W5 L12: Translation in Eukaryotes
L1:
The central Dogma
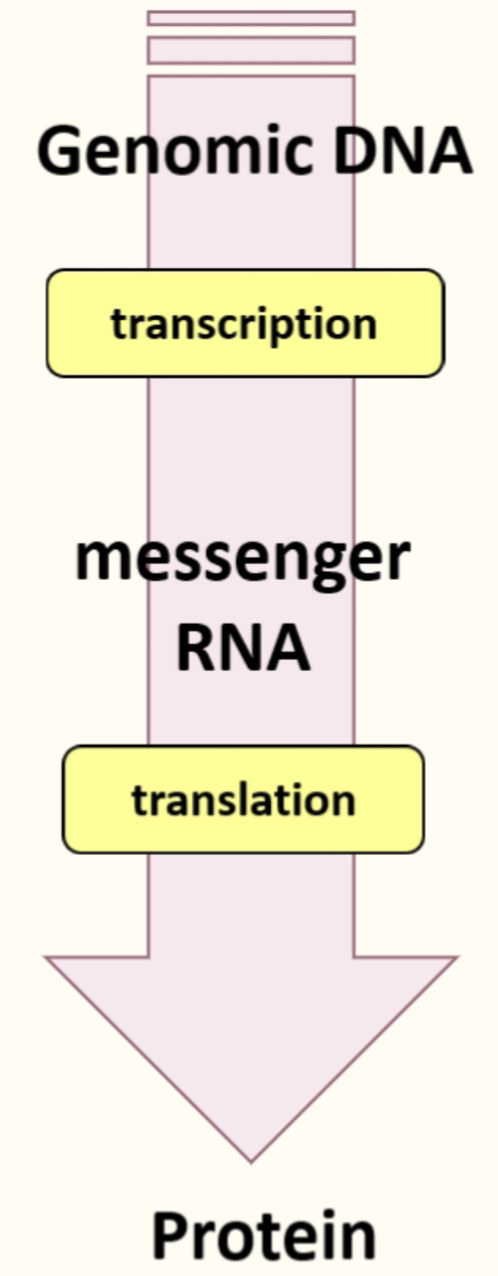
- Coding region of mRNA specifies what protein is going to be produced
- Ribosomes translate mRNA into proteins, translation factors promote this pathways & tRNA deliver the relevant AA in the right order
- Amino acids are the raw materials used in translation
- Uses ATP & GTP hydrolysis to provide E for process
- Multiple surveillance pathways to make sure protein production & reading the code is done accurately - makes sure process only occurs when needed and when E is available
Ribosome composition
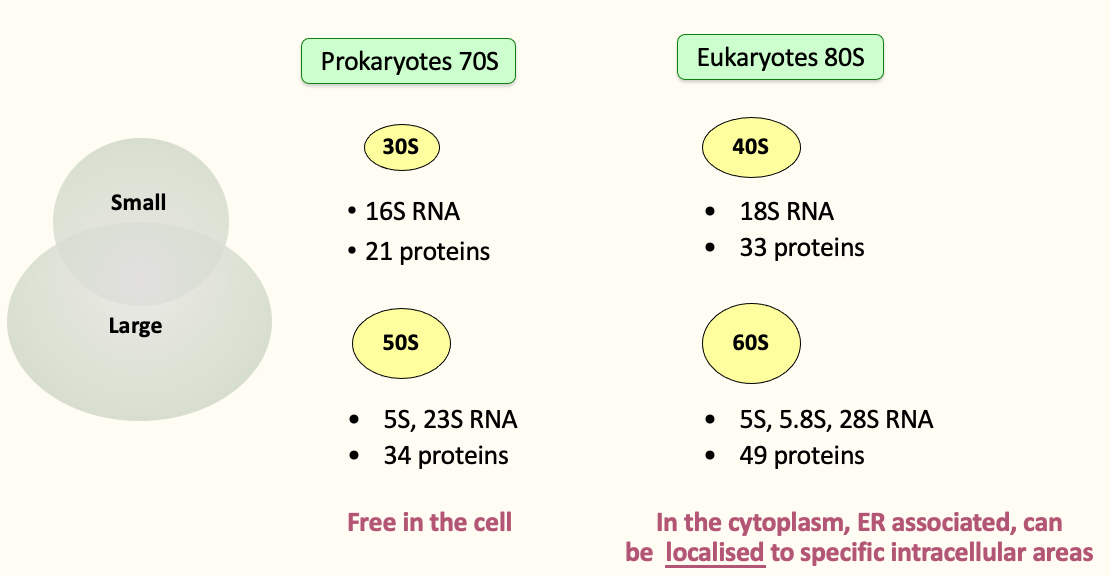
- An RNA scaffold coated by proteins
Ribosome - History & background
- 1970s → biochemical investigation of translation
- 2000 → 1st high resolution structure of ribosome (x-ray) by Venki Ramakrishnan
- 2000 onwards → dozens of structures of various functional state of the ribosome - rationalisation of more than 30 yrs of biochemical findings
- 2009 → Nobel Prize (chem.) to Yonath, Steitz & Ramakrishnan for their structural investigations of the ribosome
→ translation pathways deciphered (initiation-elongation termination)
→ antibiotics mode of action unravelled
Ribosomes
- Many RP paralogs produced in higher eukaryotes
- RPs can be post-translationally modified
- Variable rRNA modification
- Variability in composition → specialised ribosomes
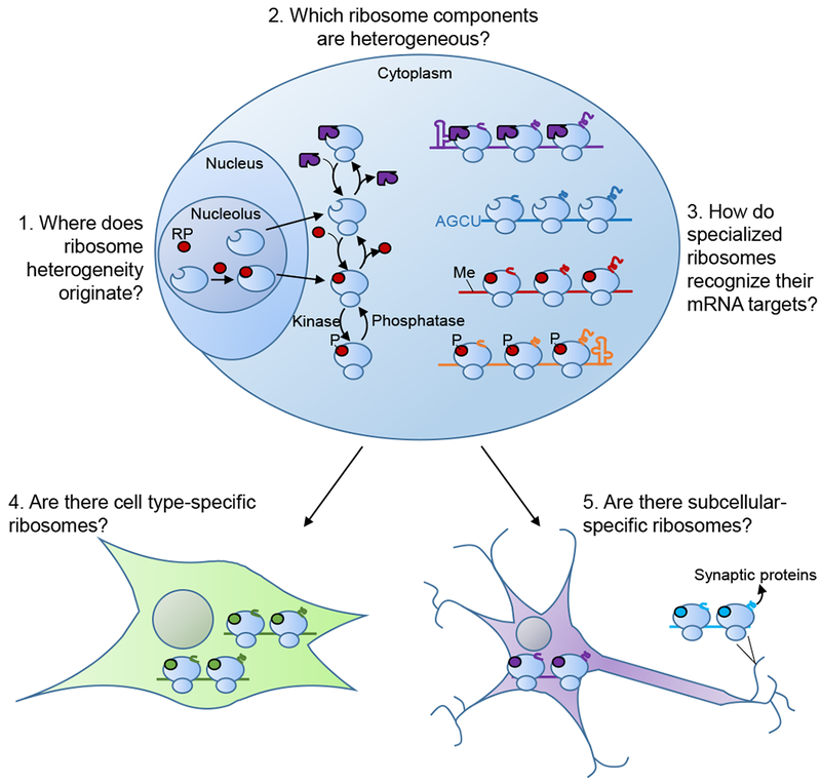
Ribosomes - protein synthesis in 3 steps
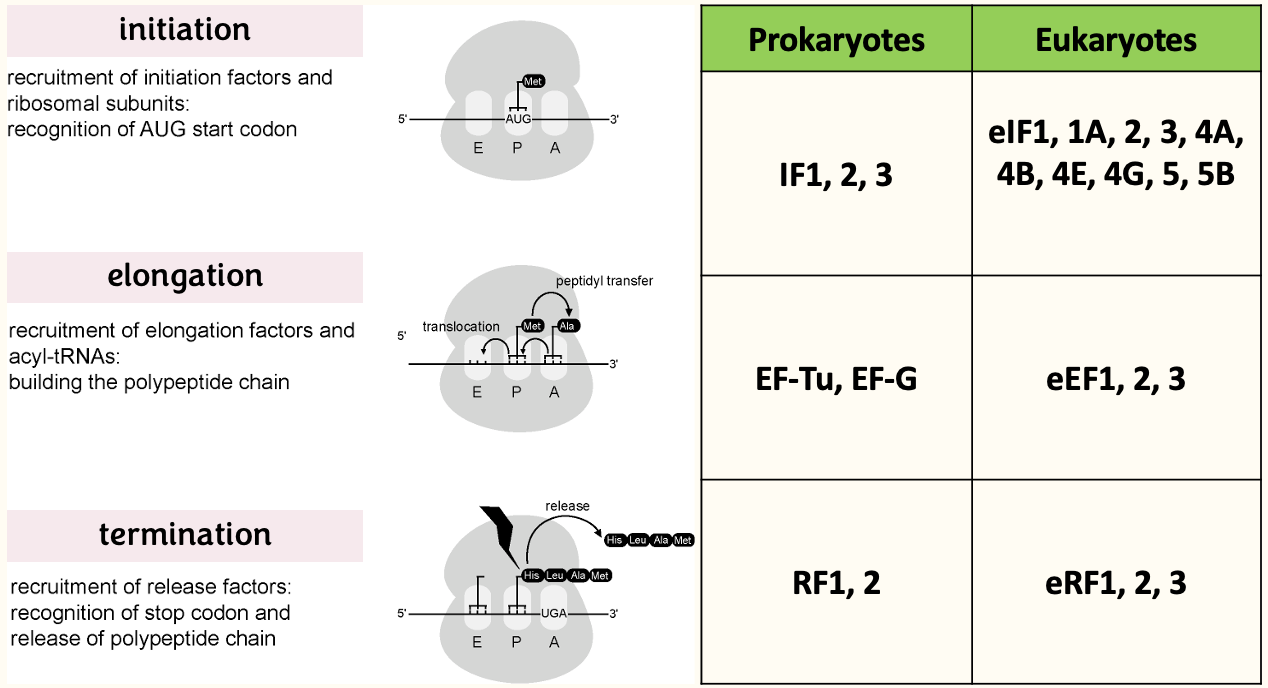
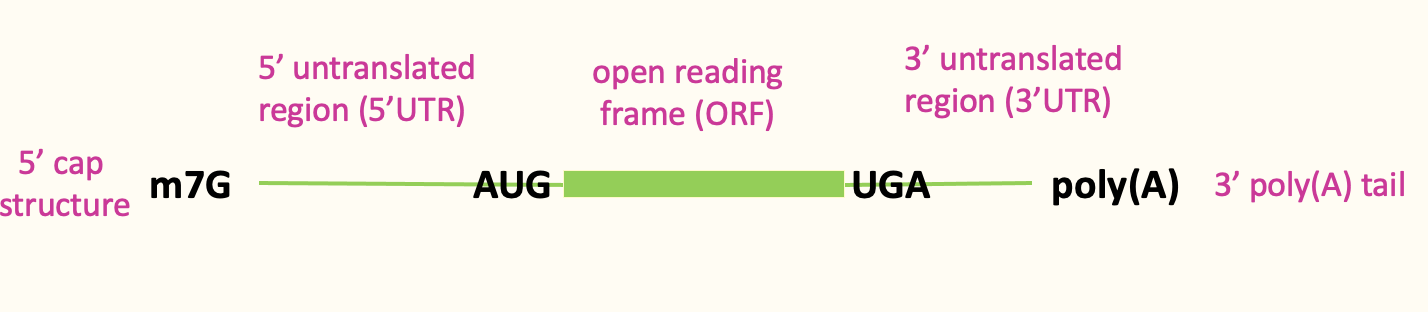
mRNA features that influence eukaryotic translation
- Eukaryotic DNA capped at 5’ end w/ m7G cap → protective for degradation & crucial for translation
- ORF contributes to protein code being produced - section from start codon to end codon
- 3’ poly(A) tail contributes to translational control
- Most eukaryotic translation is cap dependant
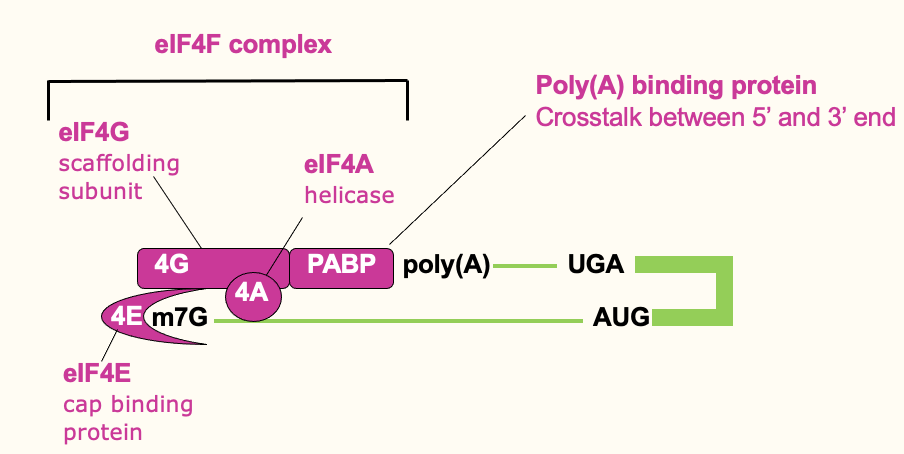
Translation initiation: cap binding
- Cap binding complex (eIF4F complex) which helps signify the 5’ end
- eIF4F complex → 3 diff. proteins that contribute to cap recognition
* eIF4E → high affinity for the cap & binds it directly
* eIF4G → bigger protein which binds 4E
* eIF4A → associated w/ 4G & its an RNA helicase - Poly(A) tail on 3’ end - bound by poly(A)-binding protein which interacts w/ eIF4G & helps stabilise 4F complex & binding at the 5’ end
