BIOC E Note
1.Membrane lipids/Organization
Biological membranes = plasma and organelle membranes
Functions of cellular membranes
- Permeability barrier/compartmentalization
- Communication
- Energy Conversion
- Surface recognition
Structure = 45% lipid, 50% protein, 5% carbohydrate
Amphipathic (hydrophobic and hydrophilic properties
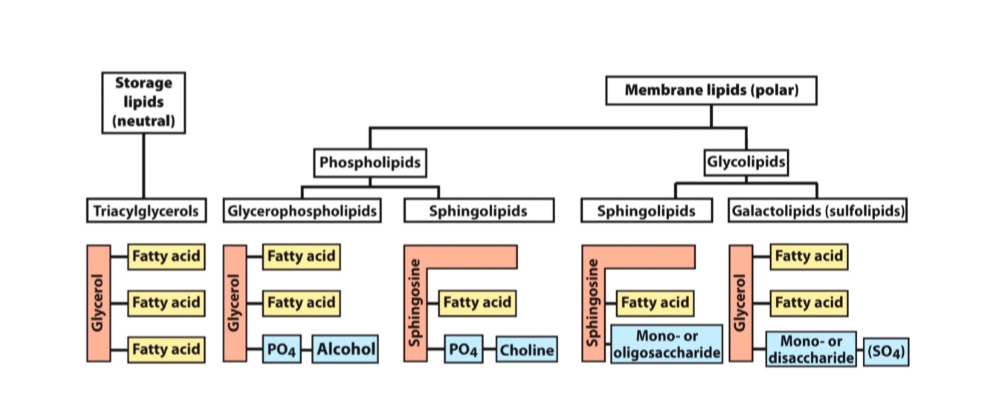
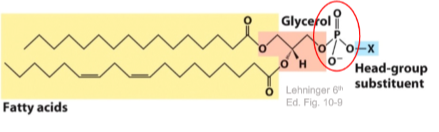
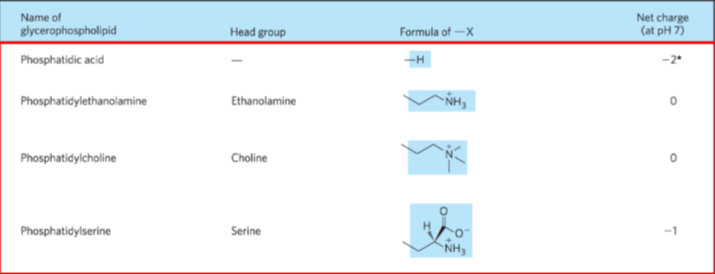
Phosphoglycerides
Glycerol backbone
2 fatty acids in an ester link
Alcohol head group
Sphingolipids
Sphingosine backbone
1 fatty acids in an amide link
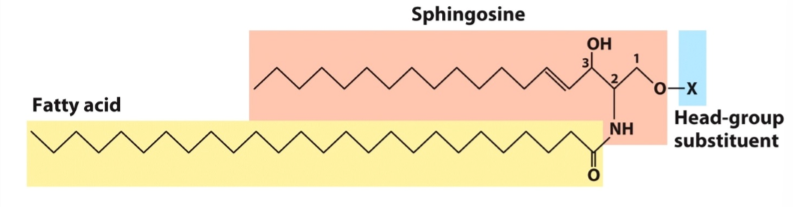

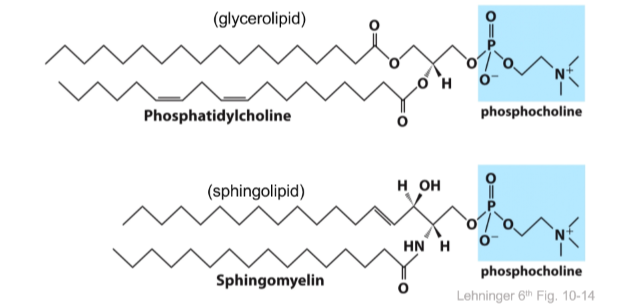
Similar in shape to glycerolipids (glycerol + one fatty acid)
Glycolipids
- sphingolipids with carbohydrate headgroups
- glycosphingolipids = part of the ABO blood type
- Blood compatibility
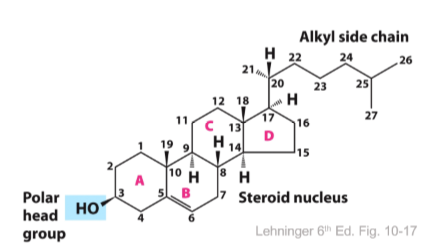
Cholesterol
major animal sterol
4 fused rings
alkyl side chain in the steroid
polar group is a simple OH
Membrane composition based on function
- Different membrane types have different compositions
- phosphatidylcholine is a major lipid in all membranes
- cholesterol is a major component of plasma membrane
Monolayers (artificial)
- form at the air-water interface
- the polar head group interacts with aqueous solution
- hydrophobic tails interact with the air
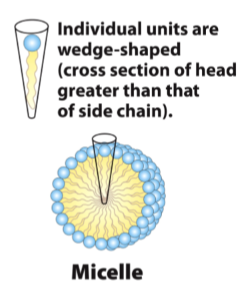
Micelles
detergents and lipids with one acyl tail
head group is wider than the tail so it curves
lipid tails interact with tails and heads interact with water
Bilayer
- nonpolar tails associate in the interior
- ~3mm thick (stable)
- basic structure of all biological membranes
- impermeable to ions and polar molecules
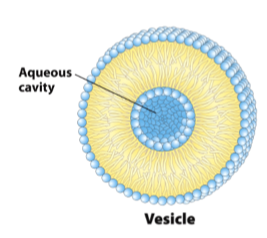
Liposomes/vesicles
resemble bilayers
center is an aqueous solution
artificial liposomes allow the study of membrane transporters
2. Membrane Proteins
Functions
- transporters and channels
- receptors
- structural components
- adhesion proteins
- surface antigens
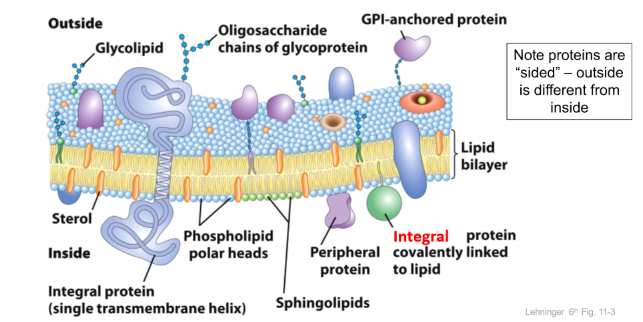
Fluid Mosaic model
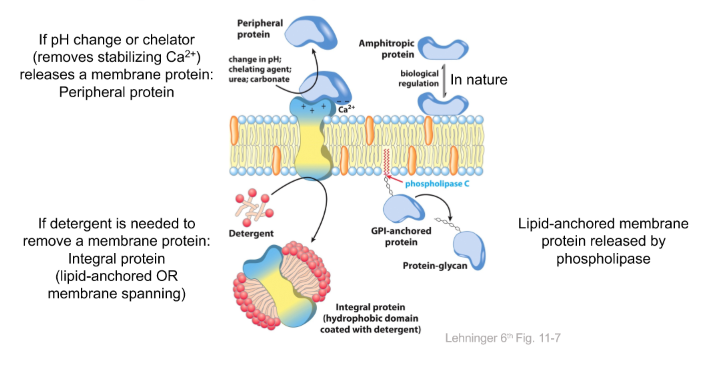
pH or chelator releases a membrane protein = PERIPHERAL
