Chapter 3: Proteins
Proteins:
- Proteins constitute most of the cell’s dry mass.
- Cell’s building blocks and also execute the majority of cell’s functions.
- Heteropolymer of amino acids.
- After water, proteins are the major components of protoplasm.
- Peptide bond is present.
- Most abundant protein on earth: Rubisco
- Most abundant protein in mammals: Collagen
- Proteins embedded in the plasma membrane form channels and pumps that control the passage of small molecules into and out of the cell.
- Proteins from a chemical point of view are very complex and functionally sophisticated molecules.
- The location of each amino acid in the long string of amino acids that forms a protein determines its three-dimensional shape.
Shape and Structure of Proteins:
- 20 different amino acids.
- A protein molecule is a long un-branched chain of these amino acids.
- Proteins are called polypeptides.
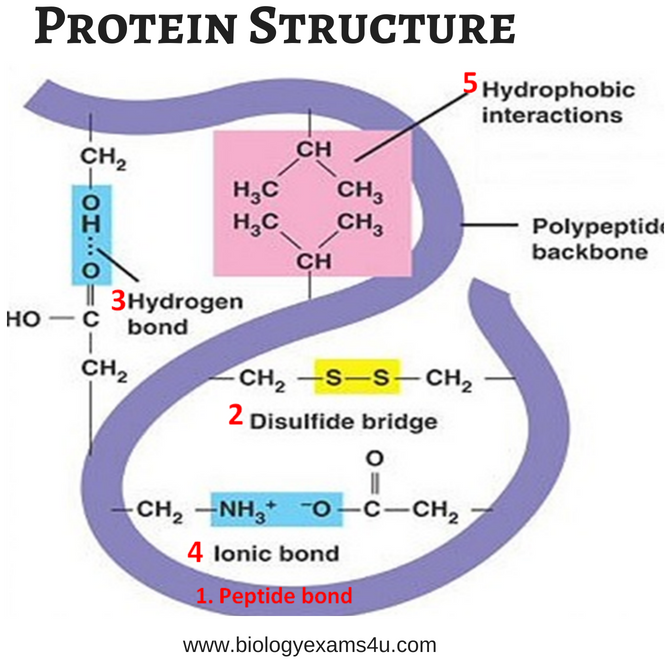
- It consists of:
* Peptide bond
* Disulphide bridges
* Hydrogen bond
* Ionic bond
* Hydrophobic interactions
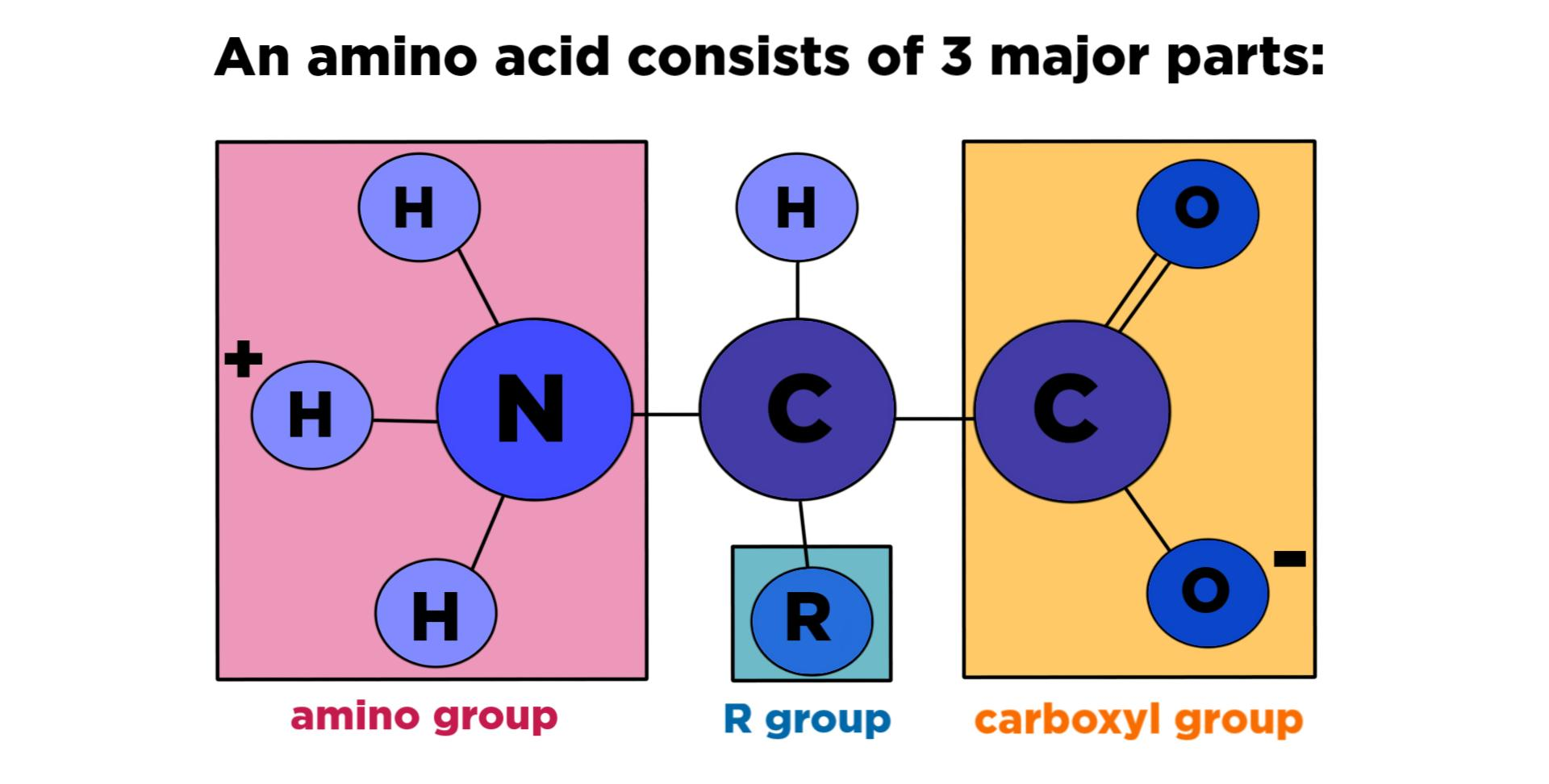
Amino Acids:
- Micro molecules/acid soluble pool.
- Monomer of protein/building of protein.
- Substitute of methane.
- Amino acids consist of:
* Amide group: basic group, positively charged.
* R: variable group, decide name, nature, and properties of amino acid.
* COOH: carboxylic acid, acidic group, acidic nature, negatively charged.
* C: chiral carbon or alpha carbon.
Properties of Amino Acids:
- Configuration of protein.