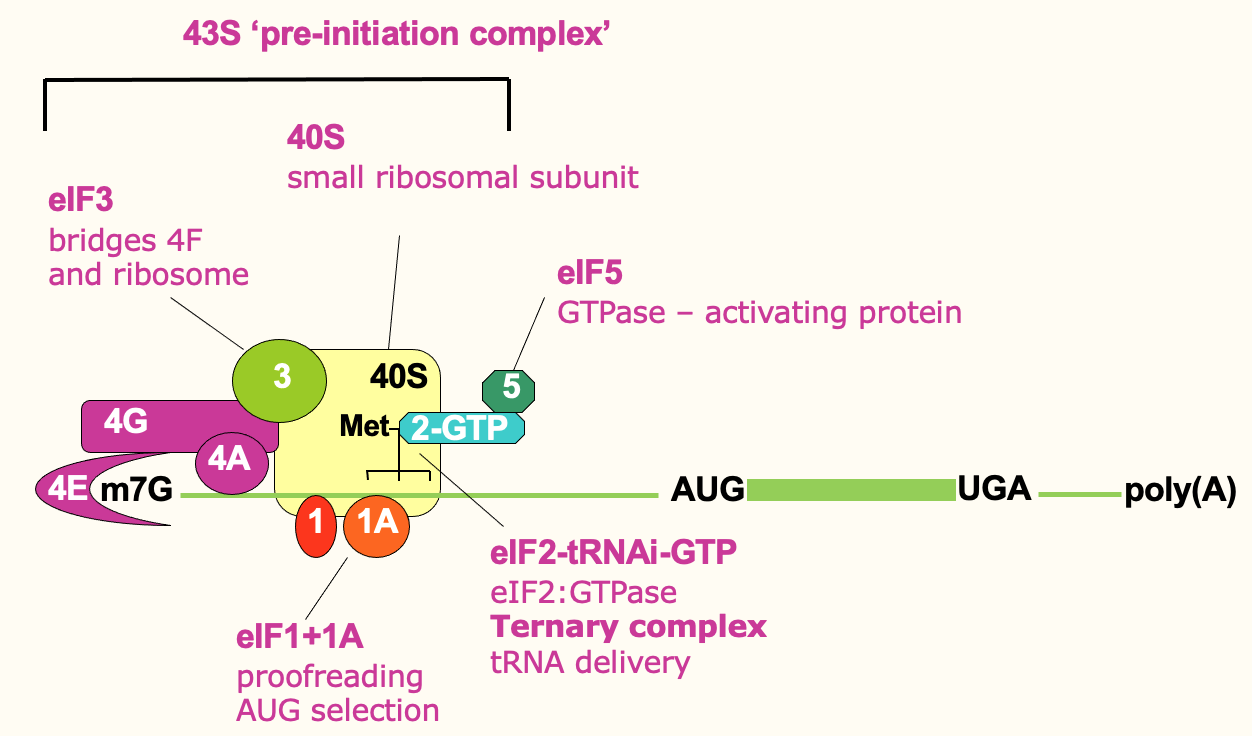
Translation initiation: ribosome recruitment
- Small ribosome (40s) forms a pre-initiation complex w/ other initiation factors
* eIF3 directly binds 4F & the ribosome → help bridge & bring that ribosome in
* eIF1 & 1A have proofreading roles
* eIF2 part of a Ternary complex which comes in & delivers initiator tRNA (tRNA loaded w/ initiator methionine) & comes in w/ eIF5 → provides GTPase activity (release E) - 43S initiation complex brought in to 5’ prime end of mRNA through bridging interactions of eIF3
Translation initiation: scanning
- To find the correct start codon, it scans along the mRNA, 5’ to 3’ until it finds AUG
- eIF4A is helicase used to unwinds RNA structure to be scanned properly
- Use E through ATP hydrolysis
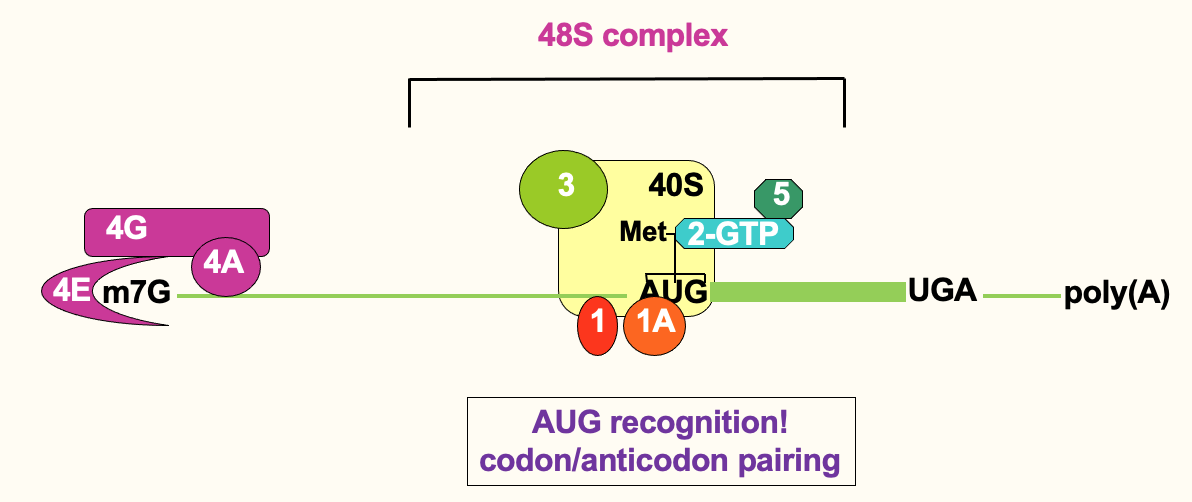
- Recognition of AUG is evident through base pairing
* anticodon of initiator tRNA base pairing w/ AUG codon - Once scanning ribosome has located AUG → 48S complex
Translation initiation: AUG recognition
- This triggers release of eIF5 & eIF2 - using ATP hydrolysis
- eIF1 & 1A - important for locating correct AUG → to avoid frame shifts
* by sitting at correct interface b/w tRNA anticodon & mRNA
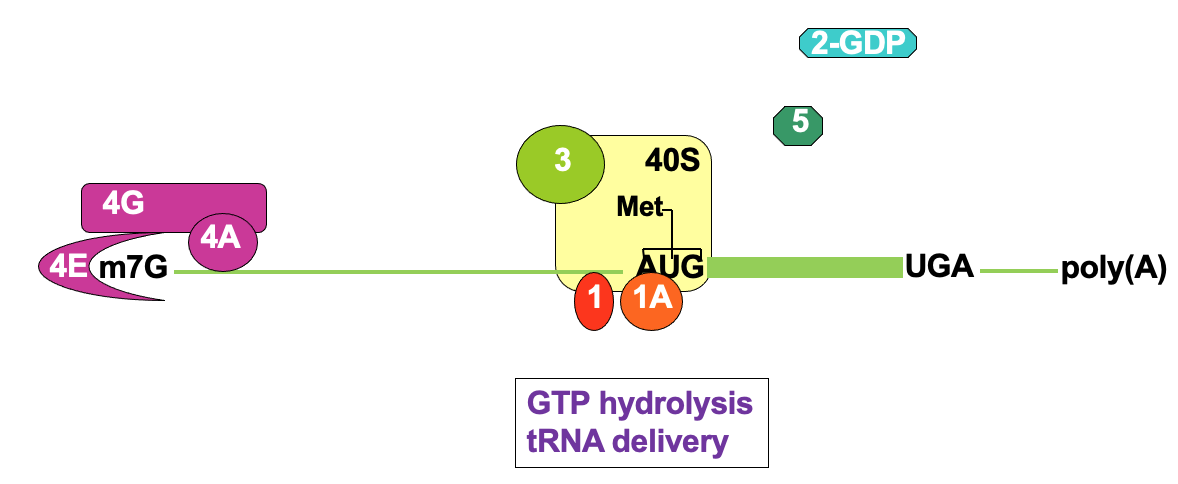
Translation initiation: 80S assembly
- Joining of 60s large subunit w/ 40s→ facilitated by eIF5B
* using GTP hydrolysis - Ejection of remaining initiation factors - not needed anymore
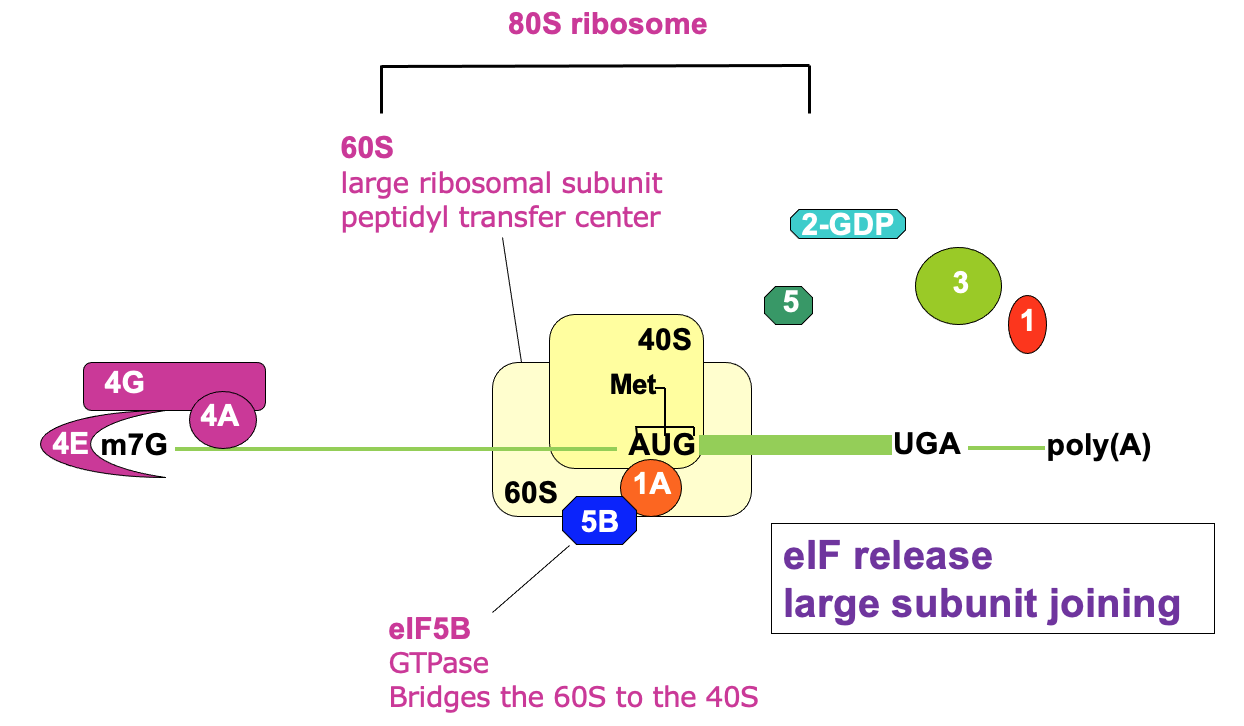
Translation initiation complete
- 80s ribosome also known as monosome → ready to elongate
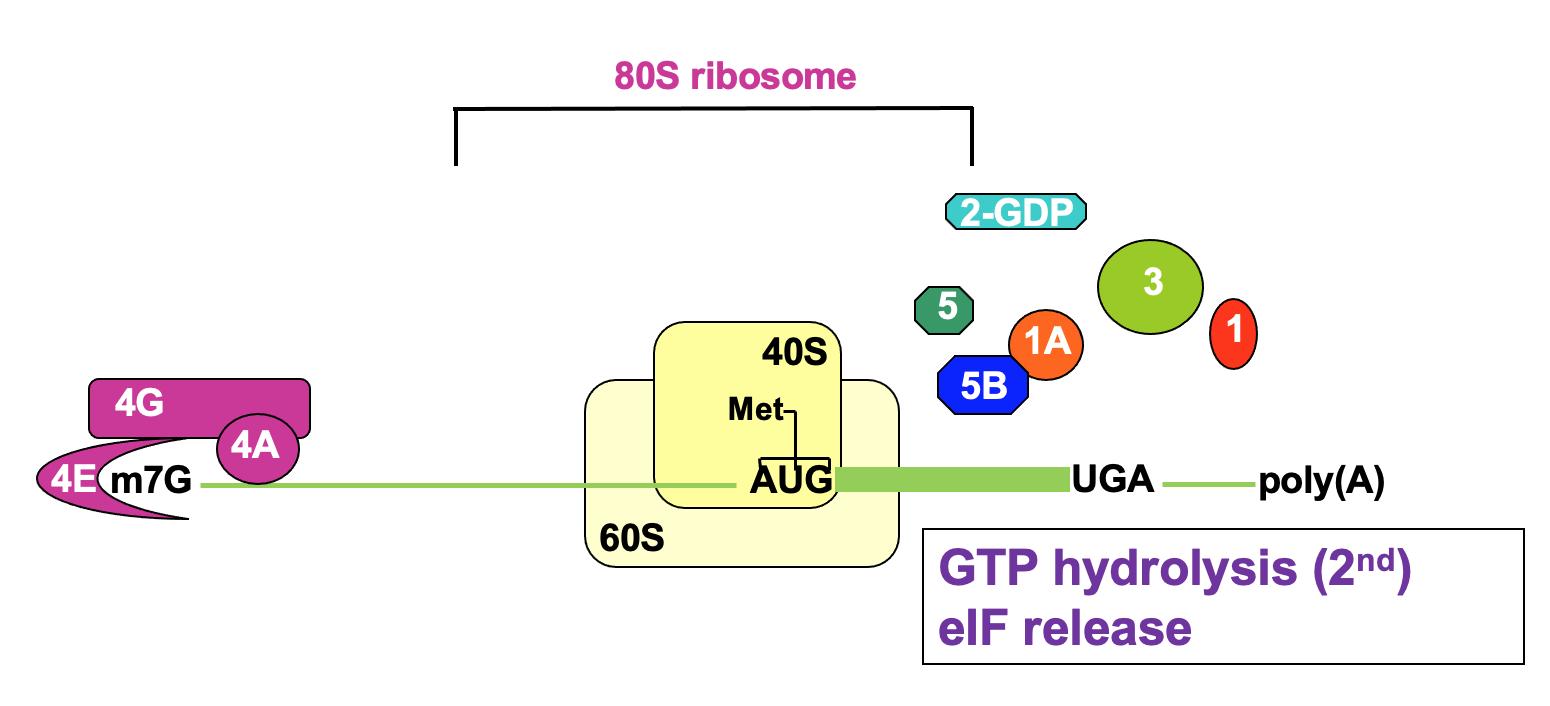
Comparison of eukaryotic & prokaryotic translation
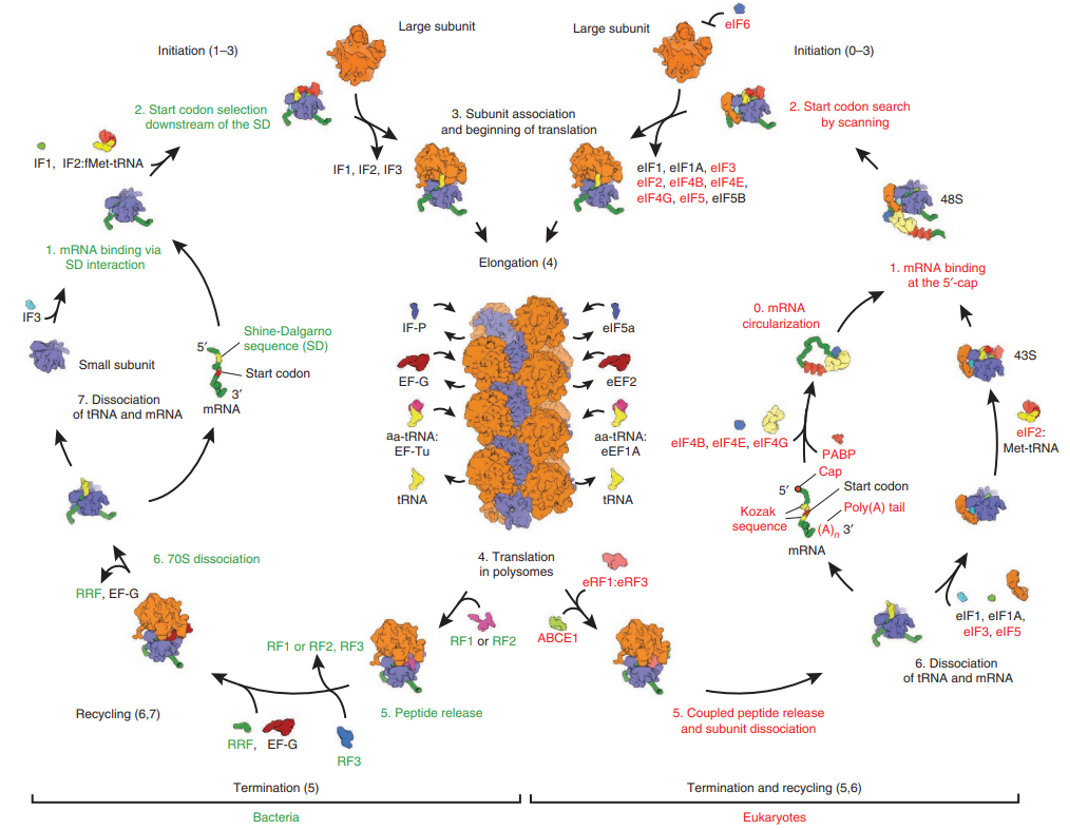
L2:
Translation control & disease
- Rate limiting steps are key targets for regulation & dysregulation
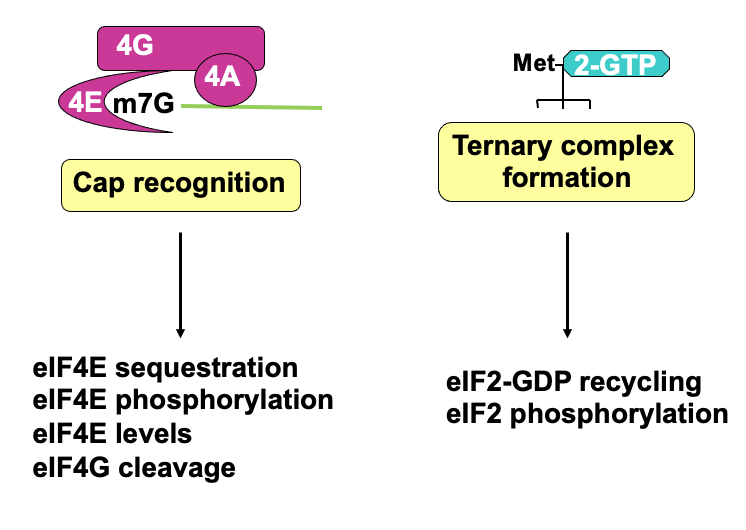
eIF4G structure & regulation during apoptosis
In normal cells, eIF4F complex helps to recruit 43s ribosome pre-initiation complex - through interactions that eIF4G has w/eIF3
eIF4F complex also interacts w/ poly(A)-binding protein which is bound at the 3’ end
* communication b/w 5’ & 3’ helps stabilise the 4F complex association w/ mRNA & promote translationWhen all interactions proceed, cap-dependant translation occurs
Apoptosis→ important in development - to get rid of cells that are no longer needed & in normal tissue homeostasis to eliminate infected cells