Detergent releases a membrane protein = INTEGRAL
Phospholipase = lipid-anchored
1. Peripheral
- interactions between peripheral proteins and polar head groups of membrane lipids
- electrostatic (cation or charged side chains)
- hydrogen bonds
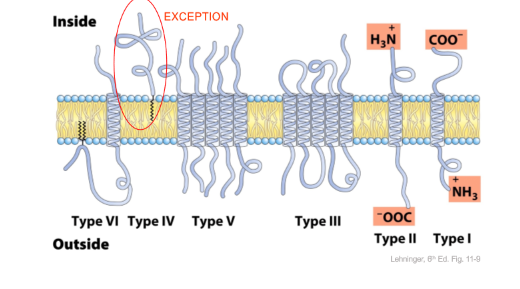
2. Lipid Anchored
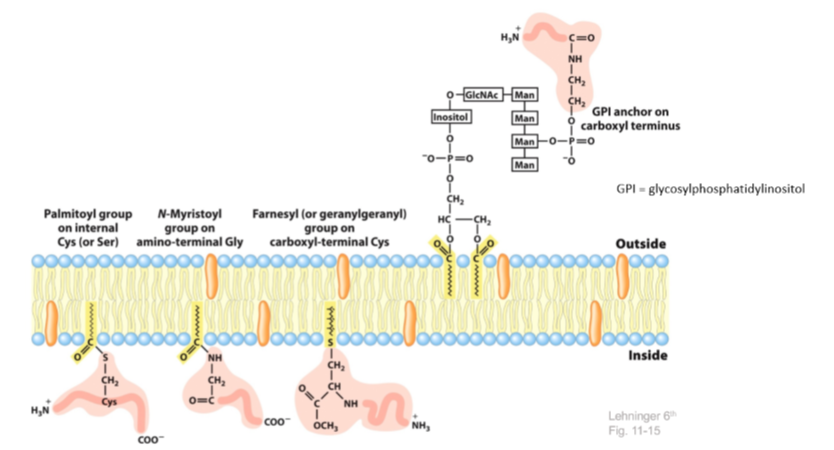
- Palmitoylaiton = addition of a palmitoy group on the cys or ser
- Myristoylation = addition of a myristoyl group on amino-terminal glycine
- Farnesylation = addition of an isoprenoid to the c terminal of a cysteine residue
- GPI anchor = 2 phosphates and an inositol group (protein tether)
- fatty acids (that do not want to be in the aqueous environment of the cell)
3. Membrane Spanning
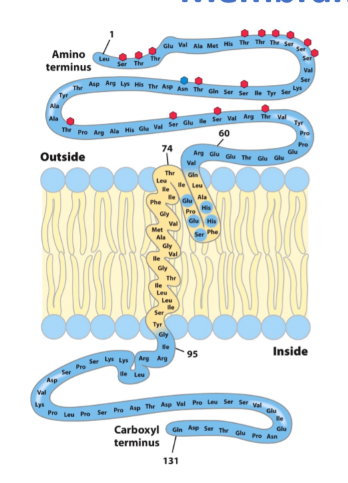
glycophorin A = single spanning transmembrane
- glycosylated
highly hydrophobic regions in yellow
less hydrophobic regions in blue
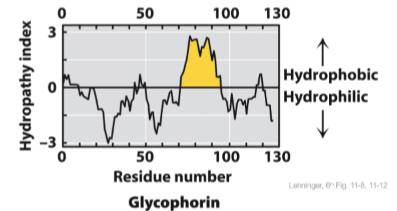
Hydropathy plots predict transmembrane helices
average hydrophobicity of a protein segment
if >20 successive residues have a high hydropathy index, possibly a transmembrane segment
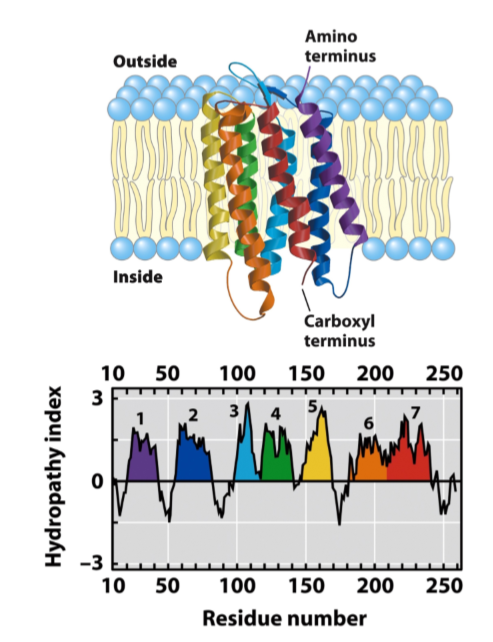
Bacteriorhodopsin - multispanning transmembrane
GPCR (g protein receptor family)
7 transmembrane segments, but short extracellular loops
Annular Lipids
crystal structures of membrane proteins often show a layer of well-ordered lipids
head groups of these annular lipids interact with the hydrophilic extracellular loops
Fatty acid tails interact with the transmembrane helices
Lipids look similar to the bilayer
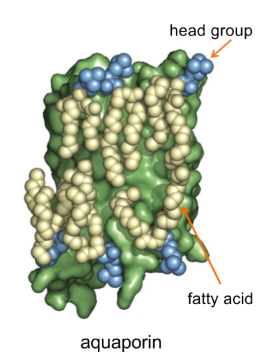
B-Barrel integral membrane proteins
bacterial and mitochondrial outer membrane proteins are B-barrels
Backbone hydrogen bonds between strands
Strands can be seven residues → don’t show up on hydropathy plots
amino acid side chains alternate hydrophobic/hydrophilic
- hydrophobic side chains oriented towards hydrophobic lipid tails
- hydrophilic side chains oriented towards aqueous environment of the pore
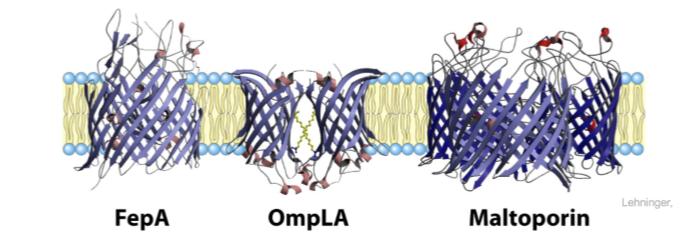
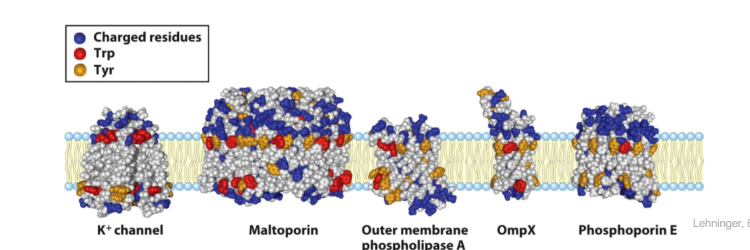
Residues at the membrane interface
Tyrosine and tryptophan are concentrated where polar head groups meet acyl chains
