BIOB10 - Cell Biology - Midterm
Table of Contents
Cells & Genomes
Protein Structure
Protein Transport
- Endoplasmic Reticulum
- Mitochondria
Enzymes
Intracellular Membrane Traffic
Golgi Apparatus
Endocytic Pathway
Cells & Genomes
Universal Features of the Cell
==DNA - Deoxyribonucleic acid==
- Stored information in the double helix
- Nucleotide made up of a sugar phosphate (sugar + phosphate) and a base (A, C, T, G)
- Sugar phosphate back bone. Bases connect, not actually stuck together.
==DNA replication==
Making a complimentary strand from the template strand.
==Templated polymerization==: using another strand to make another strand. using a template to make a complimentary strand.
==Hydrolysis== aids (addition of water)
- GTP (three phosphates) → high energy bonds break → GDP (two phosphates) + energy
- ==Nucleotide==: nucleoside with extra one or more phosphate group attached (ex. monoguanosine)
- ==Nucleoside==: the sugar and the backbone (ex. guanosine)
Following hydrolysis, the monophosphate attached (monomer)
==Transcription==
- Cell needs to make sense of information
- Take one strand, strand is used as template to form RNA strand
==Translation==
- “What amino acid do I add?”
- Create a protein basically
==Plasma membrane==
- Cell is closed in
- Selective barrier → how things move in and out, semi-permeable
- Amphiphilic → hydrophobic tails face inward, hydrophilic heads face towards water.
Prokaryotes and Eukaryotes
Prokaryotes
- Spherical or log shape
- Way smaller
- Cell wall so it is more rigid
- No nice compartments
==Eukaryotes==
- House genetic information in the nucleus (double membrane)
- Bigger genome, difference in cell structures
- Plant cells are eukaryotes (have specific organelles)
Gene expression
==Transcription==
- DNA → RNA
- Only using one strand to make RNA (template strand)
- Complimentary base pairings
- In RNA T does not exist, it’s U
- Read 1 at a time
- RNA strands called a transcript, use for protein
==Translation==
- RNA → Protein
- Read three nucleotides at a time (codon)
- 1 codon : 1 amino acid
- Ribosome does the readings
Macromolecules
Subunits / monomers - Amino acids, nucleotides
Macromolecules / polymers - Proteins, nucleic acid
Monomers do not float around loose that often, not as abundant as polymers.
==Macromolecules== fold into an energetically favourable shape/confirmation, whatever makes the most sense.
Protein Structure
- Gene expression → same throughout whole transcription process, same between all eukaryotes
- Proteins can be enzymes
- ==Enzymes==: proteins that facilitate certain reactions
- DNA Polymerase: hydrolysis happens, taking that nucleotide away.

- Exonuclease - cutting from end
- Endonuclease - snipping in the middle
- Proteins can be used to build stuff in the cell
- Build structural components
- Act as a molecular motor
- A motor protein - do the moving in cells
Protein structure
- ==Polypeptide chain==
- multiple amino acids connected by peptide bonds
- Polypeptide backbone - each amino acid has a part that forms the polypeptide backbone, where peptide bonds occur
- Side chain - parts where amino acids stick out
- N-terminus is front end, C-terminus is back end
- N-terminus is a free amino group
- C-terminus is a free carboxyl group
==Hydrophobic clustering==
Dropping protein into an aqueous watery environment causes protein to cluster together to protect parts from water. Non-polar parts are hydrophobic, and are hydrophobic clustering.
- Hydrogen bonds: a bunch of weak attractive bonds with H and something else (like Oxygen)
Protein folding
Protein tries to be in lowest energy conformation (happy state)
Proteins can denature or renature
- ==Denature:== treat proteins with a solvent to mess up bonds (unfolds)
- ==Renatures==: remove solvent and protein folds again
==Molecular chaperones:== Proteins that help proteins fold properly. Folding helpers.
- Proteins can fold based on interactions with side chains
- There are exposed hydrophobic parts that are meant to be folded inward
- ==Off-pathway conformations==: exposed hydrophobic regions can result in off-pathway conformations. Desperately going around and binding to stuff and making a mess.
- Final 3D shape determined by specific amino acid sequence. Chaperones just help form it.
==Common structures==
- Coiled-coil
- All hydrophobic regions racing each other, and they form this helix shape. Doesn’t want to interact with other stuff.
- Protein complex
- Protein subunits join together (clump looking thing)
- Dimer
- 2 polypeptide chains bind to each other in a head-to-head arrangement forming a symmetric complex.
- Held together by interaction between two identical binding sites
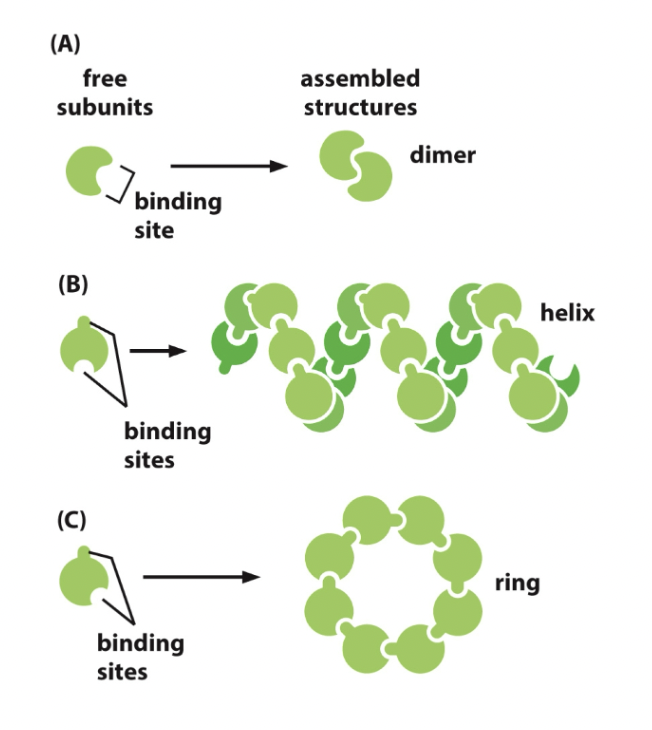
Binding site: Any region of protein surface that interacts with or binds with another molecule
==Proteins can be==
- Fibrous
- Long proteins, coiled-coil. double or triple helix.
