Chapter 8: Analyzing Cells, Molecules, and Systems
Isolating cells and growing them in culture:
Tissue Dissociation:
- Tissues can be dissociated to obtain a single-cell suspension for cell culture.
- Mechanical methods such as mincing, grinding, or shearing can be used to disrupt tissues.
- Enzymatic digestion with proteolytic enzymes like trypsin or collagenase can help release cells from the extracellular matrix.
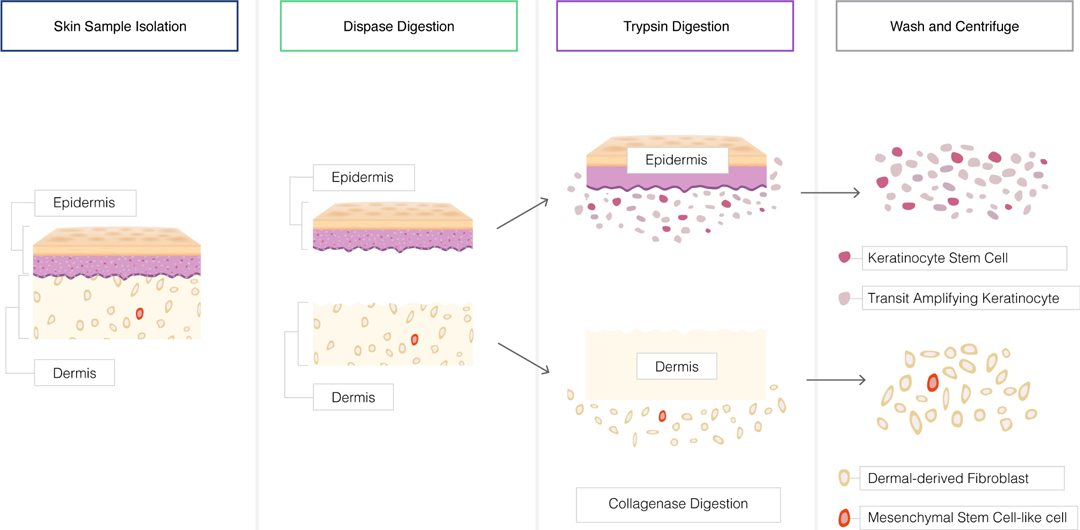
Cell Separation:
- If a specific cell population is desired, cells can be separated based on their physical or molecular properties.
- Techniques such as fluorescence-activated cell sorting (FACS), magnetic-activated cell sorting (MACS), or density gradient centrifugation can be employed for cell separation.
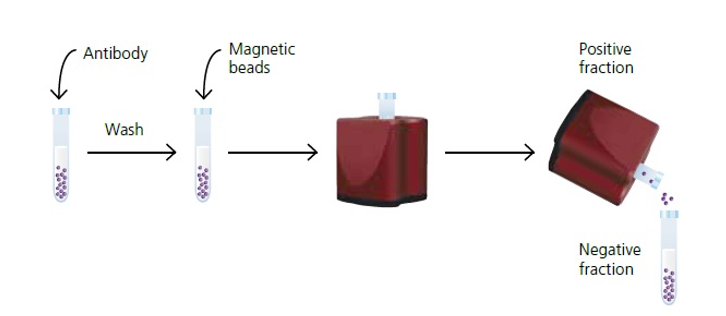
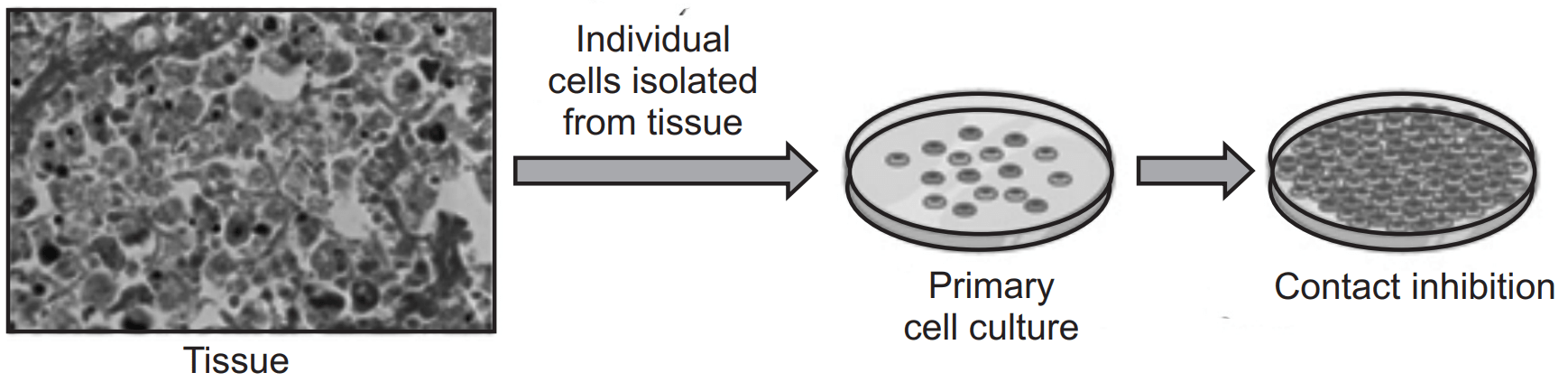
Primary Cell Culture:
- Primary cells are isolated directly from tissues and have a limited lifespan in culture.
- Cells are seeded onto culture dishes or plates coated with extracellular matrix proteins to promote cell attachment.
- Culture medium containing essential nutrients, growth factors, and supplements is provided to support cell growth and proliferation.
- Primary cells can be used for short-term experiments or expanded through serial passaging.
Immortalized Cell Lines:
- Immortalized cell lines are derived from primary cells but have undergone genetic modifications to overcome replicative senescence.
- Immortalized cells can be cultured indefinitely and are widely used in research.
- Common examples include HeLa cells (derived from cervical cancer) and HEK293 cells (derived from human embryonic kidney).
- Immortalized cell lines can maintain specific characteristics or be genetically modified to mimic disease conditions.
Cell Culture Conditions:
- Cells require a controlled environment for optimal growth in culture.
