AP BIO Unit 6 Review - Gene Expression and Regulation
<<Transcription<<
Vocab
%%Transcription%%: The synthesis of RNA using a DNA template
Complementary/
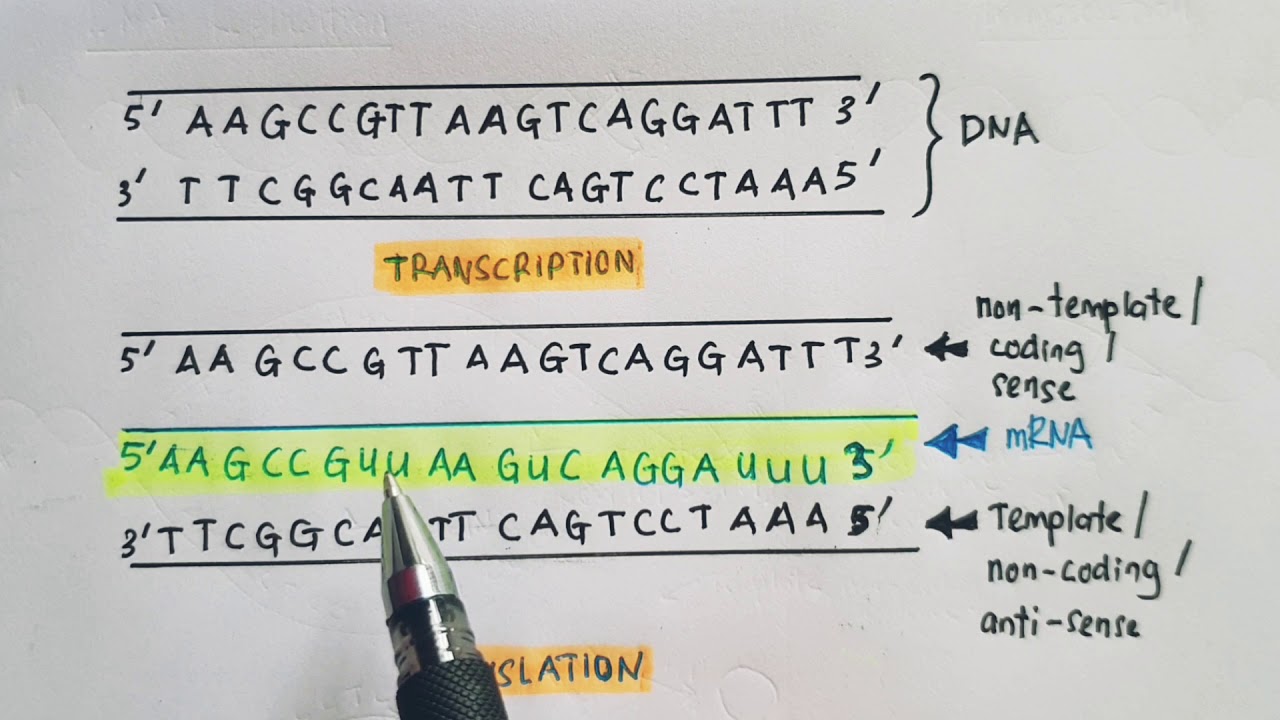
%%Coding Strand%%: The DNA strand whose base sequence is similar to its primary transcript (RNA)
%%Template Strand (non-coding/antisense)%%: The strand that is used during transcription to produce RNA
- the RNA synthesized using the template strand will have the same base pairs as the coding strand (except for T and U)
%%origin of replication%%: Site where the replication of a DNA molecule begins, consisting of a specific sequence of nucleotides
%%replication fork%%: A Y-shaped region on a replicating DNA molecule where the parental strands are being unwound and new strands are being synthesized
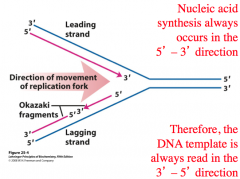
\ Nucleic acid synthesis occurs in the __ direction
- 3’ → 5’
DNA is read from __
- 5’ → 3’
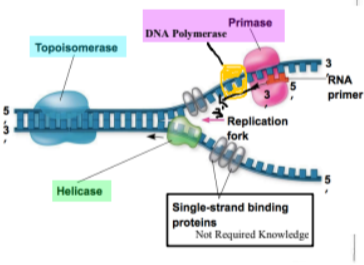
{{Enzymes:{{
- ==Helicase== * unwinds strands (zipper)
- ==polymerase== * synthesizes new strands of DNA (builder) * limitations * requires RNA primers * discontinuous on the lagging strand
- ==Ligase== * joins fragments (glue)
- ==Topoisomerase== * prevents overwinding * relives stress from unoiling
- ==Primase== * creates primers where DNA synthesis is initiated
- ==single strand binding proteins== * stops the single strands from joining and reforming
Other Vocab
^^Leading Strand:^^ The new complementary DNA strand is synthesized continuously along the template strand toward the replication fork in the mandatory 5′ → 3′ direction
^^Okazaki Fragments^^: A short segment of DNA synthesized away from the replication fork on a template strand during DNA replication. Many such segments are joined together to make up the lagging strand of newly synthesized DNA
^^Lagging Strand^^: A discontinuously synthesized DNA strand that elongates using Okazaki fragments, each synthesized in a 5′ → 3′ direction away from the replication fork
<<Translation<<
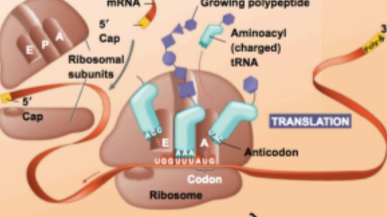
mRNA is translated as codons into amino acids