- Globular protein
- Big globby thing, DNA polymerase
- Disorder in regions
- Nuclear pores
- Gates have disordered regions that control what comes in and goes out.
Protein binding:
- Tight or weak
- Have specificity
==Ligand binding:== referring to something that is doing the bonding (like enzymes, often first step of what an enzyme does)
- Fits perfectly with binding site
- ==Ligand:== surface contours of the molecule fit very closely to the protein.
- Folds help with forming the ligand.
==Protein binding interfaces==
- Surface-string
- Helix-helix
- Surface-surface
Protein Transport
Three main means of transportation
==Gated transport==
- Pores where things can go in and out. Referring to nuclear pore complex
- Between cytosol and nucleus
==Protein translocation==
- Proteins need to get to a destination. Transmembrane proteins.
- The protein uses transmembrane protein to snake through.
==Vesicular transport==
- Take a vesicle that transports from one place to another
==Signal sequences==: Sorting signal. Sequences on the N-terminus or in the middle of the polypeptide chain.
- I==n Endoplasmic reticulum== - signal is on N-terminal
- Protein targeting - using signal sequences to get protein, some proteins have no signal sequence
- ==Cytosolic protein== - no signal sequence, just in the cytoplasm
Endoplasmic reticulum
- RNA transcript leaving nucleus to go get ribosomes in cytoplasm
- Ribosome reads three at a time (codon) and builds protein chain.
- ER sequence is coded and tells it to go to endoplasmic reticulum.
- Ribosome sits on ER and finishes making protein (rough endoplasmic reticulum forms)
==Co-translational translocation==
- Cells move proteins into the ER before completion of polypeptide synthesis
- ER sequence shows up and protein synthesis continues on ER as chain gets snaked into ER.
- ==Post-translational translocation== - finishes making protein and then goes to where it needs to be. Translocation after translation.
==Signal recognition==
- ==ER signal sequence==: Guided to the ER by two main parts
- ==Signal-recognition particle (SRP)==
- Small ribosomal subunit and large ribosomal subunit
- Little SRP binds to the ribosomal subunits on the signal sequence of growing polypeptide.
- ==SRP receptor==
- In rough ER. SRP attaches to it.
- Allows ribosome growing polypeptide to make it’s mark.
Process
SRP binds to ER signal sequence
SRP bends and blocks elongation factor site (pauses translation)
SRP receptor binds SRP and it moves whole junction of ribosome and SRP down to do so.
SRP receptor bonding.
Protein elongates through protein translocator, while ribosome is released.
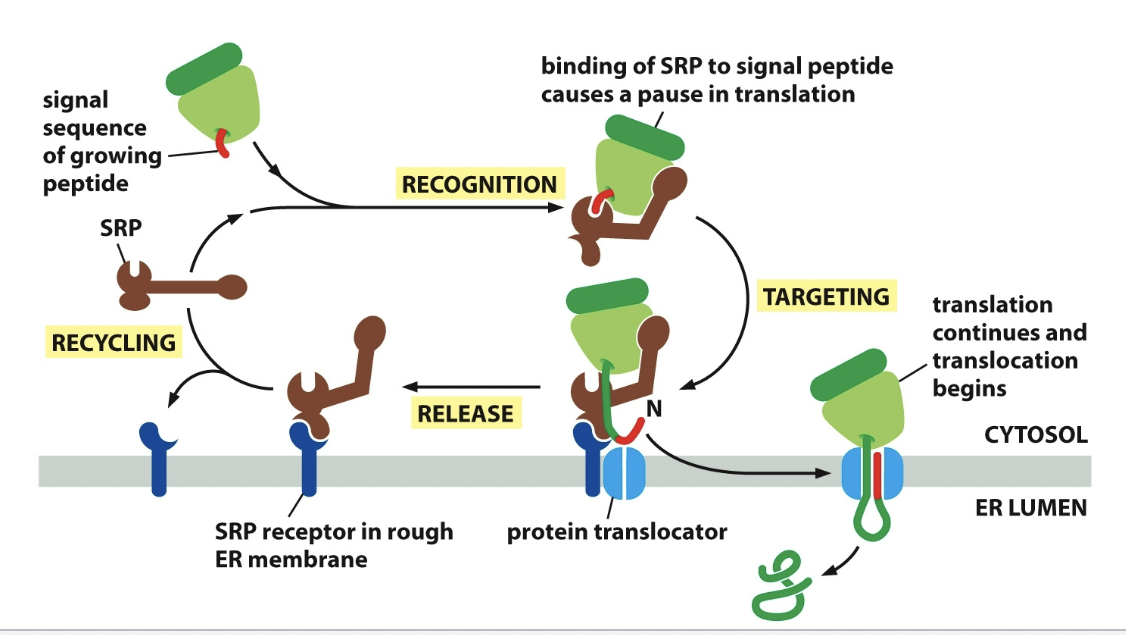
ER signal sequence - ER captures 2 types of proteins from cytosol
- Transmembrane protein
- Water soluble protein (in the lumen)
Why go to the ER?
- Some protein work in the ER so they go there
- It uses ER as a pitstop
- ==Secretory pathway==: where stuff is going to get closer and closer to exterior of cell
Ribosomes
- ==Membrane bound ribosomes==
- Physically on ER, doing translation on the ER.
- Free ribosomes
- Polyribosomes
- ==Free polyribosomes==
- Transcript with 5’ and 3’ end (read from 5’)
- Ribosome keeps reading and moving down
- Second and third (etc.) don’t need to wait till the first one is done, they’ll just go when there’s enough room.