- Amino acids are amphoteric in nature.
- All amino acids are officially active, and they show optical isomerism-except glycine.
- Zwitter ions: Dipolar ions
* at low ph (acidic) = positive charge
* at high ph (basic) = negative charge
Classification of L-alpha amino acid:
- Acidic amino acid:
* It contains an extra COOH group.
* Aspartic acid, Glutamic acid. - Basic amino acid:
* It contain extra NH2 group.
* Histidine, Lysine, Arginine. - Neutral amino acid:
* It contains one NH2 group and one COOH group.
* Asparagine, serine, tyrosine, etc.
Classification of amino acids on the basis of functional group
- Amino acid with aliphatic group: GAVIL
* Glycine, Alanine, Valine, Isoleucine, Leucine - Amino acids containing hydroxyl (-OH) groups: ST
* Serine, Threonine - Sulphur containing amino acids: CM
* Cysteine, Methionine - Acidic amino group: AAGG
* Aspartic acid, Asparagine, Glutamic acid, Glutamine - Basic: LAH
* Leucine, Arginine, Histidine - Aromatic: PTT
* Phenylalanine, Tryptophan, Tyrosine - Imino: Proline
- Non-polar amino acids: They have no charge on the “R-group”.
- Polar amino acids: Have charge on the “R-group”.

Classification of amino acids (on the basis of synthesis in the body)
- Essential amino acid:
* Not synthesized in our bodies.
* Need to be taken in our diets. - Non-essential amino acids:
* Synthesized in their body cannot be taken in diet. - Semi-essential amino acids:
* Produced at a very slow rate can be synthesized by the adult body but not in growing children.
