Chapter 22: Single Nucleotide Polymorphism Profiling
22.1: Basic Characteristics of SNPs
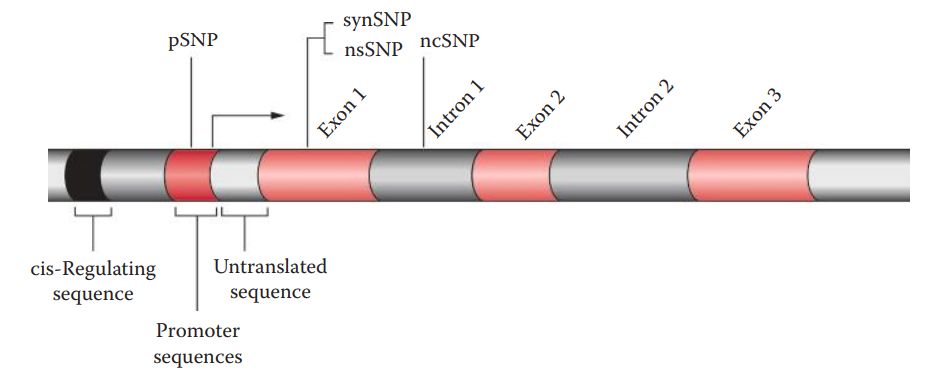
- Single Nucleotide Polymorphism (SNP): It constitutes a single-base-pair change originating from a spontaneous mutation that can be a base substitution, insertion, or deletion at a single site.
- If an SNP originating from a spontaneous mutation occurs in the germ line, it can be inherited by offspring and can spread in the human population.
- Advantages of using SNP loci as markers for forensic application: * SNPs are abundant within the human genome; therefore, a sufficient number of SNP loci that are suitable for human identification can be selected. * In SNP analysis, amplicons are usually 50–100 base pairs in length and are smaller than that in STR analysis. * SNP loci have lower mutation rates than STRs and are, therefore, useful for specialized human identification such as paternity testing. * It is possible to achieve high-throughput SNP analysis by utilizing multiplex SNP assays.
22.2: Forensic Applications of SNP Profiling
HLA-DQA1 Locus
- HLA-DQA1 gene: A member of the human leukocyte antigen (HLA) family, which contains a large number of genes involved in the immune response in humans.
- DQα AmpliType® kit: The first commercial kit, developed in the late 1980s by Cetus Corporation in Emeryville, CA.
- Polymarker systems are: * Capable of analyzing a small amount of DNA sample. * Capable of analyzing degraded DNA samples which is not possible in VNTR analysis.
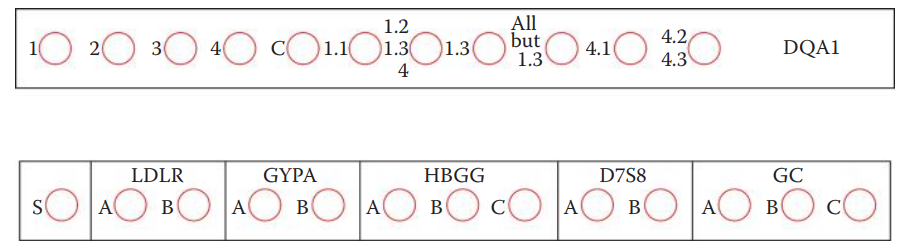
- Both DQα AmpliType® and Polymarker kits utilize the allele-specific oligonucleotide (ASO) hybridization technique.
* ASO technique analyzes single-nucleotide variations, such as SNPs, at a given locus.
* It is based on the principle that ASO probes, usually 14–17 bases in length, hybridize to their complementary DNA sequences to distinguish known polymorphic alleles.
\
Current and Potential Applications of SNP Analysis
Forensic Applications of SNP Profiling
| Testing | Candidate SNP Loci | Application |
|---|---|---|
| Identity | Autosomal SNPs | Human identification and Human Identification via degraded DNA. |
| mtDNA SNPs | Human identification | |
| Y chromosome SNPs | Paternity testing | |
| Biogeographical origin | Ancestry informative markers (AIMs) | Physical characteristics |
| Biogeographical origin | Ancestry informative markers (AIMs) | Physical characteristics |
| P (gene has role in pigmentation) | Eye color identification (investigative lea | |
| Pathology | KCNH2 (cardiac potassium channel gene) | Determining cause of sudden death from cardiac arrhythmia long QT syndrome |
| SCN5A (encodes cardiac sodium channel gene) | Determining cause of sudden death from cardiac arrhythmia long QT syndrome | |
| Toxicology | CYP2C19, CYP2D6, CYP3A4, CYP2E1 (drug metabolizing enzyme genes) | Investigation of drug overdose (including death) |
\
22.3: SNP Techniques
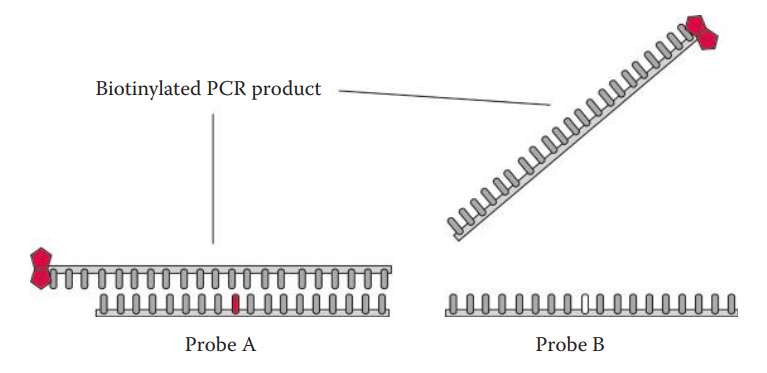
- In allele-specific hybridization, allele discrimination is based on an optimal condition allowing only the perfectly matched probe–target hybridization to form.