During apoptosis - there is activation of caspases (enzymes that can cut other proteins)
* they proteolytic cleave eIF4G in 2 positions → cuts separate the parts of the proteins that interact w/ protein interactors that promote translation initiation
* left w/ middle fragment (p76) that interacts w/ eIF4E, but not w/ poly(A)-binding protein & impaired in ability to promote translation initiation
* ∴ disruption of cap-binding complex activity & overall deregulation of translation in apoptosising cells
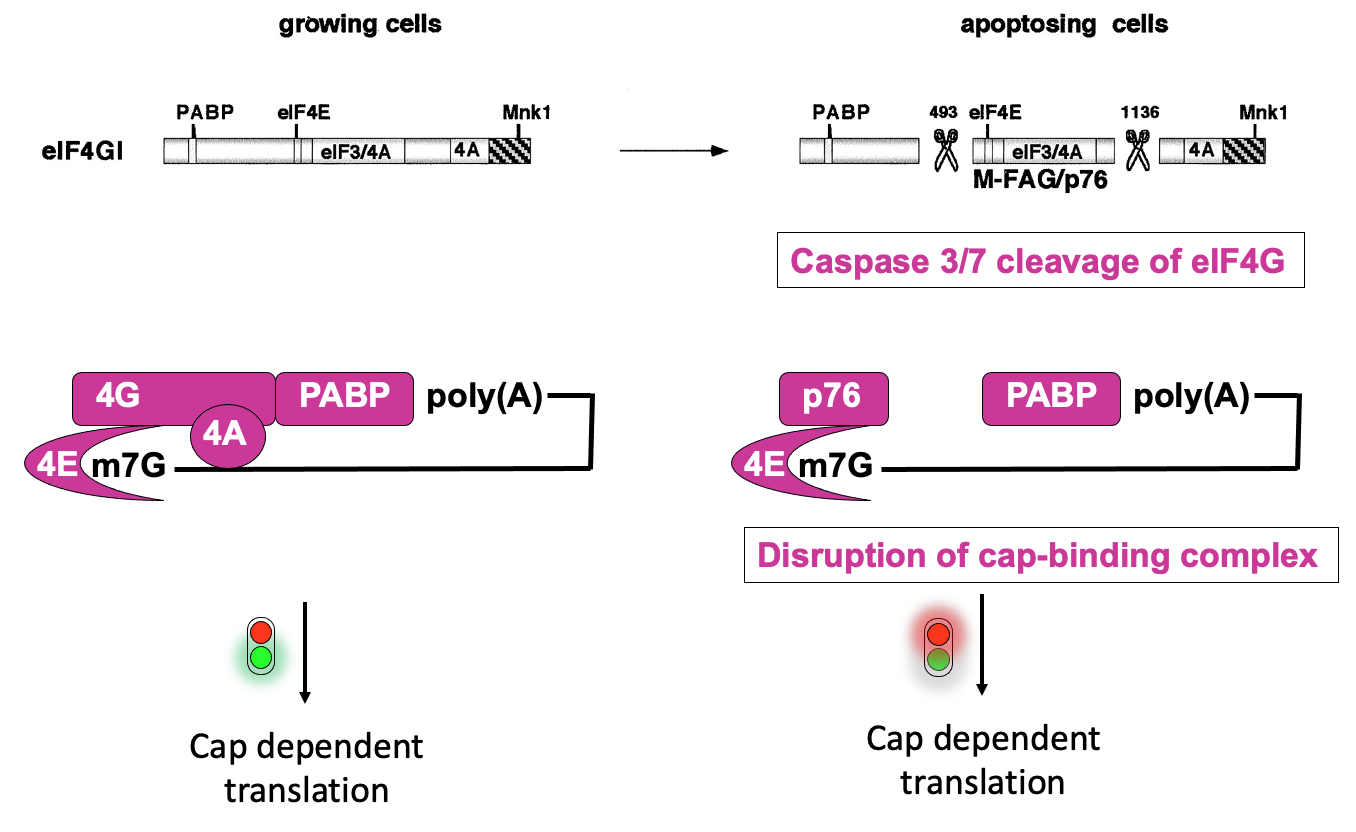
- Some specific messages that are required to proceed w/ apoptosis, need to be produced
* through alternate mechanism → cap-independent translation by internal ribosome entry site (IRES) - e.g. XIAP (inhibitor of apoptosis)
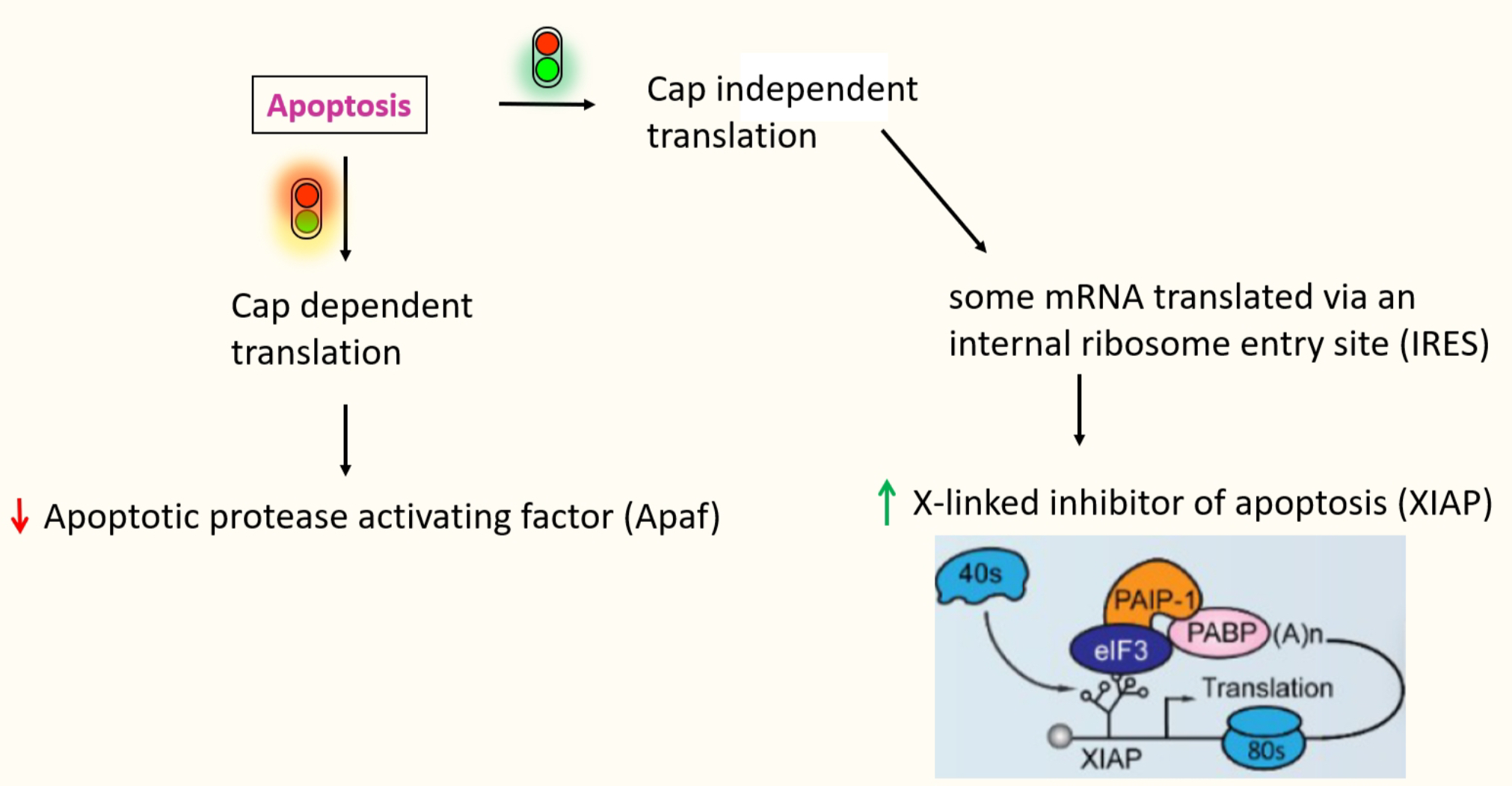
IRES dependant translation
- IRES → structure RNA domain that recruits the ribosome
* facilitates recruitment of a ribosome internally to the mRNA - rather than recruiting a ribosome directly to the 5’ cap - No need for the cap recognition & scanning - don’t need eIF4E binding for the cap
- Require only few eIFs, specific subset varies w/ each IRES
- Contains stem loops → highly stable & helps recruit in these proteins
- e.g.XIAP → IRES recruits in eIF3 directly
* to allow positioning of 40s ribosome directly onto the AUG
* mediates 5’ -3’ communication the same way 4G does - Present in both cellular mRNAs & viral RNAs
* evolved in viruses to produce their proteins even when the cell is apoptosing
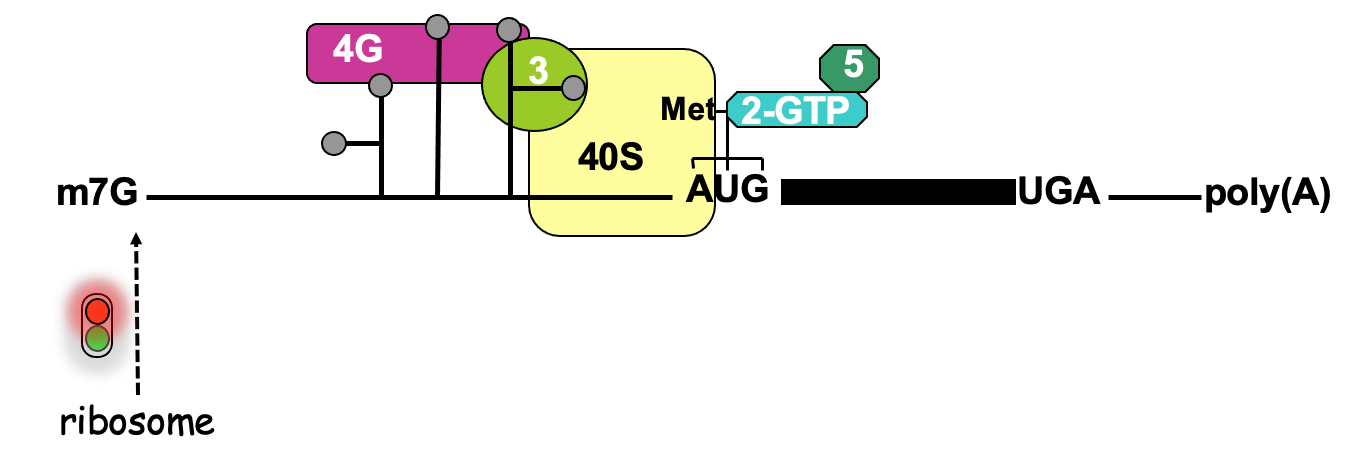
IRES dependant translation
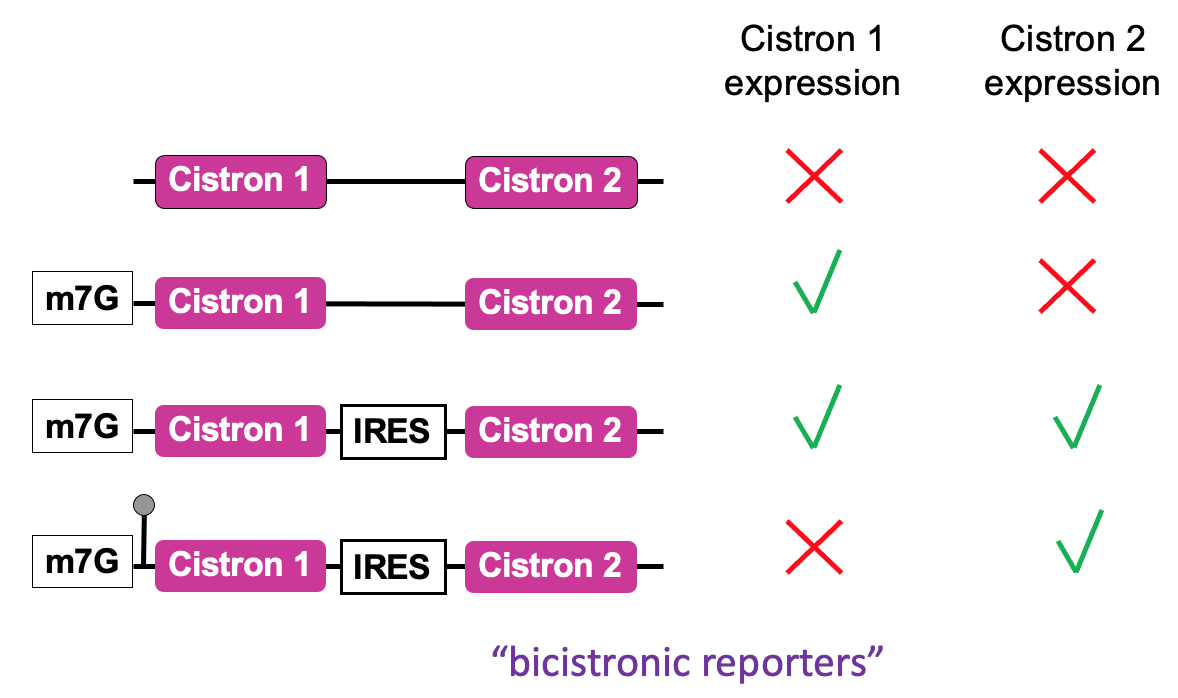
- An assay used to define the function of an IRES - in place of cistrons - fluorescent proteins of diff . colours
- 2 cistrons (open reading frames that code for reporter proteins) are added to an mRNA
- Positioning 2 open reading frames next to each other in mRNA that’s capped
- In eukaryotic cell via cap-dependant translation
* 1st cistron will be translated to produce the product & at end of translation, ribosome will terminate & ejected from mRNA protein being made
* no translation of downstream 2nd cistron - To test whether a seq. identified in RNA is functioning as an IRES - the sequence is put b/w 2 cistrons
* 1st cistron - translated in cap-dependant manner → after cap binding complex has recruited the ribosome at cap & allowed 1st cistron to be translated