- @@codon@@ - A sequence of three consecutive nucleotides in a DNA or RNA molecule that codes for a specific amino acid
Translation starts in the ribosome
- when rRNA interacts with mRNA
- AUG start codon
Stops at a stop codon
- UAG, UAA, UGA
- signal the end of the polypeptide chain during translation
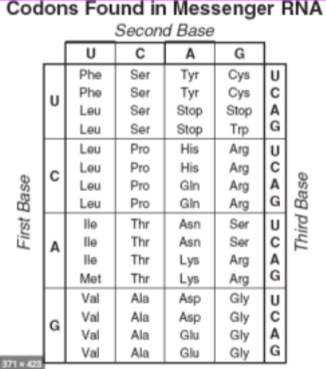
\ Many amino acids are encoded by more than one codon
- @@wobble@@ - the redundancy in the genetic code such that the same amino acid may be encoded by multiple codons
\
{{Pro vs Eu{{
In prokaryotic cells, transcription and translation happen simultaneously in the cytoplasm
in eukaryotic cells, transcription happens in the nucleus, and mRNA must be exported
\ intron and exon spicing occurs in eukaryotic cells
- alternative splicing → different variation
in eukaryotic cells, a poly-A tail and gtp cap are added
- prevents degradation from hydrolytic enzymes
- facilitate the export of mRNA from the nucleus
- help attach to ribosomes
}}Regulation and Operons}}
\ Operons - https://www.youtube.com/watch?v=F7wRwHV_J5Q
{{Lac operons{{
when lactose is present, the repressor becomes inactive
- genes are expressed to break down lactose
{{Trp Operon{{
When trp is not present, the repressor stats inactive
- genes are expressed to synthesize trp
\ ^^Histone Acetylation^^ - Loosens chromatin structure promoting the initiation of transcription
^^Methylation^^ - can condense chromatin and lead to reduced transcription
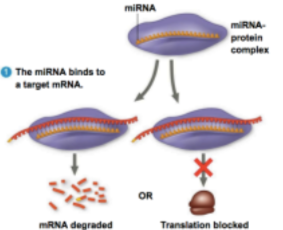
\ %%RNA Interference%% - Inhibition of gene expression by RNA
%%MicroRNA%% - small, single-stranded RNA molecules that bind to complementary mRNA sequences
- degrade mRNA
- block its translation
%%Small Interfering RNA%% - a double-stranded RNA molecule that is non-coding operating in the RNA interference pathway
\ \
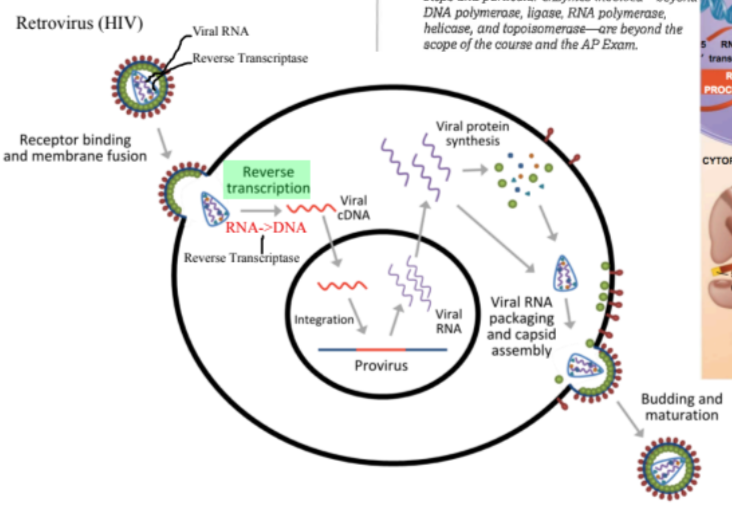
[[Reverse Transcriptase[[
<<Reverse transcriptase<<
- A reverse transcriptase is an enzyme used to generate complementary DNA from an RNA template, a process termed reverse transcription
the normal sequence of information
- DNA → RNA → Protein
Sequence in retrovirus
- RNA → DNA → RNA → Protein
\