Chapter 9: Detection Methods
9.1: Direct Detection of DNA in Gels
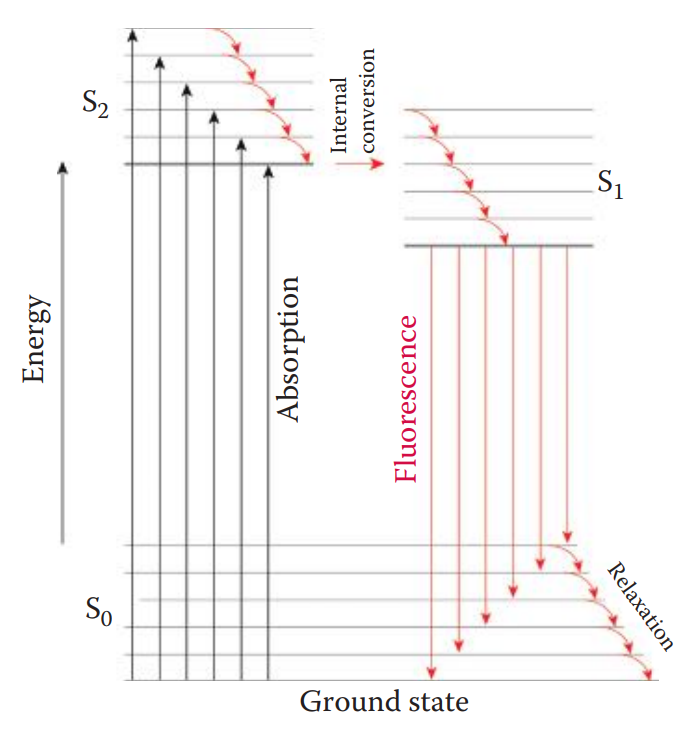
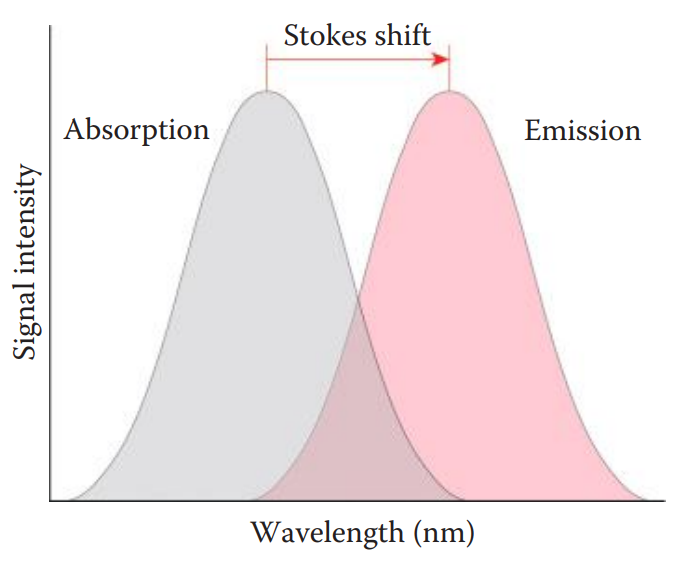
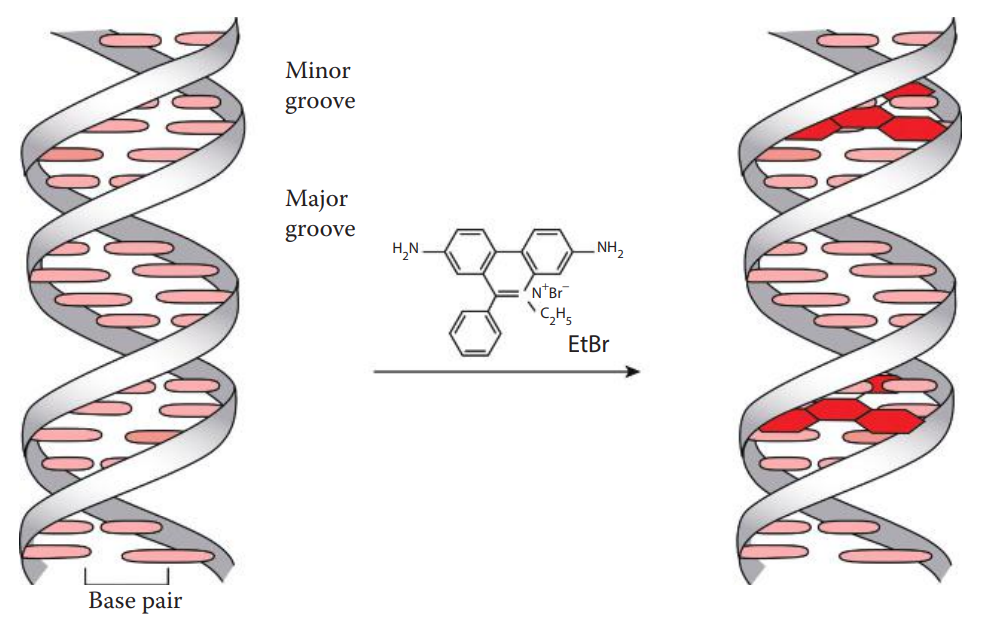
Fluorescent Intercalating Dye Staining
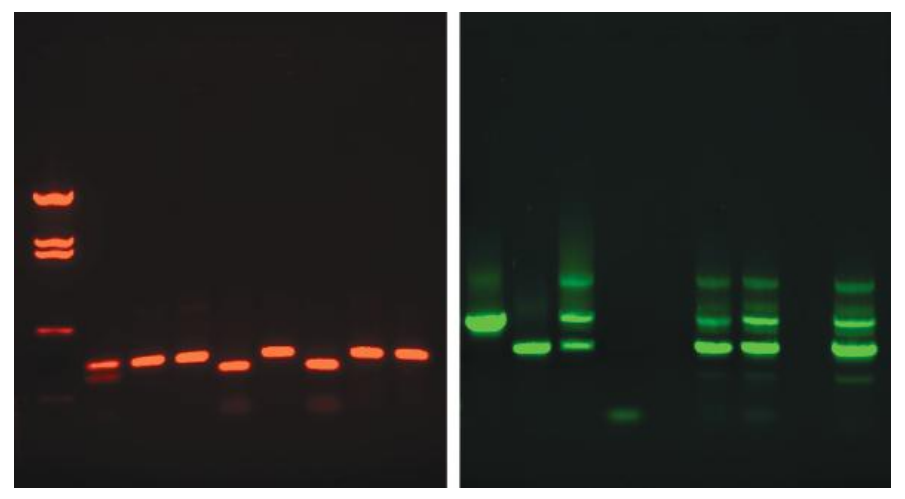
- These techniques allow the detection of DNA bands as small as 10 ng in agarose gels.
- Staining of DNA in agarose gels can be achieved by including an intercalating dye in the gel or staining the gel after electrophoresis in a