Chapter 11: Human Genetic Variation (and Chapter 15.2 Chromosomal Abnormalities)
]]Origins of DNA Sequence Variation]]
@@What types of sequence variation exist (single nucleotide to chromosomal level)?@@
- What is the difference between variant and mutation?
- same thing
- {{SNP Point mutations????????{{
- substitution
- deletion/ addition
- Copy number variation (CNV)
- polyploidy
- Aneuploidy
- Mixoploidy
- Recombination errors
- chromosome cross-over during mitosi
- Chromosomal
- Translocations
- Duplications
- Inversions
- Indels/Delins
@@What are the most common damages that occur to a DNA molecule as the result of endogenous sources, including chromosomal abnormalities (Ch. 15.2)?@@
- DNA replication errors:
- replication slippage → deletion or expansion ( especially during very repetitive strands
- causes Huntington disease ( accumulation of toxic debris) → CAG repeats → Q( glutamine accumulation)
- Base damage
- reactive oxygen species(ROS) proudes during normal metabolic processes can damage DNA base
- Strand breaks
- single strand breaks
- double-strand breaks
- Chromosomal abnormalities
- Chemical damage
- hydrolytic most common damage to DNA → depurination the most common hydrolic damage of DNA
- reactive oxygen species(ROS)
- Methylation
@@What is aneuploidy and polyploidy (15.2)?@@
- → an extra set of chromosomes ( lethal/ not viable)
- → abnormal number of chromosomes ( a single chromosome is multiple or missing)
- example of aneuploidy where the people survive
- trisomy 21
- super male (XYY)
- chromosome 18
- → some cell have mutations that result in different number of chromosome sets → cause mosaicism (lethal)