Charged residues (ARG, LYS, GLU, ASP) are found exclusively in the aqueous phase (almost exclusively)
Glycoproteins (STRUCTURE)
- N-linked carbohydrate chain
- ASN side chain (-CO-NH2)
- N-acetylglucosamine (GlcNAc)
- O-linked carbohydrate chain
- SER or THR side chain (-OH)
- N-acetylgalactosamine (GalNAc)
- Sugar groups of glycoprotein and glycolipids
- Contribute to cell surface recognition
- function as receptors
3. Membrane Dynamics
1. Fluidity
Membranes are dynamic
- Change shape without loss of integrity or becoming leaky
- Fluid-mosaic model
- Allows lateral movement and “Flipping/Flopping/Scrambling”
Lipid bilayer state changes
- Gel phase:
- All motion of bilayer is constrained
- Lipids ordered in a paracrystalline state
- Lipid-ordered state (physiological)
- intermediate thermal motion of acyl chains and atoms
- lateral movement in the plane of the bilayer is allowed
- Lipid-disordered state (fluid state)
- hydrocarbon chains are in constant motion, no regular organization
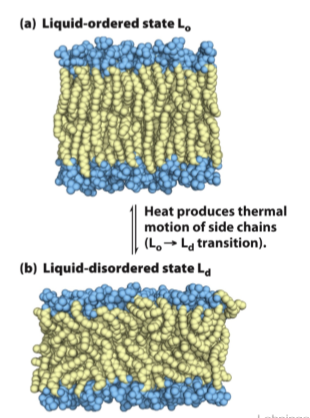
Membrane composition affects fluidity
- At physiological temperatures
- Long chain saturated FA (C16:0, C18:0) pack well into liquid-ordered state
- Unsaturated and shoter chain fatty acids favor liquid-disordered state
- Sterols (cholesterol) reduce fluidity
- Cells regulate lipid composition to achieve a constant membrane fluidity
- Bacteria synthesize more unsaturated fatty acids and fewer saturated ones when cultured at low temperatures
2. Cholesterol
- Long saturated fatty acids = INCREASE fluidity
- Cholesterol interferes with acyl chains
- Unsaturated, cis fatty acids: decrease fluidity
- cholesterol allows efficient packing of kinked chains
- High temperatures: decrease fluidity
- rigid cholesterol interacts with flexible acyl chains
- Low temperatures: increases fluidity
- cholesterol prevents acyl chains from interacting
3. Diffusion/movement in membrane
At physiological temperatures, transbilayer motion or “flip flop” occurs very slowly
Requires that the polar or charged head group leaves its aqueous environment and moves through the hydrophobic interior of the bilayer
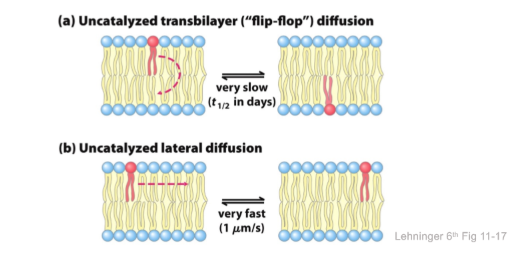