dye-containing solution, followed by a washing step known as de-staining to reduce nonspecific staining.
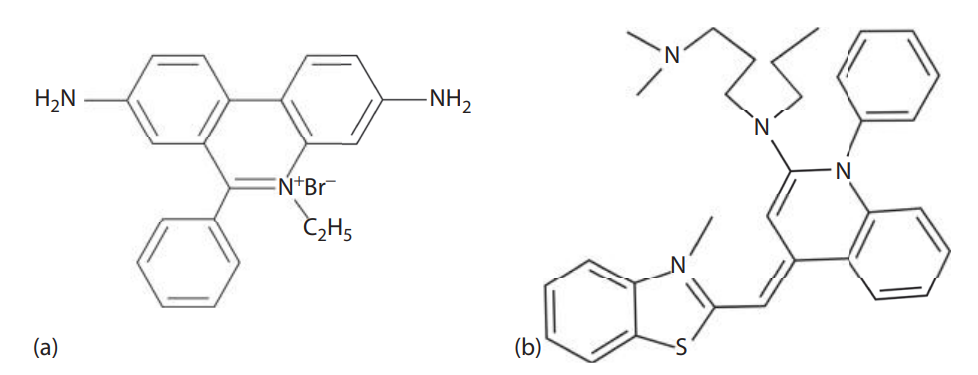
- Historically, ethidium bromide-stained gels were photographed with a standard ultraviolet transilluminator at approximately 300 nm using a camera with an orange filter, whereas gels can currently be documented using digital techniques.
- When examining a DNA-containing gel under a UV lamp, the eyes and the skin should be protected from UV exposure.
