6.4 DNA Replication
<<Semi Conservative Replication<<
- ^^Matthew Meselon and Franklin Stahl^^ in 1958
- DNA is double stranded
- 2 parent strands are seperated
- complementary strand is made from each parent strand
- results in 2 double stranded molecules, each containing one old strand and one new strand
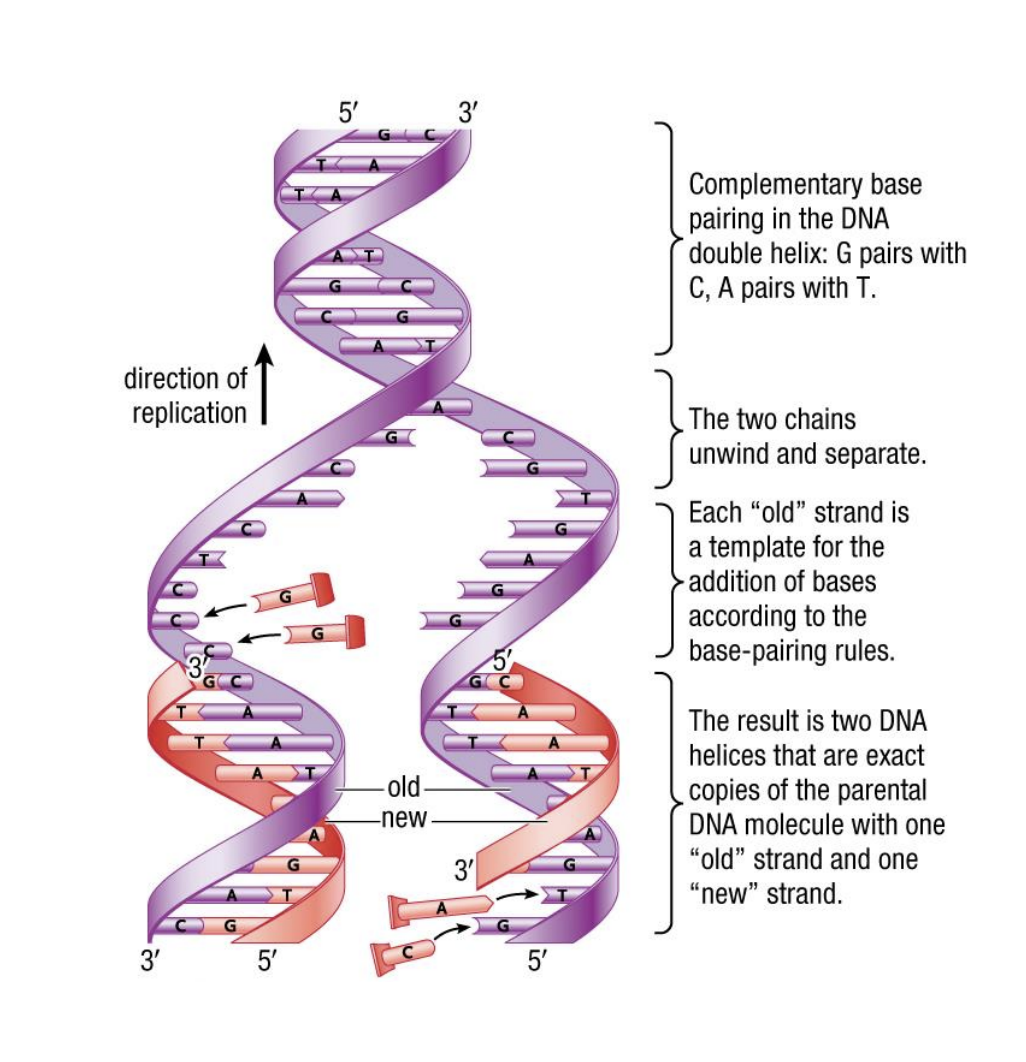
<<Step 1: Separation<<
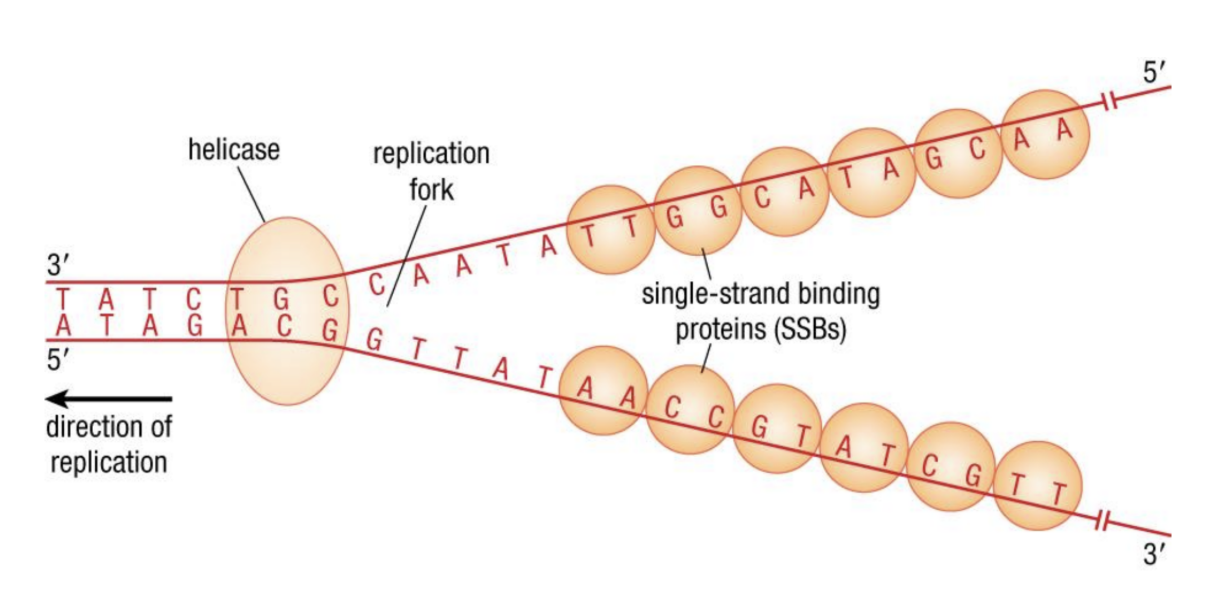
- Strands are unwound from ^^replication origins^^ by %%DNA helicase%%==(1)==.
* Forms a ^^replication fork^^ - %%Topoisomerases%%==(2)== cut and rejoin strands to keep them from tangling (supercoils)
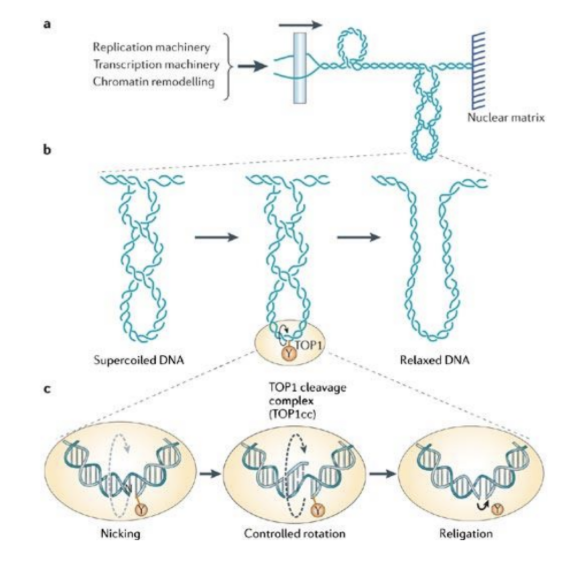
- %%Single-strand binding proteins%%==(3)== attach to unwound strands to prevent them from rejoining (annealing)