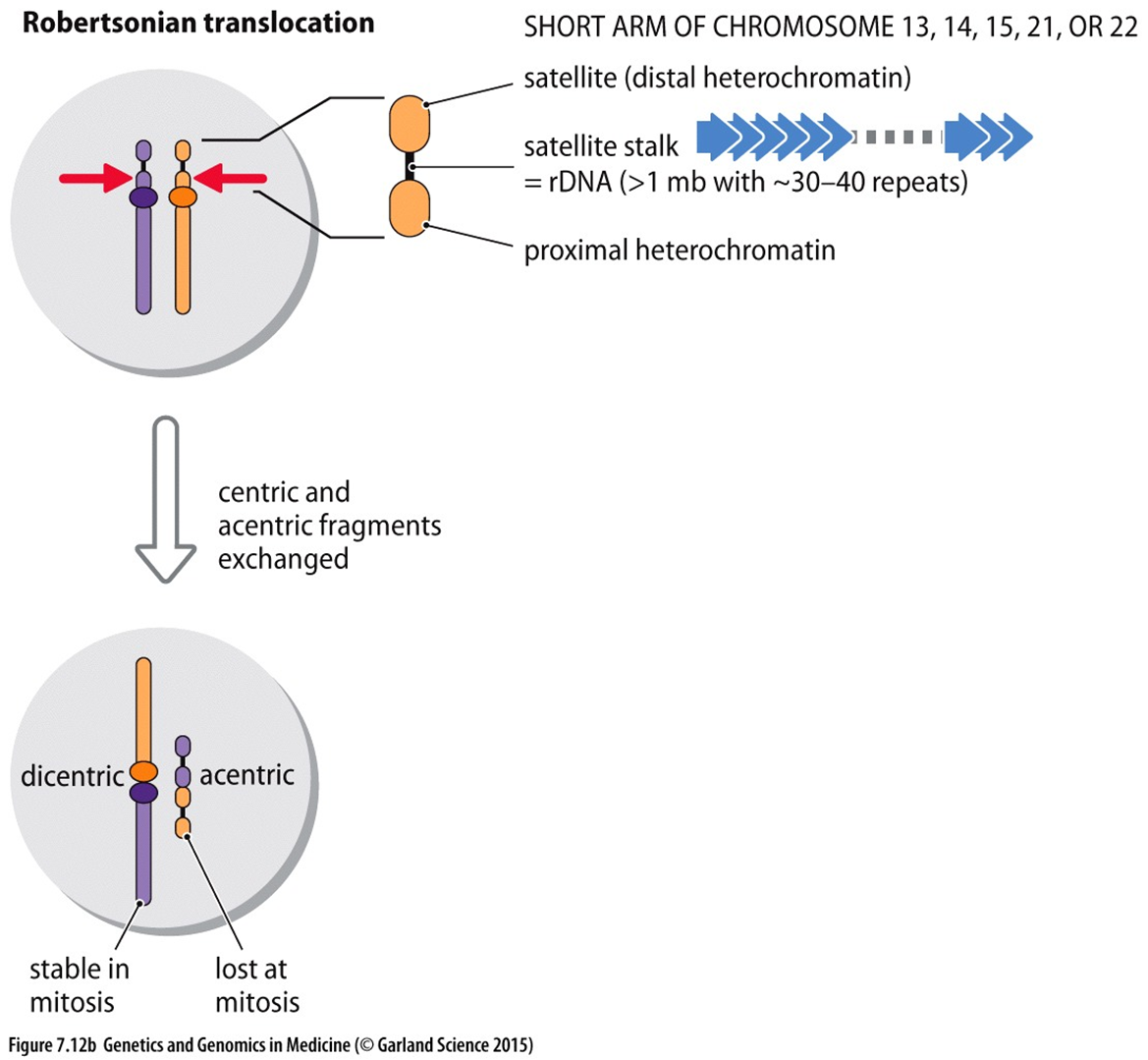
@@What is a Robertsonian Translocation and how does it occur?@@
- Robertsonion Translocation (centric fusion)
- translocation in chromosomes but two parts of the chromosome that attach have two nearby centromeres. Both centromeres fuse together and don’t cause a problem during further mitosis → genetic information is not lost
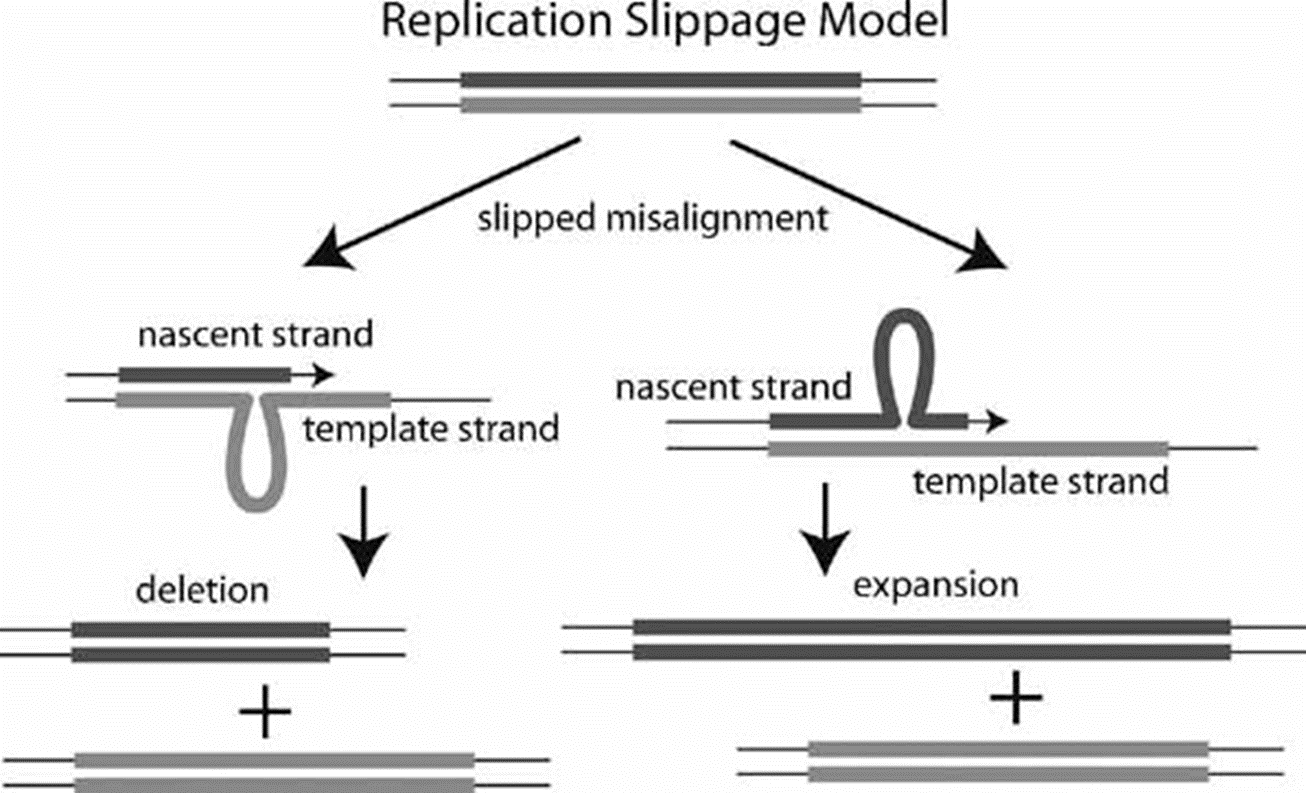
@@What is replication slippage and why are short repetitive sequences more vulnerable to this?@@
- occurs during DNA synthesis when the leading strand is misaligned with the leading strand template
- Polymerase can slip back wars or forward, causing an insertion or deletion
- all of this can cause egntic disorders and diseases
@@What genetic variations may result in Huntington Disease or Fragile X syndrome? Why?@@
- Fragile X syndrome is a genetic disorder caused by an expansion of the CGG repeat in the FMR1 gene on the X chromosome. This results in reduced or absent production of the FMR1 protein, leading to a range of cognitive and behavioral problems. It is an X-linked dominant disorder, affecting males more severely than females because female shave another X and, therefore FMRI1 gene.
@@What are the most common exogenous sources of damage? What types of damage are created?@@
- Radiation
- UV
- Xrays
- chemicals
- cigarette smoke
- nitrate and nitrate preservatives
- benzoyl peroxide
- Infectious agents
- HPV
- Heliobacter pylor
@@What are the significance of CpG islands and why are they?@@
- Cp(phosphate)G island
- CGCGCGC
- found in promoters but not actively transcribed
- when the C get methylated( which it easily done C→T) then there is no transcription