- Ethidium bromide is a mutagen and a potential carcinogen. It should be handled according to the Material Safety Data Sheet and safety protection wear should be used while handling the chemical.
- Ethidium bromide waste is usually disposed of as hazardous waste and is handled in accordance with laboratory guidelines.
Silver Staining
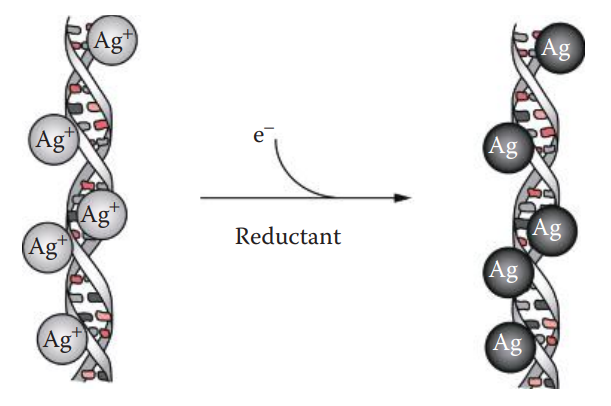
- Silver staining of polyacrylamide gels has been used for the amplified fragment length polymorphism method of variable number tandem repeat profiling.
- The sensitivity of silver staining is approximately 100 times higher than that obtained with ethidium bromide, and silver staining is less hazardous than ethidium bromide detection.