- Once done, ribosome disassembles into individual subunits
- ==Membrane-bound polyribosomes==
- ER signal sequence, SRP bounding to ER membrane like from earlier example
- It continues elongating as protein snakes through
- As soon as there is space, one comes and binds and does the same, attaching next to first ribosome.
- Only initial ribosome needs to read it before it attaches.
Translocator (seam and plug)
==Sec61 complex== (core of translocator)
- Has a plug, when it recognizes the sequence, the plug moves aside and allows protein to enter.
- Signal peptide is hydrophobic and cannot snake through.
==Accessory proteins==
- Work with translocator to help with the process
- ==BiP== - binding protein in the ER
- ATP to ADP through hydrolysis. Happens in order for BiP to pull the protein through.
==Translocon==
- Translocator + accessory proteins
==Targeting== (part of SRP cycle)
- ==Start-transfer sequence== - it is recognized by ribosome, it opens up the translocator and enters.
- Signal sequence doesn’t need to stay and can be cut off
- Signal peptidase cuts it off so things can snake in until stop transfer.
- ==Stop transfer== - get out and the little protein is embedded in the membrane
- Transmembrane protein means you’re in the membrane
==Single-pass transmembrane protein== (in the middle)
- Goes in too, doesn’t get cleave off, serves as the anchor
- With or without stop transfer sequence
- It is stuck in membrane
==Multi-pass transmembrane protein==
- Rest of protein snakes in, no stop transfer, just goes straight through and it’s done.
- Double pass if there is a stop transfer.
==ER lumen proteins==
- If they wanna be right inside the ER
- One signal sequence, cleaved off by signal peptidase.
- Snakes right through.
==ER resident proteins==
- BiP is an ER resident protein cuz it stays in the ER and works in there
- Have ER retention signal
==Protein quality control==
- ER must make sure proteins are folded correctly
- ==Oligosaccharides== added to proteins
- Add 3 sugars to proteins - protein glycosylation (the process) - adding the oligosaccharide chain to the protein at the ER.
- Oligosaccharyl transferase (glucosyl transferase) catalyzes
- ==ER chaperone== proteins help proteins fold properly
- Lectins (calnexin and calreticulum)
- Bind to oligosaccharides on incompletely folded proteins and retain them in ER
- ==Oligosaccharide processing==
- Adding 3 glucose to unfold protein. Glucosidase trim these off one at a time
Process
- Glucose trimming
- Protein now with one glucose left
- Calnexin has high affinity for this glucose (they bind)
- Glucosidase trims glucose and thus releases protein from calnexin
- Tries to fold into shape it is supposed to be. Checked by glucosyl transferase.
- Fold correctly → it can leave ER
- Fold incorrectly → glucosyl transferase adds a glucose right back on, goes to calnexin and redoes process. (may only do it once or twice, otherwise it’ll destroy it)
==Retrotranslocation==
- If protein folding process doesn’t work out second time, it’ll normally just get cut.
- Retrotranslocation is the cutting process
- ==Mannosidase== removes a mannose on the core oligosaccharide tree
- Not gonna get glucose added back
- Mannose is removed
- Protein without mannose is gonna be destroyed
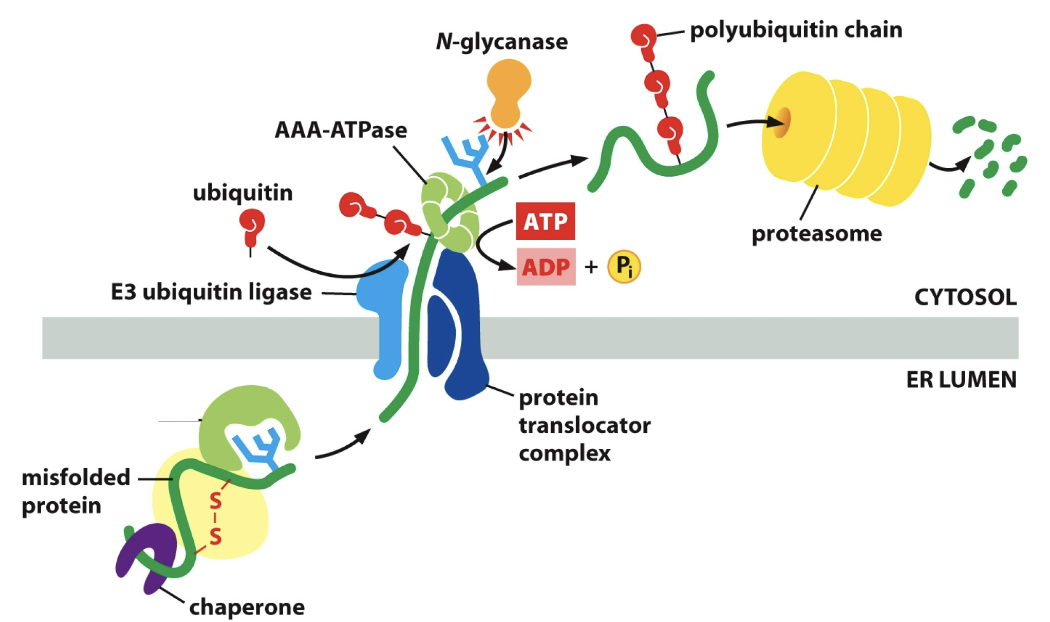
- Misfolded ones sent through special translocator
- ==E3 ubiquitin== (enzyme) stick onto the protein and protein translocator complex. Stamps it to be destroyed.
- Protein in cytosol has polyubiquitin chain
- Goes through proteasome and gets cut up. Hydrolase cuts proteins
Parts of retrotranslocation
==AAA-ATPase:== like BiP, helps pull protein out. Does this by hydrolyzing ATP
==N-glycanase==: enzyme, still has the ubiquitin chain. It saves the chain.
==Heat-shock response==: wants more chaperones to be made, needs to go all the way back to the DNA to get that coding sequence.
- Misfolding with a lot of heat
- Stimulates transcription of genes encoding cytosolic chaperones that help refold the protein.