- Culture conditions include temperature, humidity, and a specific gas composition (typically 5% CO2 and 95% air).
- Culture medium is supplemented with amino acids, vitamins, salts, and serum (or serum-free alternatives) to provide essential nutrients and growth factors.
- pH is maintained within a physiological range (usually around 7.4) using a buffering system.
Cell Adhesion and Substrates:
- Cells require a suitable substrate for attachment and growth.
- Culture vessels can be coated with extracellular matrix proteins such as collagen, fibronectin, or gelatin to enhance cell adhesion.
- Synthetic substrates or hydrogels can also be used to mimic the extracellular matrix environment and provide specific cues for cell behavior.
Cell Proliferation and Passage:
- Cells in culture proliferate and expand over time, forming a monolayer or three-dimensional structures.
- As cells reach confluence, they can be detached using enzymatic (e.g., trypsin) or non-enzymatic methods.
- Passage refers to the process of transferring cells to new culture vessels to maintain cell viability and prevent overgrowth.
- Cells are typically subcultured at a defined ratio to maintain their characteristics and avoid replicative senescence.
Contamination Control:
- Maintaining sterility is crucial to avoid contamination in cell culture.
- Aseptic techniques, including working in a laminar flow hood and using sterile equipment and reagents, are employed.
- Antibiotics and antifungal agents can be added to the culture medium to prevent microbial contamination.
- Regular monitoring and periodic testing for contaminants are essential to ensure cell culture quality.
Quality Control and Characterization:
- Cell lines should be routinely authenticated and characterized to ensure their identity, purity, and functionality.
- Authentication techniques include DNA fingerprinting, short tandem repeat (STR) analysis, and karyotyping.
- Characterization involves assessing cell morphology, growth characteristics, surface markers, and functional assays.
Purifying proteins:
Cell Lysis and Homogenization:
- Cells are lysed to release their contents, including the target protein of interest.