- Silver staining involves processing a gel followed by exposure to a series of chemicals.
9.2: Detection of DNA Probes in Hybridization-Based Assays
Radioisotope Labeled Probes
- Radioisotope probe labeling was used for early versions of VNTR testing and DNA quantitation.
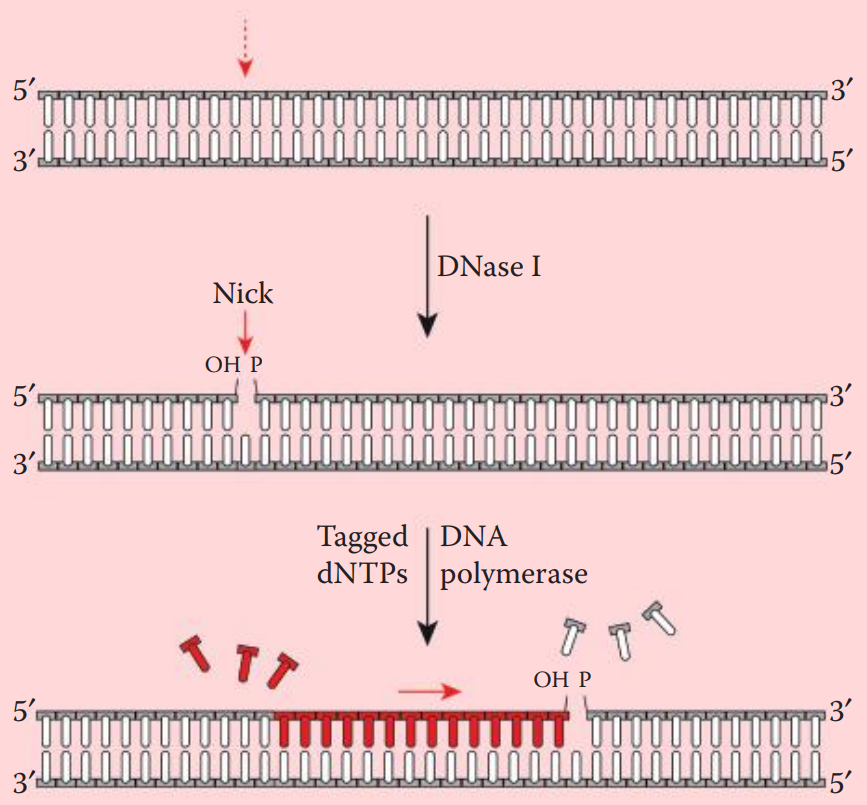
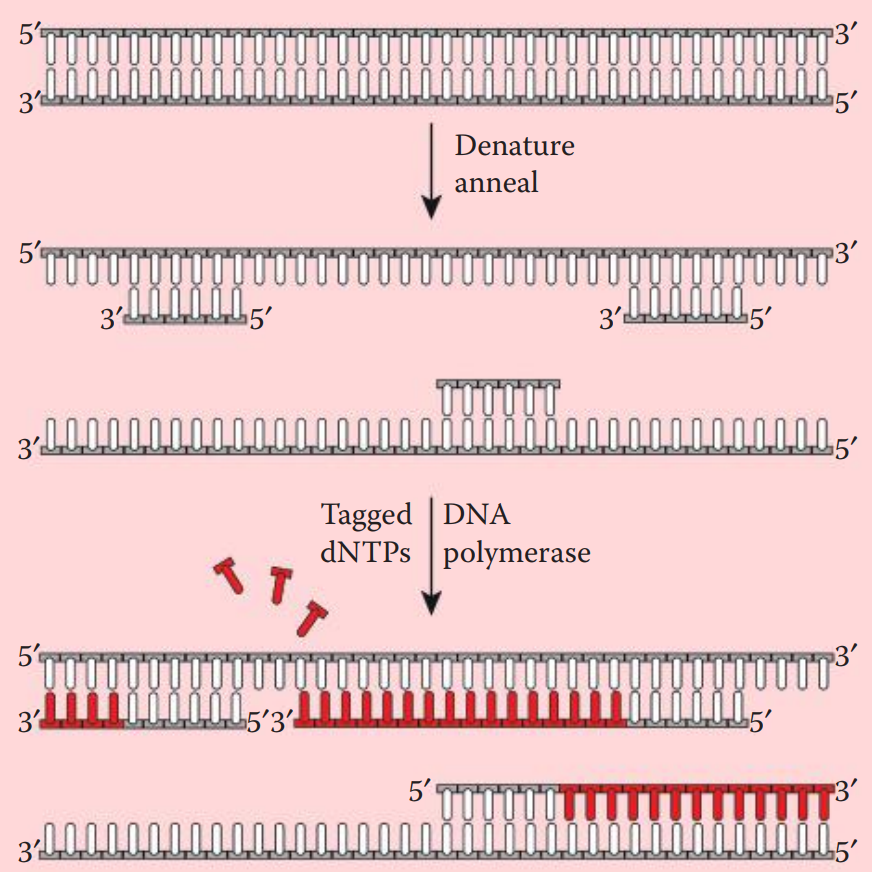
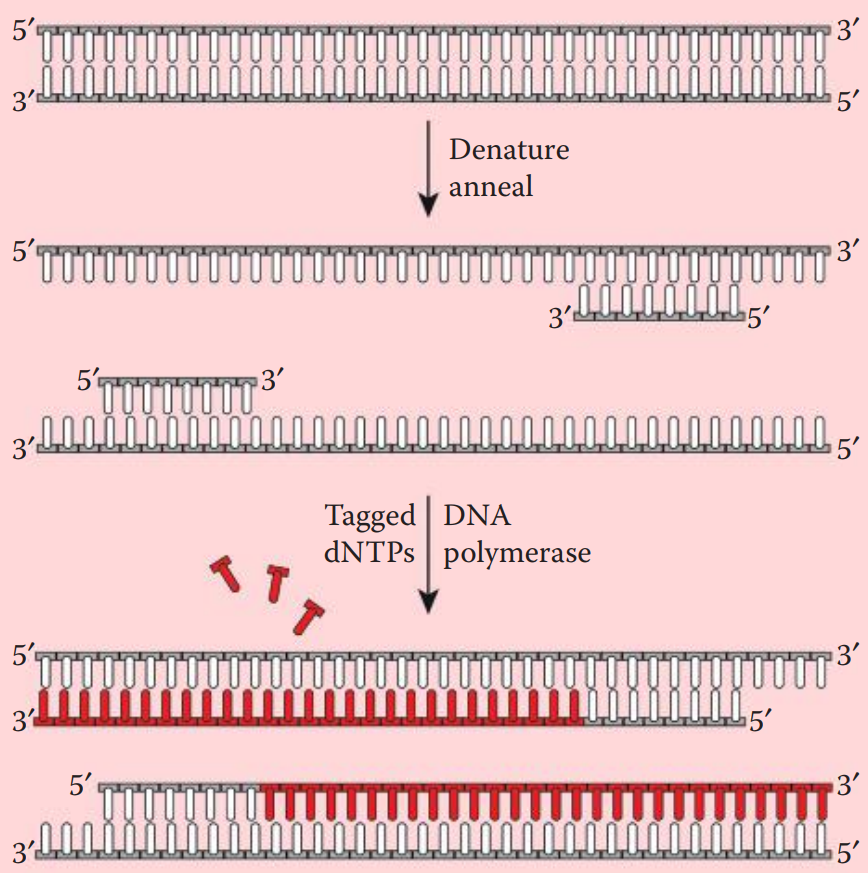
- Nick translation incorporates labeled deoxyribonucleotides (dNTPs) into double-stranded DNA. * A DNase is utilized to create nicks along a DNA strand. Tagged nucleotides (one or more than one of the dNTPs) are then incorporated into the DNA strand using a DNA polymerase. * DNA synthesis extends from the 5′ to 3′ end and the original strand is degraded.