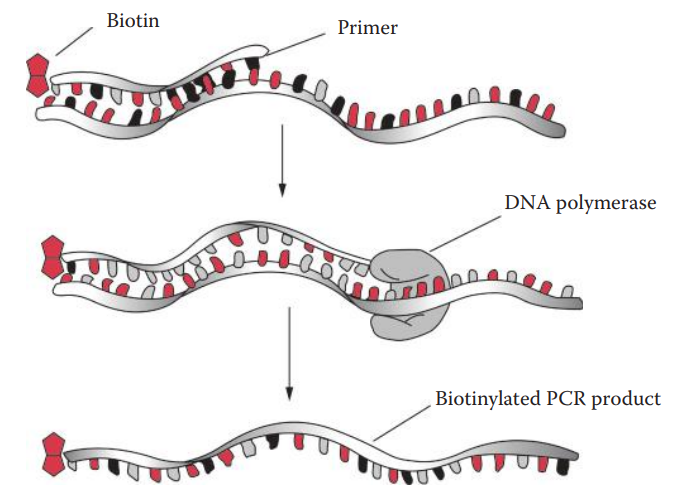
- Primer extension methods: These are based on the ability of DNA polymerase to incorporate specific deoxynucleotides (dNTPs) complementary to the sequence of the template DNA.
- Allele-specific oligonucleotide ligation: It is based on the condition that only the allelic probe perfectly matched to the target is ligated.
- In the invasive cleavage method, allelic discrimination is based on DNA sequence-specific cleavage by endonucleases.
\
Next-Generation Sequencing Technologies
- In de novo genome sequencing, uncharacterized genomes or characterized genomes with substantial structural variations are sequenced.
- In resequencing applications, characterized genomes are sequenced. Sequence reads are assembled against an existing reference sequence to identify sequence polymorphisms.
- Target resequencing is a useful method of resequencing that can be utilized for forensic applications. The genomic regions of interest from a DNA sample are selectively isolated through a method known as enrichment.
\
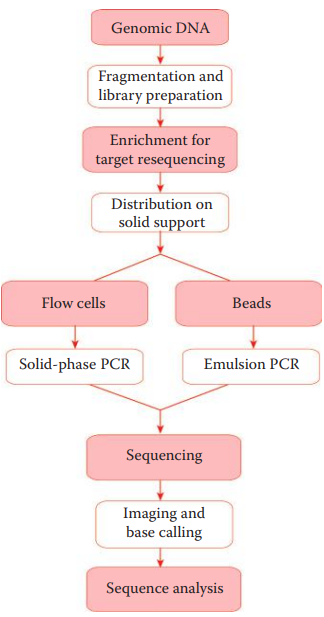
DNA Samples, Sequencing Library, and Template Preparation
- Human autosomal genome sequencing usually requires converting a DNA sample into a sequencing library.
- Two types of sequencing libraries are usually used: * Mate-pair library: It is often used in de novo sequencing applications. * Fragment Library: It is often used for resequencing and forensic applications.
- DNA templates are amplified using either emulsion or solid-phase PCR. * In the emulsion PCR, single-stranded DNA templates are bound to beads. * In solid-phase PCR, both forward and reverse primers are attached to a slide.
\
NGS Chemistry
- Pyrosequencing Technology: During this method, each nucleotide substrate is introduced one at a time. Only the correct nucleotide corresponding to the template is incorporated, and pyrophosphate is released.
- Cyclic reversible termination: In this method, chain terminators are used to extending a primer sequence complementary to the template DNA.
\
NGS Coverage
- In NGS, sequencing reads are usually not distributed evenly over the genomic regions of interest.
- The average number of times that each nucleotide in the genomic regions of interest is sequenced is known as the coverage or the sequencing depth.
\

\