- Various methods can be used, such as sonication, freeze-thaw cycles, or mechanical disruption.
- Lysis buffers containing detergents, salts, and protease inhibitors are used to maintain protein stability and prevent degradation.
Fractionation:
- Cell lysate is subjected to fractionation techniques to separate cellular components based on their size, charge, or solubility.
- Common fractionation methods include differential centrifugation, ultracentrifugation, and filtration.
- This step helps remove cell debris, organelles, and other macromolecules, allowing for the enrichment of the target protein.
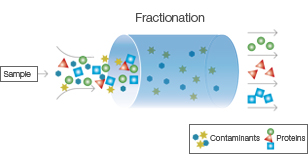
Protein Extraction:
- Proteins can be extracted from cellular fractions using various techniques, depending on their physicochemical properties.
- Salting out: Precipitation of proteins by adding high concentrations of salts (e.g., ammonium sulfate).
- Solvent extraction: Partitioning of proteins based on their solubility in organic solvents or detergents.
- Chromatography: Separation of proteins based on their affinity for specific ligands or physical properties.
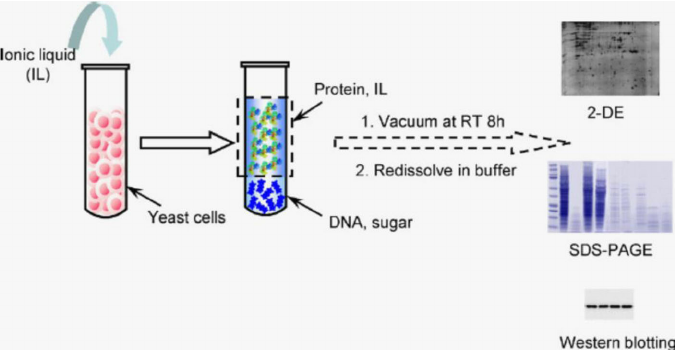
Chromatography Techniques:
- Chromatography is the most commonly used method for protein purification, allowing for high resolution and specificity.
- Different chromatographic methods can be employed at different stages of purification, including:
- Affinity chromatography: Exploits specific interactions between the target protein and an immobilized ligand (e.g., antibody, metal ions, or receptor).
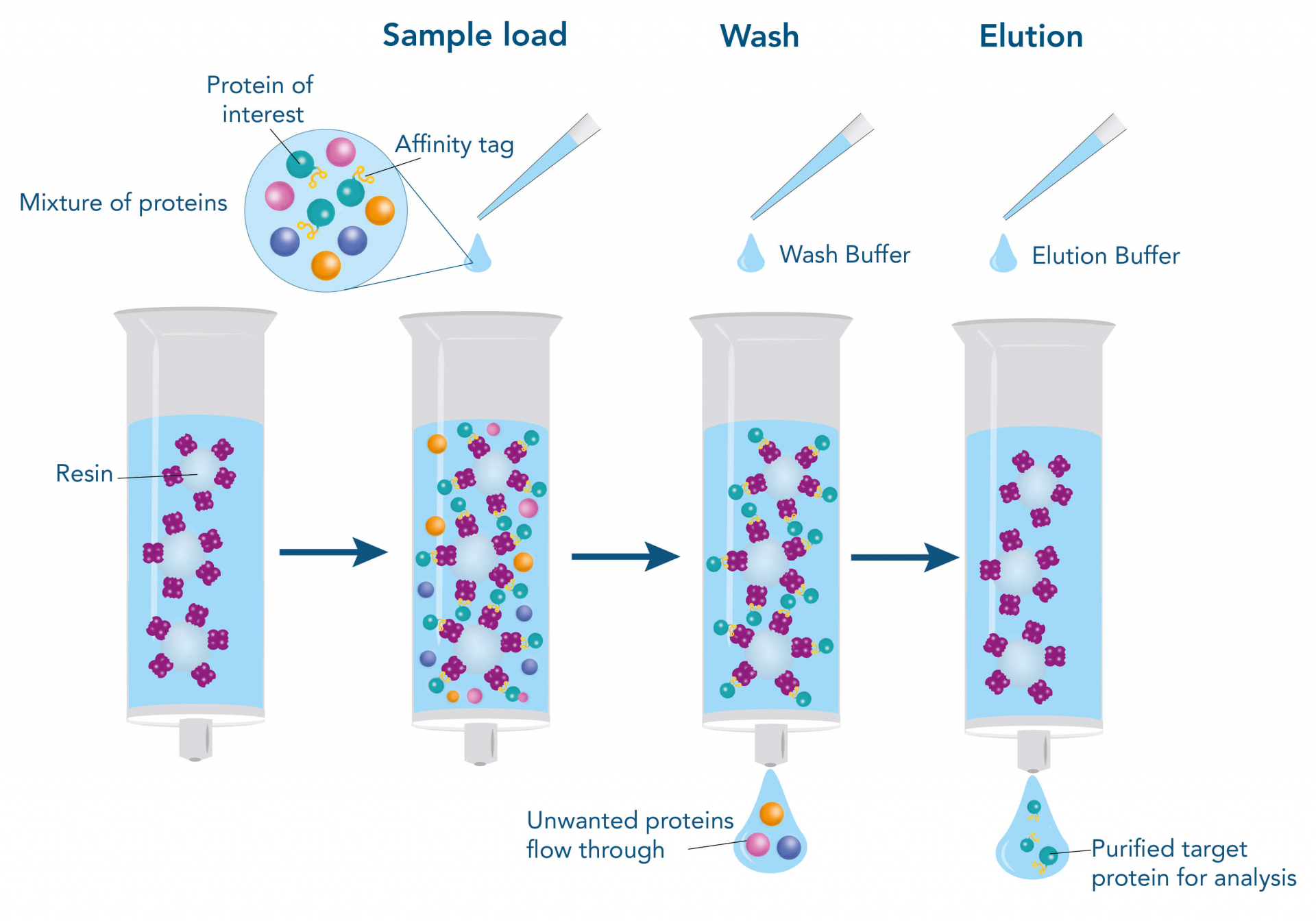
- Ion-exchange chromatography: Separates proteins based on their net charge and affinity for charged resin.
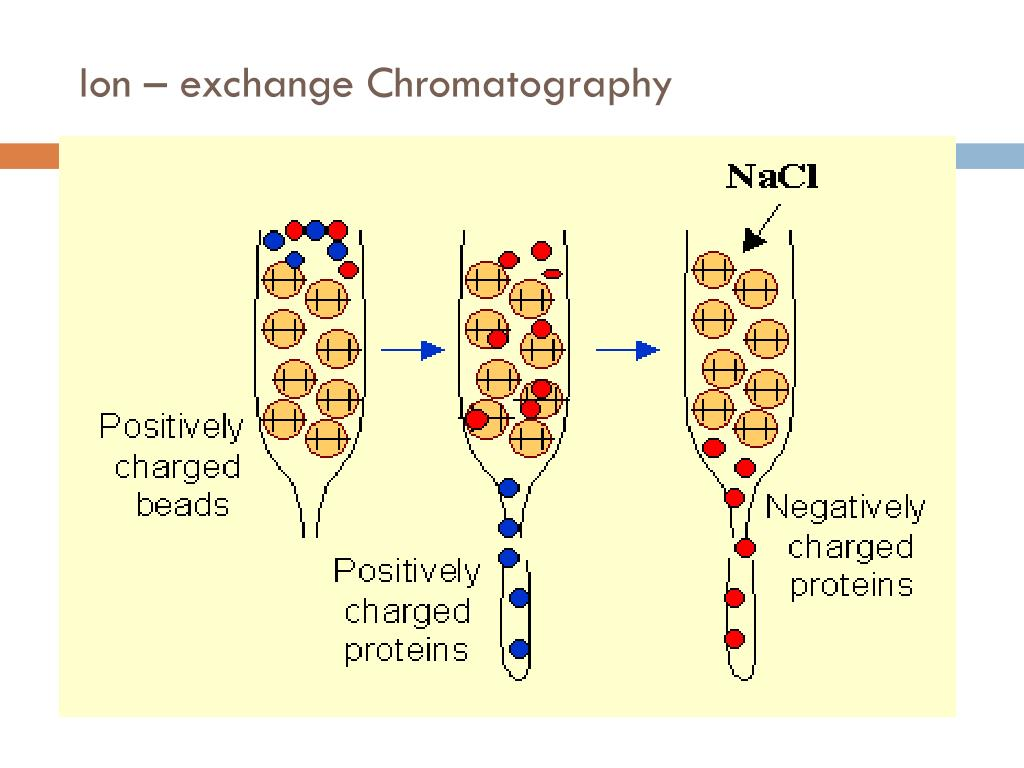
- Size-exclusion chromatography: Separates proteins based on their size and shape, allowing for the removal of contaminants.
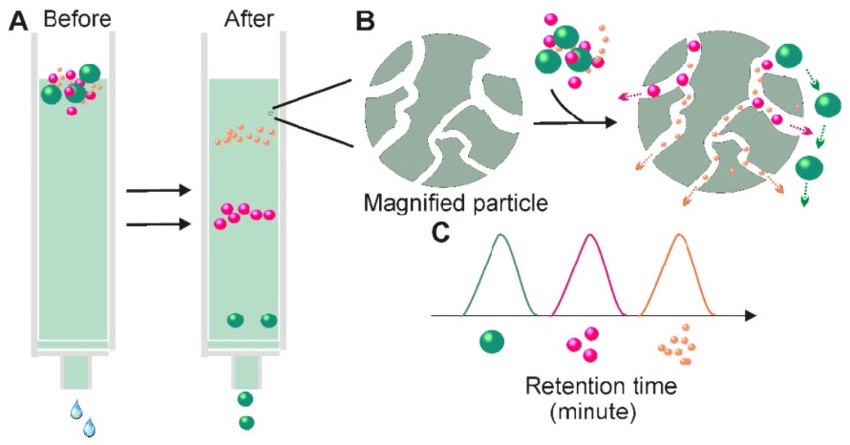
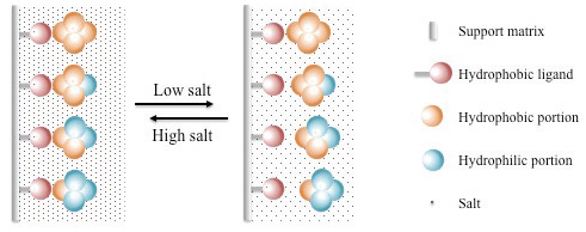
- Hydrophobic interaction chromatography: Separates proteins based on their hydrophobicity, utilizing the interaction between the protein and a hydrophobic resin.
