nm
LONGEST
UV-B
280-315nm
UV-C
100-280nm
MOST DANGEORUS TO BACTERIA
Most dangerous wavelengths for bacteria (3 important ones)
265= MOST DAMAGING to DNA (emitted by our UV lamps)
WHy?
Most efficient for cell penetraiton and pyrimidien dimers
280nm = protein concentration determined
Proteins absorb UV light at this wavelength MAXIMALLY
260 = wavelength that DNA and RNA concentration is determined
High absorbance from DNA bases
How does UV light cause bacterial death
UV light causes DNA damage in form of PYRIMIDINE DIMERS
Thymine dimer = most common
COVALENT bonds between thymine ===== distort DNA helix + blocks replication genreally because cant read DNA
CANNOT be repaired by organisms DNA repair mechanism
UV Lights distance from plate
5 inches
Which bacteria = ENDOSPORE FORMER and what does this do
Bacillus subtills = ENDOSPORE FORMER
Can withstand longer exposures
Will have some growth rather than none
E.coli + S.epidermidis = no growth at extended exposure (2.5-5)
Once conditions return to normal, how does b.subitlis function
During incubation/ normal ocndtion, spore can GERMINANTE and new VEGETATIVE CELL will grow