catalyzed transbilayer translocations
- Flippase = P-type ATPase
- moves PE and PS from outer to cytosolic leaflet
- Floppase = ABC transporter
- phospholipids from cytosolic to outer leaflet
- Scramblase
- moves lipids in either direction toward equilibrium
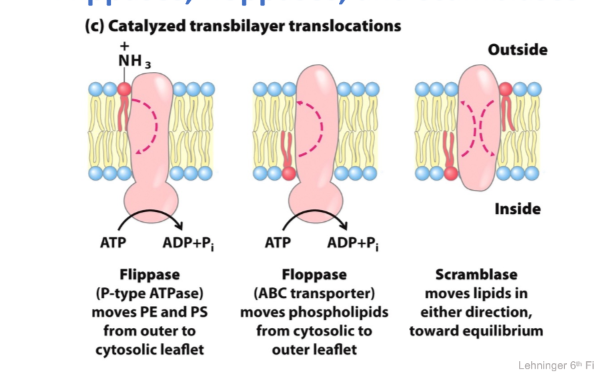
Lateral Diffusion by hops
- Single particle tracking = follows single lipid molecule on short time scale
- traces show that lipids diffuse rapidly and freely within a restricted region
- More rarely they “hop” into a new region
- Lipids behave as through they are corralled by fences, which they occasionally escape
Motion is restricted by SPECTRIN
- proteins are tethered in aggregates or patches
- Spectrin is part of the cytoskeleton
- it links to membrane proteins through ankyrin
- may act as the corral, keeping lipids from diffusing freely
4. Microdomains
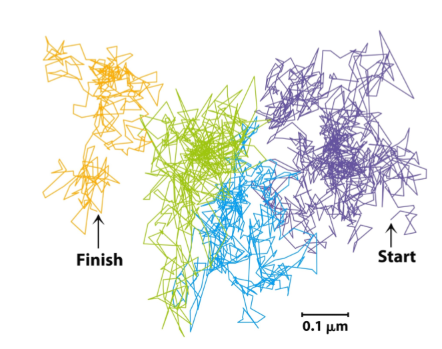
Lipid rafts in plasma membranes
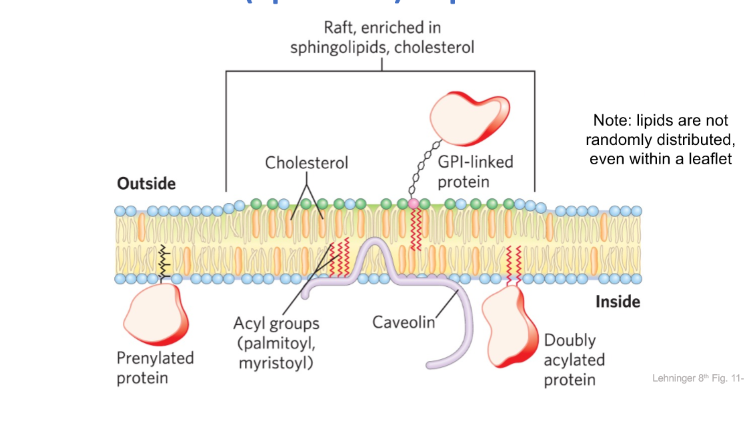
4. Intracellular membrane trafficking
Mediates:
- Reorganization of membrane-bound compartments
- exchange of membrane and “cargo” between compartments
- internalization/recycling/degradation of material from plasma membrane
Trafficking is a complex, multi-stage process
Budding (fission) of the vesicle from the parent membrane
Transport of the vesicle
Tethering/docking at target membrane
- Fusion of vesicle and target membrane
SNAREs
mediate membrane fusion during many key cellular processes
- Insulin secretion
- up-regulation of glucose transporters
- Transport between ER and Golgi
- Phagocytosis
- neurotransmitter release
Soluble N-etylmaleimide-sensitive factor attachment protein receptor
v-SNARE and t-SNARE are single-spanning transmembrane proteins
they also have an extended helical domain (60AA)
helical domains can interact to form a coiled-coil structure with SNAP25
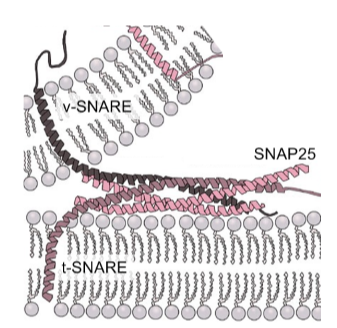
SNAREs mediate neurotransmitter release
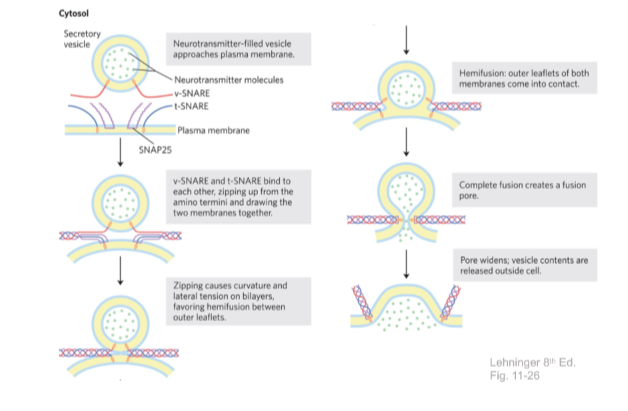
5. Membrane permeability/transport
Permeability
- Ions and polar molecules cannot cross (essentially impermeable)
- Na+, Cl-, sugars, amino acids
- Small uncharged molecules can cross slowly
- glycerol, ethanol