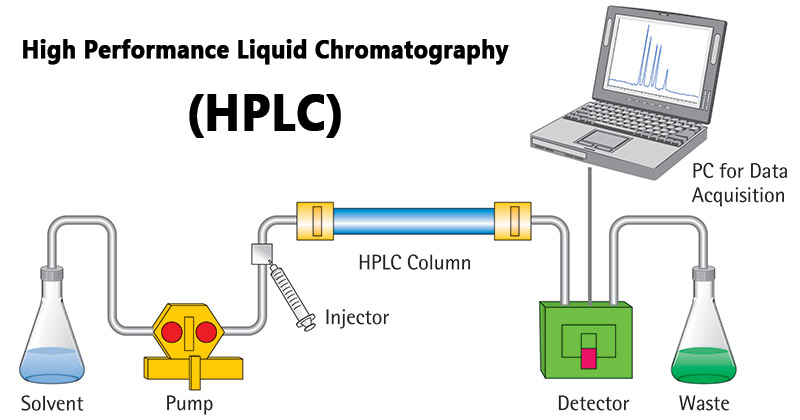
- High-performance liquid chromatography (HPLC): Utilizes advanced liquid chromatography techniques for higher resolution and efficiency.
Protein Analysis:
- Throughout the purification process, protein fractions are analyzed to assess purity and quantity.
- Common methods include sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE), western blotting, and enzyme activity assays.
- Spectrophotometry and fluorometry can be used to measure protein concentration and determine purity.
Protein Refolding:
- If a purified protein has lost its native conformation and activity during purification, refolding may be required.
- Refolding techniques involve carefully controlling the protein's environment, including buffer conditions, temperature, and the presence of additives or chaperones.
- Optimization of refolding conditions can restore the protein to its functional form.
Storage and Preservation:
- Purified proteins should be stored under appropriate conditions to maintain their stability and activity.
- Storage temperatures, buffer composition, and the addition of cryoprotectants (e.g., glycerol) or stabilizers can help prevent degradation and denaturation.
- Aliquots of purified protein can be stored at ultra-low temperatures (e.g., -80°C) or lyophilized for long-term preservation.
Analyzing Proteins:
Protein Quantification:
- Protein quantification methods determine the concentration of proteins in a sample.
- Common techniques include the Bradford assay, Lowry assay, bicinchoninic acid (BCA) assay, and spectrophotometric absorbance at 280 nm.
- Quantification is important for ensuring accurate loading of proteins in downstream experiments and comparing protein levels between samples.
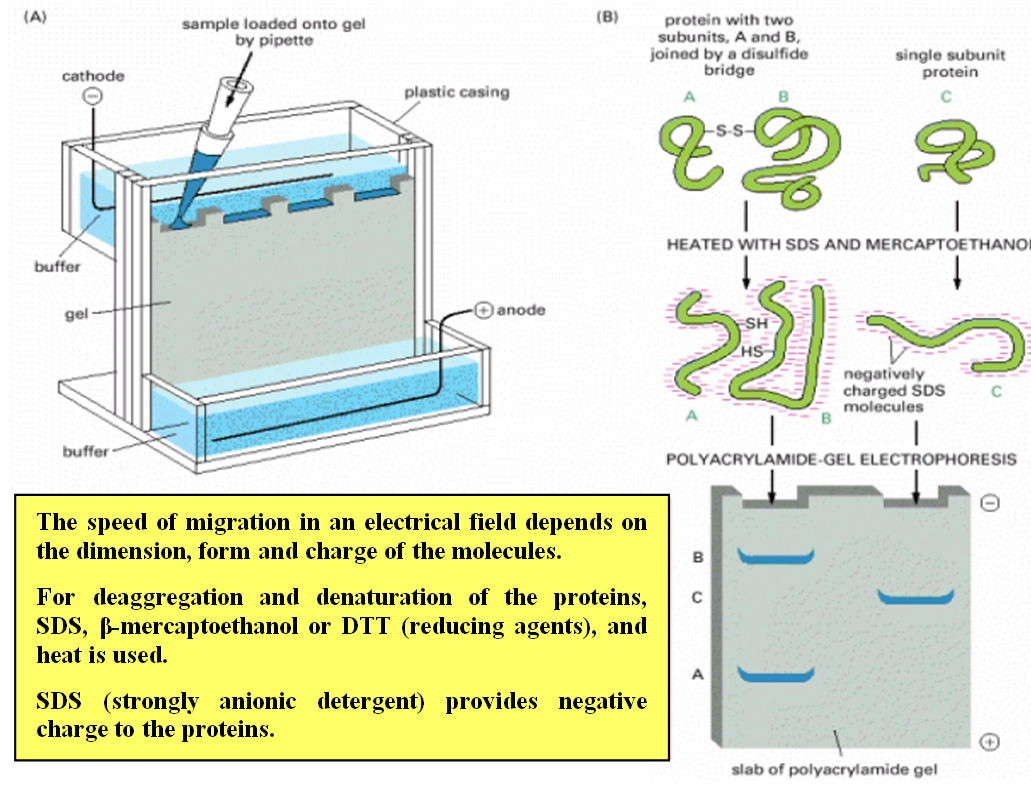
Sodium Dodecyl Sulfate-Polyacrylamide Gel Electrophoresis (SDS-PAGE):
- SDS-PAGE separates proteins based on their size.
- Proteins are denatured and coated with SDS, a detergent that imparts a negative charge to the proteins.
- The proteins are then loaded into a polyacrylamide gel, and an electric field is applied.
- Smaller proteins migrate faster through the gel, while larger proteins migrate more slowly.
- SDS-PAGE allows visualization of protein bands and estimation of molecular weight using protein size markers.