* direct recruitment of a ribosome to IRES seq. - if functioning will allow production of 2nd protein → finds out whether seq. can function as IRES & what the requirements are e.g. what eIFs are important for functioning of IRES - Cap-independent translation → no translation of either cistrons
- Highly stable stem loop added to 5’ UTR → prevents unwinding activity of eIF4A → ∴ no translation from 1st cistron but effective translation from 2nd cistron using IRES
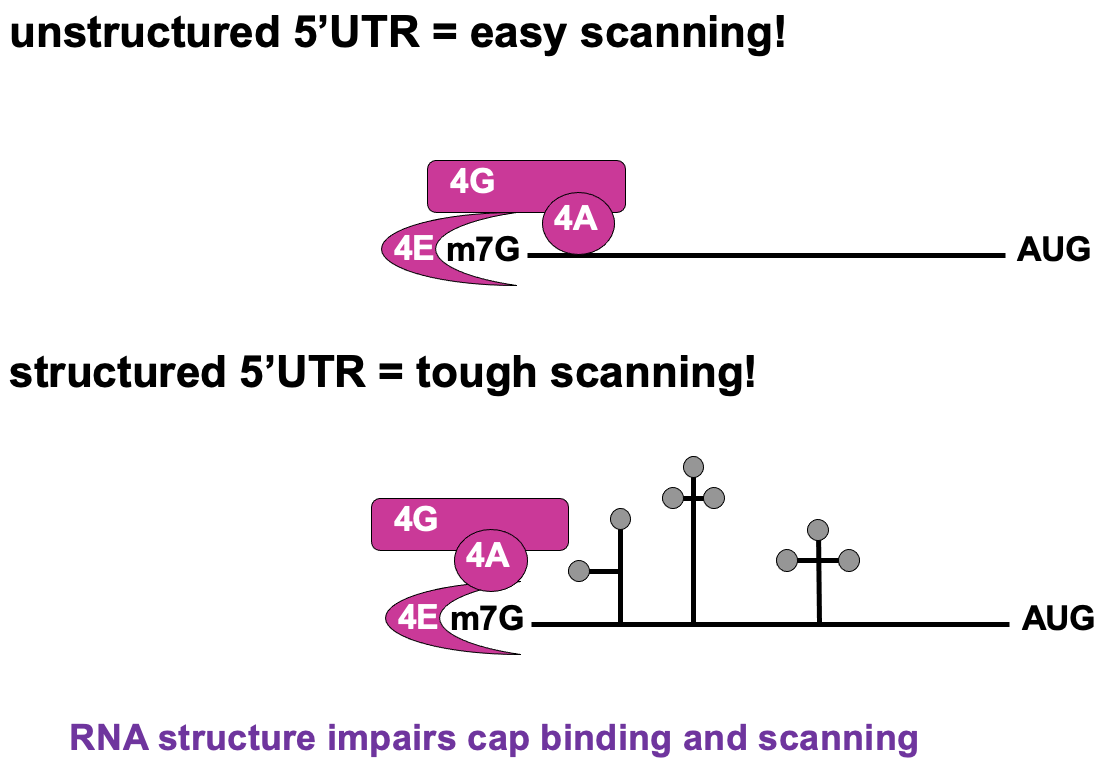
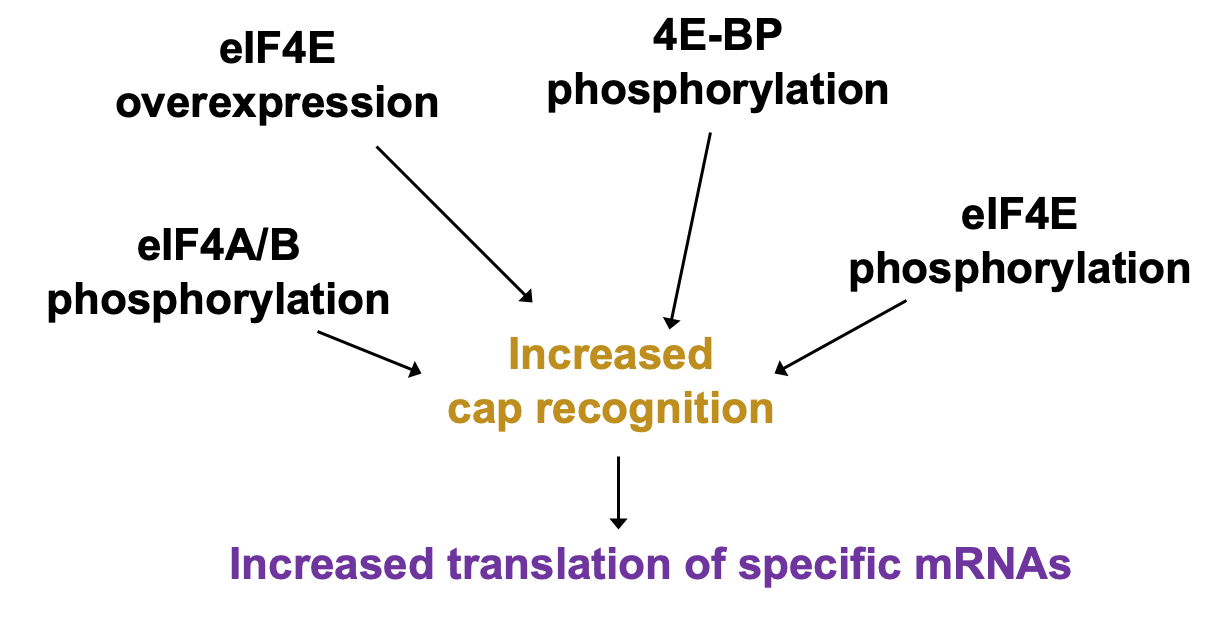
Regulation of cap recognition
- Unstructured 5’UTR → less dependant on 4A activity ∴ not using as much E
- Structured 5’UTR → e.g. stem loops present - require lots of E from 4F complex to unwind & allow translation to occur → poorly translated in cap-dependant translation
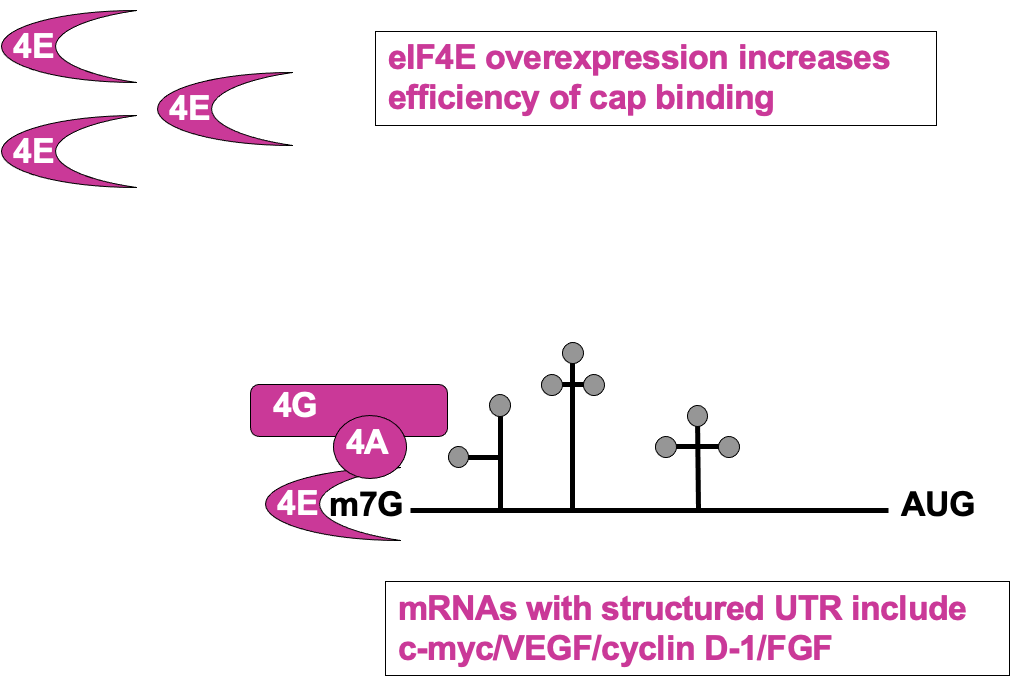
Regulation of cap recognition: eIF4E levels
- UTR containing mRNAs disproportionately benefit when 4E levels are higher
* higher 4E levels → help translate poorly translated mRNAs
* pro-proliferative factors e.g Cyclin D1 & C-myc contain these - upregulation of their expression may cause uncontrolled growth of the cells
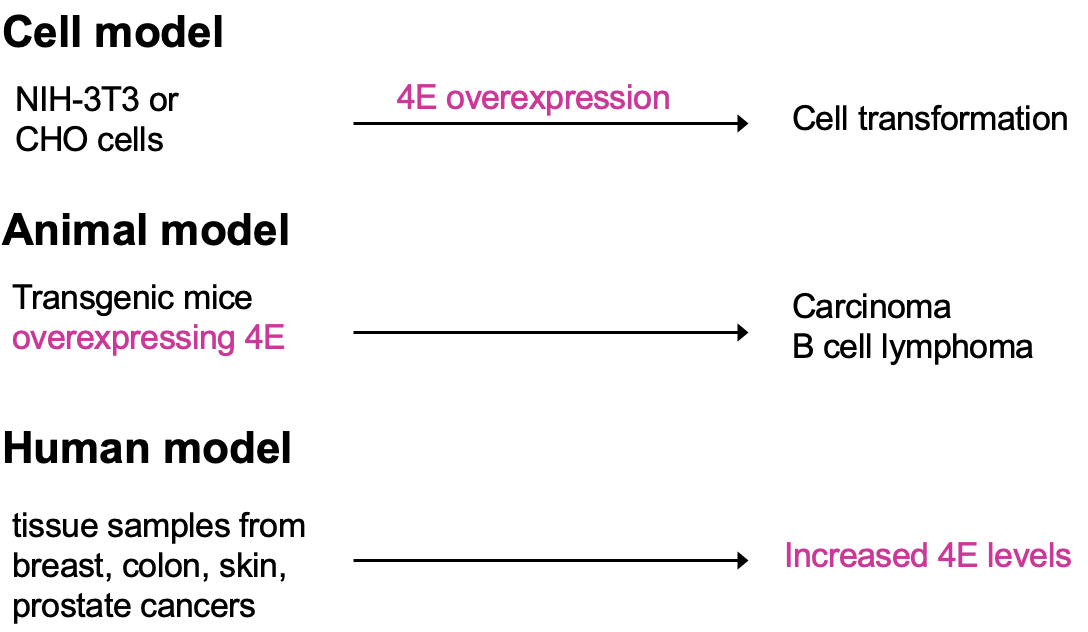
- Cultured cells that have overexpressed eIF4E observe cell transformation (transformation to phenotype resembling cancer)
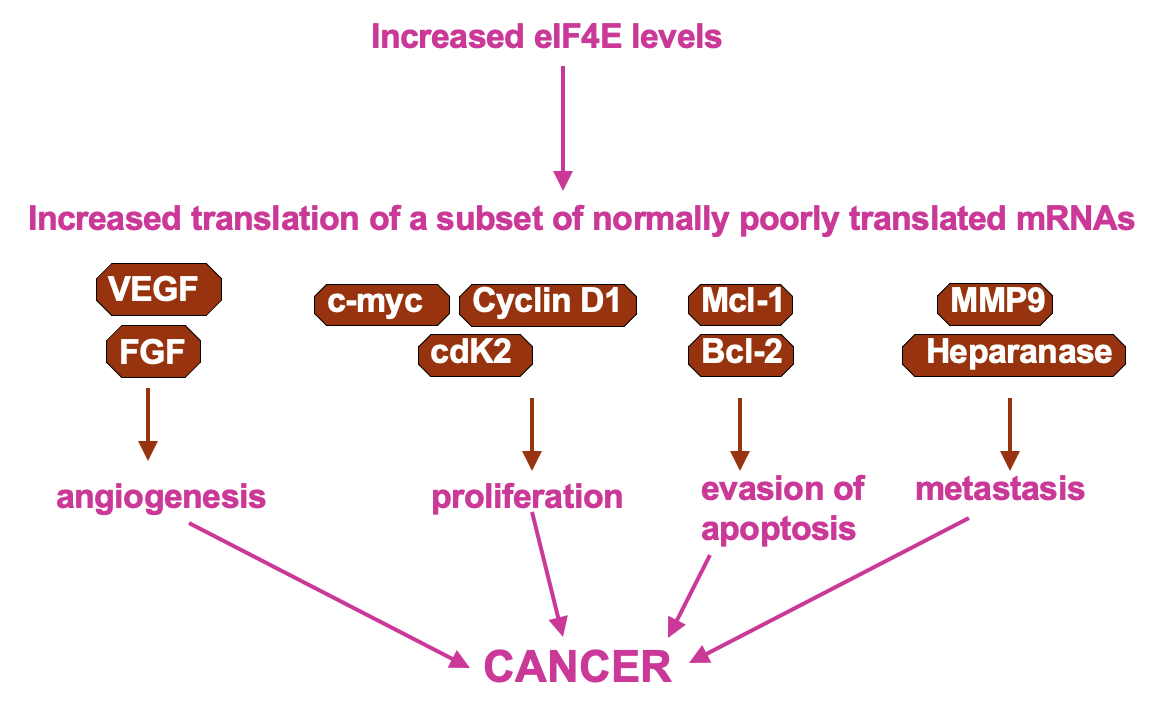
Regulation of cap recognition: eIF4E sequestration
- Δ availability of eIF4E in cell can be regulated in its activity
* 4E bound by eIF4E binding protein (4E-BP) - 4e no longer able to bind cap & participate in translation initiation
* 4e-BP phosphorylation releases 4E for efficient cap binding
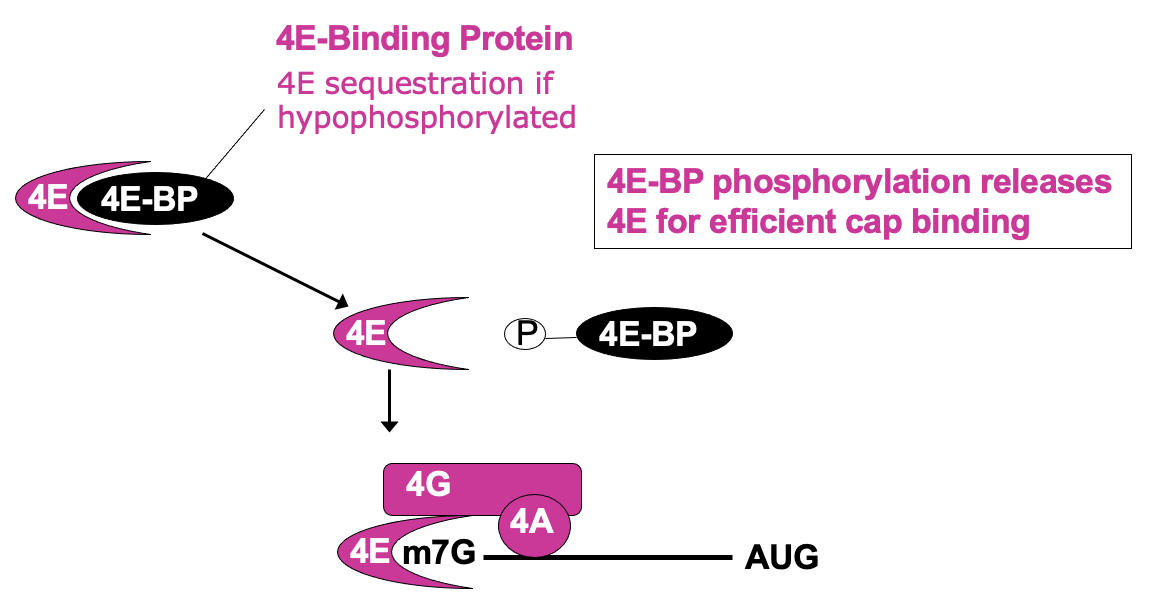
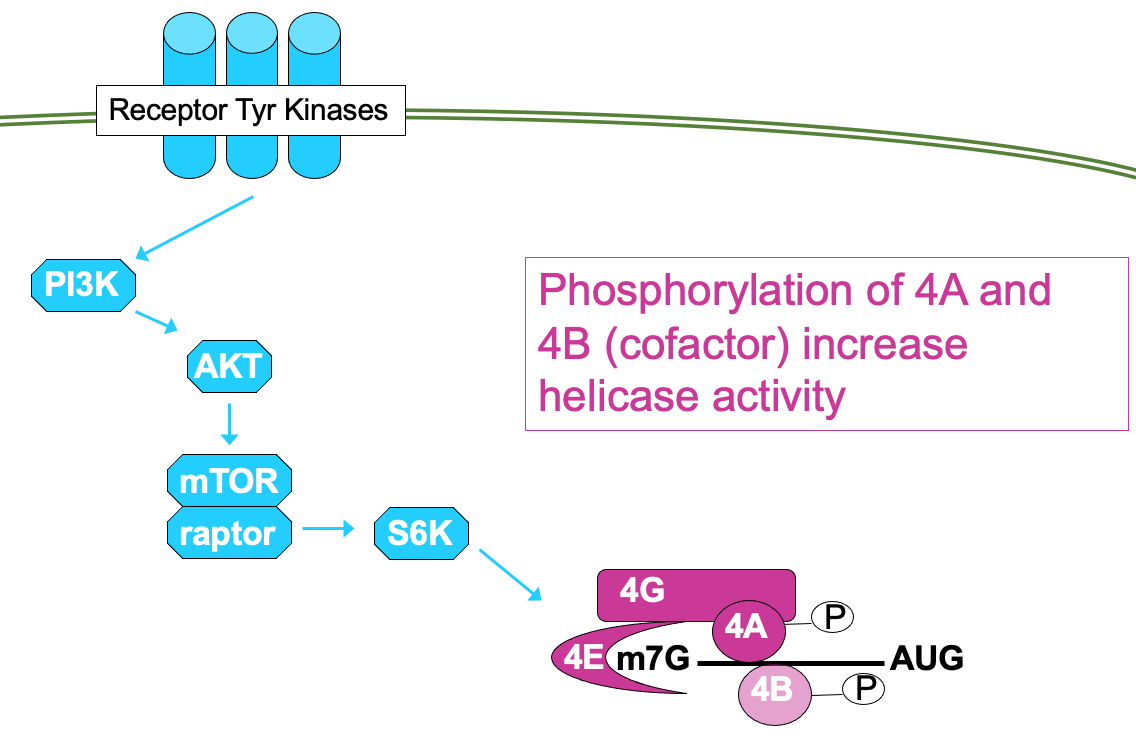
- This is regulated through signalling cascade - mTOR signalling pathway →
* processes mitogenic signals from cytokines, hormones etc. that are interpreted by receptor Tyr kinases
* through cascade of phosphorylation, culminating in activation of mTOR → results in phosphorylation of 4E-BP activating translation
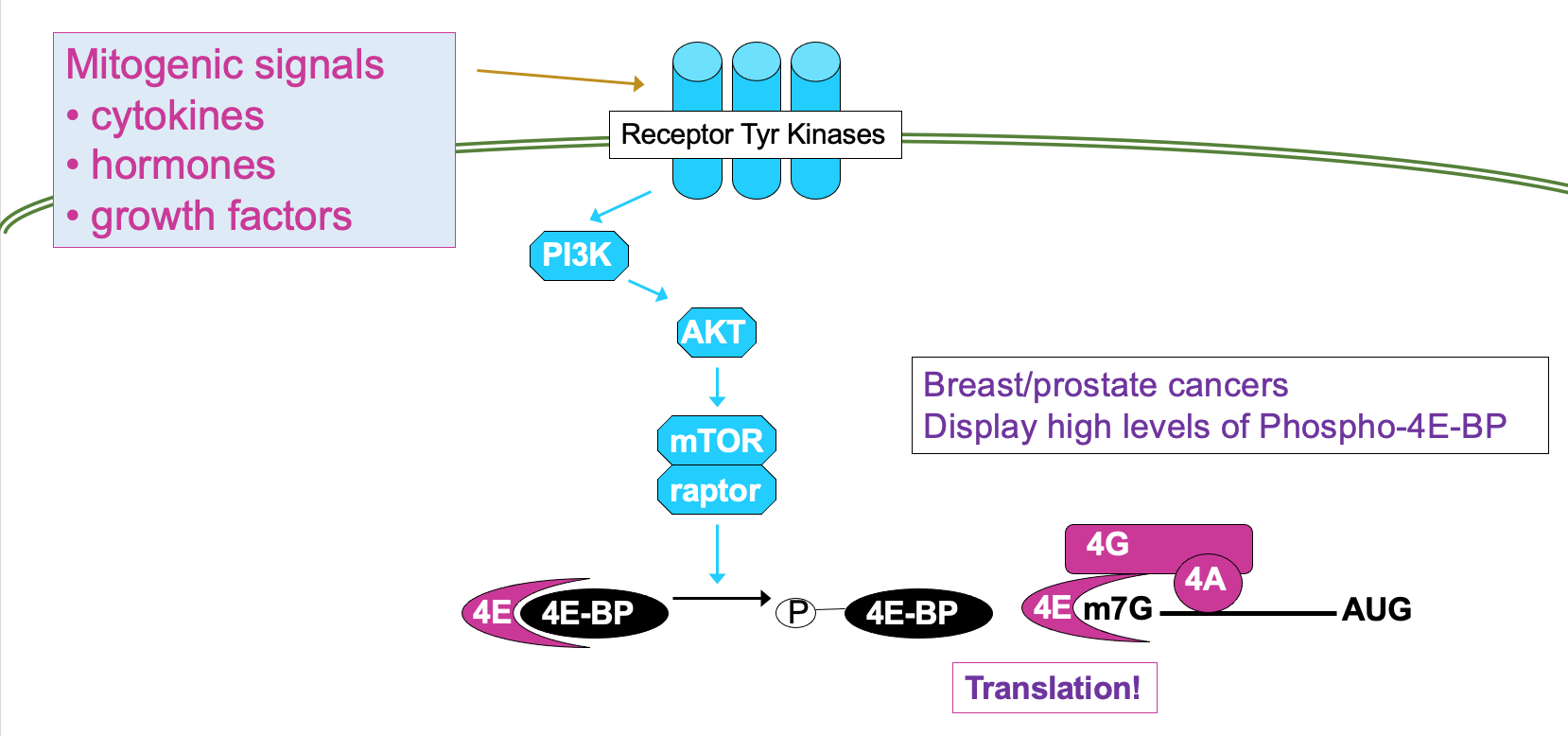
- 4E can be directly phosphorylated by kinase Mnk → ↑ affinity for cap so functions better
- Mnk activated through MAP kinase pathway - also activated by mitogenic signals
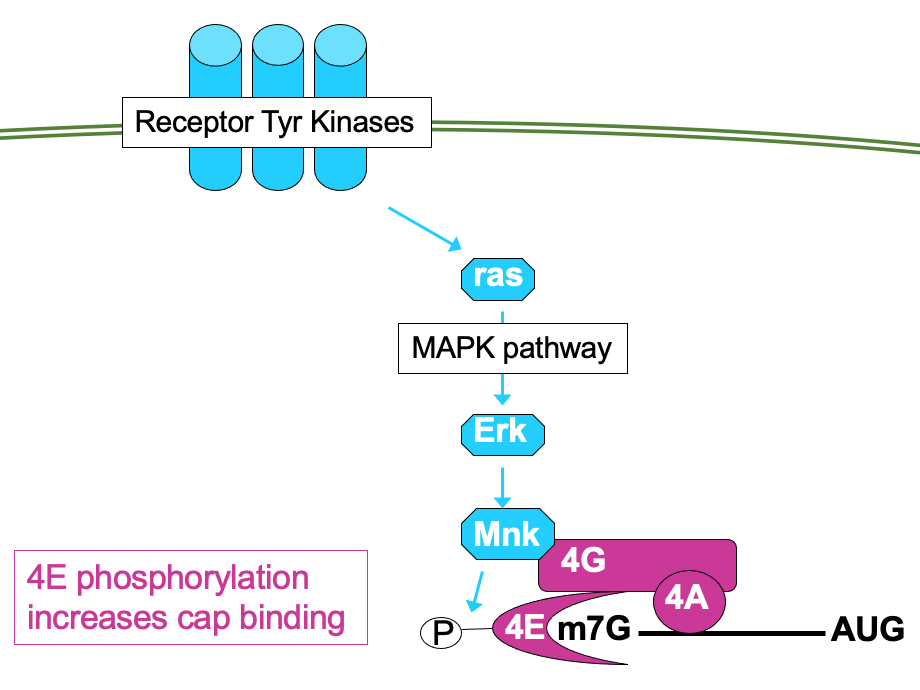
- Signalling pathways can impact the phosphorylation activity of other eIFs e.g.