- DNase I is used to introduce single-strand nicks within the DNA fragment to be labeled.
- DNA Polymerase I recognizes the nicks and replaces the preexisting nucleotides with new strands containing labeled dNTPs.
- Prior to hybridization, the probe is denatured into single-stranded fragments by boiling for a few minutes followed by rapid cooling on ice.
- After the hybridization process, these probes can be visualized by exposing the DNA-containing membrane to an x-ray film.
- The energy released from the decay of the radioisotopes is absorbed by the silver halide grains in the film emulsion and forms a latent image.
- A chemical development process amplifies the latent image and renders it visible on film.
\
Enzyme-Conjugated Probe with Chemiluminescence Reporting System
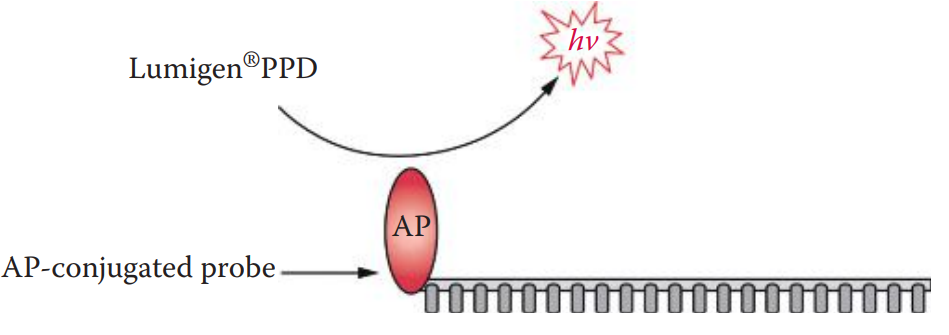
- The use of alkaline phosphatase (AP)-conjugated probes with chemiluminescent substrates comprises a highly sensitive non-radioisotopic detection system.
- Alkaline phosphatase can cleave the phosphate groups from a variety of substrate molecules.
- Its enzymatic activity can be measured using dioxetane-based chemiluminescent substrates such as Lumigen® PPD.
- The Lumi-Phos Plus kit of Lumigen Inc. contains this substrate and can serve as a detection system for slot blot assays for DNA quantitation and RFLP assays for VNTR profiling.
- AP catalyzes the cleavage of the phosphate ester of Lumigen® PPD, resulting in the release of a photon.