Proteins classified on the basis of chemical nature and stability:
- Simple protein:
* Made up of amino acids.
* Protein part:
* Globular: spherical/oval shaped.
* Fibrous: Collagen, Kinetin, Actin - Conjugate protein:
* Made up of protein + nonprotein part. - Derived protein:
* Primary: Due to denaturation of protein.
* Secondary: formed due to digestion. - Protein are also divided as:
* Complete protein: All 20 essential amino acids present.
* Incomplete protein: One/two essential amino acids lacking. - Monomeric protein: Made up of one polypeptide chain.
- Oligomeric protein: Made up of two/more polypeptide.
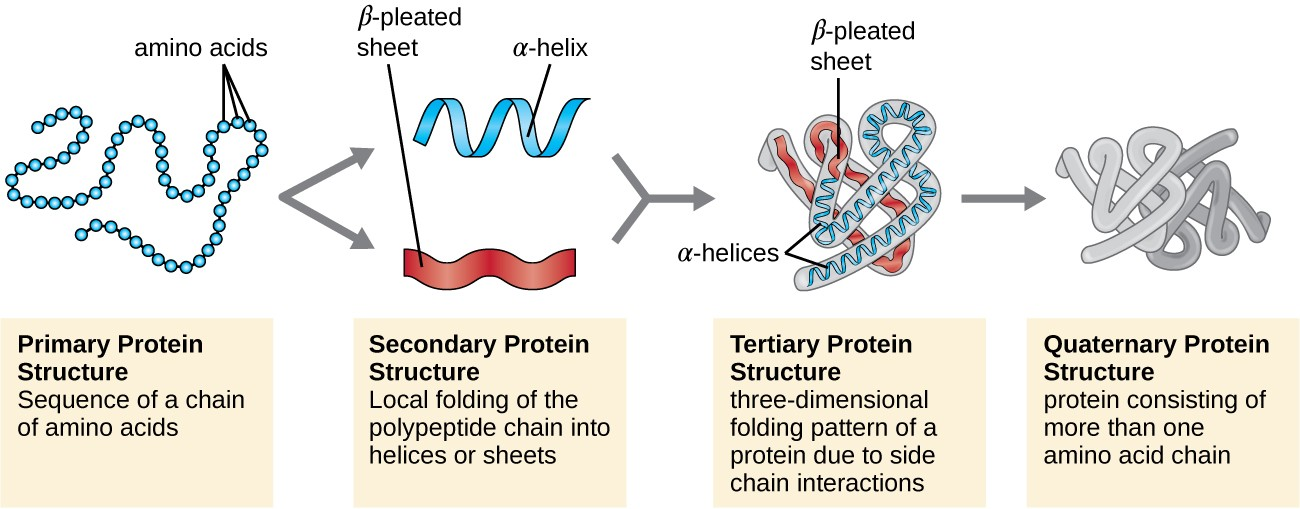
Structure of protein
- Primary structure:
* It is a linear chain of amino acids linked by peptide bonds.
* It is a newly formed protein on the ribosome.
* This structure of a protein is highly unstable/not functional but decides the fate of protein. - Secondary structure:
* It comprises of alpha helix and beta plated sheet.
* The folding of linear polypeptide chains in a specific coiled structure is called secondary structure.
* A new bond is formed: Hydrogen bond.
* 2 bonds: hydrogen + peptide
* Alpha helix:
* a most common type of secondary structure and rigid rearrangement of polypeptide chain.
* stable configuration.
* right-handed helix.
* bonds: intramolecular h-bonding, peptide bond.
* Beta-plated sheet:
* made up of 2 or more polypeptide chains are held together by intermolecular-H bonding.
* zig-zag shape.
* protein of secondary structure insoluble in water and fibrous in nature. - Tertiary structure:
* protein of tertiary structure are highly folded and globular in nature.
* soluble in water.
* more folded than secondary.
* bonds:
* peptide bond
* H-bond
* disulfide bond
* hydrophobic interactions
* ionic bond
* most of the proteins and enzymes show tertiary structure in protoplasm. - Quaternary structure:
* it is made up of two or more than two polypeptide chain.
* oligomeric protein in which R-group close to each other.