* mTOR pathway can also lead to phosphorylation of eIF4A - helicase in eIF4F translation initiation complex & phosphorylation of cofactor 4B
* phosphorylation of both factors is associated w/ ↑ activity
Targeting translation as cancer therapy
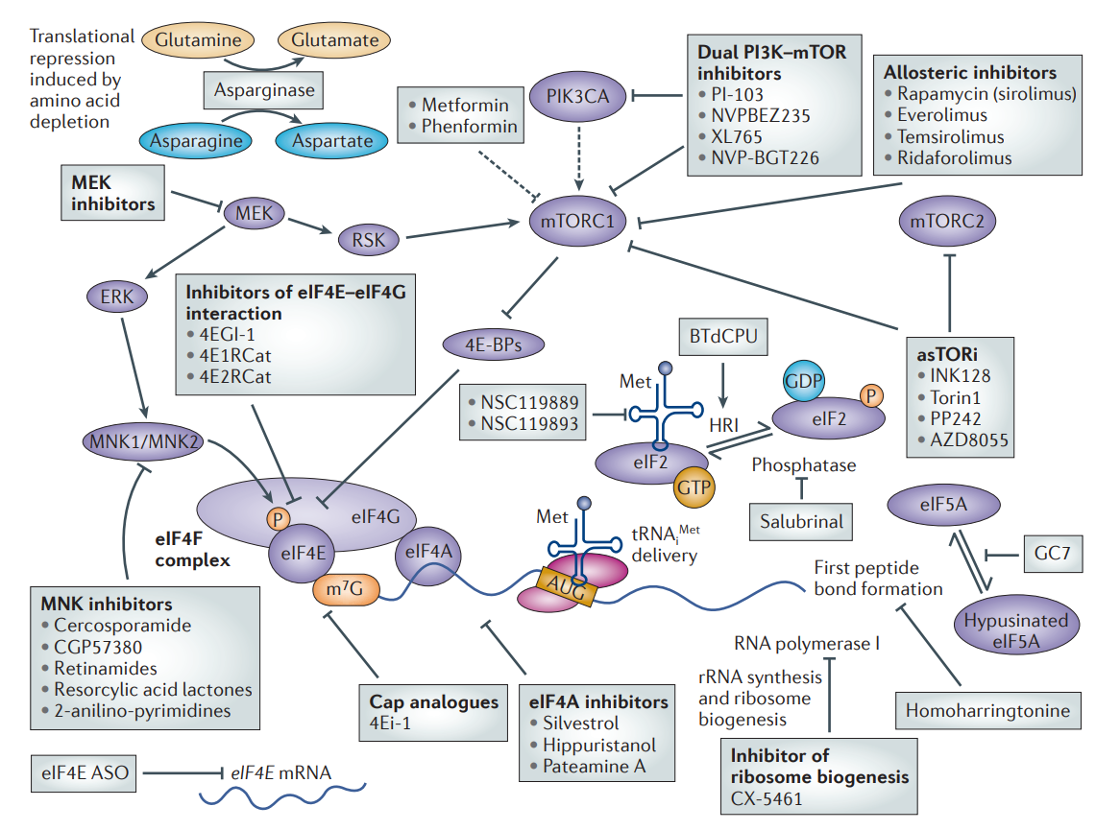
Translational control & disease
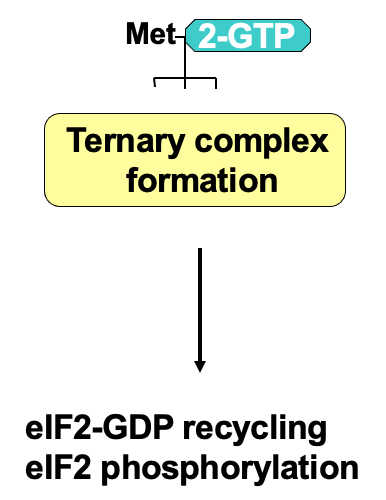
Regulation of ternary complex formation
- At the end of translation, eIF2 associates w/ GDP due to GTP hydrolysis during initiation
- eIF2 has 3 subunits: α, β, γ
* γ is bound to GTP or GDP - GTP required for translation initiation - associates w/ initiator tRNA & the ternary complex can participate in translation
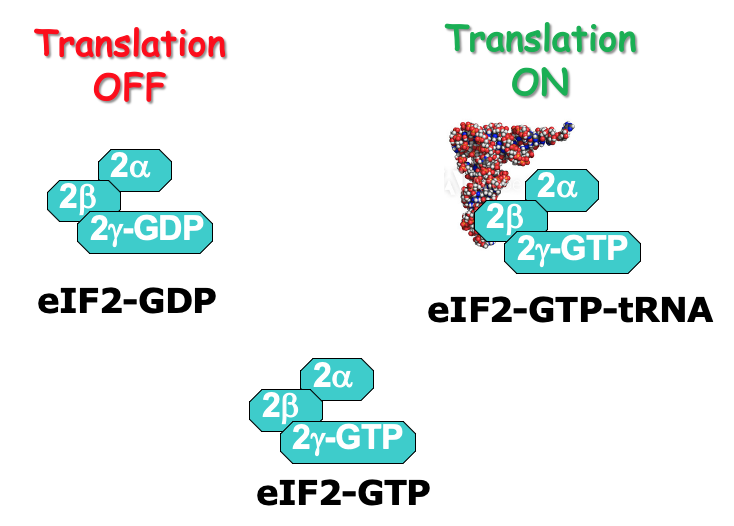
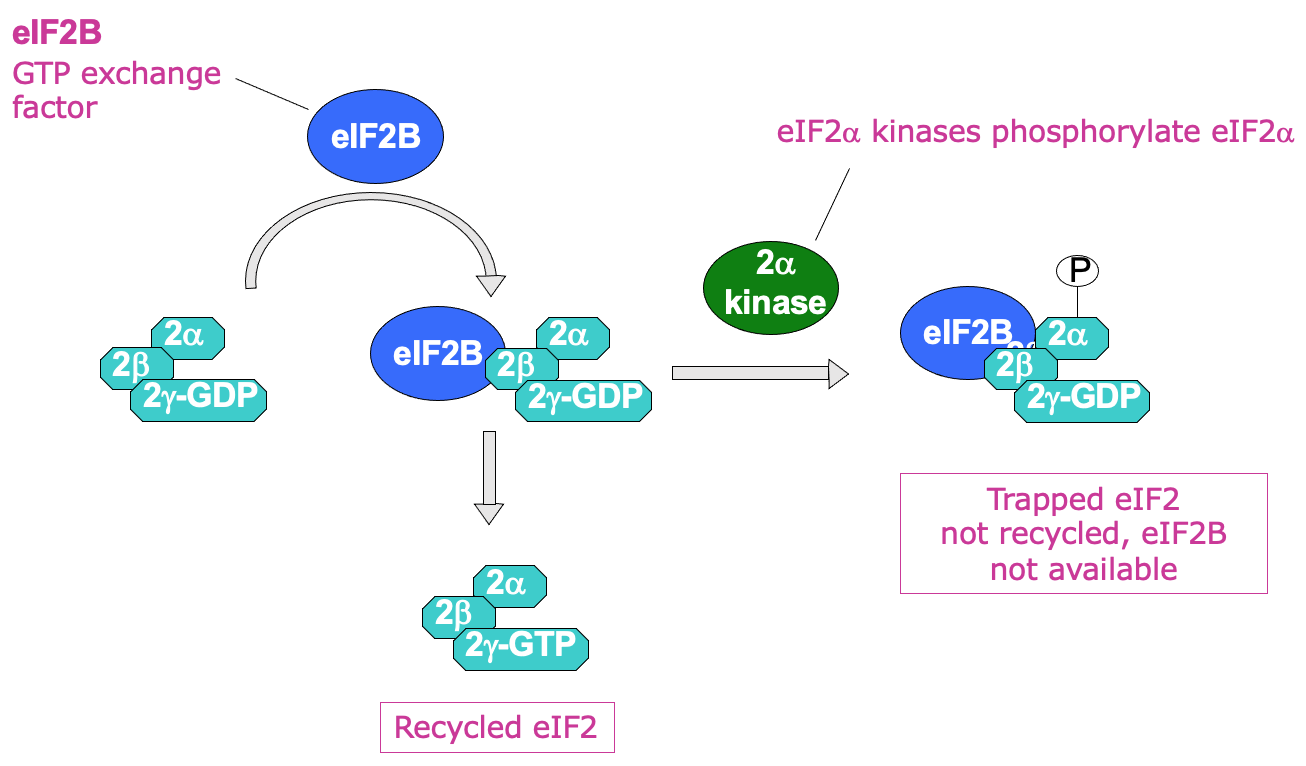
- eIF2 recycling → eIF2B (GTP exchange factor) required to help promote transfer of GDP to become GTP
* by directly binding eIF2 - Process of recycling can be regulated by phosphorylation of eIF2 α subunit using eIF2 α kinases
* when phosphorylated GDP bound eIF2 can’t be recycled into GTP - Phosphorylated eIF2 binds eIF2B v/ tightly → issue due to less eIF2B in the cell than eIF2
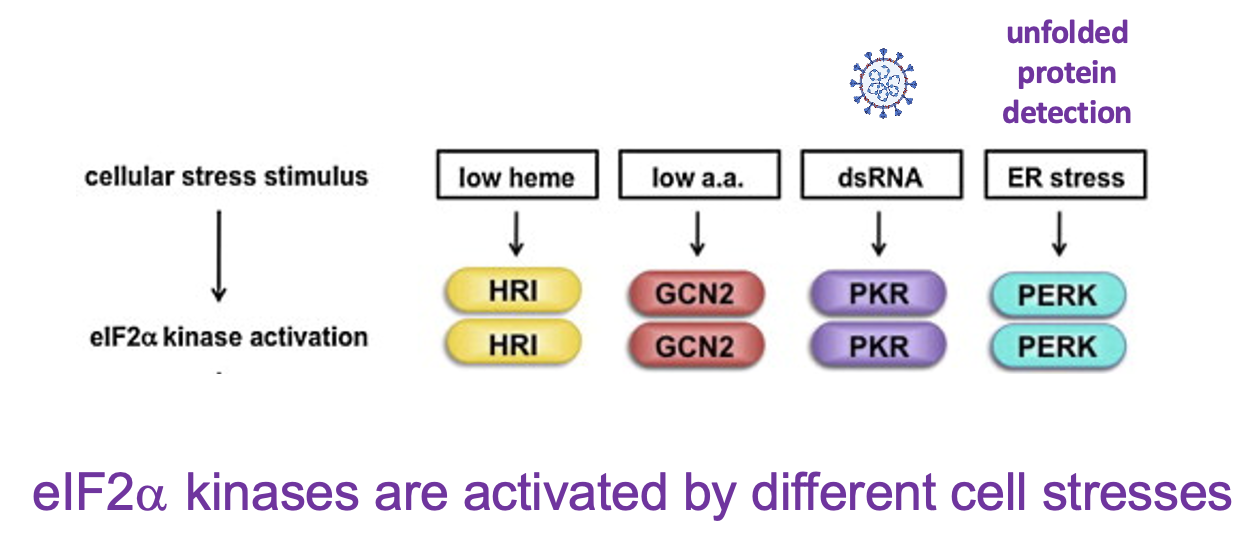
- 4 diff. types of eIF2 alpha kinases in mammalian cells → each processes diff. stress responses:
* HRI → sense low haem levels & inhibits translation of proteins to reduce production of globins
* GCN2 → sense low amino acid levels
* PKR → activated by dsRNA → hallmark of virus infection
* PERK → senses ER stress - response to presence of unfolded proteins in ER - ]]Q: why shut off translation when cells are stressed?]]