- Hydrophobic molecules, gases cross quickly
- steroid hormones, O2, CO2, N2
Simple and Facilitated Diffusion
Ion Gradients across membranes
- ionic composition of the cytosol is different from the extracellular environment
- Concentration gradients
- across the plasma membrane
- across organelle membranes
- Ion gradients are actively maintained by cell, at the cost of ATP
- Ion gradients can do work
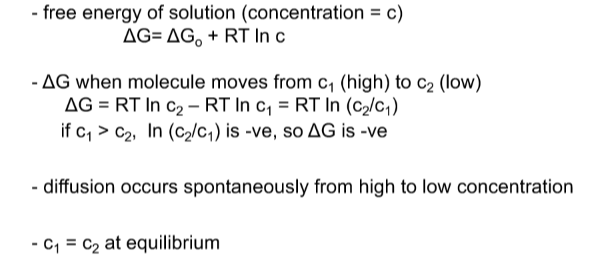
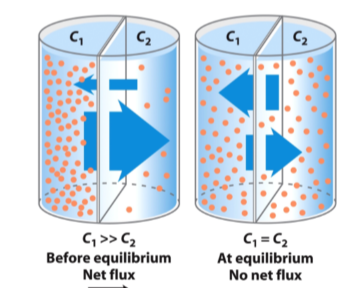
Simple Diffuison
Movement of a neutral solute across a permeable membrane
- neutral compound = no charge implications
- HIGH TO LOW C until equilibrium is reached
Transport of hydrophilic solutes
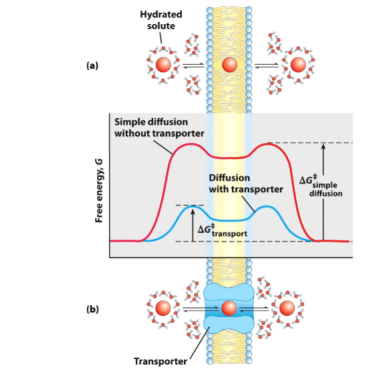
- Loss of energy is required to pass to the other side and regain the hydration sphere
- Adding a transport protein = transported through a transporter
- she hydration sphere to enter the transporter
- Transport protein provides an environment for the molecule to move through without entering hydrophobic area of the lipid bilayer
1. Channels
- Donut-like pore spans bilayer
- Solutes flow through RAPIDLY (diffusion)
- Rate of transport is not saturable
- Can be gated: open and close in response to stimuli
- Many types, highly selective
- Na+ channels, K+ channels, Cl- channels, H2O channels
Aquaporins are water channels
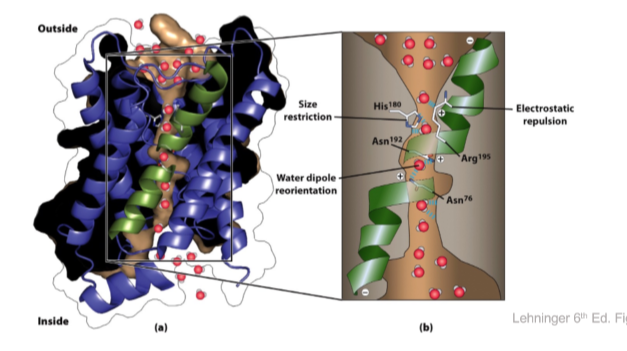
- Purple and black = aquaporin
- Green helices only go halfway
- Size Restriction = HIS 180
- His 180 creates an opening to the pore that restricts what molecule can enter based on size
- Electronic repulsion (ARG 195)
- protons present in the water can only associate with H30+
- Change in pH in the inside of the cell moves protons from the inside to the outside
- Repulses H3O, only letting H2O enter
- Water dipole reorientation (ASN 192, ASN 76)
- when liquid, water is H2O, sometimes these protons dissociate and jump from water to water
- 2 individual H+ are exchanged with other waters to make it through
- 2 ASN residues hold water in specific orientation so that it cannot pass protons
Ion Channels
- Difference between channels and transporters
- Rate of flux
- Saturability
- Channels are gated
- Ion channels