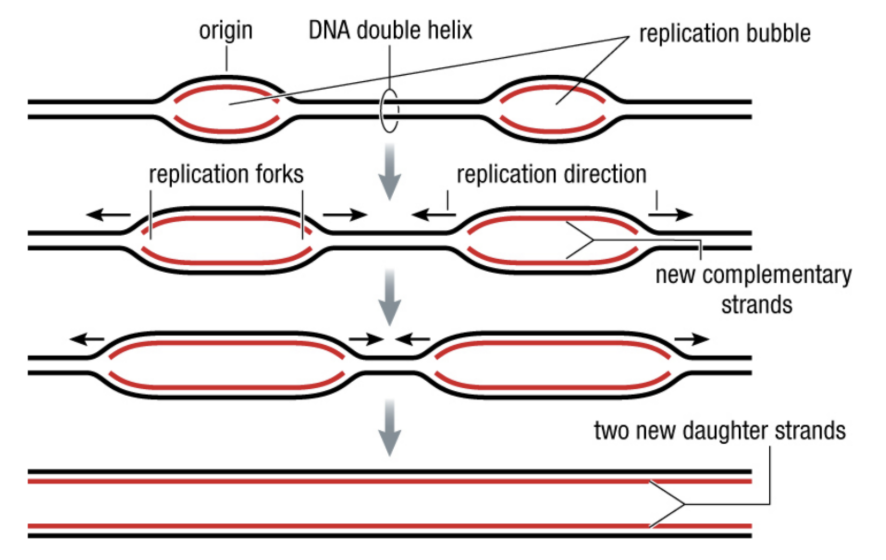
<<Replication Bubbles<<
- because there are many replications going on, especially in Eukaryotic DNA
- Helicase unwinds DNA in both directions from each origin, producing replication bubbles
- Complementary strands are synthesized as the forks continue (step 2)
- Bubbles meet and merge, producing 2 separate daughter strands
<<Step 2: Synthesis / Replication<<
- %%Polymerases%% build complementary strands in 5’ → 3’ direction
* can only add to 3’ carbon
* which means its reading 5’
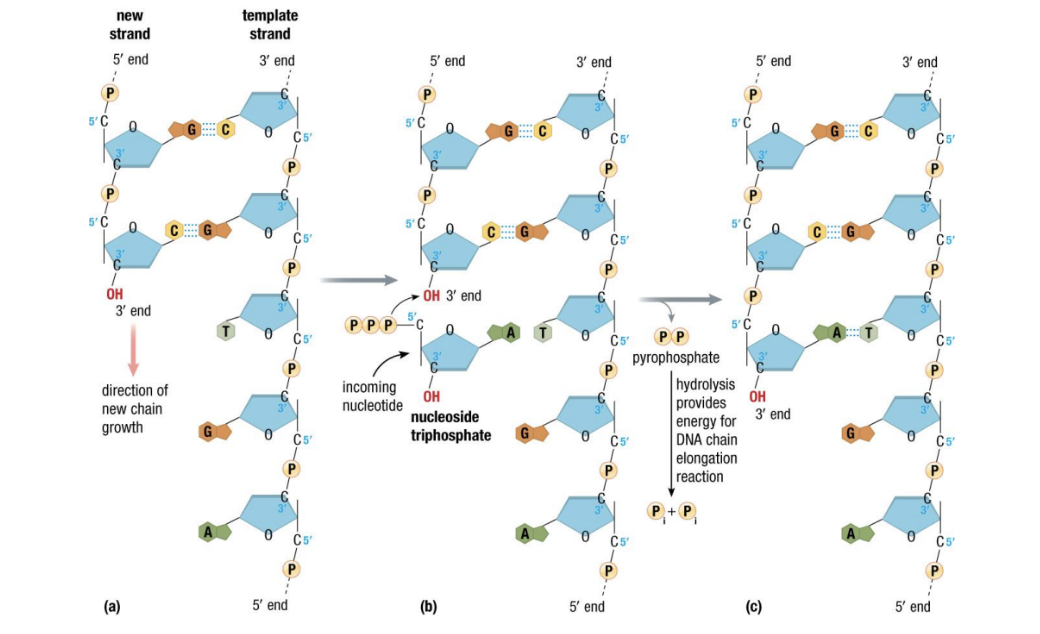
* new strands are always 5’ → 3’ - Nucleotides are added as ^^nucleoside triphosphates^^