- turn off translation when AA are low
* prevents the wrong AA from being translated
* prevents E from being wasted on wrong resources - turn off translation if cells infected w/ virus
* viruses that infect cells contain own RNA/DNA or enzymes → so stops virus infection by preventing viral cells from using host ribosomes & stopping protein synthesis of viral proteins
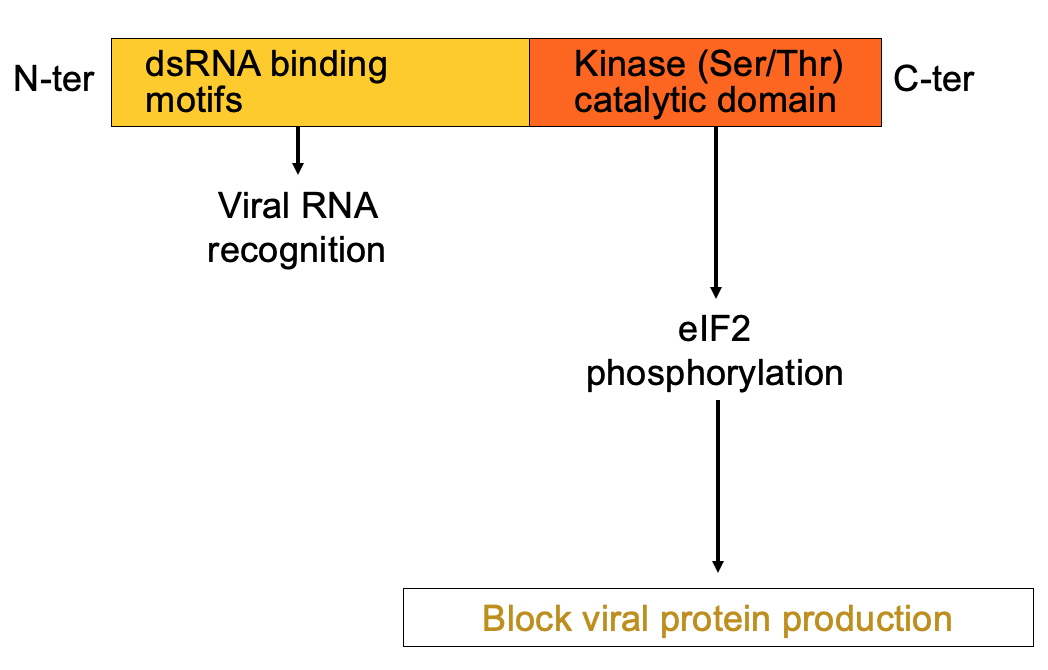
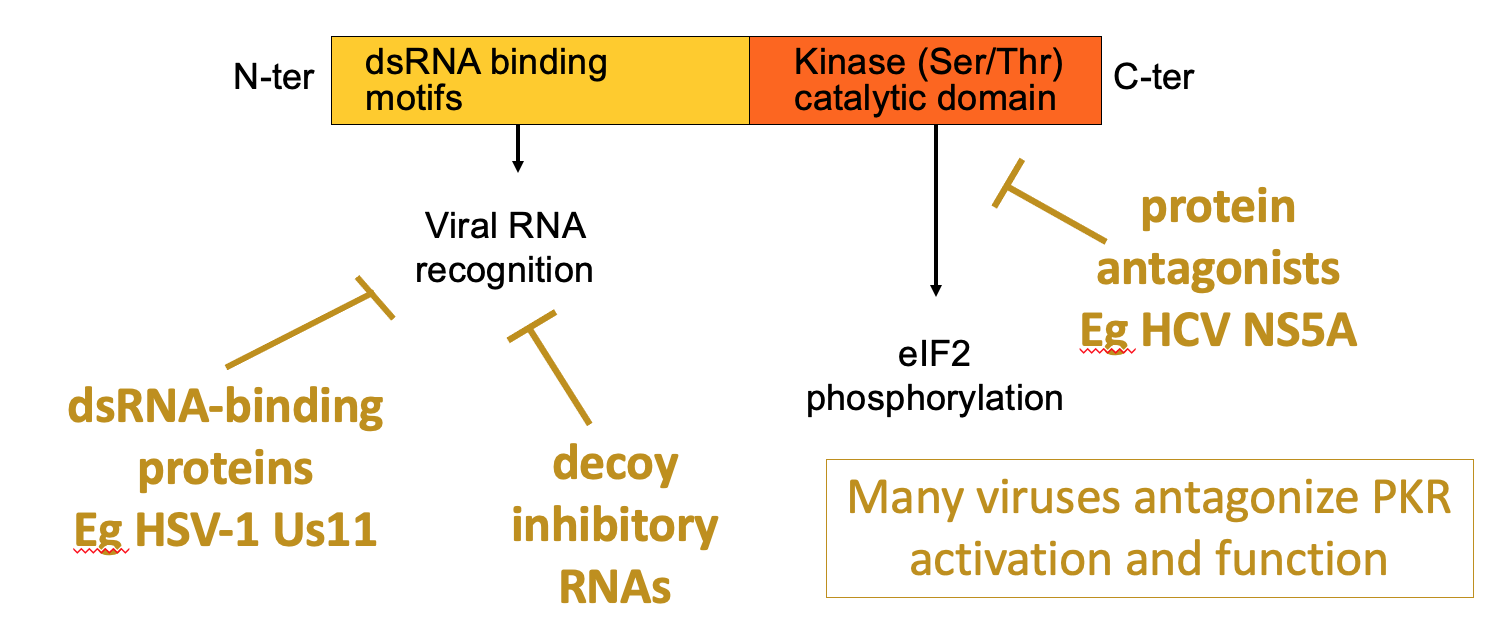
PKR structure, function & regulation
- dsRNA is hallmarker for viral infection
- PKR has 1 domain that helps bind dsRNA & kinase domain at C terminus that’s activated when PKR binds the dsRNA → leads to eIF2 alpha phosphorylation → blocks viral protein production
- Viruses have evolved ways to antagonise PKR
* e.g. viruses e.g. HSV1 Us11 - produce dsRNA binding proteins - shield recognition of dsRNA
* some viruses may decoy RNAs which are bound by PKR but don’t activate it
* HCV NS5A - make antagonists of kinase activity of PKR - so they directly stop activity or kinase domain
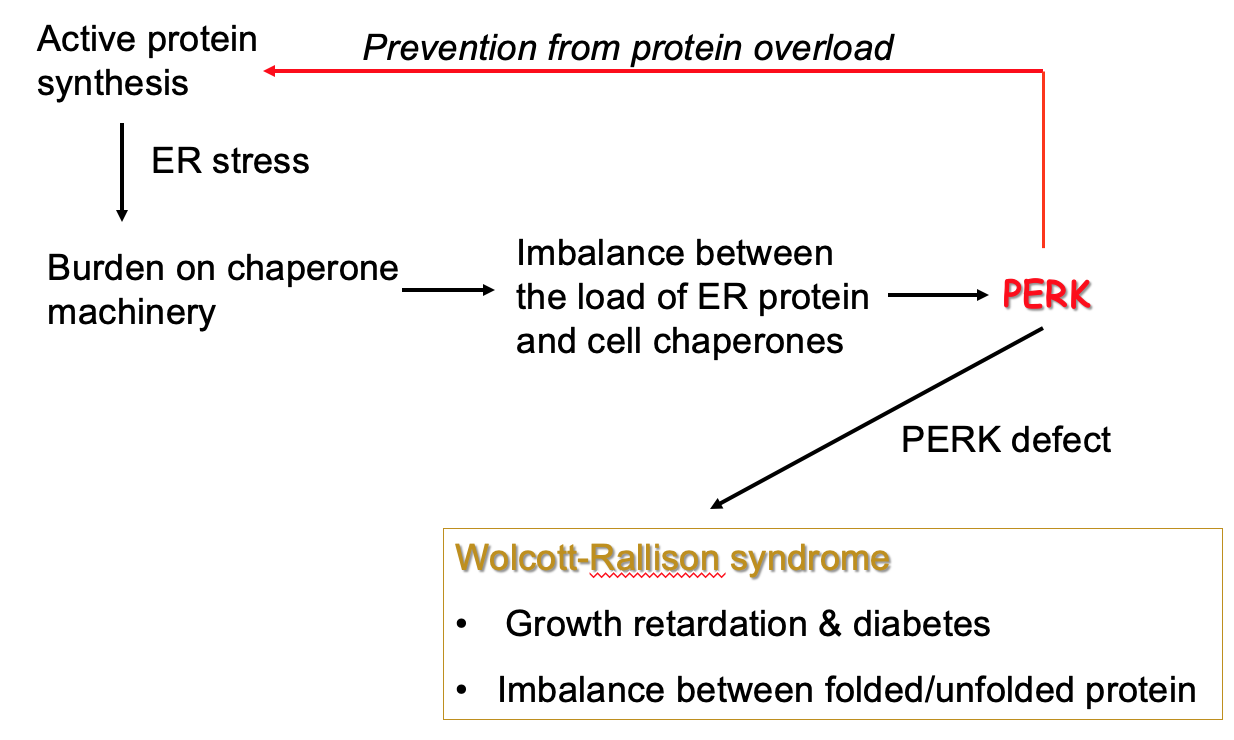
PKR-like ER localised kinase (PERK)
- Sense unfolded protein buildup on ER → burden on machinery that folds proteins into active form & causes imbalance
- Unfolded proteins may be toxic - need to stop build up
- PERK inhibits protein synthesis so that backlog is reduced
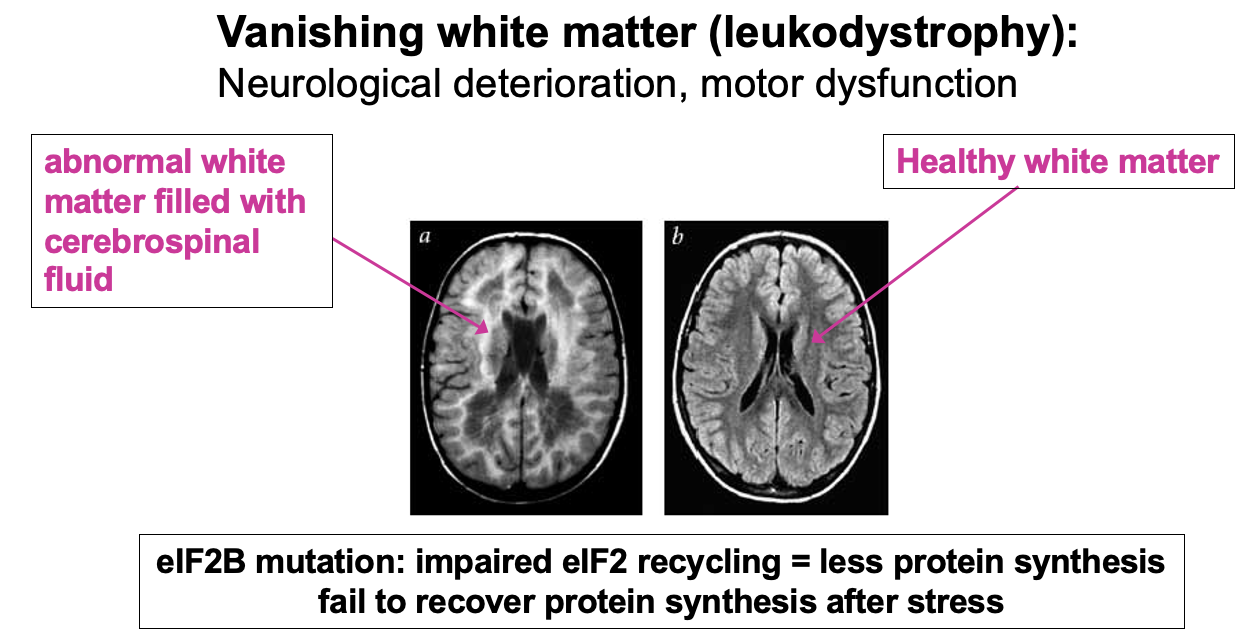
eIF2B mutations cause disease
- Example of failure to fully recycle eIF2 & maintain normal levels of ternary complex is in the disease Vanishing white matter (leukodystrophy)re
* specifically affects myelinated cells (white matter)
L3:
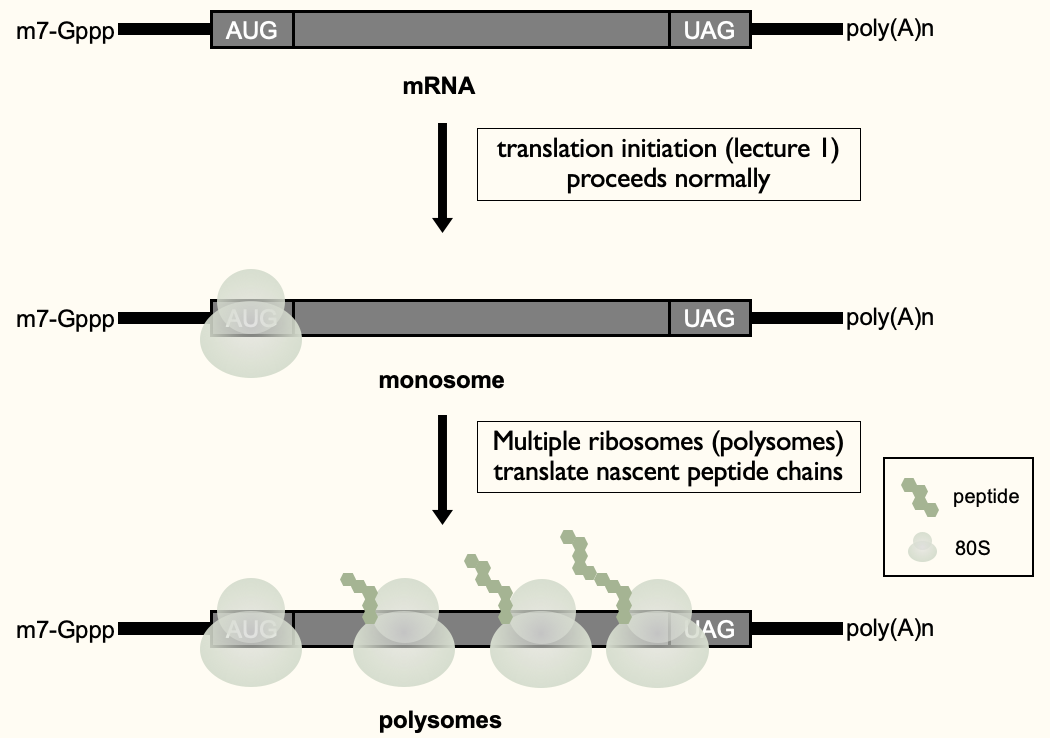
When translation goes right…
- Recruited ribosome to 5’ end (monosome) & then further rounds of recruitment of ribosomes to 1 mRNA - to maximise production of proteins from 1 mRNA (polysomes)
- Functional protein produced
When translational goes wrong…
- **Premature termination codon (PTC)**→ when a termination codon is present in middle of open reading frame - introduced by transcription error in nucleus or mutation in DNA
* only makes 1/2 a protein ∴ non-functional & waste of E
* toxic truncated proteins → e.g. in transcription factors - 2nd part of protein (not coded) may have contained a regulatory domain which helped keep control of enzyme
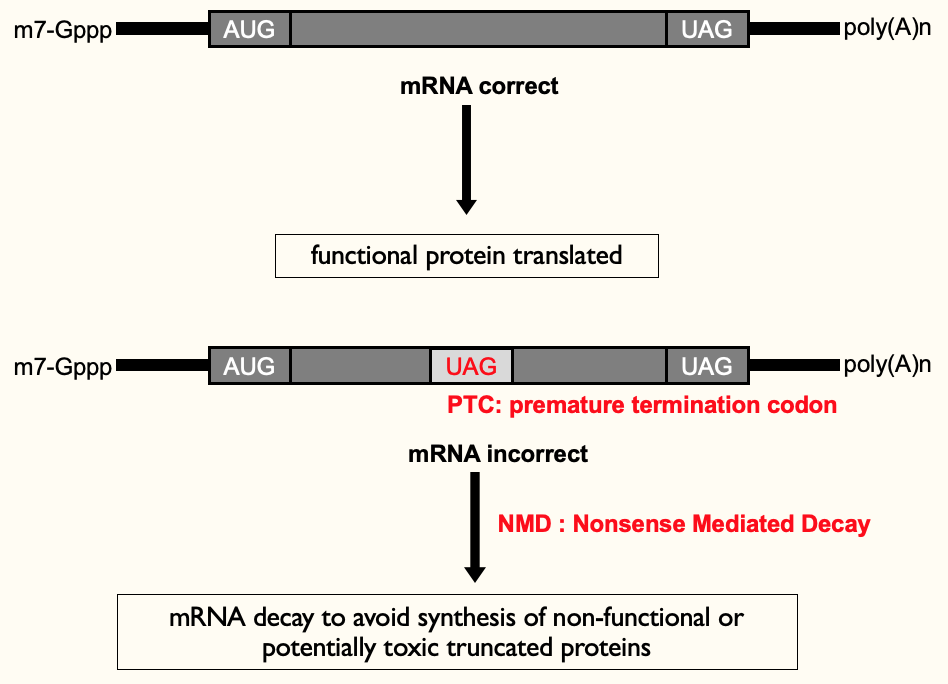
Nonsense Mediated Decay (NMD)
Features in mRNA that help to identify premature termination codon (PTC):
- **Exon-junction complex (EJC)**→ at exon-exon boundary
* during pioneer round, ribosome is recruited & as it elongates through open reading frame, it displaces the EJCs along the length of mRNA
* after pioneer round, no more EJCs & mRNA can be translated efficiently
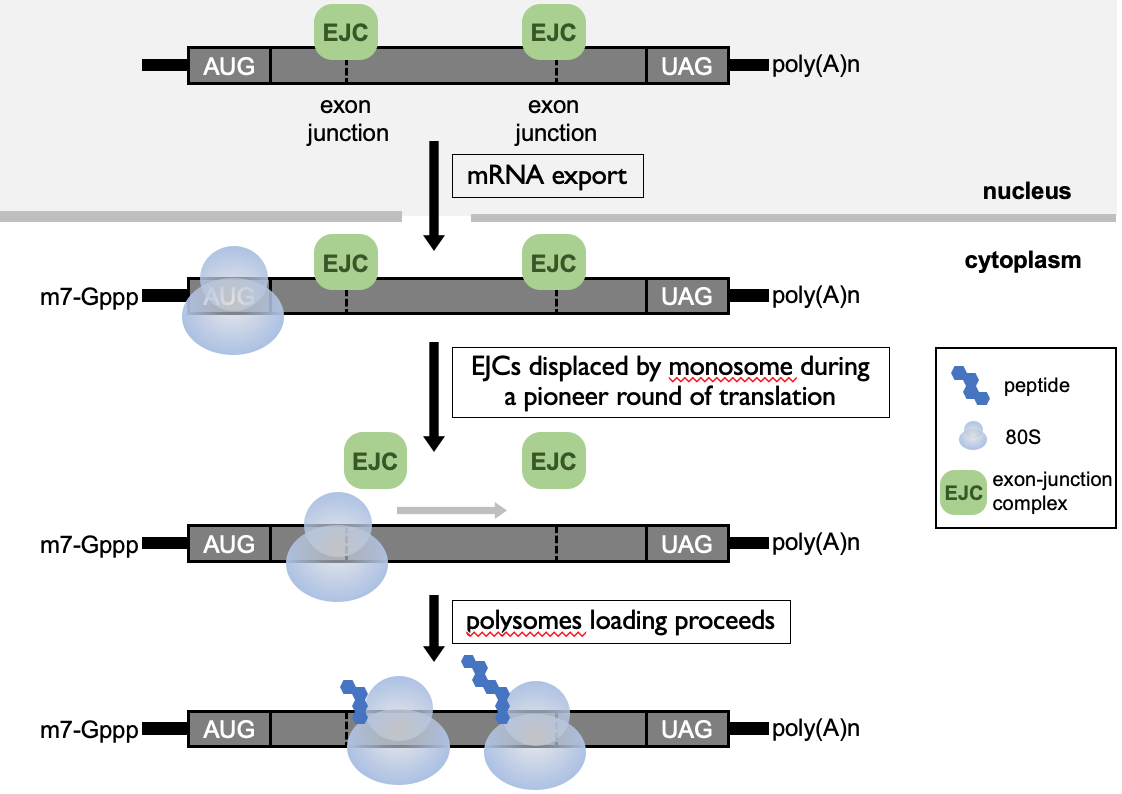
- When PTC is present - translating ribosome starts elongating along 1st portion of open reading frame & knocks off EJCs - then stops at termination codon to terminate
* downstream, still EJC present → alarm to ribosome that termination codon isn’t in correct place
* SURF complex formed due to paused ribosome in proximity of EJC → adaptor to bring in RNA decay effectors that dispose of mRNA that is detected as abnormal
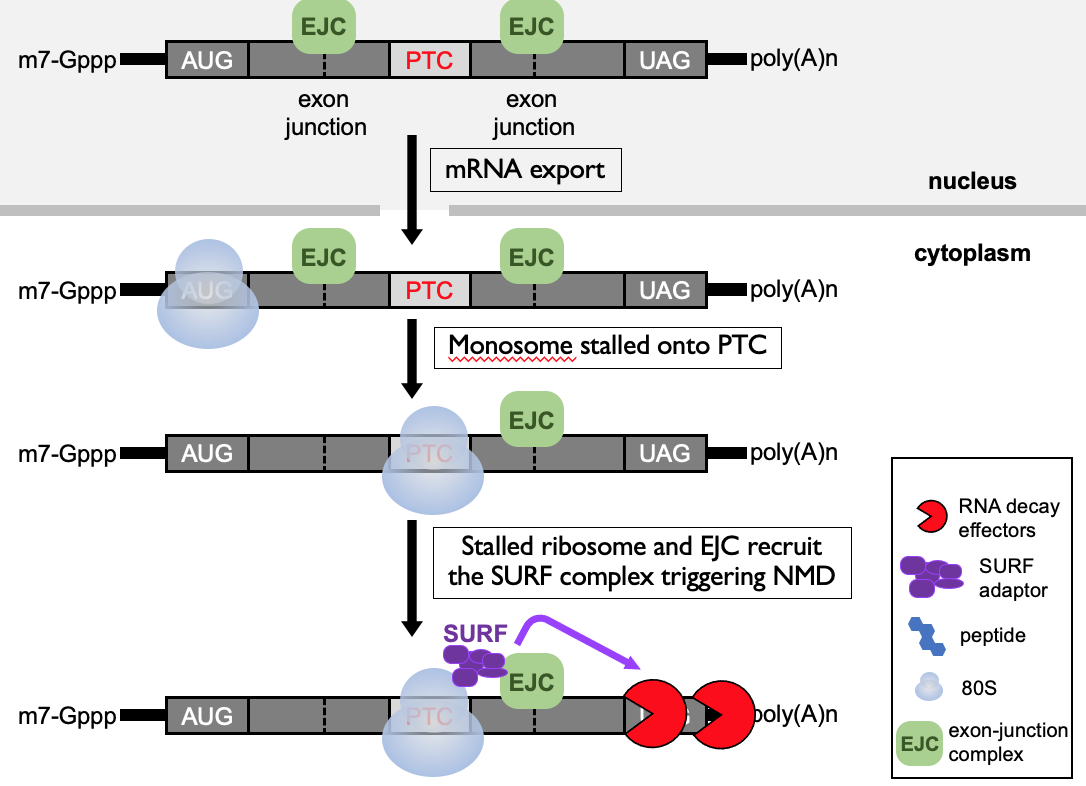
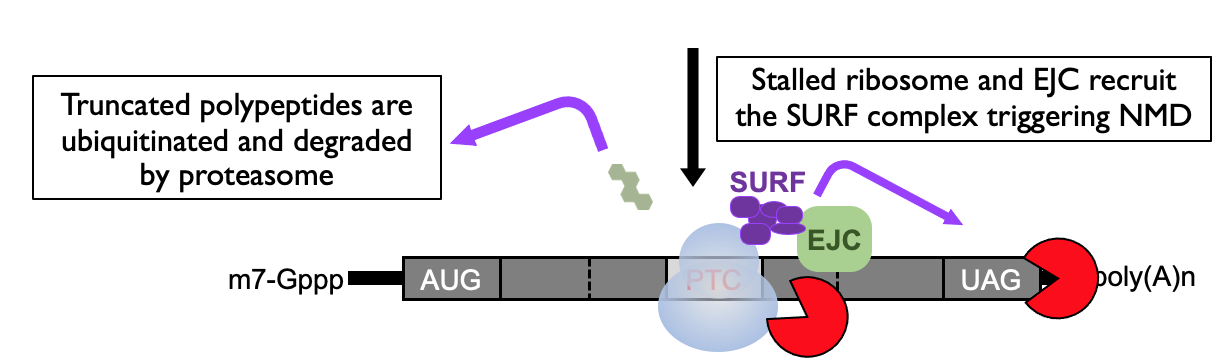
- Triggers a safety net to degrade the truncated polypeptide that’s been generated
- Decay that’s initiated is an endoribonucleolytic cleavage - cuts in middle of RNA
