Polymerase Chain Reaction
Keywords:
Primers (forward and Reverse)
Antiparallel DNA
Temperature (heating and cooling)
3 stages (Denaturation, Annealing and Elongation)
Taq Polymerase (DNA polymerase) - Phosphide bonds
Amplify - lots of copies (quickly)
Thermal Cycler
Free DNA nucleotides
Buffer
DNA of interest / target DNA
talking about:
- amplification of DNA using polymerase chain reaction
PCR Purpose and Product
like a molecular photocopier
Used to rapidly amplify (make millions of copies) a small length of DNA )between 100bp<x<40,000bp)
Can be applied in
Forensics
Parentage testing & Diagnosis of hereditary diseases
Gene Cloning and Genotyping
DNA sequencing (genome sequence)
Construction of DNA-based Phylogenesis (how closely related some species are to each other U4)
How PCR works
PCR amplifies a specific region of DNA (also known as the region of interest)
Step 1) Identify a 25-base sequence at the 3’ end of each strand bracketing the region of interest
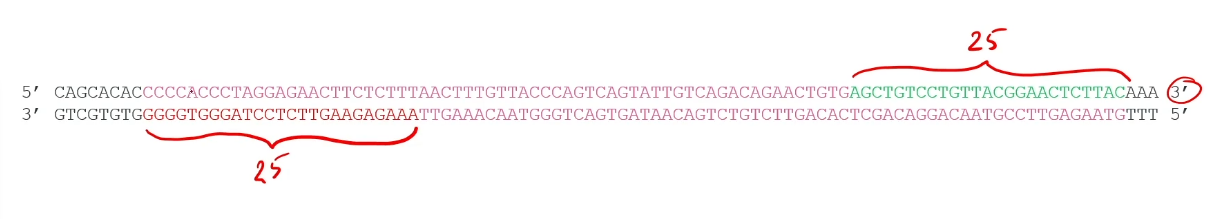
Step 2) Make some DNA primers that are complementary to that 25-base sequence. (Made using a DNA synthesizer machine)
on the lagging side of DNA, the first thing that has to happen is an enzyme called Primase builds a little RNA primer as a starting point for DNA polymerase then extends that primer, and then another DNA polymerase comes and replaces that RNA primer.
Point: DNA POLYMERASE CAN'T JUST START ADDING NUCLEOTIDES AGAINST THE TEMPLATE DNA OUT OF THIN AIR. There has to be something there to add to the three prime ends. That is what the primer does. They’re the starting point.
Denaturing:
Temperature is heated up to 95 degrees
The strands of DNA seperate because they denatured
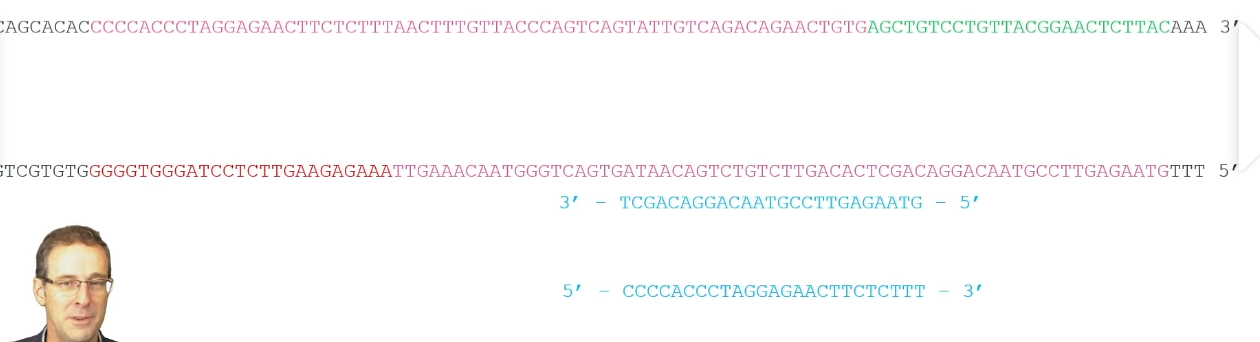
Annealing:
Temperature is lowered to 55 degrees
The primers anneal to the complementary region of interest on the DNA
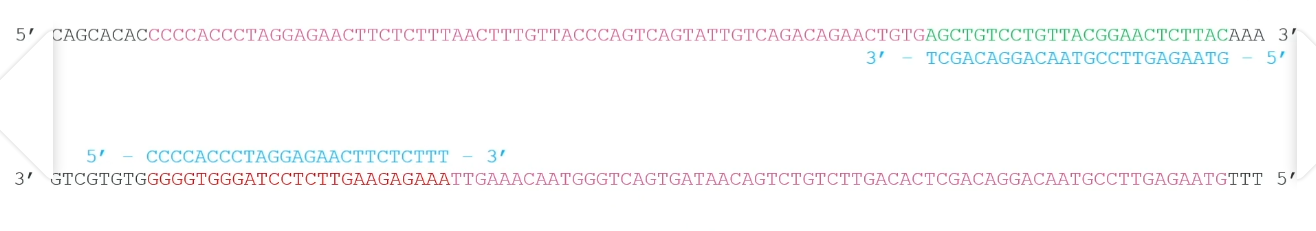
TAQ polymerase
Temperature risen to 72 degrees
Extend the primers by adding nucleotides to the 3’ end in both directions
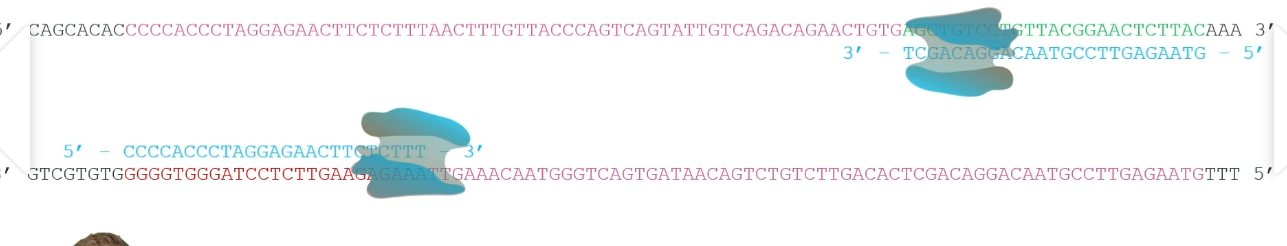
PCR Machine
A Machine capable of ‘ramping’ between temperatures very quickly as it is programmed to heat and cool DNA in a cycle between three temperatures.