\
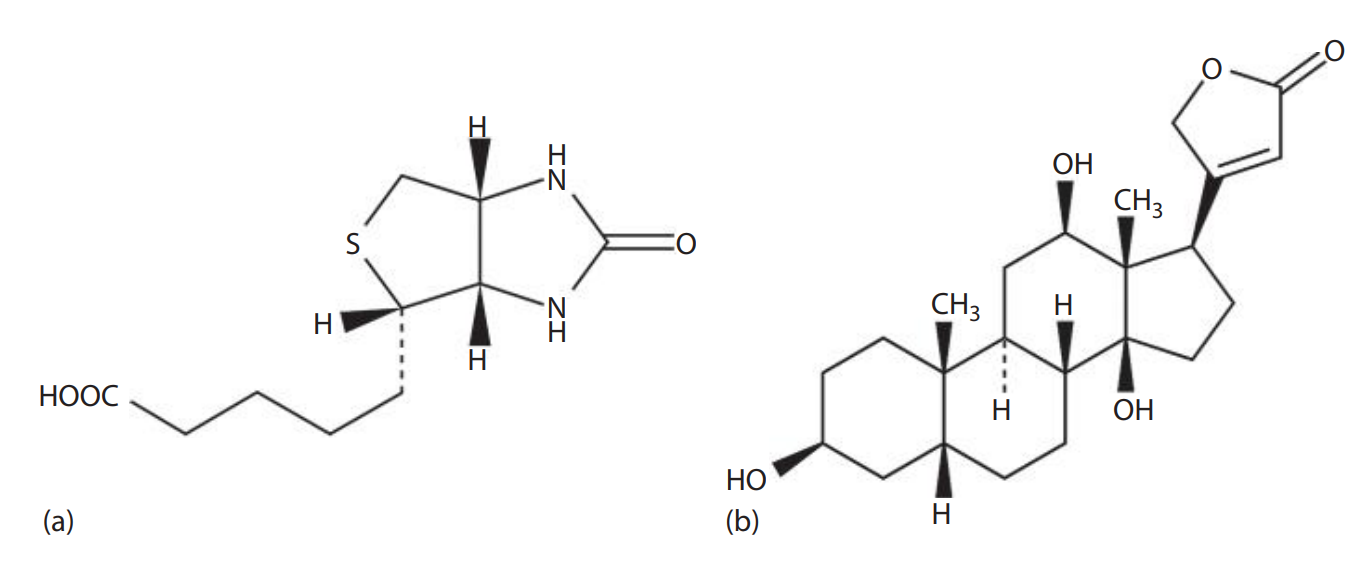
Biotinylation of DNA with Colorimetric Reporting Systems
- Biotin: Also known as Vitamin H, a water-soluble molecule found in egg yolk. * It can be incorporated into oligonucleotide probes without interfering with the ability of probes to hybridize because of its small size. * Signals from a biotinylated probe can be detected with an enzyme-conjugated avidin system. * Two steps are required to detect biotin-labeled probes. * First, an avidin conjugate consisting of a reporter enzyme is added. * Then, the reporter enzyme is assayed with substrates.
\
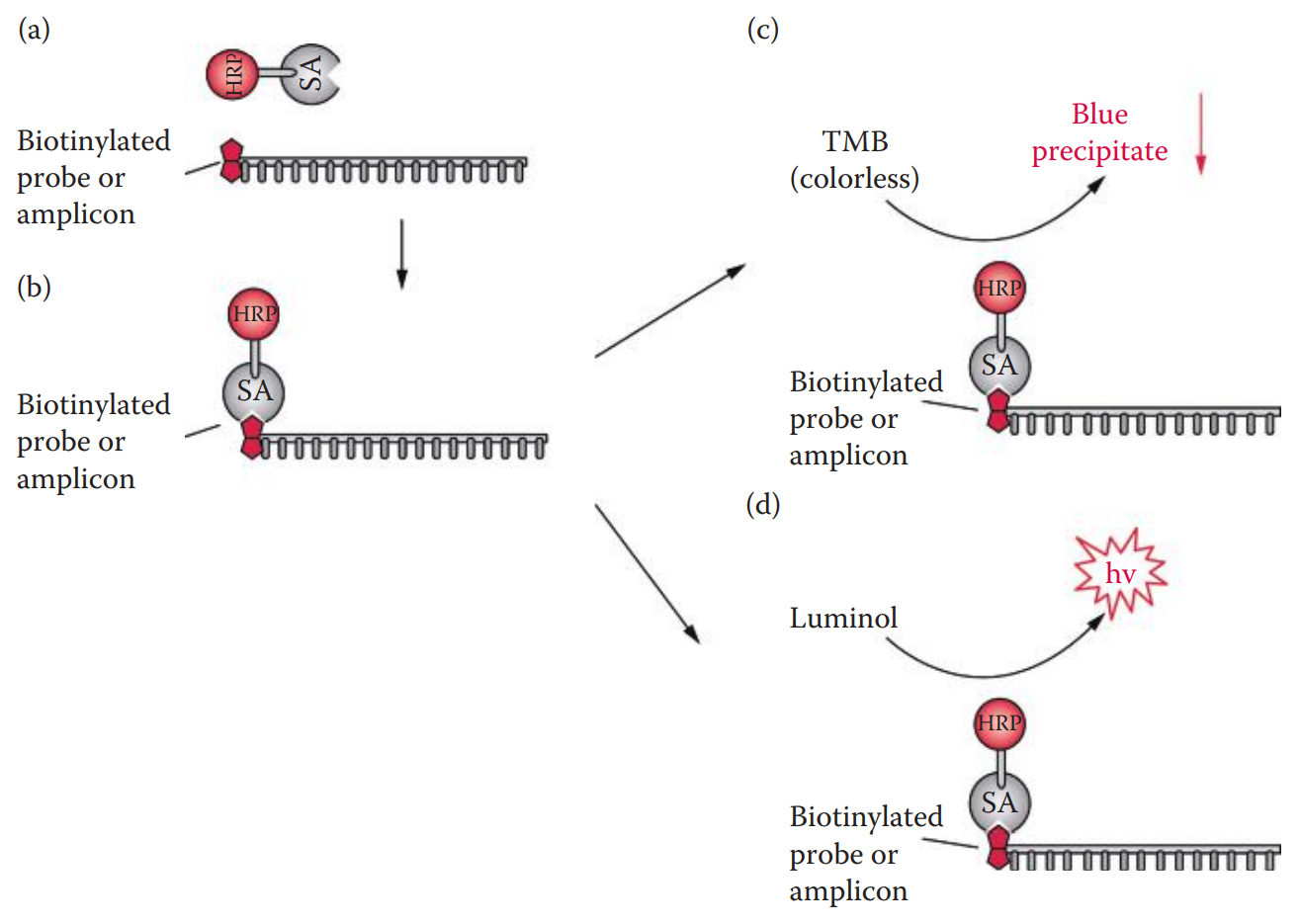
- Enzyme-Conjugated Avidin * Avidin: A glycoprotein found in egg white; it binds to biotin with extremely high affinity. * Avidin detection has a high background due to nonspecific binding. * The nonspecific binding can be reduced by replacing avidin with its streptavidin counterpart from Streptomyces avidin.