* complex also recruits SMG6
Other quality control pathways
- Nonsense-mediated decay (NMD) → (premature termination codons) quality control to eliminate problematic mRNA & limit production of toxic proteins in cytoplasm
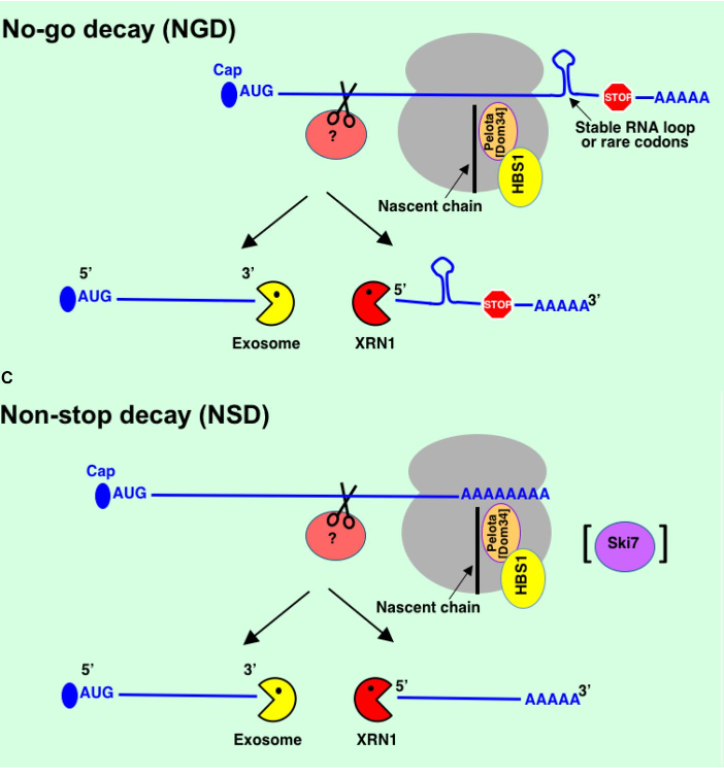
- No-go decay (NGD) → ribosome stalled in translation mRNAs e.g. it hits a stable RNA loop, rare codon present in mRNA or damage to RNA
* to rescue stuck ribosome - Pelota & HBS1 are recruited - Non-stop decay (NSD) → mRNAs w/o natural stop codons e.g. mistake in transcription, mutation in DNA or transcription termination hasn’t happened fully at end of gene ∴ added poly(A) tail to middle of seq. → ribosome starts translating into poly(A) tail
Specific mRNA translation regulation
- Most regulation occurs through 5’ & 3’ UTRs
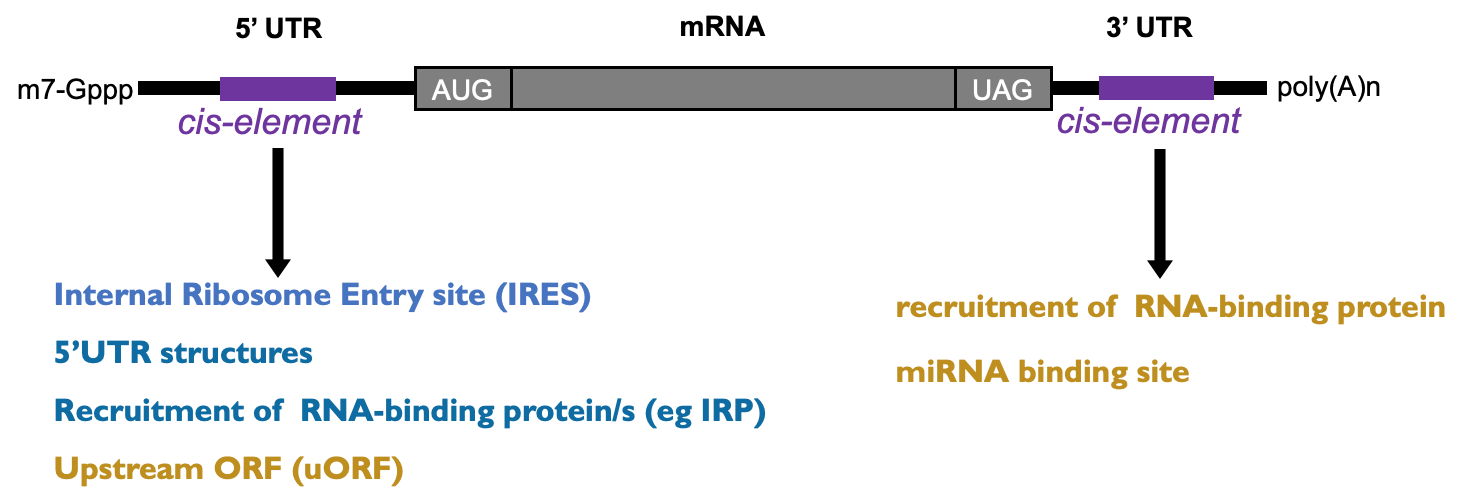
- When global mechanisms have specific consequences (uORFs)
- How miRNAs & RBPs can regulate translation from 3’ end
- how epitranscriptomic changes can influence translation
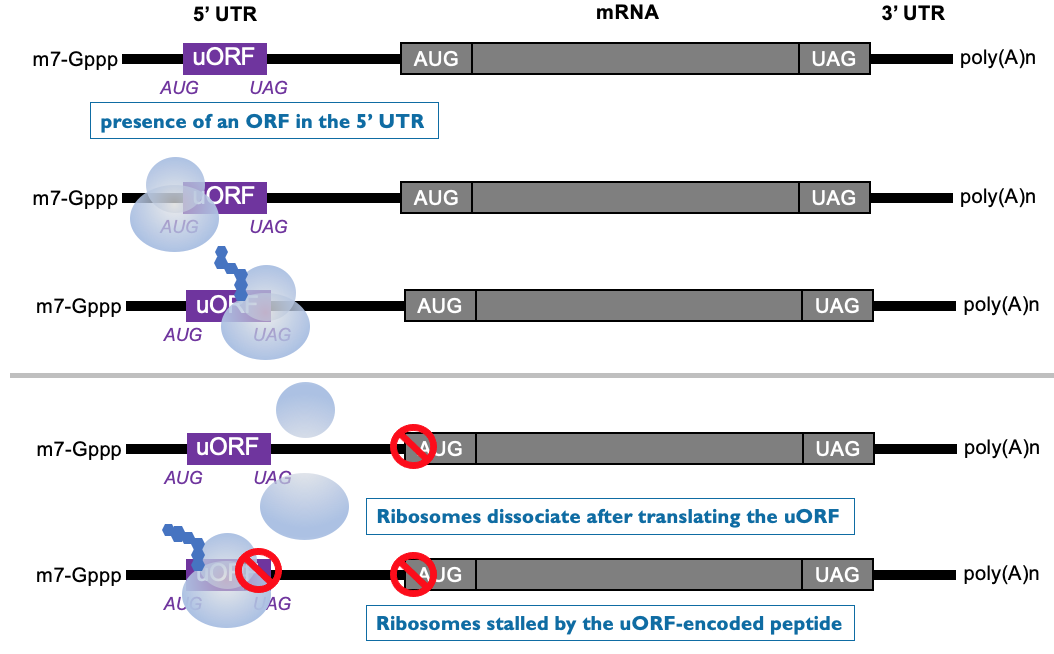
Translational repression by a uORF (upstream Open Reading Frames)
- Translation from central ORF is dependant on scanning from 5’ cap of a ribosome & initiating at AUG
- Some mRNAs have short ORF in 5’ UTR - encode for small peptides
* scanning ribosome initiates at upstream ORF to produce short peptide - at end of translation, ribosome terminates & leaves mRNA
* downstream ORF isn’t translated ∴ poorly translated as ribosomes won’t reach downstream AUG
* beginning translation at upstream ORF, inhibits initiation at downstream ORF
* peptide encoded by uORF can specifically interfere w/ translation at ribosome
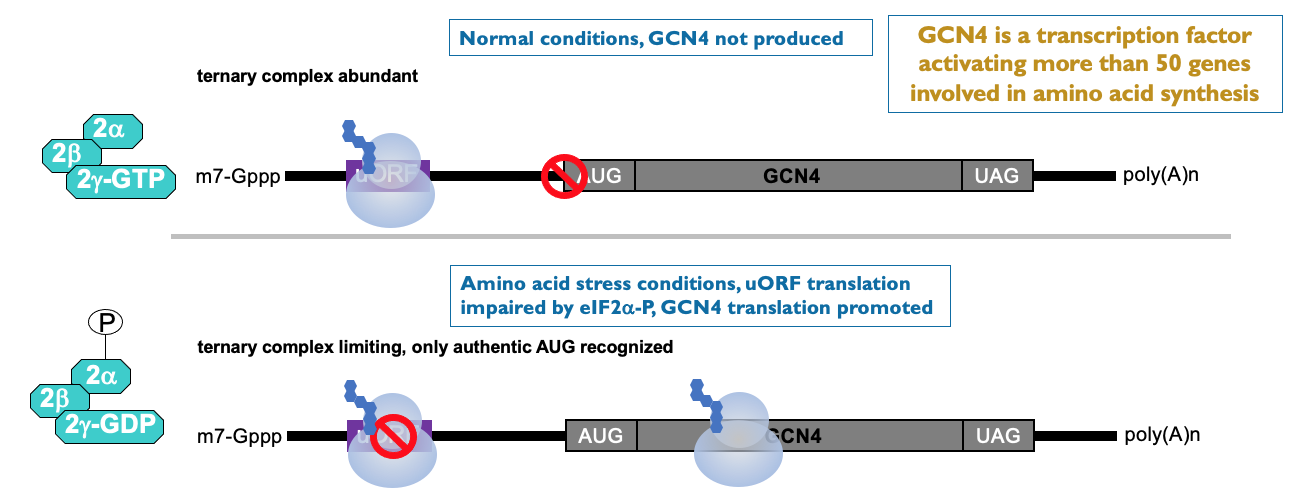
Translational control by a uORF (upstream ORF)
- mRNAs w/ upstream ORFs are generally encoding products that post-production needs to be v. tightly controlled & strongly inhibited most of the time
- These examples contain ORF & help ↑ translation when eIF2α is phosphorylated:
* e.g. GCN4 → transcription factor - activates large no. of genes involved in AA synthesis → most of time - no GCN4 produced as translation inhibited by upstream ORF
* low AA & eIF2α kinase has lead to phosphorylation of eIF2 complex → ↓ in ternary complex availability that delivers initiator tRNA to ribosome → ribosome initiated on GCN4 mRNA - scans along UTR but misses upstream ORF due to low availability of initiator tRNA to start translation
* translation improves on uORF containing mRNA from low base level
* ↑ production of GCN4 helps synthesis AA to get out of cell stress situation
* e.g. ATF4 → helps produce chaperone proteins to help resolve ER stress → improves translation
* GADD34 → helps guide phosphatase to dephosphorylate eIF2α
Translational control from 3’ end - RNA-BPs
- RNA binding protein binding to cis-acting element in 3’ UTR can impact on initiation at 5’ cap
- Cis-acting element→ element in RNA which acts on same RNA (on itself)
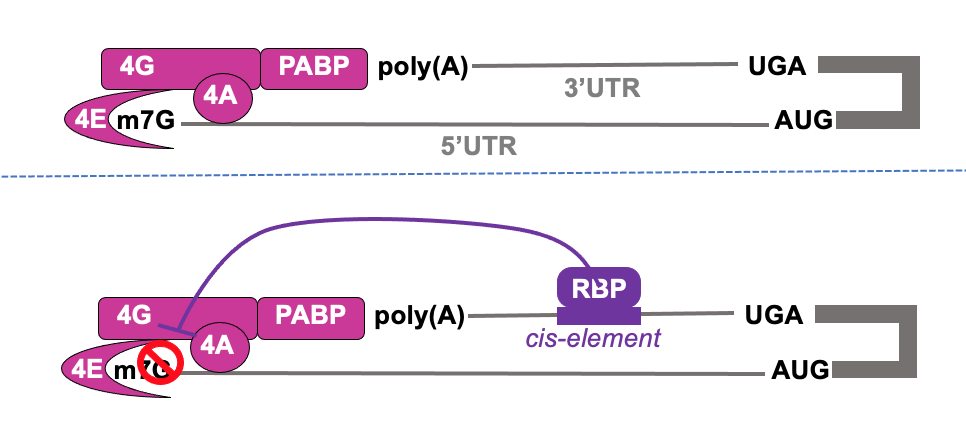
Translational control from 3’ end- e.g. caudal & bicoid
- Patterning needs to occur so that v. early on in development, embryo knows which end in head & which is tail
* done molecularly by gradients of proteins (morphogens) that influence this patterning - communicate their fate to cells during embryogenesis
* e.g. bicoid → gene is selectively expressed at head end (anterior)
* transcription factor that’s important for production of other patterning mols. & acts as translational regulator on mRNA called caudal