- Present in plasma membrane of all cells
- With ion pumps, define the permeability of membrane to ions
- Rapid movement of ions across membranes (10 to the negative 7/8 ions per channel molecule)
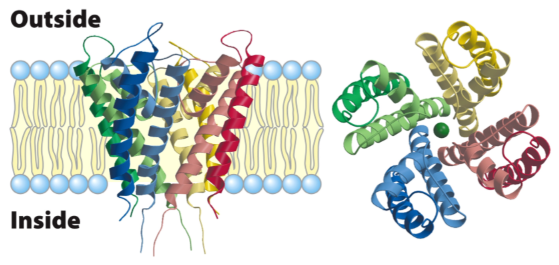
Gated K channel
tetramer: each subunit contains 2 transmembrane helices, a shorter helix and a selectivity filter connecting the short helix to one of the long filters
Outer helices in each subunit interact with a bilayer
inner helices in each subunit contribute to the inner pore
10,000 fold more selective for K+ v.s. Na+
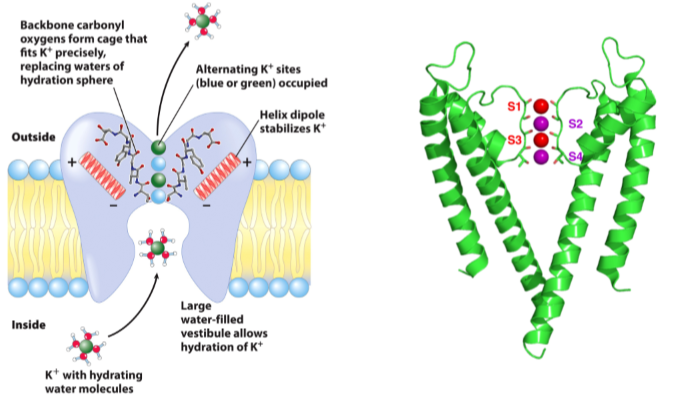
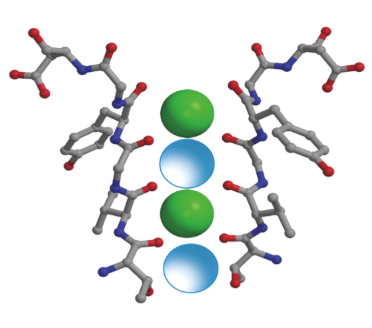
Selectivity
Size
Partial negative charges on C=O Gly-Tyr-Gly-Val-Thr
- Carbonyl oxygen coordinate with unhydrated K+
Gating of ion channels
This means that
- By default, they are closed, and let nothing past
- They open in response to a specific stimulus
- An inbuilt timer closes them again after a short delay, even if the stimulus is still present
The type of stimulus needed to open divides channels into:
i. Ligand-gated
- acetylcholine receptor ion channel
ii. voltage-gated
respond to changes in the voltage potential across the membrane
Voltage-gated Na+ channel organization
- Na+ channels differ from K+ channels predominantly in having a narrower specificity pore (Na+ is smaller)
- Alpha helix 6 is the pore-forming helix, 5 faces the membrane
- The voltage gating mechanism requires the addition of four additional alpha helices (1-4)
- The four separate chains are fused into a single polypeptide, serving as domains (I-IV)
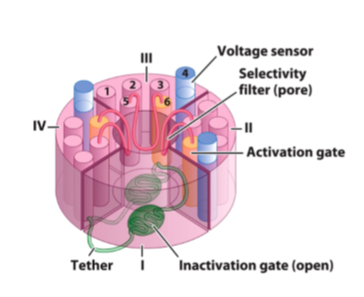
Voltage-gated Na+
the four pore-forming helices are arranged around the pore
the voltage sensing helix 4 (blue) can move in response to changing membrane voltage
the pore lining helix (6) is also called the activation gate
the inactivation gate is a small soluble domain (green) that connects domains III and IV
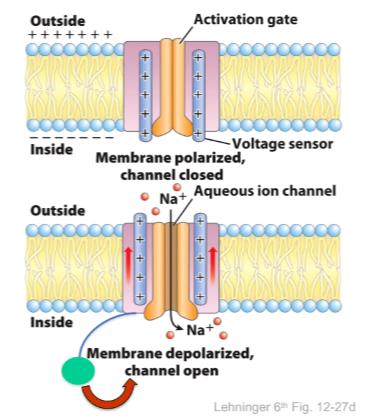
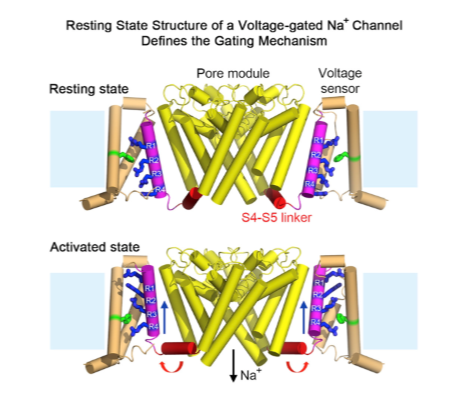
Opening the voltage gated Na+ channel
Helix 4 has a high net positive charge, and is sensitive to membrane voltage
The net negative charge inside the cell puts it inward
When membrane is depolarized helix 4 relaxes, and moves towards outside