==Unfolded protein response==: making more proteins involved in retrotranslocation, protein degradation in the cytosol, and ER chaperones.
Mitochondria
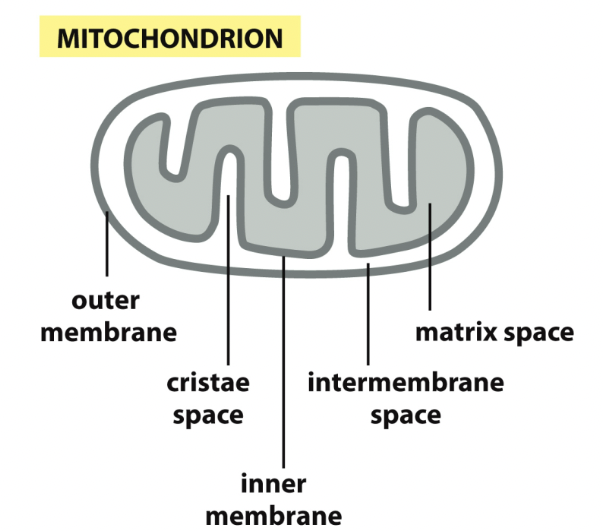
If you’re not an ER protein, you may have a sequence that states where you must go
- No pausing or finishing up in that area
- Post-translation translocation mechanism
Mitochondrial protein is made in cytoplasm and then ER surface helps direct it.
==Protein translocators==
- Doors/gates that allow protein to pass
- Mitochondrial
- ==TOM complex== - transfer proteins across the outer membrane
- Initial recognition
- ==TIM complex== - transfer proteins across inner membrane
- TIM22. TIM23.
- Continue journey down, help insert protein down
- ==SAM complex== - once protein enters TOM, SAM helps them fold properly
- ==OXA complex== - some proteins can be made in mitochondria and it does that
Protein folding
- Binding onto protein
- Gonna keep going
- Little signal sequence is cleaved off
- Exists in the matrix space
Import to matrix
- ==Cytosolic hsp70== - Interacting with protein and gonna keep it unfolded. Helps that interaction.
- Recognized by translocator - Cytosolic hsp70 comes off as it snakes through, this occurs with ATP hydrolysis
- Goes into TIM23 complex
- Interact with mitochondrial hsp70. Through ATP hydrolysis. Constant binding and releasing helps fish it through.
==Mitochondrial hsp60==
- Chaperone protein
- Interacts with protein, and once it comes through, hsp60 helps it fold through ATP hydrolysis
- hsp70 then helps it
Pore-forming beta-barrel proteins
- Comes in and it’s not folded, interacts with a bunch of chaperones
- Get back out, SAM helps it fold.
- Through TOM, to SAM
Import to inner mitochondrial membrane (either:)
- Transmembrane protein
- Talking about protein getting stuck in inner membrane
- Stop transfer sequence - there’s the cleaved part and then the stop that stops it from transferring to matrix any further.
- Matrix targeting signal sequence
- OXA complex helps it snake through
- Get’s cleaved in matrix.
- Confusion and OXA will help it and stick it in the inner membrane
==Chloroplast==
- Like mitochondria:
- Processes occur post-translationally
- Uses translocation complexes
- Requires energy (ATP hydrolysis)
- Unlike mitochondria
- Chloroplast uses TOC and TIC
- Added thylakoid
Process
- Recognize
- TOC helps it through
- TIC helps it through
- Cleavage of signal sequence
- “Go to thylakoid” signal sequence is now readable
- Goes to thylakoid space, and that signal sequence is cleaved off
Enzymes
Help carry out reactions. Bind to a substrate. Act as a catalyst. Make reactions happen.
Enzyme + Substrate → ES → EP → Enzyme + Product
- enzyme substrate complex and enzyme product complex
- More substrate, higher rate at which product forms, reaches max rate
- ==Turnover number==: where the reaction rate will max out
Intermediate state
- Transition state: middle state is in (unstable)
- Activation energy: energy required to get to unstable intermediate state
==Active site of an enzyme==
- When a substrate binds to an enzyme, bonds in the substrate are often distorted, changing substrate shape
- Changes drive a substrate towards a transition state
==Lysozyme==
- Enzyme that cut cell wall’s of bacteria, cell bursts.
- Needs to be bound by the enzyme and then does its business
- It can only do this IF the substrate is in its transition state
Enzyme-catalyzed reaction
- Product of one reaction can become substrate for next enzyme-catalyzed reaction
- Rxn rate limited by amount of substrate and enzyme
- Bunch of products together → multienzyme complexes. Easier for enzyme two to now start.
Cells regulate enzyme activity
- Regulating the expression of the gene that encodes the enzyme. Have more enzyme made.
- Confining enzymes and substrates to particular subcellular compartments
- Protein destruction; want to stop reaction
- Molecule binding to enzyme; control it’s activity
==Regulatory sites==: Small molecules bind to regulatory sites. Separate from active sites.