* all types of bonds like intra, inter-H bonding, ionic bonding, covalent bond, hydrophobic interactions etc, are formed.
* these protein play important/significant role in the regulation of metabolism and cellular function.
ENZYMES
- Enzymes enhance the rate of biological chemical reaction by lowering down activation energy.
- It is a biological catalyst.
- Enzymes are biological middlemen.
- All enzymes are proteinaceous except ribozyme and ribonuclease.
- Enzymes show tertiary and quarternary structure and very specific for biological activity.
- Maximum enzymes are found in mitochondria.
- Small enzyme: Peroxidase.
- Largest enzyme: Catalase
Characteristics features of enzymes:
- Enzymes do not disturb reaction equilibrium.
- Turn over (The number of substrate molecules transformed per min/per sec by one enzyme molecules)
- Turn over no. depends on:
* number of active sites of an enzyme.
* fastest reaction
* separation of product. - Active site catalytic is directly proportional to turn over number.
- Maximum turn-over number: Carbonic anhydrase.
- Minimum turn-over: lysozyme
- Reversibility in nature:
* Substrate + Enzyme → ES complex - Very specific in nature:
* temperature specific:
* high temperature: denaturation
* low temperature: inactivation
* ph specific - Molecular weight is high.
- Amphoteric in nature.
Nomenclature and Classification of Enzymes
- Nomenclature: suffix= ase
- Source of extraction: from where it is extracted.
- 6 classes of enzymes:
* OTHLiL
* Oxidoreductase:
* enzymes involved in oxidation-reduction reaction.
* alcohol dehydrogenase, cytochrome oxidase.
* Transferase:
* Enzyme that catalyze reactions the transfer of functional group.
* e.g.: hexokinase, trans-aminase.
* Hydrolase:
* Enzyme catalyzing hydrolysis of ester, ether, peptides etc.
* These enzyme breaks large molecules into smaller molecules by the introduction/presence of H2O molecules.
* Lyases:
* They break specific covalent bonds and remove a group without hydrolysis, oxidation etc.
* e.g. Aldolase, fumarase.
* Isomerase:
* Rearrangement of molecular structure to form isomers.
* Ligases:
* Enzyme catalysing the synthetic reaction where two molecules are joined together.
Types of Enzymes:
- Simple enzyme: consist of only proteins and catalyze their substrate specific reactions.
- Conjugate enzyme/Holo enzyme: Made up of protein and non-protein parts.
* Protein part: Apoenzyme
* Non-protein part: Co-factor
* Organic:
* Coenzyme: A coenzyme is a loosely bound/organic co-factor. It can be easily removed.
* Prosthetic group: A prosthetic group is tightly bound organic co-factor.
* Inorganic: They form coordination bond with side-chain at the active site and the same time for one/more coordination bond with substrate.
Mode of enzyme action
Mostly enzymes are protein in nature.
The hypothesis regarding the mode of enzyme action
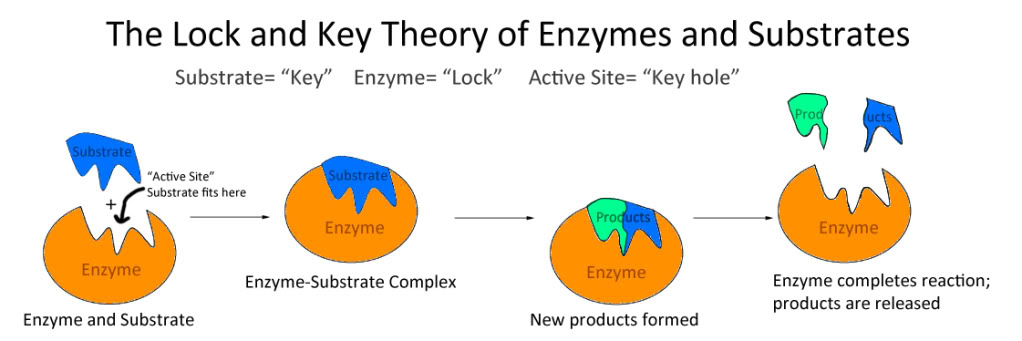
- Lock and Key Hypothesis:
* According to this theory:
* Enzymes are rigid and pre-shaped.
* Substrate fit to the active site just as a key fit into a proper lock.