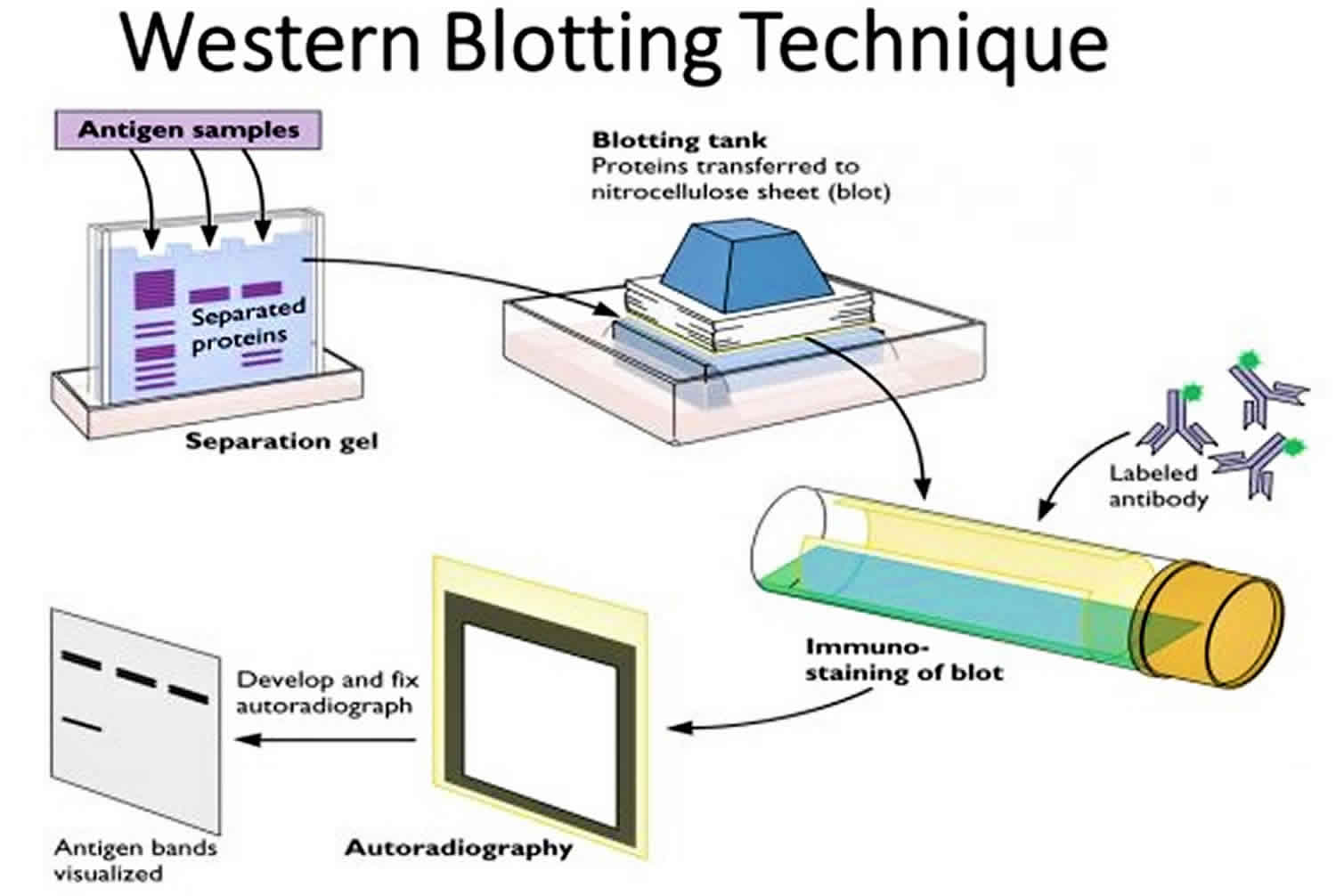
Western Blotting (Immunoblotting):
- Western blotting detects and characterizes specific proteins within a complex mixture.
- Proteins separated by SDS-PAGE are transferred to a membrane (typically nitrocellulose or PVDF).
- The membrane is incubated with primary antibodies that recognize the target protein.
- Detection is achieved using secondary antibodies conjugated to enzymes or fluorophores, generating a signal that can be visualized.
- Western blotting allows quantification of protein expression levels and identification of post-translational modifications.
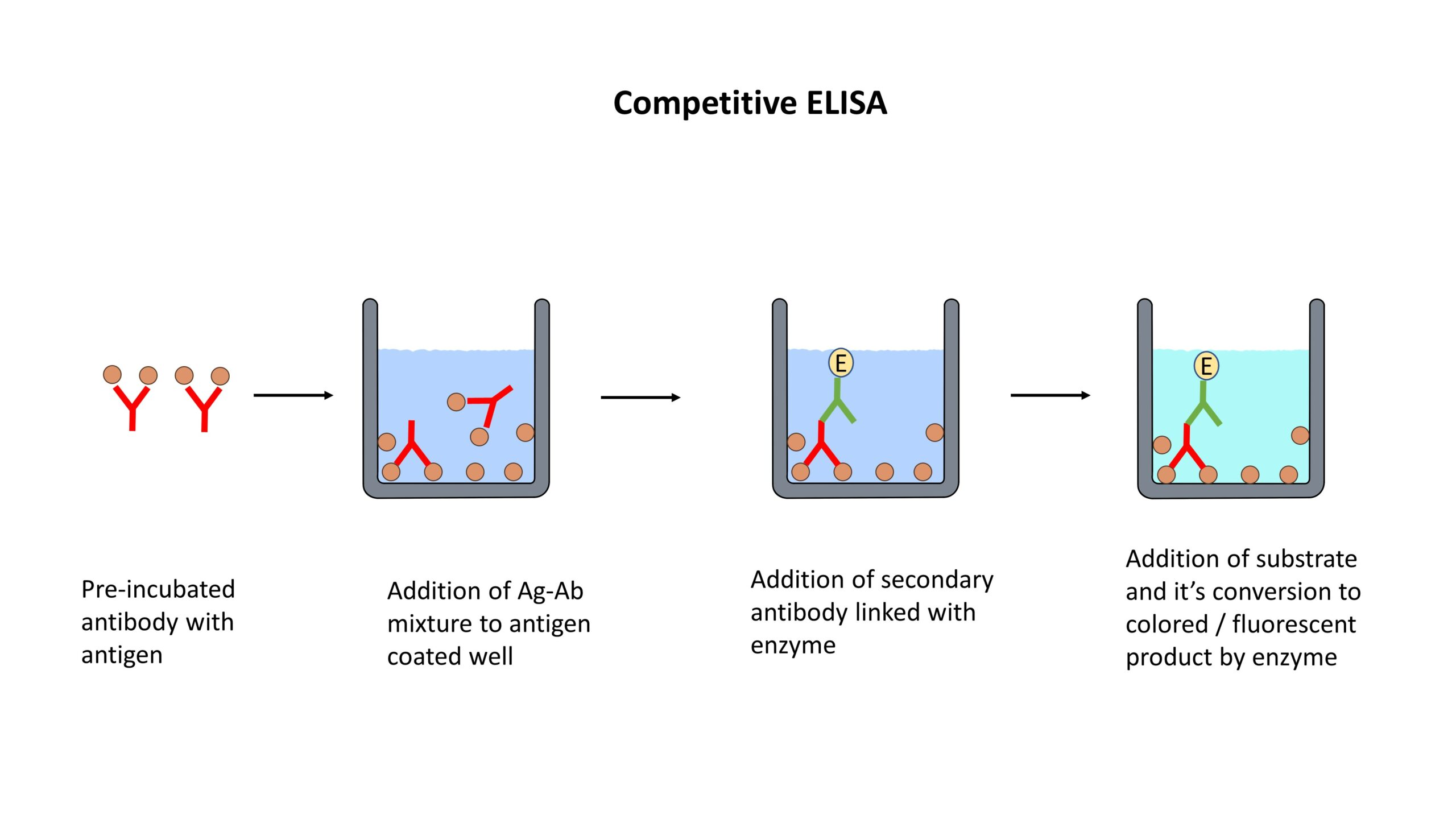
Enzyme-Linked Immunosorbent Assay (ELISA):
- ELISA detects and quantifies specific proteins using antibodies.
- The target protein is immobilized on a solid surface, such as a microplate.
- The immobilized protein is then incubated with a primary antibody specific to the target protein.
- Detection is achieved using a secondary antibody conjugated to an enzyme, which produces a colorimetric or fluorescent signal.
- ELISA is widely used for protein quantification, biomarker analysis, and immunological research.
Mass Spectrometry (MS):
- Mass spectrometry analyzes proteins by measuring their mass-to-charge ratio.
- Proteins are digested into smaller peptides using proteolytic enzymes (e.g., trypsin).
- The resulting peptides are ionized and introduced into the mass spectrometer.
- Mass spectrometers detect and measure the masses of the ions, allowing identification and quantification of the peptides.
- Protein identification is achieved by comparing the obtained peptide masses to protein databases.
- MS can also be coupled with liquid chromatography (LC-MS) to improve peptide separation and identification.
Protein-Protein Interactions:
- Studying protein-protein interactions helps understand protein function and cellular processes.
- Techniques such as co-immunoprecipitation (co-IP), pull-down assays, and yeast two-hybrid screening can identify interacting proteins.
- Co-IP involves immunoprecipitating a target protein along with its interacting partners using specific antibodies.
- Pull-down assays use affinity tags or bait proteins to capture interacting partners from a complex mixture.
- Yeast two-hybrid screening utilizes the yeast cell's transcriptional activity to identify protein-protein interactions.
Structural Analysis:
- Determining protein structure provides insights into function and interactions.
- X-ray crystallography and nuclear magnetic resonance (NMR) spectroscopy are commonly used methods.
- X-ray crystallography involves crystallizing the protein and analyzing the diffraction pattern produced by X-rays.
- NMR spectroscopy analyzes the interactions between atomic nuclei in the protein to determine its structure in solution.
- Cryo-electron microscopy (cryo-EM) is a powerful technique for visualizing protein structures at near-atomic resolution without the need for crystallization.
Proteomics:
- Proteomics involves the large-scale analysis of proteins and their modifications.
- It includes techniques such as 2D gel electrophoresis, protein profiling, and shotgun proteomics.
- Proteomic approaches provide comprehensive insights into protein expression, post-translational modifications, and protein networks.
Analyzing and Manipulating DNA
DNA Extraction:
- DNA extraction is the first step in analyzing and manipulating DNA.
- Common methods include cell lysis to release DNA, followed by purification steps to remove contaminants such as proteins and RNA.
- DNA extraction can be performed from various sources, including cells, tissues, blood, or environmental samples.