Allosteric enzymes: have two or more binding sites
==Active site==: recognize the substrate
Regulatory site: recognizes a regulatory molecule
- Change it to active site’s confirmation, change so it can or can’t bind
Binding changes
- Change active site to easily bind to a ligand, or so it no longer wants to bind to a ligand
==Positive regulation==
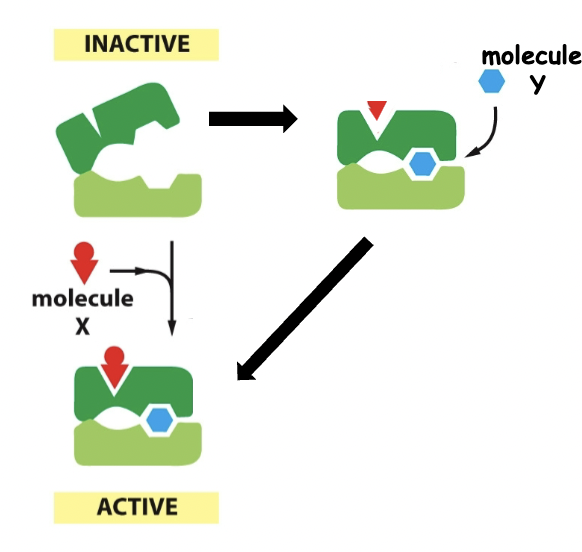
==Negative regulation==
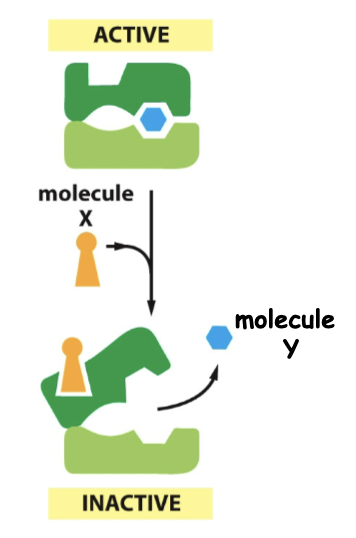
==Cooperative allosteric transitions==
Start: Enzymes in a closed mouth confirmation
New: Attached to another enzyme, subunit one and two, make up this enzyme. Multi subunits.
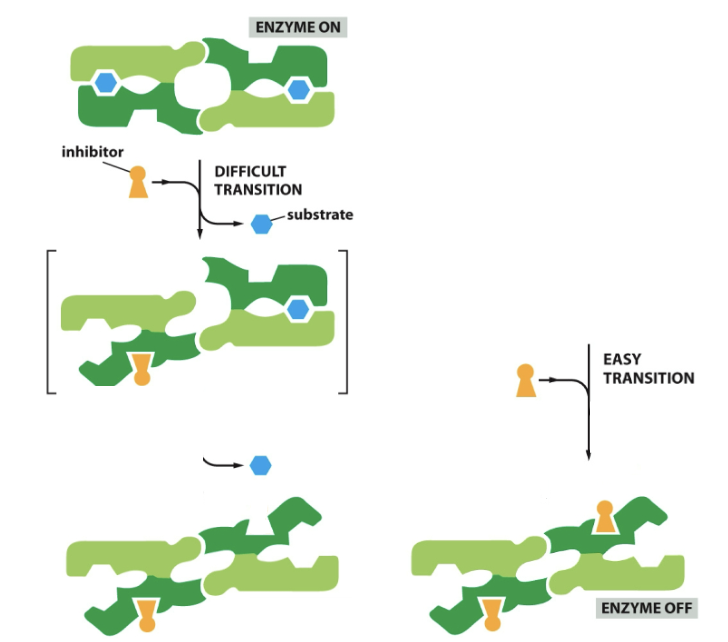
Intracellular Membrane Traffic
==Endocytosis==: pinching in of plasma membrane. Removes plasma membrane components and delivers them to internal compartments called endosomes.
- ==Endocytic pathway==: leads inward from plasma membrane
==Exocytosis==: Secretory pathway delivers new proteins to plasma membrane or extracellular space.
- ==Secretory pathway==: leads outward from ER towards Golgi and cell surface (or side route to lysosome)
Transport vesicles
==Coated vesicles==: cage of protein covering cytosolic surface, discard cote before fusing. Coat has 2 layers.
==Clathrin-coated== → Golgi to Plasma Membrane
Clathrin major protein component
3 heavy chains, 3 light chains (polypeptide) that form outerlayer.
- Called triskelyons
- Make coated pits or buds
- Clathrin cage - cage like assembly
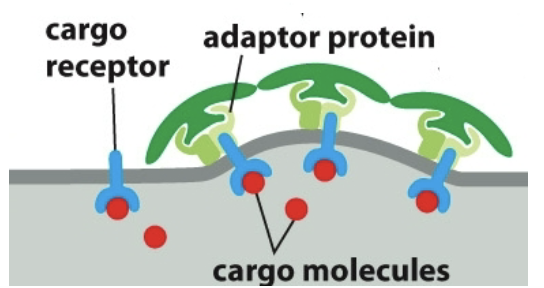
Adaptor proteins: bind to clathrin coat to the membrane and trap various transmembrane proteins. Bind to cargo receptors which bind to cargo molecules.
Adaptor proteins and cargo receptors bind with specificity
==PIPs== example: bind to AP2s and change adaptor proteins conformation.
- can become phosphorylated at different regions and then bind to APs and change them. Then it is possible to bind to receptors.
==COPI==-coated → Golgi to ER
==COPII==-coated → ER to Golgi
==Membrane bending proteins==: have crescent shaped domains (BAR domains), help bend membrane. Bind to and impose shape, help form circular bud.
- Dynamin: cytoplasmic protein, tethers to the membrane. Non cytosolic parts of the membrane, help squeeze together and bud off. It calls over other protein to neck of the bud.
Uncoating membrane
==Release from membrane==: PIP that is packed into vesicles leave, this weakens adaptor protein’s bind. Adaptor proteins uncoat, as soon as they pinch off coat is useless.
- Uncoating ATPase - ATP hydrolysis helps peel off coat
Coat formation
COPI - Arf involved
COPII - Sar1 Involved
Arf and Sar1 are coat-recruitment GTPases
- Molecular switch work with the two GTPases
- They can switch proteins from being turned to ON or OFF.
- ==GEFs== - GDP → GTP (activate protein) (GEF = guanine nucleotide exchange factor)
- ==GAPs== - GTP → GDP (inactivate protein) (GTPase activating protein)
- ==Sar1==: Inactive bound to GDP, needs to work with molecular switch. GEF takes inactive protein and activates it.
- Sar1-GEF turns Sar1 protein on. GDP is gonna leave, in comes GTP. Exchanges them.
- Active membrane bound, ready to initiate buding.
- ==Arf==: Active, bound to GTP. Works with GAPs to inactivate.
- PIPs control when it is active and inactive (can’t have coat when inactive)
Coat disassembly (cont.)