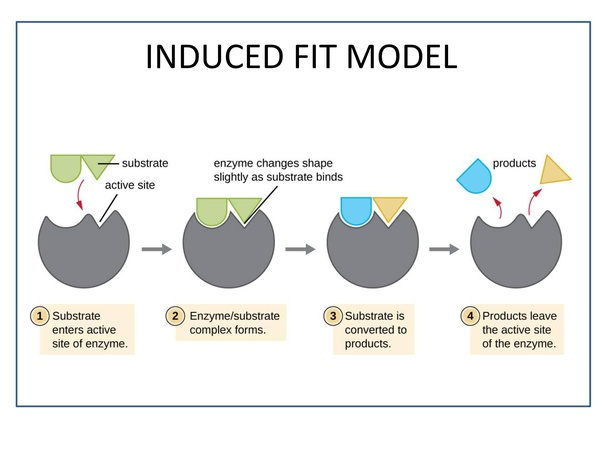
- Induced fit hypothesis/ theory:
* Proposed by Kosh land.The monomer
* Most accepted hypothesis on the basis of enzyme action.
* Enzymes are not rigid and pre-shaped.
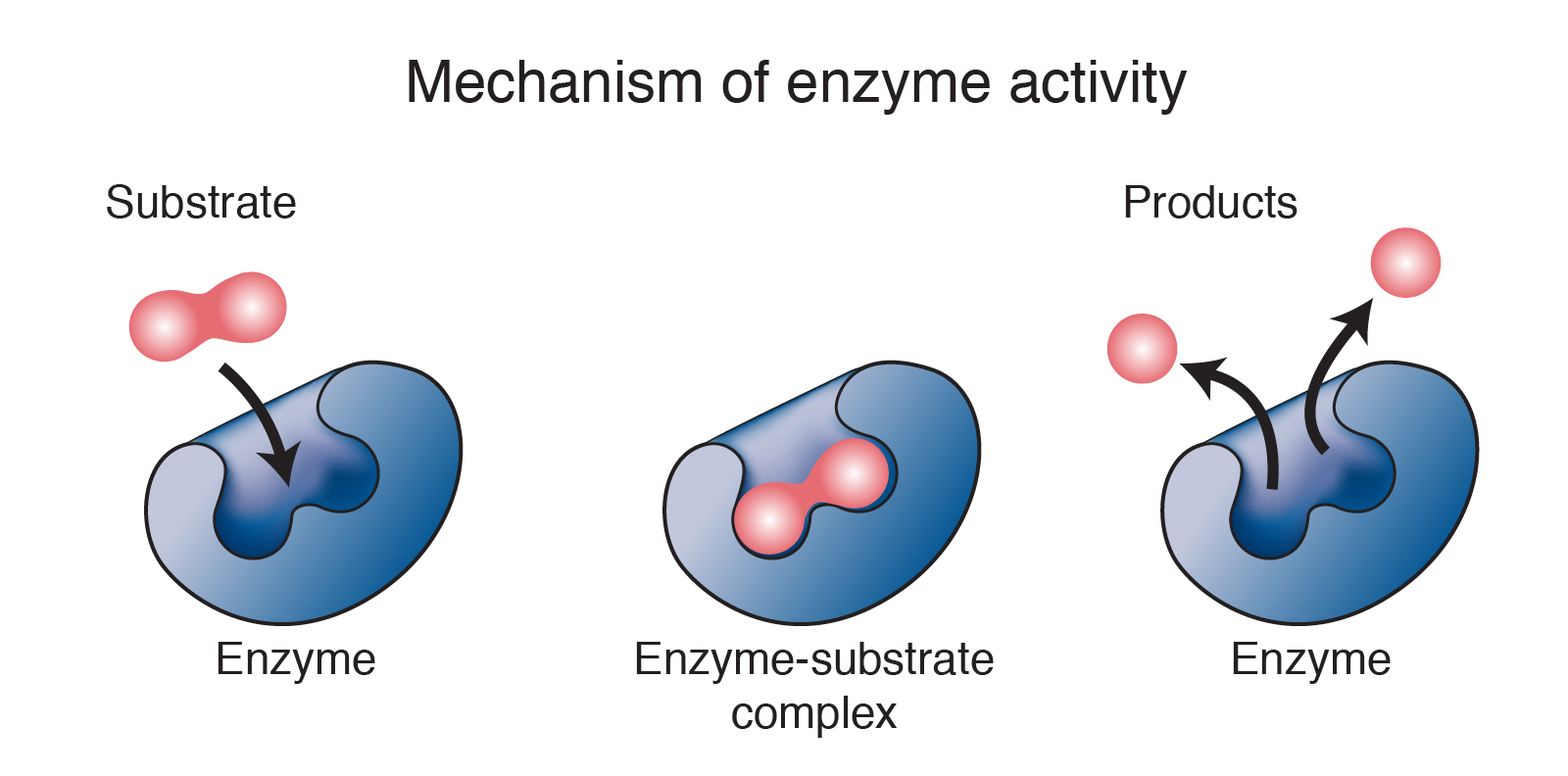
Mechanism of enzyme action:
- Substrate → Product
- Lowering down of activation energy.