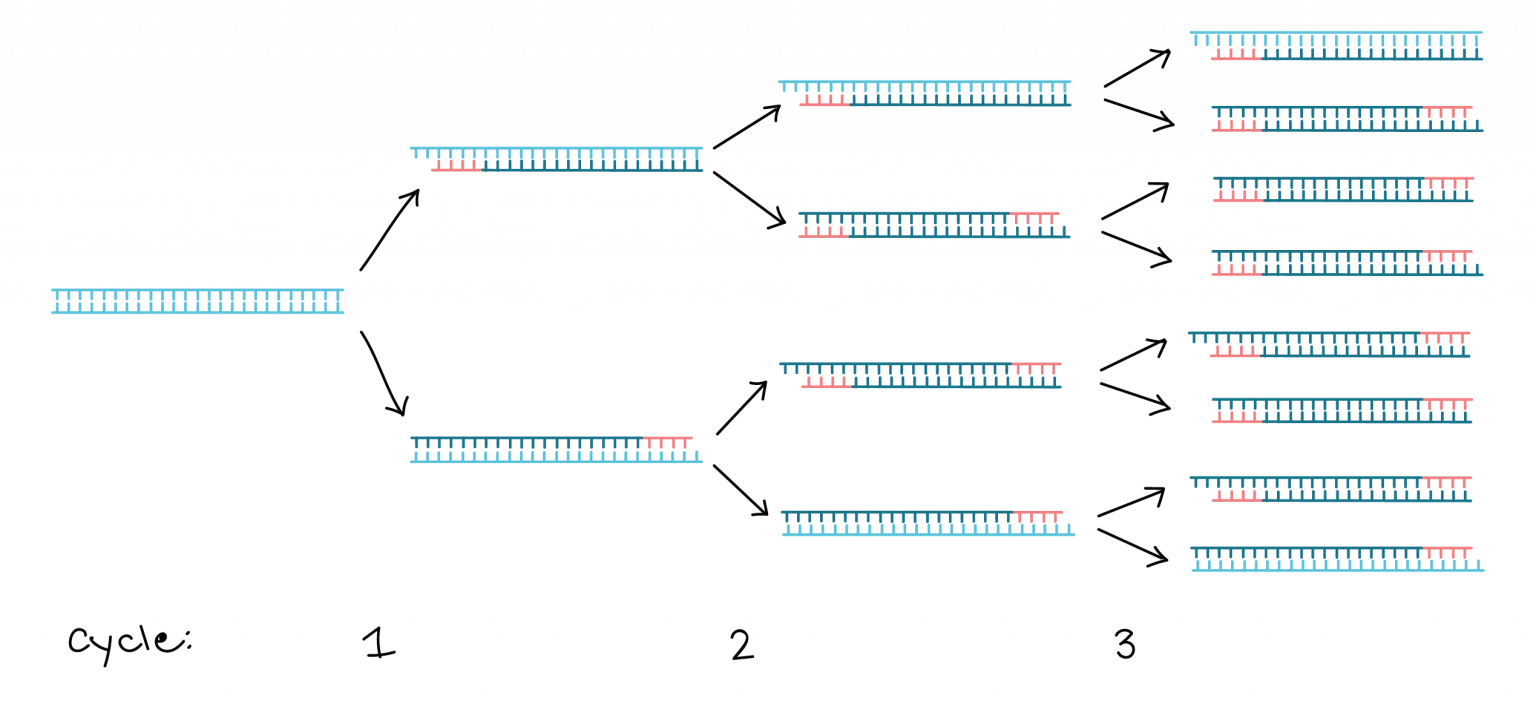
Polymerase Chain Reaction (PCR):
- PCR amplifies specific DNA sequences in vitro.
- It involves a series of temperature cycles that denature the DNA, anneal primers specific to the target sequence, and extend the DNA using a DNA polymerase enzyme.
- PCR enables the amplification of a small amount of DNA into millions of copies, allowing for further analysis or manipulation.
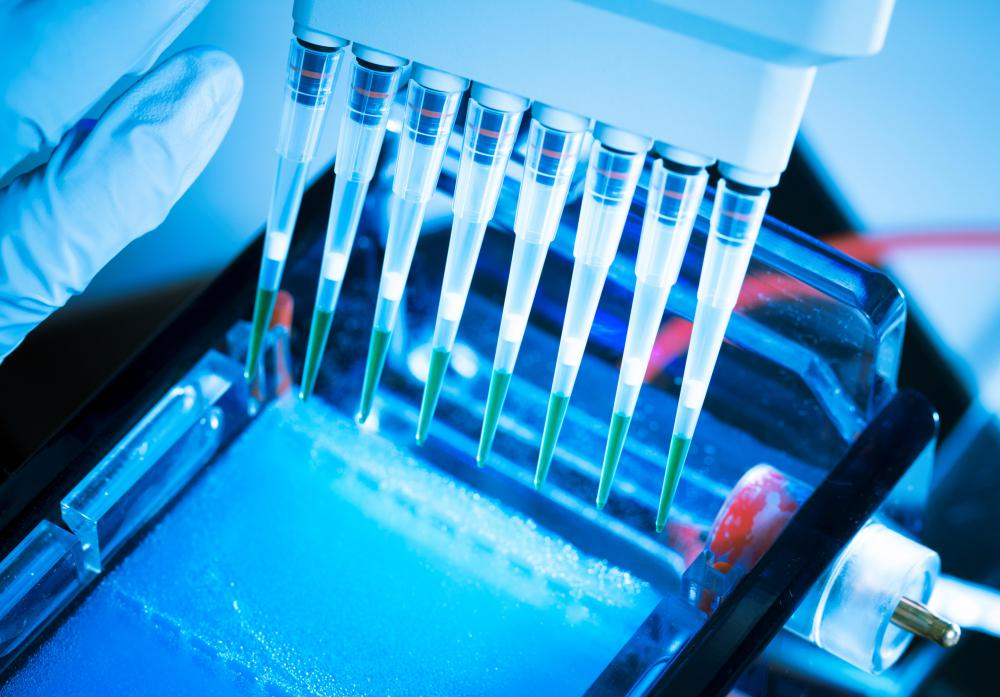
Gel Electrophoresis:
- Gel electrophoresis separates DNA fragments based on their size and charge.
- DNA samples are loaded into wells of an agarose or polyacrylamide gel and subjected to an electric field.
- Smaller DNA fragments migrate faster through the gel, while larger fragments migrate more slowly.
- DNA bands can be visualized using fluorescent dyes, intercalating agents, or specific DNA stains.
DNA Sequencing:
- DNA sequencing determines the order of nucleotide bases in a DNA molecule.
- Sanger sequencing, also known as chain-termination sequencing, was the first widely used method.
- Next-generation sequencing (NGS) techniques, such as Illumina sequencing, enable high-throughput sequencing of DNA.
- DNA sequencing is crucial for studying genetic variations, identifying mutations, and understanding genomic structures.
DNA Cloning:
- DNA cloning involves the replication of a specific DNA fragment and its insertion into a vector for replication in a host organism.
- The DNA fragment of interest is typically inserted into a plasmid or a viral vector.
- Cloning allows for the production of large quantities of DNA, studying gene function, and generating recombinant proteins.
Restriction Enzymes:
- Restriction enzymes, also known as restriction endonucleases, cleave DNA at specific recognition sites.
- They recognize short DNA sequences and cut the DNA, generating fragments with sticky ends or blunt ends.
- Restriction enzymes are widely used in DNA cloning, genetic engineering, and molecular biology techniques.