- COPI & Clathrin shed immediately
- COPII sheds closer to target membrane
==Rab-proteins==: GTPases, plays a role in vesicle arriving at correct target membrane
Rab-GDP is inactive, when inactive it is bound to GDI (Rab-GDP dissociation inhibitor)
Float around while inactive
Membrane bound GEFs activate proteins
Rab-GEF activates Rab. GDP → GTP bound.
The scrunched up tail unfolds and gets into membrane itself. (Rab-GTP is active and anchored to membrane)
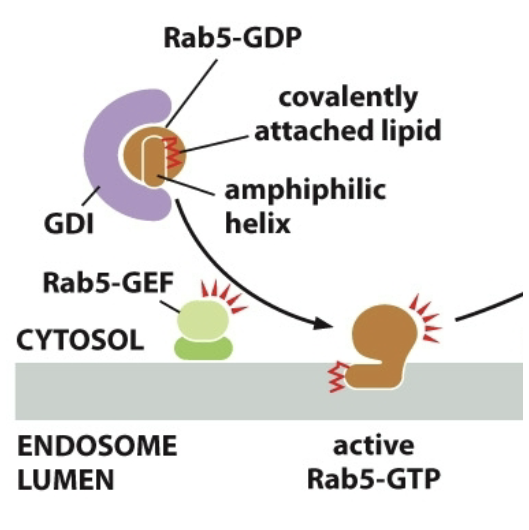
Rab effectors: Rab-GTP works with them, they’re tethers, catch vesicles and bring em in. Tethering proteins. Long-threadlike
==SNARE Proteins==: Responsible for vesicles completing membrane fusion once tethered.
Membrane fusion: bringing lipid bilayers close to each other so they can merge and dump the contents.
- v-SNARES → found on vesicle membranes (1 protein needed to get it stuck in plasma membrane)
- t-SNARES → found on target membrane (3 proteins to get vesicle stuck in it)
- v- and t-SNARES wrap around to form a stable 4 helix bundle → ==trans-SNARE complex==; locks the 2 membranes together.
Process
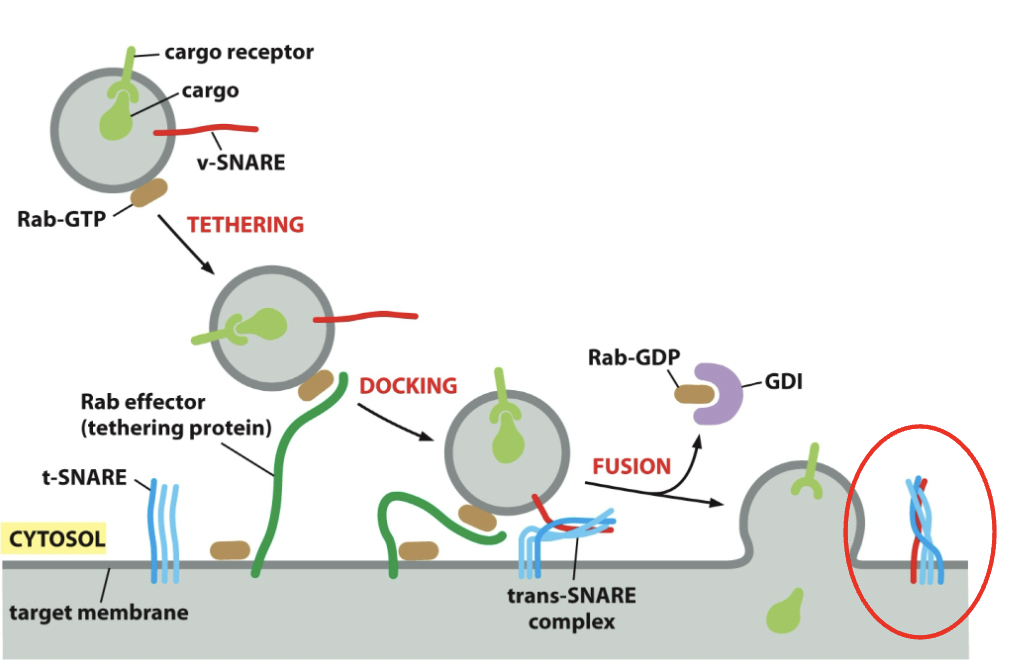
- Once vesicle fuses, cargo discharged; it’s possible to have one v-SNARE vesicle, but you can have more. NSF protein works with accessory proteins and is able to cycle membranes and cytosol. Uses ATP (→ADP energy) to pry apart trans-SNARE complex. The v-SNARE will return to the original compartment and get reused.
Transport from ER to Golgi
- Transport can happen from A→B or B→A
Protein leaving ER
- Protein transported in COPII coated transport vesicles
- Exit in ER exit sites (transitional ER)
- Can be membrane or soluble cargo proteins
- Cargo membrane proteins - display exit signals on cytosolic surface (COPII recognizes)
- Soluble cargo proteins - (in ER lumen) have exit signals attach them to transmembrane cargo receptors
- Vesicular tubular clusters - two vesicles have their trans-SNARE complexes separated by NSF. Two vesicle’s clusters bind together and then have membrane fusion.
Protein going back to ER
- ==Retrograde (retrieval) Transport==
- COPII vesicles going from ER to Golgi
- They're fusing into tubular cluster. A large one.
- Things need to go back to ER as cluster or individual
- Could be ER resident proteins or receptors that got stuck on
- COPI-coat from Golgi to ER, and it goes back to ER.
- ==Retrieval pathway==
- ER resident proteins have retrieval signals if they’re accidentally sent out
- Soluble ER proteins (BiP) have a C-terminal signal
- KDEL sequence → recognized by KDEL receptor and the BiP is now the cargo being brought back to the ER.
- KKXX Sequence for transmembrane
- Retention mechanism: ER resident proteins bind to each other and form complexes that are too big to enter.