[[Repair[[
==What are the primary mechanisms that cells use to repair DNA damages? How does DNA polymerase “proofread”?==
- if bases are mismatched, then they don’t fit together and the following bases are also wonky
- The polymerase also notices and pauses replication and uses exonuclease activity to remove the incorrect nucleotide.
==What are the general steps in Base Excision Repair (BER), Nucleotide Excision Repair (NER), Mismatch Repair (MMR), Homologous Recombination (HR) and Non-Homologous End Joining (NHEJ)?==
==What is translesion synthesis? When and why is it initiated?==
- only used in emergencies because there is no proof reading
==What are the primary diseases/outcomes seen in individuals with defective DNA repair mechanisms?==
- cancer
- apoptosis
%%The Scale of Human Genetic Variation%%
}}What are the varying scales of genetic variation? What is structural variation? How does balanced structural variation differ from unbalanced structural variation? What is aneuploidy?}}
balanced variation → all of the gentic martial is still there
- NO net loss or gain
- Translocations
- Inversions
- Point mutations
unbalanced → not all genetic material
- Change in copy number
- Abnormal chromosome segregation
- Point mutations
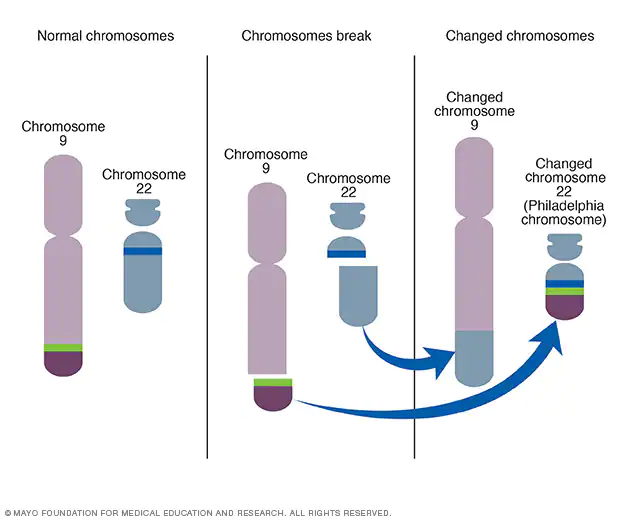
Example: Philadephia chromosome
- when chromosome 9 and chromosome 22 break and exchange portions
- causes Leukemia
}}What is satellite DNA? Why is replication slippage more common in arrays of tandem repeats?}}
- Satellite DNA: highly repetitive noncoding region if the genome, located near the centromeres and telomeres region of chromosome
- why is replication slippage more common?