* a building block and energy source for replicating DNA
* very similar to nucleotides in the finished DNA
* DNA phosphate needs energy - Hydrolysis of extra phosphates provides energy for synthesis (from ATP)
- makes the pair grow on the 3 prime side w ATP
- the extra %%phosphates%% make up the phosphodiesther bonds on the sides
- the released energy drives DNA synthesis
RNA Primase
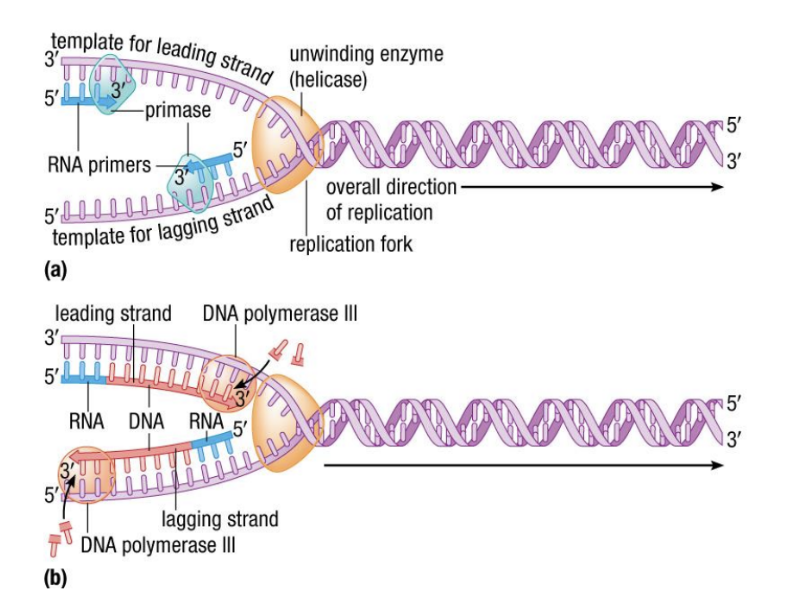
- %%RNA primase%% ==(4)==builds small complementary RNA primers at the replication fork on its own, these short pieces are called ^^RNA primers^^
- %%DNA polymerase III%% ==(5)== ( first one, last one discovered) synthesizes complementary strands in opposite directions (5’ → 3’), can only add to previously existing nucleotides