* e.g. Caudal → important for patterning the posterior end of embryo, but it’s mRNA is expressed throughout the embryo - bicoid stops protein being where bicoid is present - Patterning the early drosophila embryo:
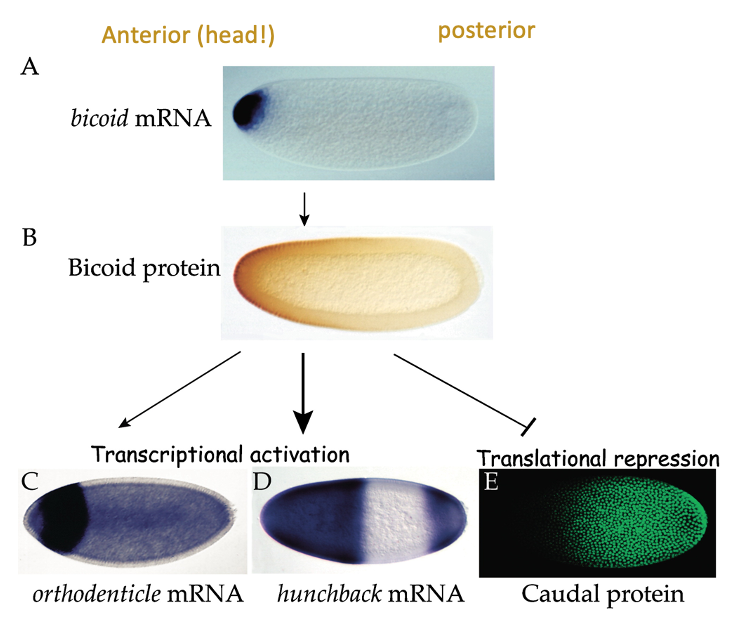
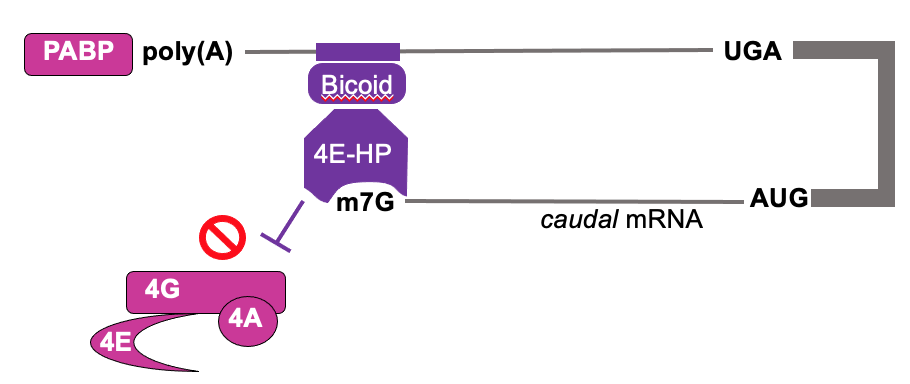
- Caudal is an important patterning mol. during embryogenesis
- Bicoid (RBP) is RNA binding protein that interacts w/ 3’ UTR of caudal mRNA
- Bicoid recruits 4E-HP (4E homologous protein) to bing caudal cap but not initiate translation
- 4E-HP blocks eIF4F binding & translation of caudal mRNA
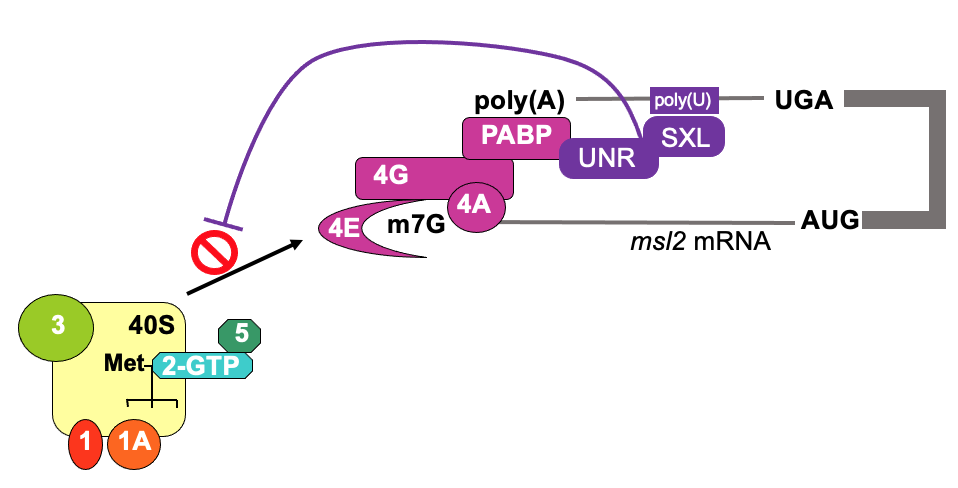
Translational control from 3’ end - e.g. msl2 & SXL
- In drosophila, to achieve dosage compensation of X-linked genes, in male flies, the dosage compensation complex (feat. msl-2) is required for hypertranscription (massively upregulate) of single X chromosome
* mediated by protein msl-2
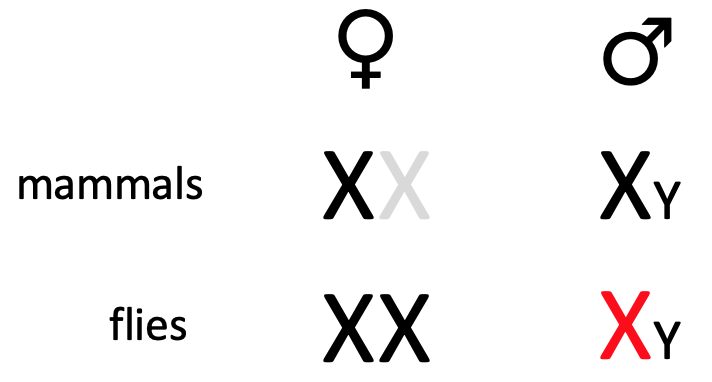
- msl2 mRNA is transcribed in both, but translated in male but not female during development - to repress production in female flies
- SXL (“sex-lethal”) is only expressed in females & is a binding protein
- SXL & UNR bind to 3’ UTR of msl-2 & interact w/ poly(A) binding protein & influences eIF4F complex to stabilise it’s interaction w/ pol(A)
* becomes too stable - prevents 4F to recruit 43s ribosome - translation is inhibited on msl2 in female flies - In early embryogenesis, mutation of SXL will compromise establishment of sex
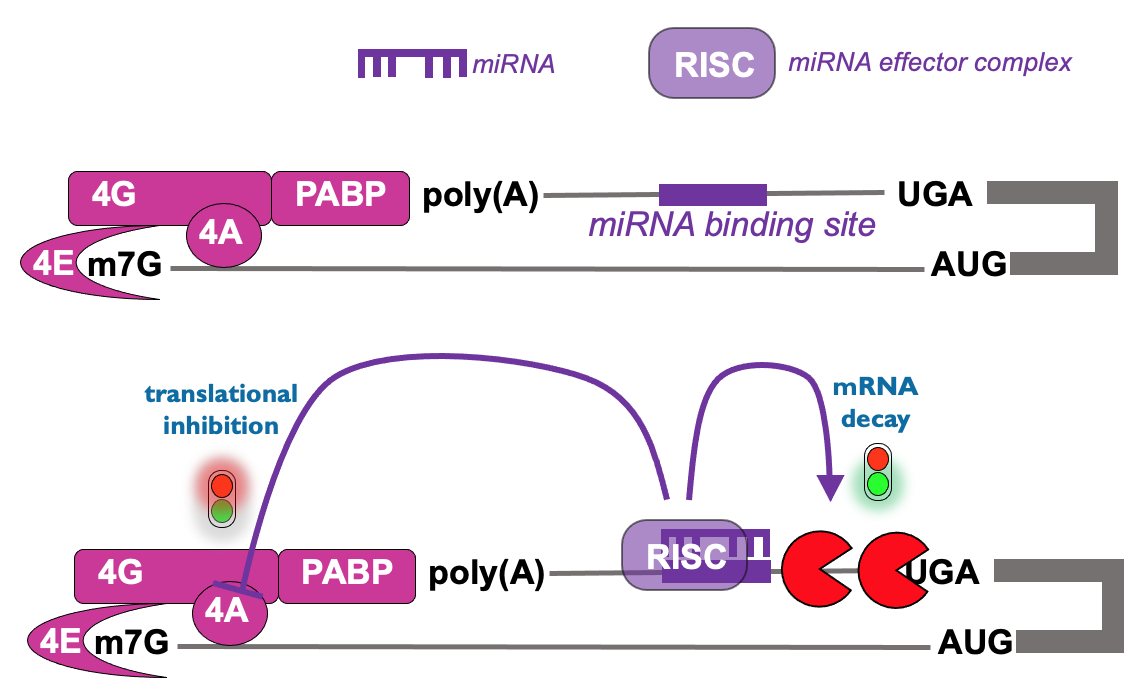
Translational control from 3’ end - microRNAs
- Regulatory RNA seq. in 3’ UTR that mediates binding of RNA complex & leads to inhibition of translation initiation → occurs prior to mRNA decay & also occurs on microRNA targets
- other example of mech. of control from 3’ UTR
Translational control - beyond the genetic code
- RNA bases can be chemically modified in a way that doesn’t Δ the code → the epitranscriptome
* influences mRNA fate
* e.g. methyl-cytosine or methyl-adenosine
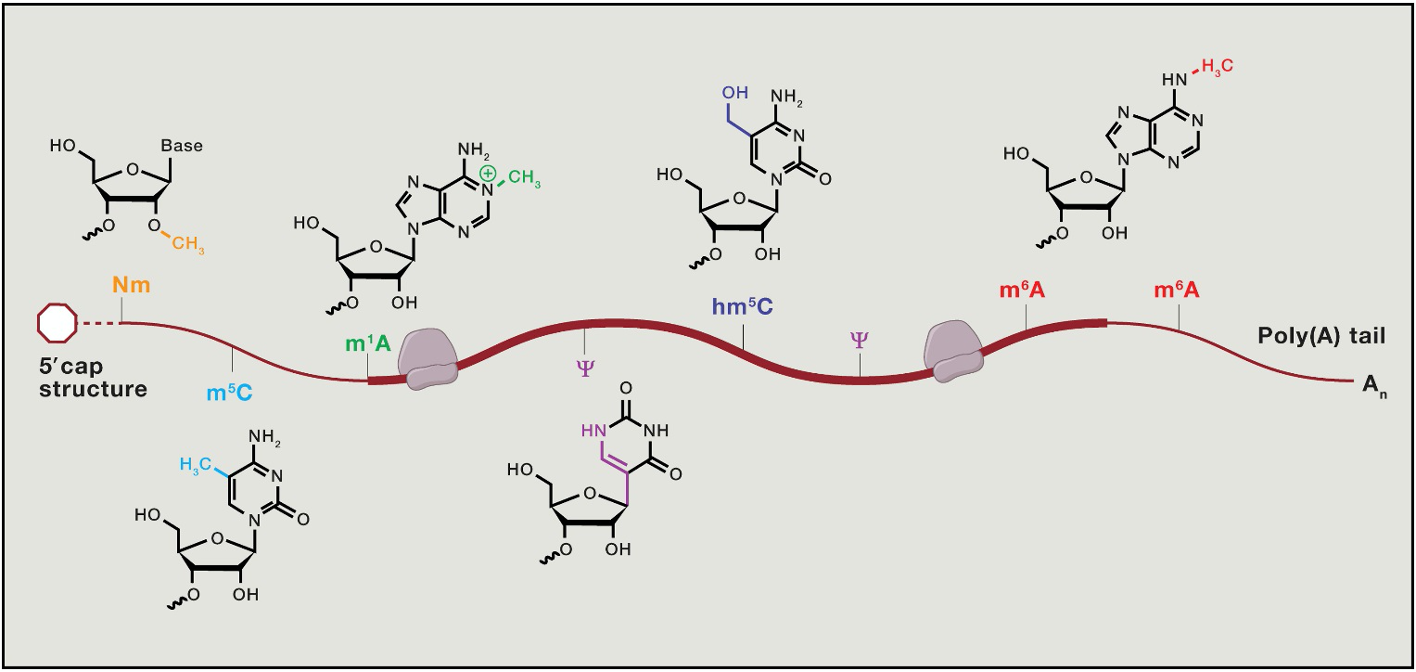
- mRNA modifications correlate w/ mRNA features
* most freq. modification - m7 cap → methyl 7 on guanosine cap
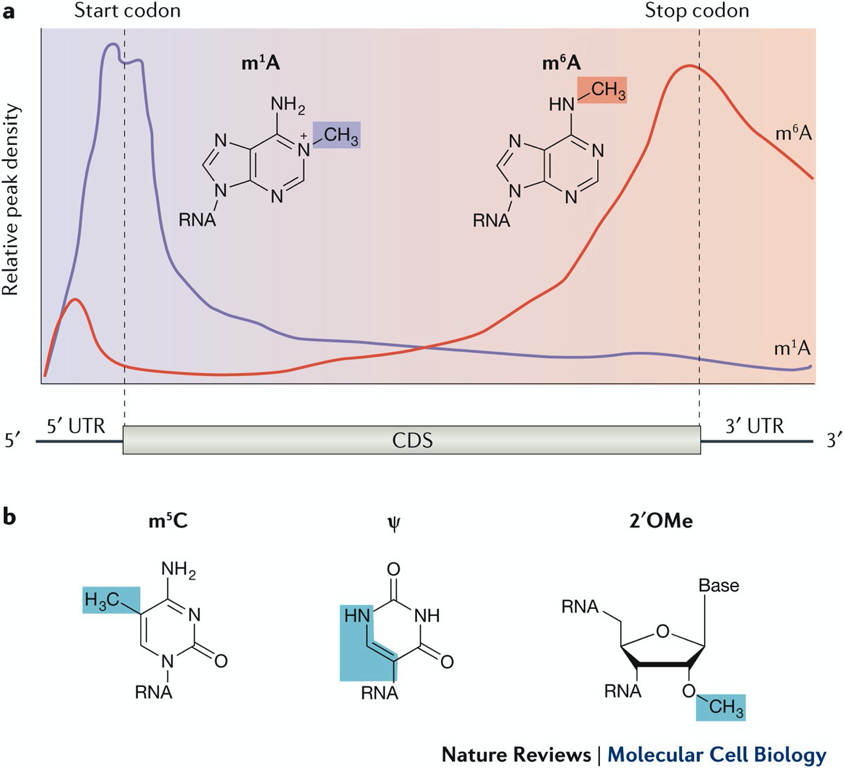
Translational control - N6-methyladenosine (m⁶A)
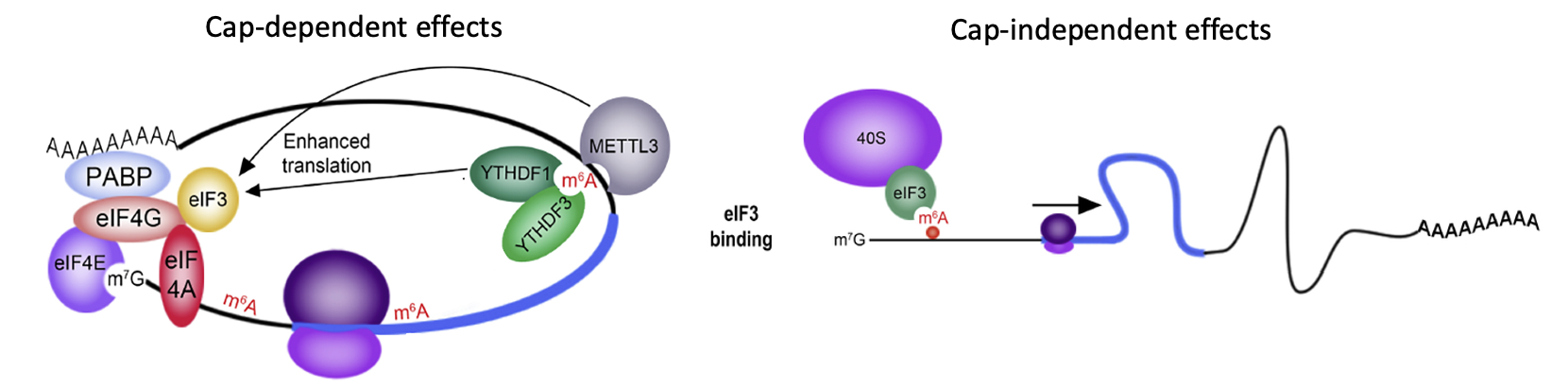
- m⁶A promotes mRNA translation by multiple mechanisms
- “reader proteins” can bind m⁶A in 3’ UTR & enhance translation initiation e.g YTHDF1 & 3
- METTL3 → helps install m⁶A & adds on m⁶A → promotes translation of message m⁶A is on (cap-dependant translation)
- When m⁶A is present on 5’ UTR it can function like IRES → translation factor eIF3 can be recruited by m⁶A directly
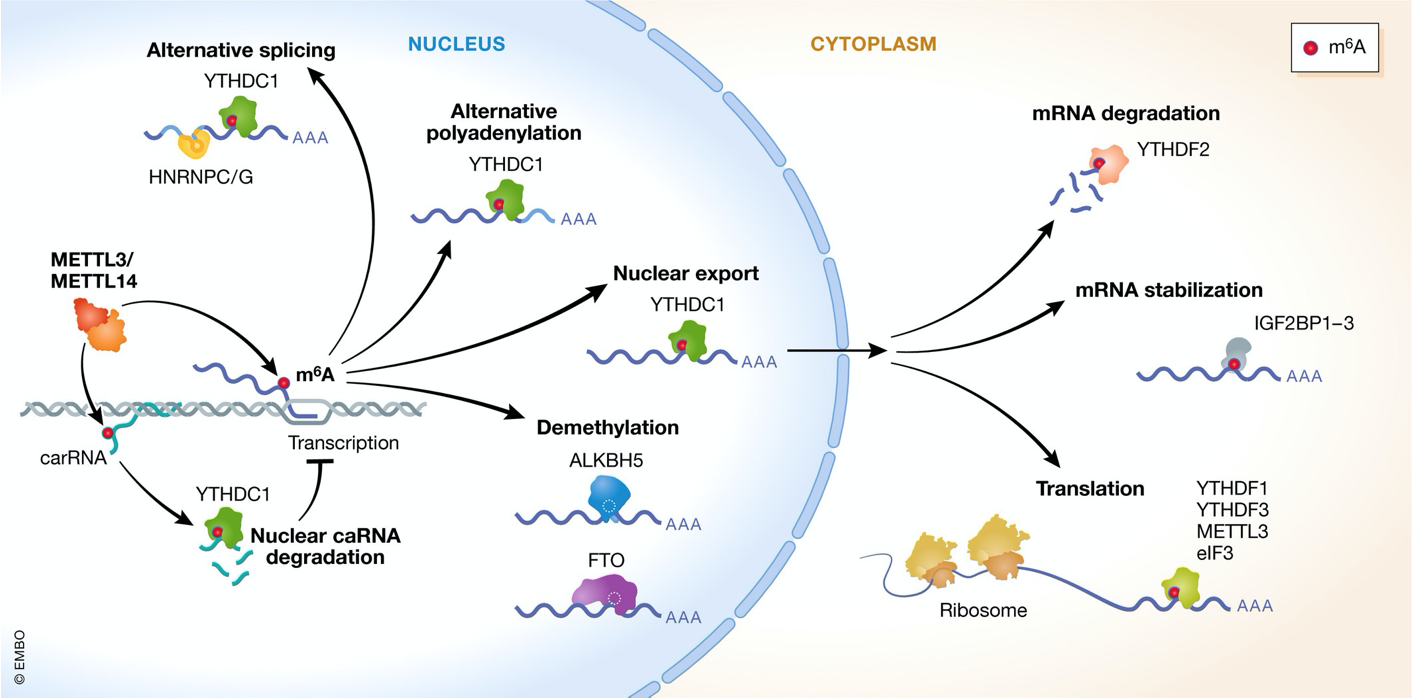
m⁶A controls multiple RNA-regulatory processes
- m⁶A implicated in control of RNA stability & degradation
- Can impact nuclear export & splicing
- Functions are dictated by diff. kinds of BPs that binds the m⁶A modification
- can be demethylated later on → turned on or off → epitranscriptomics
Questions & themes to consider
- What mechanisms exist to ensure accurate translation of error-free mRNAs?
- Which control mechanisms target general translation factors but selectively affect the translation of a subset of mRNAs?
- Which mutations or changes in eIFs are associated w/ disease?
- Which control pathway normally prevent disease?
- How might a circular RNA, lacking a 5’ cap, be translated?