- when the DNA polymerase enzymes slips from the DNA strand and adds or deletes on e or more repeats
- if slippage occurs pieces still fit together and polymerase is unable to detect if they made a mistake
}}Know the similarities and differences between Single Nucleotide Variation, Single Nucleotide Polymorphisms (SNP), “indels”, Copy Number Variants (CNV), Variable Number Tandem Repeats (VNTR).}}
| }}Type of Variation}} | }}%%Definition%%}} | }}%%Similarities%%}} | }}Differences}} |
|---|---|---|---|
| SNV | A single nucleotide change in the DNA sequence | Can occur anywhere in the genome; can be a germline or somatic mutation | Only involves a change in a single nucleotide base |
| SNP | A type of SNV that is present in at least 1% of the population | Can occur anywhere in the genome; can be a germline or somatic mutation | Must have a minor allele frequency of at least 1%; used as genetic markers in association studies |
| Indel | A small insertion or deletion of DNA sequence | Can occur anywhere in the genome; can be a germline or somatic mutation | Involves the addition or removal of one or more nucleotides |
| CNV | A large segment of DNA that is duplicated or deleted in the genome | Can occur anywhere in the genome; can be a germline or somatic mutation | Involves the gain or loss of a large segment of DNA; can be variable in size |
| VNTR | A region of DNA where a short sequence of nucleotides is repeated in tandem | Can occur anywhere in the genome; can be a germline or somatic mutation | Involves the gain or loss of a variable number of tandem repeats; can be used as genetic markers in forensics and population genetics |
Note: Germline mutations occur in the DNA of sperm or egg cells and are passed on to offspring, while somatic mutations occur in non-reproductive cells and are not passed on to offspring.
}}What is population based genome sequencing? What is the 1000Genomes project. What is the T2T Consortium? What did they find?}}
- 1000Genome project: indicated in 2008 in 26 different population
- goal: search for de novo mutations
- found that African populations had the most variations in genome and Europe, east Asia and South Asia least variation