- Reporter Enzyme Assay * This technique has been used for forensic DNA testing such as slot blot assays for DNA quantitation. * A chemiluminescent substrate such as a luminol-based reagent can also be utilized with HRP. * The peroxidase catalyzes the oxidation of luminol to form a chemiluminescent product.
9.3: Detection Methods for PCR-Based Assays
Fluorescence Labeling
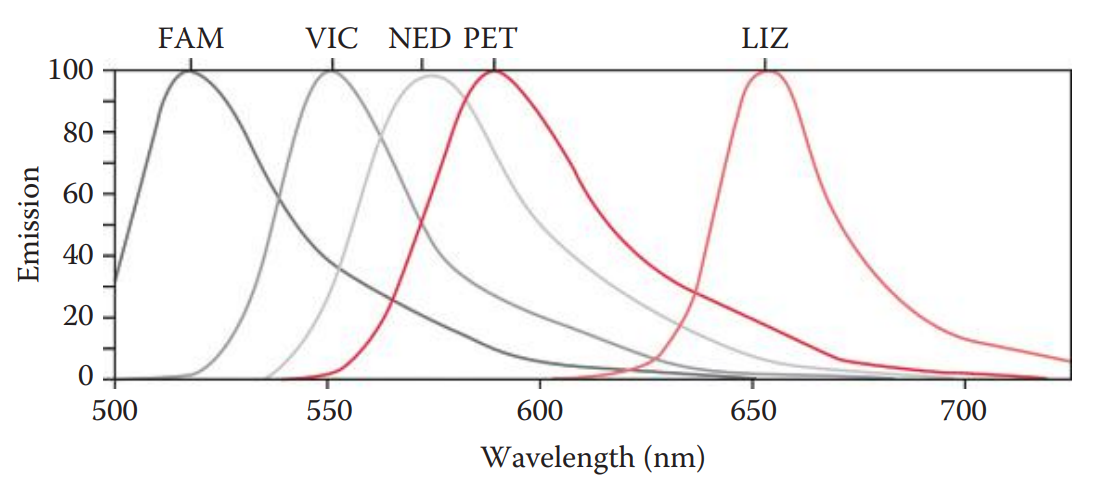
- Fluorescent Dyes * The advantages of fluorescence detection methods include a higher sensitivity and broader dynamic range than comparable colorimetric detection methods. * They have the capacity for simultaneous analysis of complex samples such as multiplex PCR products with different fluorescent labels allowing the distinction of various amplicons.