Tubes placed in the thermocycler contain
Buffer with DNA (prevents pH from changing DNA is an acid and if you’re making lots of acids it’s going to lower the pH which will then affect the way that enzymes work)
The Primers
TA polymerases
MgCl2 (co-factor of DNA polymerase)
DNA nucleotides
Temperature | Effect |
|---|---|
95 degrees | DENATURE: DNA is denatured as the hydrogen bonds between complementary nucleobases breaks |
55 degrees | ANNEALING: The primers anneal to the DNA ( more) |
72 degrees | TAQ polymerase extends the primers because it’s at optimal temperature |
PCR from Edrolo Workbook
Prerequisite Knowledge & Future applications
Prerequisite Knowledge:
Lesson 2C: Taq polymerase acts similarly to RNA polymerase, except that it synthesizes a DNA strand rather than a strand of RNA.
Lesson 3A: Enzymes are organic catalysts that speed up chemical reactions. Taq polymerase is an enzyme that functions optimally at 72 °C.
Lesson 4A PCR amplifies small samples of DNA using a polymerase enzyme.
Future Applications:
Lesson 4D: Often PCR is performed before gel electrophoresis, to increase the amount of
DNA available.
Lesson 10A: PCR can be used to amplify a DNA sample from some fossils, such as from teeth or bones, to provide an insight into the relatedness between organisms.
Purpose of Polymerase chain reaction
the polymerase chain reaction amplifies a sample of DNA by creating additional copies. It is a multistep process that involves thermal cycling
Theory Details:
PCR is a DNA manipulation technique that amplifies DNA by making multiple identical copies used by scientists when there is an insufficient amount of a DNA sample for testing and is often used in paternity and forensic testing and gene fragments for genetic diseases.
When undertaking a polymerase chain reaction cycle, scientists do not usually copy the entire genome. Instead, they focus on certain genes through the use of primers or restriction endonucleases to make the process more efficient. After each cycle of the polymerase chain reaction, the amount of DNA present is doubled (Table 1).
Process of the polymerase chain reaction
Overview:
The process of the polymerase chain reaction involves manipulating temperatures to cause denaturation and annealing in a four-step process.
Theory details
The polymerase chain reaction requires the following materials to take place:
• a DNA sample that subsequently gets denatured and amplified through the polymerase chain reaction
• Taq polymerase is required in the elongation stage to bind complementary nucleotides to the single-stranded DNA
• nucleotide bases must be constantly available for Taq polymerase to create a new strand that is complementary to the single-stranded DNA
• sequence-specific DNA primers join to the 3’ end of single-stranded DNA by complementary base pairing to form the first segment of double-stranded DNA, allowing Taq polymerase to attach and begin extending the DNA strand.
To begin the polymerase chain reaction, a mixture of the above materials is placed into a thermal cycler (Figure 1), where it undergoes the following processes:
1 Denaturation – DNA is heated to approximately 90–95 °C to break the hydrogen bonds between the bases and separate the strands, forming single-stranded DNA.
2 Annealing – the single-stranded DNA is cooled to approximately 50–55 °C to allow the primers to bind to complementary sequences on the single-stranded DNA.
3 Elongation – the DNA is heated again to 72 °C, which allows Taq polymerase to work optimally. Taq polymerase binds to the primer, which acts as a starting point, and begins synthesising a new complementary strand of DNA.
4 Repeat – the cycle (steps 1–3) is repeated multiple times to create more copies of DNA
Forwards and reverse primers
In the polymerase chain reaction, there are two different DNA primers needed. This is because, during denaturation, the double-stranded DNA molecule has been separated into two single-strands – the template strand and the coding strand. The two primers are:
the forward primer, which will bind to the start codon at the 3’ end of the template strand. This causes Taq polymerase to synthesise a new DNA strand in the same direction that RNA polymerase would function.
the reverse primer, which will bind to the stop codon at the 3’ end of the coding strand. This causes Taq polymerase to synthesise a new DNA strand in the reverse direction that RNA polymerase would function.