Golgi Apparatus
Parts
- Cisternae: Flat Golgi stuff, what Golgi is made up of
- Golgi apparatus: What the tubular connection are called.
- Cis side - interconnected tubular structure - Cis Golgi Network (CGN)
- Beginning of cycle
- Trans side - Interconnected tubular structure - Trans Golgi Network (TGN)
- Where sorting happens
- Golgi sorting station: Where they need to go next (cell surface, outer-cell, earlier compartment, etc.)
- ==Pathway==
- CGN → cis cisterna → medial cisterna → trans cisterna → TGN → lysosomes
==Secretion== (leaving TGN)
Destined for lysosomes (have digestive enzymes, they digest macromolecules)
Exocytosis (getting sent to cell membrane)
- Immediate delivery to cell surface
- Destined for secretory vesicles
==Lysosomes==
Membrane enclosed organelles filled with enzymes that digest macromolecules. There is proteases and nucleases within them.
==Acid hydrolases==: enzymes in lysosomes need an acidic environment to function. Controls them from going and destroying everything. They’re physically contained and the pH isn’t suitable for them.
- Golgi → specific process → vesicles off Golgi → attach to endosome → mature → destroy (?)
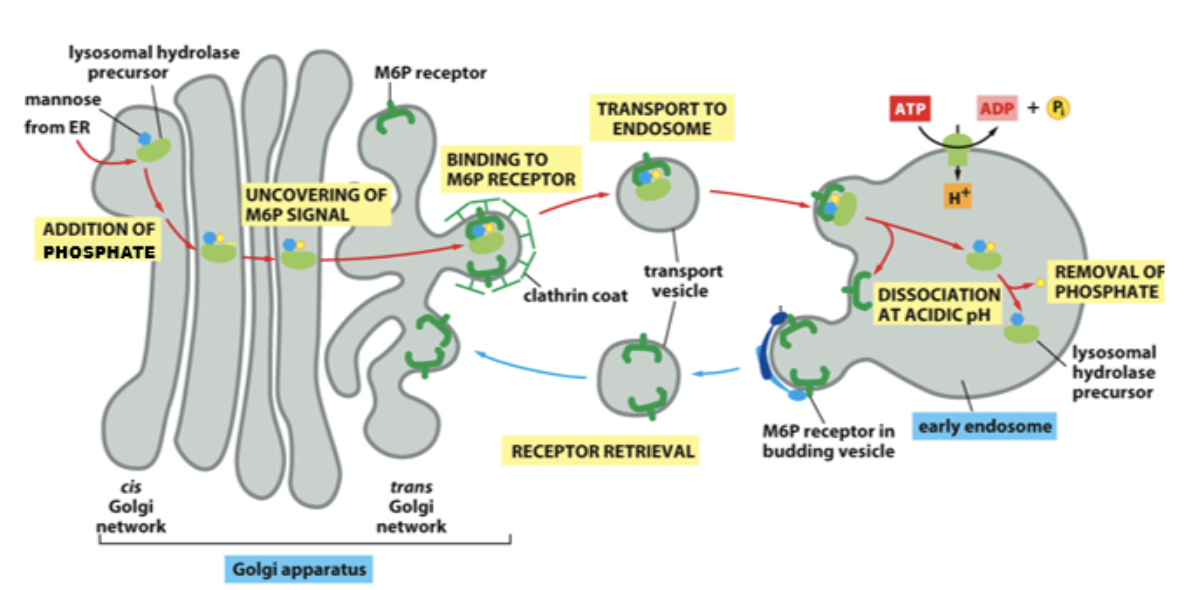
Protein: ==lysosomal hydrolases==. Precursor with mannose attached. Mannose is phosphorylated (doesn’t enter Golgi that way) and this signal brings it to lysosome.
- Route 1
- Proteins that needs to be destroyed or are a lysosomal hydrolases (does the destroying)
- Proteins have M6P tag to mark it to bring to lysosome (the phosphorylated mannose)
- Signal patch: a sequence that gives a location to where it needs to go.
- Transmembrane M6P receptors (on TGN)
- Recognize group
- Bind to the lysosomal hydrolase and to adaptor proteins to form a clathrin-coated vesicles
At TGN the M6P receptor recognizes M6P. Binds to adaptor proteins. Disassociates from the receptor. Fusion occurs. Receptors go back to Golgi.
In lysosome after receptors go back, and it’s like almost endosome, phosphate is removed and it becomes a mature/late endosome with stuff ready to be destroyed.
 Endosome steps
Endosome steps
Early endosome → late endosome → proteins destroy or get destroyed
Late endosome + lysosome = endolysosme
- Digestion completed in the lysosome
- Hydrolase destroys stuff
- Lysosome with nothing but hydrolase at the end
Late endosome can also fuse with endolysosome, doesn’t need to be a lysosome
Final product: Reuse or excrete products (like amino acids)
Exocytosis
Fusion of the vesicles with the plasma membrane.
==Constitutive secretory pathway==: operates continuously in all the cells. Immediate delivery to cell surface as default pathway (doesn’t need a signal), otherwise it goes back to ER or is retained in Golgi.
==Regulatory secretory pathway==: soluble proteins and other substances are stored in secretory vesicles for later release by exocytosis. Wait till things are called. Specifically for things that release rapidly like a neurotransmitter
- Held in vesicles until their time, the vesicles form and wait.
- Extracellular signal summons the product and then the vesicle fuses with cell membrane.
- Cells that are specialized for secreting some of their products rapidly on demand, store these products in regulated secretory vesicles.
- ==Dense-core secretory vesicles==: leave the TGN, they don’t just fuse right away, they wait for signal. They’re called dense because they’re densely concentrated.
- Some vesicles fuse (large immature secretory vesicles) and send excess membrane back to the TGN → they become tightly concentrated
- Membrane recycle immature secretory vesicle’s, bring back with clathrin coated retrieval vesicle
- After fusing, stuff is discharged and membrane becomes part of plasma membrane
Regardless of pathway → plasma membrane does not just get bigger. There’s equal amount of pinch off (endocytosis) as there is sending stuff (exocytosis)
Endocytic pathway
Endocytosis: process by which cells will take up stuff. The material to be ingested is enclosed by a small portion of the plasma membrane, which first invaginates and then pinches off to form an endocytic vesicle.