- Do not alter the equilibrium.
- Enzymes are biocatalyst.
Factors affecting enzyme action:
- Temperature:
* at high temperature: denaturation
* at low temperature: inactivation
* optimum temperature: 25-40 degrees Celsius for enzymatic activity. - pH:
* optimum pH = enzyme activity very high.
* enzymes:
* endoenzyme (inside cell)
* exoenzyme (enzymes are synthesized inside in the cell but secreted from the cell to work externally). - Substrate concentration:
* Enzyme is larger in size and bears several active sites with the increase in substrate concentration the velocity of the reaction rises in first and the reaction reaches a maximum velocity. (Vmax)
* The velocity is not exceeded by any further rise in the concentration of substrate.
* Michalis Menten Constant (Km):
* It is a mathematical derivation/constant which indicate concentration of substrate at which reaction velocity reaches half of Vmax.
* Km indicate affinity of the enzyme for its substrate.
* A high Km indicate low affinity of enzyme and low Km indicate high affinity.
* Km is inversely proportional to turn over number.
* Allosteric enzymes do not obey Km.
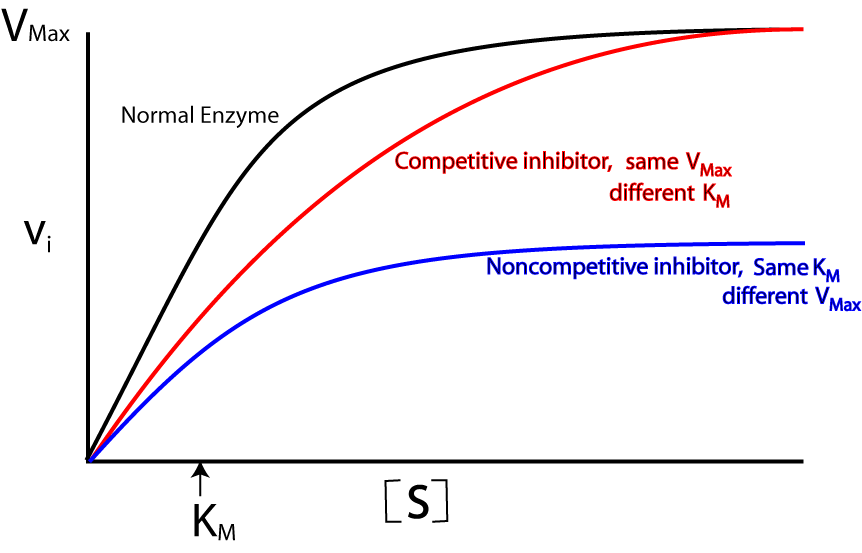
Inhibitors:
- It is chemical molecules inhibit enzyme activity.
- Inhibitors are of two types:
* Competitive inhibitors:
* Inhibitors are structure similar to substrate.
* They favor lock and key hypothesis.
* Reversible in nature.
* Km increase but Vmax remain constant. - Non-competitive inhibitors:
* Some inhibitors do not compete for active site of enzyme but destroy the structure of enzyme, the physical structure of enzyme is altered as a result and do not form enzyme-substrate complex.
* They favor induced-fit theory.
* Irreversible in nature.
* Km remain constant but Vmax change.