Leading & Lagging Strands
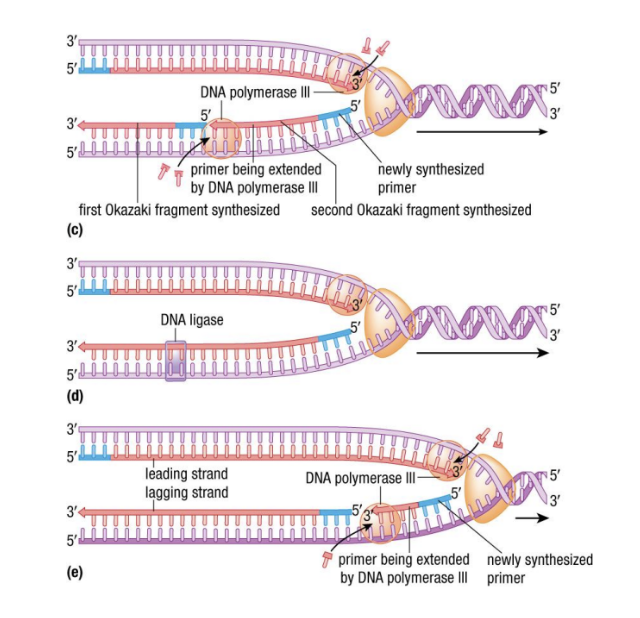
- Leading strand is elongated toward the fork; efficient (5’)
- ^^Lagging strand^^ is elongated away from the fork, requires multiple RNA primers: constantly being synthesized backwards (3’)
* Results in segments of RNA primers & DNA called ^^Okazaki fragments^^
* %%DNA polymerase I%% ==(6)== replaces RNA primers with DNA - purifies
* %%DNA ligase%% ==(7)== joins Okazaki fragments into one continuous strand
<<Step 3: Repair<<
- Mismatched base pairs don’t bond properly and distort shape of the DNA molecule
- %%DNA polymerase II%% ==(8)== and other enzymes search for distortions, removes portion of the stand with mismatch and fills in the gap.