Pathway
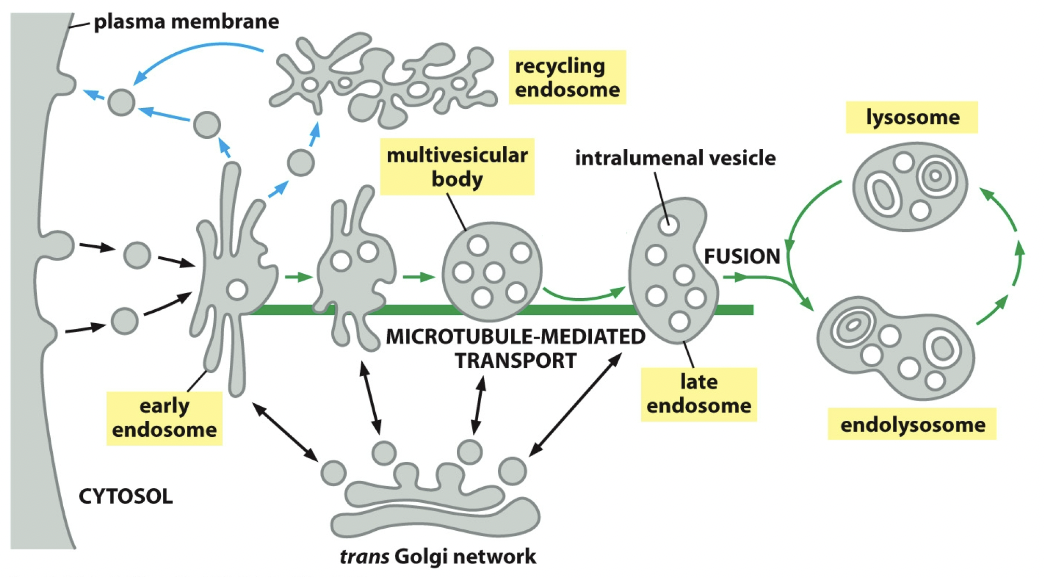
- Early endosome
- Formation of vesicle
- Endocytic vesicles fuse with receiving compartment
- Retrograde transport: recycling endosome; fuse with it, into vesicles, into plasma membrane
- Late endosome
- Pinched off membrane, cargos get more and more concentrated
- Collab with endosome or endolysosome (for destruction)
- Stuff can come from Golgi or outside Golgi to add to late endosome
- Retrograde transport stops recycling and is gonna take it, collab and hydrolase is gonna destroy it
- Trans Golgi network
- Some materials leave TGN and enter destructive pathway, however they can be sent backwards
- The M6P receptors go back to where they started.
==Endocytic (pinocytic) vesicles==: all eukaryotic cells ingest portions of plasma membrane from small vesicles
==Endocytic-exocytic cycle==: same amount of membrane being removed by endocytosis is being added by exocytosis
==Clathrin-coated pits:== we see pinching and vesicles. Clathrin shed coat right away. It’s not going to have this coat for long and fuses with the early endosome.
==Receptor mediated endocytosis==
- Ligands need to bind to receptors, and receptors are going to catch them
- Transmembrane receptors accumulate in the coated pits
- Enter in clathrin-coated vesicles
- Ex. Cholesterol uptake via RME → transported in blood in LDLs, receptors bind to cholesterol (LDL + coat + receptor), genes transcribed, protein translated, secretory pathway, PIP is phosphorylated and receptor binds to it, low pH brings receptor back to plasma membrane, maturation happens, and free cholesterol is release.
Endosome maturation
Parts of membrane pinches inward (area with stuff bound to it), called ==intralumenal vesicles==. An endosome with multiple is called multivesicular body.
==Tubular domain==: peripheral regions, where things will pinch off, getting sent back. Where stuff will fuse.
==Vacuolar domain==: Do not participate in pinching off, remain centre throughout maturation process.
Fate of endocytosed receptors
- Return to plasma membrane
- Endocytosed ligands: upon dissociation for ligand, they’re taken back to plasma membrane.
- Early endosome is a sorting station, can disassociate from receptors or stay with receptors
- Transferrin receptors - apotransferrin is low pH endosome releasing iron not transferrin. Word for ironless transferrin. Goes and find more iron.
- Receptor down regulation
- Destroy receptors
- Mark with ubiquitin; intralumenal vesicles need that tag and help get guided into clathrin-coated vesicles
- Ubiquitin binding proteins: recognize the ubiquitin chain and help direct to clathrin coated pits.
- Ubiquitin blocks receptor recycling to the plasma membrane and directs the receptor into degradation pathway
- Attaches to intralumenal vesicle to not be destroyed
- ==ESCRT protein complexes==
- How the pinching and invagination happens (sequestration)
- Regions on endosome that pinch in one themselves and form intralumenal vesicles
- ESCRT complexes (endosome sorting) get protein receptors marked with ubiquitin in pinching off area.
- Passes protein receptor from one ESCRT to the next.
- PIP is phosphorylated and serves as binding site for ESCRT.
- ==Transmembrane protein fate==
- Recycling - straight from early endosome to plasma membrane, further to interior of the cell, and then back to plasma membrane
- Transocytosis - all the way to opposite end
- Degradation - in early endosome, into endolysosome, then gets destroyed.
- Recycling endosome importance
- Insulin receptors increase glucose production or release it.
==Phagocytosis==
Specialized form of endocytosis, where engulfing happens.
==Phagosome==: large endocytic vesicle used to ingest large particles
Phagocytic cells: protect from infection, there is specialization
Here comes bacteria, pinching in. Vesicles forms with bacteria within it. Collab with phagosome and lysosome. Phagolysosome destroys.