Coupled movements in helix 6 (lining the pore) opens the channel
After opening, channel is quickly blocked by the inactivation loop, stopping ions from passing
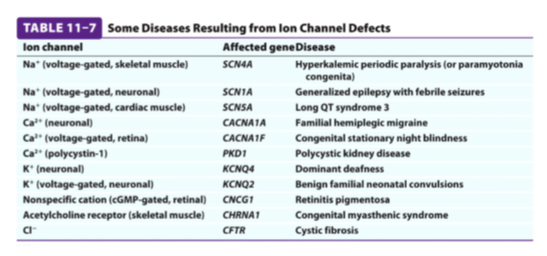
Ion Channels in disease and toxicity
Na+ channel in muscle
- channel defects result in diseases where muscles are paralyzed or stiff
- Many toxins target ion channels as the effects are fast acting and very debilitating
Na+ channel in neurons
- tetrodotoxin produced from puffer fish (fugu) binds to Na+ channels of neurons
2. Transporters
Membrane transporter classification
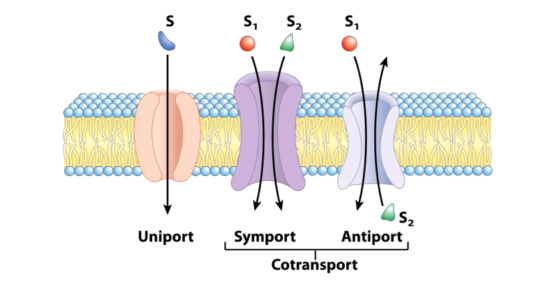
- Uniporters = transports one type of solute
- Cotransporters = transports more than one type of solute
- Symporters = transport in the same direction
- Antiporters = transport one solute in one direction and another in the opposite direction
- No statement is made about orientation in the membrane, focus on where solution is moving (no mention about passive or active transport)
Passive transporters
Transport Down a concentration (facilitated diffusion)
Highly selective (stereospecific)
Not a continuous pore through membrane, conformational change occurs to transport solute across
Transport one set of molecule(s) at a time (consistent number)
Rate of transport is regulated
- saturable number of binding site(s) for substrate
Passive Glucose Transporters
- GLUT1 in erythrocytes imports glucose
- GLUT2 in liver and intestine exports glucose
Passive transporter operation - GLUT1
- Substrate binds on one side of membrane
- Conformational change takes place
- Site opens on other side of membrane and substrate is released
- Conformational change takes place
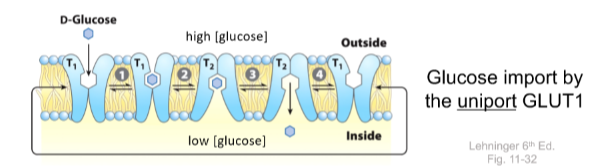
GLUT1 = Uniporter, only glucose and based on the concentration gradient, not in any one direction
Active Transporters
- Transport against a concentration gradient
- Transporters often called pump
- Many are powered by ATP hydrolysis - ion pumping ATPase
- Generate ion gradients across membranes
Ion gradients across membranes
- The ionic composition of the cytosol is different from the extracellular environment
- There are ion concentration gradients