DNA Modification:
- DNA can be modified by various methods to introduce changes in its sequence or structure.
- Site-directed mutagenesis allows for the introduction of specific mutations in the DNA sequence.
- DNA modification techniques, such as methylation or acetylation, can alter gene expression patterns and epigenetic modifications.
DNA Hybridization:
- DNA hybridization involves the pairing of complementary DNA strands.
- It is widely used in techniques such as Southern blotting, Northern blotting, and DNA microarrays.
- DNA probes, labeled with fluorescent or radioactive markers, hybridize to specific DNA sequences of interest.
CRISPR-Cas9 Genome Editing:
- CRISPR-Cas9 is a revolutionary genome editing tool.
- It utilizes a guide RNA (gRNA) to target specific DNA sequences, and the Cas9 nuclease introduces precise cuts at the target site.
- CRISPR-Cas9 allows for gene knockout, gene insertion, or gene editing with high efficiency and specificity.
- It has transformed genetic research, disease modeling, and potential therapeutic applications.
Studying gene expression and function
Transcriptomics:
- Transcriptomics investigates gene expression patterns by analyzing the complete set of RNA molecules (transcriptome) in a cell or tissue.
- Techniques such as RNA sequencing (RNA-seq) provide a comprehensive view of the transcriptome, allowing the identification of expressed genes, alternative splicing events, and noncoding RNAs.
- Differential gene expression analysis compares gene expression levels between different conditions or cell types, providing insights into gene function and regulatory mechanisms.
Gene Knockout and Knockdown:
- Gene knockout involves disrupting or inactivating a specific gene to study its function.
- Techniques such as CRISPR-Cas9 can be used to create targeted mutations in the DNA sequence, leading to loss-of-function of the gene.
- Gene knockdown utilizes RNA interference (RNAi) or antisense oligonucleotides to reduce the expression of a specific gene, allowing the assessment of its role in cellular processes.
Functional Genomics:
- Functional genomics aims to understand the function of genes and their interactions within biological systems.
- Techniques such as gene expression profiling, protein-protein interaction analysis, and high-throughput screening are used to investigate gene function.
- Large-scale studies, such as genome-wide association studies (GWAS) and CRISPR-based genetic screens, provide insights into the relationships between genotype, gene expression, and phenotype.
Reporter Assays:
- Reporter assays allow the measurement of gene expression levels and promoter activity.
- A reporter gene, such as green fluorescent protein (GFP) or luciferase, is fused to the promoter region of the gene of interest.
- The activity of the reporter gene reflects the transcriptional activity of the target gene.
- Reporter assays are used to study the regulation of gene expression, identify cis-regulatory elements, and assess the effects of mutations or regulatory factors.
Gene Expression Analysis:
- Techniques such as reverse transcription polymerase chain reaction (RT-PCR) and quantitative real-time PCR (qPCR) enable the quantification of gene expression levels.
- RT-PCR converts RNA into complementary DNA (cDNA), which is then amplified and detected.
- qPCR allows for the quantification of gene expression in real-time using fluorescent dyes or probes.
- Gene expression analysis is used to study gene regulation, validate transcriptomic data, and assess changes in gene expression under different conditions or treatments.
Functional Assays:
- Functional assays assess the biological activity of genes or gene products.
- They can include assays for enzyme activity, protein-protein interactions, protein localization, or protein function.
- Functional assays help elucidate the role of genes in specific cellular processes and pathways.
Mathematical Analysis of Cell Functions
Modeling Cellular Processes:
- Mathematical models are used to describe and simulate various cellular processes, including signaling pathways, gene regulation, metabolic networks, and cell population dynamics.
- Models capture the behavior of biological systems using mathematical equations that represent the interactions and dynamics of cellular components.
- They provide a quantitative framework to understand complex cellular phenomena and make predictions about cellular behavior.
Differential Equations:
- Differential equations are a fundamental tool in mathematical biology.
- They describe how variables change over time based on their rates of change.
- Ordinary differential equations (ODEs) model intracellular processes such as enzyme kinetics, gene expression, and signaling dynamics.
- Partial differential equations (PDEs) describe spatially distributed phenomena, such as cell movement, tissue growth, and reaction-diffusion processes.
Systems Biology:
- Systems biology aims to understand biological systems as a whole by integrating experimental data with mathematical modeling.
- It focuses on analyzing and modeling the interactions between various components, such as genes, proteins, and metabolites, to understand emergent properties and system-level behavior.
- Systems biology approaches include network analysis, dynamical modeling, and parameter estimation to study cellular processes comprehensively.
Network Analysis:
- Network analysis characterizes cellular processes as networks of interconnected components.
- Graph theory is used to model and analyze these networks.
- Network analysis can reveal important nodes (genes, proteins) and interactions, identify key regulatory elements, and predict cellular behaviors.
- It helps uncover the underlying organizational principles of biological systems and identify potential targets for intervention.
Parameter Estimation:
- Parameter estimation involves determining the values of unknown parameters in mathematical models using experimental data.
- It is crucial for model calibration and validation.
- Various optimization techniques, such as least squares fitting, maximum likelihood estimation, or Bayesian inference, are used to estimate parameter values that best fit experimental data.
Computational Simulations:
- Computational simulations involve running mathematical models to simulate cellular processes and predict their behavior.
- Simulations can reveal insights into complex dynamic behaviors, test hypotheses, and explore the effects of perturbations.
- They provide a platform for in silico experiments, allowing researchers to explore the behavior of a system under different conditions or parameter values.
Model Validation and Experimental Design:
- Mathematical models need to be validated against experimental data to ensure their accuracy and predictive power.
- Model validation involves comparing model predictions with experimental observations.
- Model-guided experimental design uses mathematical models to optimize experimental protocols, identify key measurements, and explore system behavior in an efficient and cost-effective manner.
Statistical Analysis:
- Statistical analysis is crucial for evaluating the significance of experimental results and validating mathematical models.
- It includes hypothesis testing, confidence interval estimation, regression analysis, and analysis of variance (ANOVA).
- Statistical methods help determine the reliability of experimental observations, assess model performance, and make robust conclusions.
\