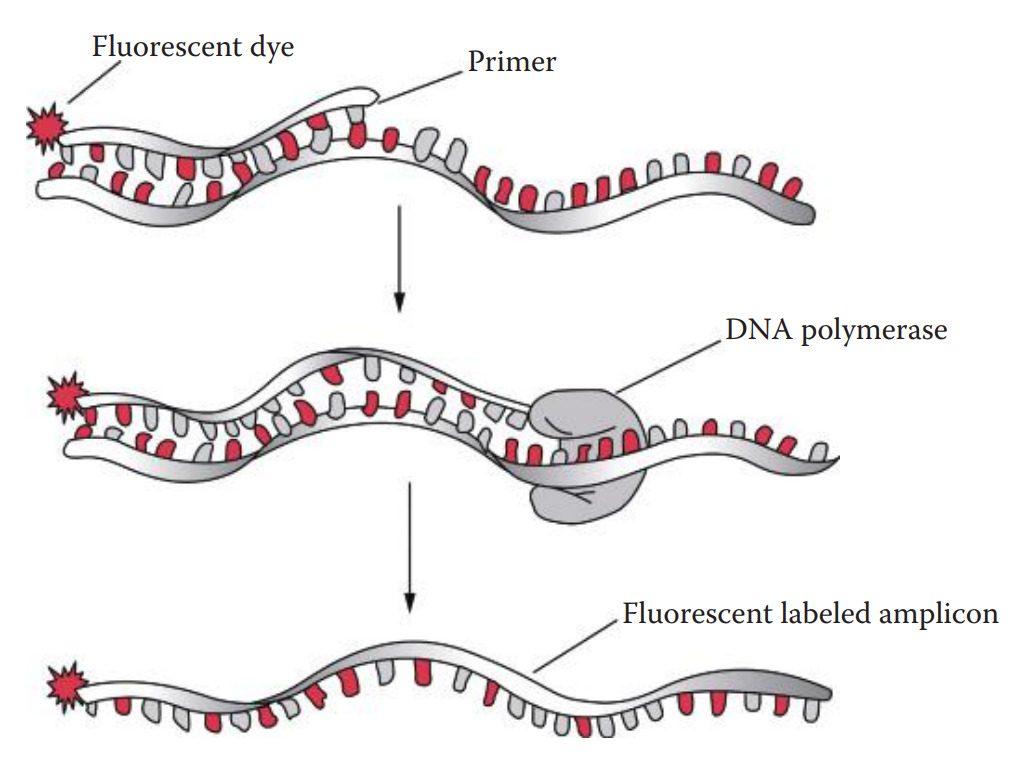
- Labeling Methods * Fluorescent dye labeling can be incorporated into a DNA fragment using a 5′-end fluorescently labeled oligonucleotide primer. * The dye-labeled primer method is usually used for STR profiling in which only one primer from each primer pair is labeled; therefore, only one strand can be detected. * The two-band pattern observed with silver staining does not appear with this method. * Dye-labeled primers allow multiplex PCR amplifications in the same tube. * Fluorescent dye labeling of DNA fragments can be carried out by incorporating fluorescently labeled dideoxynucleotides (ddNTPs) in the PCR product.
- Fluorophore Detection * Fluorophore: A component of a fluorescent dye molecule that causes the molecule to be fluorescent. * Lasers are commonly used as excitation sources because laser light emissions have high intensity and are monochromatic. * The argon ion gas laser is frequently used in applications such as fluorescence-labeled STR and mtDNA sequence analysis because the excitation wavelength of commonly used fluorescent dyes matches the wavelength of the argon laser. * Optical filters are used to filter out undesired light and to allow only one particular wavelength to pass through. * The excitation filter selectively transmits light from an excitation source. * The light is then directed by the dichroic beam splitter to DNA molecules labeled with fluorescent dyes. The light emitted from a fluorophore is also transmitted by the dichroic beam splitter toward the detector. * The emission filter selectively blocks undesired light, thus transmitting a specific wavelength of the emitted fluorescence. * The signal from the fluorophore is collected and converted to an electronic signal expressed in an arbitrary unit such as a relative fluorescence unit (RFU).