- across the plasma membrane
- across organelle membranes
- These ion gradients are actively maintained by the cell, at the cost of ATP
- These ion gradients can do work
Charged solute across a permeable membrane
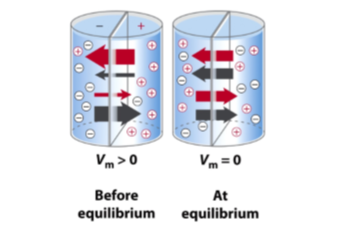
- Some ions move in the opposite direction, but net flow will be negative to the right and positive to the left
- Electrochemical gradient
Membrane potential
when a charged molecule is moved across membrane it results in a CHARGE IMBALANCE
Membrane potential (in volts)
- Typical plasma membrane (Delta Y = -60mV) negative inside

Free energy of charged species is different on each side of membrane

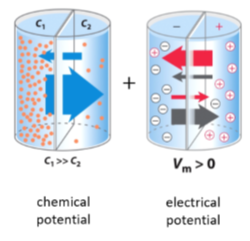
Movement of a charged solute across a permeable membrane
For a charged solute, the energy of moving the solute in the chemical and electrical gradient is ADDITIVE
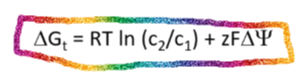
Maintaining Ion Gradients = Na+ K+ ATP
generates gradients of Na+ and K+
moves three Na+ ions out and two K+ ions in
This results in a net negative charge inside the cell
Hydrolysis of one ATP provides energy
Both ions move up their respective concentration gradients
ATPase uses 1/4 of your ATP when at rest
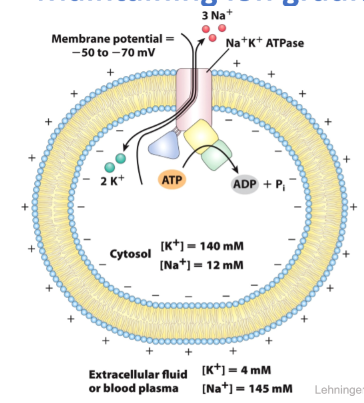
Na+ K+ ATPase: an ion-pumping ATPase
- Generates gradients of Na+ and K+
- Tetramer (alpha2, beta2); alpha performs transport
- Functions:
- Control cell volume
- drive active transport of other species
- render nerve cells electrically excitable
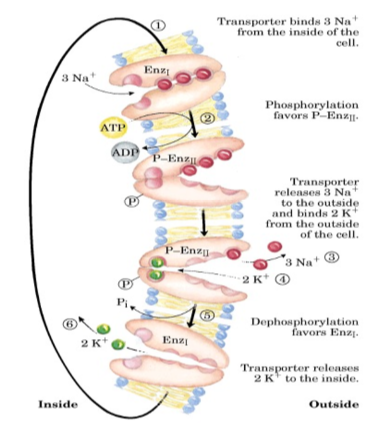
Na+ K+ ATPase transport cycle
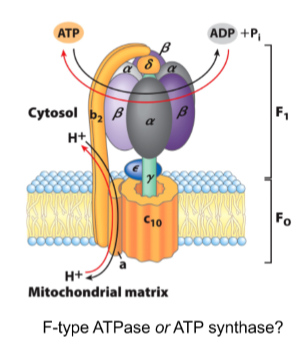
ATP is generated by reversing F-type ATPases
protons can be pumped against the concentration gradient at the expense of ATP
Alternatively, ATP can be generated at the expense of protons flowing down their concentration gradient
Ion Gradients can power transport
Ion gradients serve the cell as reservoir of available free energy
Transport of one ion down its concentration gradient can drive transport of another solute up its gradient
This is secondary active transport
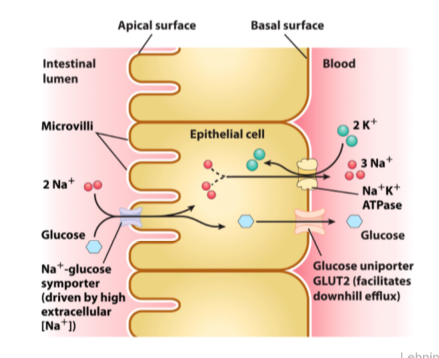
Na+ glucose transporter
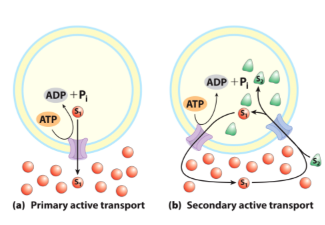
Na+ Glucose Symporter