- → founders effect
- Telomere to telomere consortium: focused on identifying the first complete assembly of human genome with euchromatic and heterochromatic region8 coding and non-coding)
}}What percent of our genome is thought to be under functional constraint? Explain why this percentage is greater than the 1.2% (the genome that is encoding for proteins). Be able to explain multiple reasons why. Compare this idea of functional constraint to findings by the ENCODE Consortium.}}
- Functional constraint: parts of DNA genome is silenced through methylation and phosphorylation
- Most protein variation have a neutral effect and not necceraly cause a a dise( example blood groups)
- functional constrains 1.2% of DNA can´t change the protein because they are so important
- but more because of regulation
- promoter regions and enhancers
}}Be able to explain how post-zygotic mutations create individuals that are genetic mosaics. What is a chimera?}}
{{{{
- %%blood group%%
- Cytogenetics: branch of genetics and biology concerned with chromosomal behavior
- FISH: a molecular cytogenetic technique that allows the localization of a specific DNA sequence or an entire chromosome in a cell
- for Philadelphia Chromosome: tag both chromosome with different fluorescence molecules and take multiple pictures of translocation
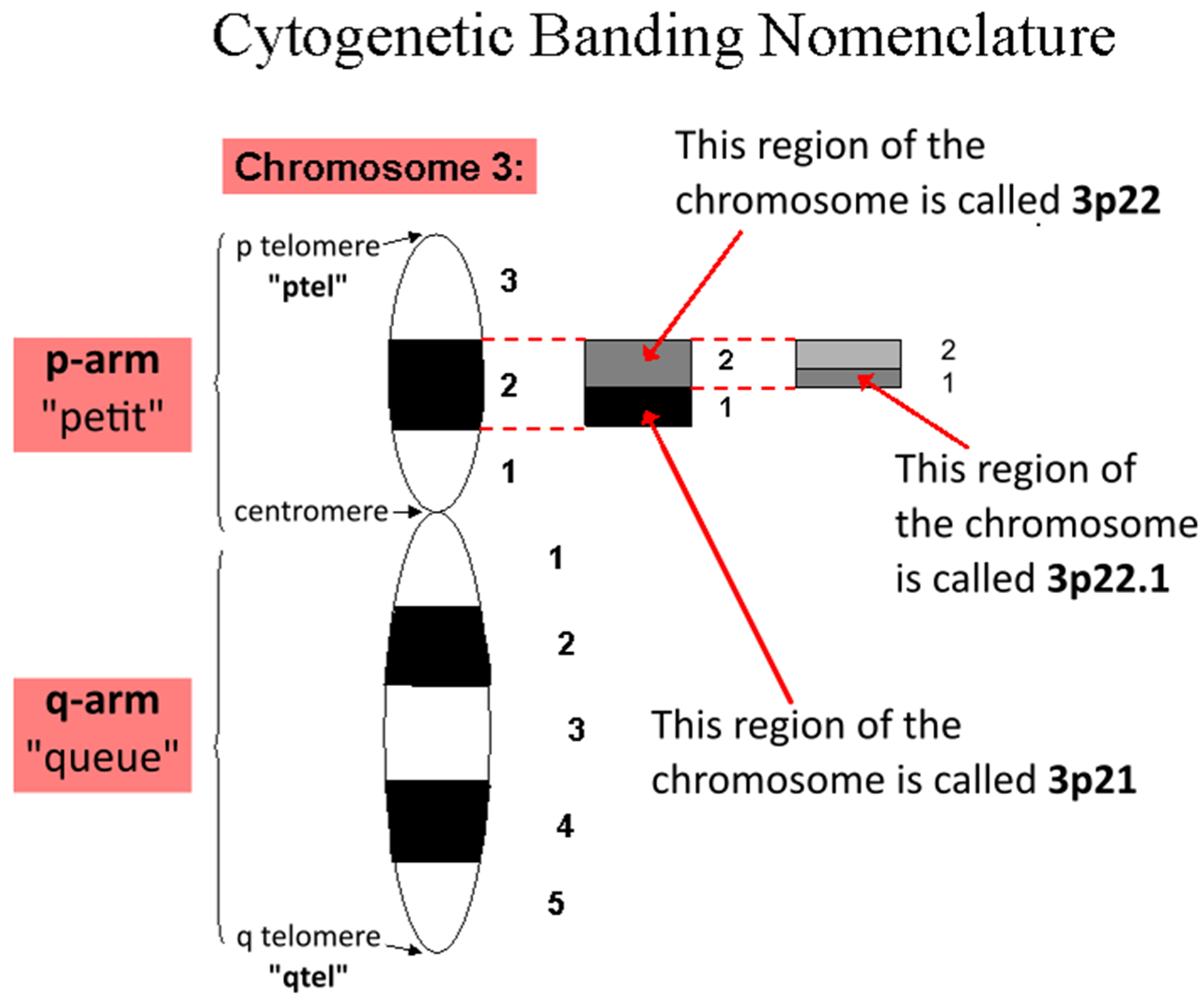
^^BOX 15.1 NOMENCLATURE OF CHROMOSOME BANDS^^
<<What is “G-banding”? In which stage of the cell cycle are chromosomes extracted and stained (mitotic or interphase)? Which parts of the chromosome stain darker and lighter?<<
- G-banding: A type of standing with Giemsa stain
- DNA from white blood cells during prometaphase ( as condenst)
- dark color= AT-rich → gene poor→ hetero chromatin
- light color= GC rich → gene-rich
<<What are the names of the two arms of a chromosome?<<
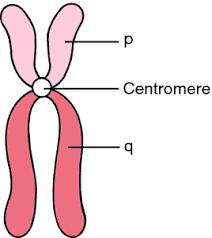
<<What does it mean to be Metacentric, Submetacentric, Acrocentric or for a chromosome to be acentric?<<
position of centromere
- Metacentric (center)
- Submetacentric (between middle and telomere)
- Acrocentric (near telomere)
- penetrance
- anticpation
- repication slippage
- dynamic
- 18
- 5 open
- 1 hardy weinberg
- bonus