Chapter 23: Mitochondrial DNA Profiling
23.1: Human Mitochondrial Genome
- Mitochondria: Are cellular organelles that serve as the energy-generating components of cells.
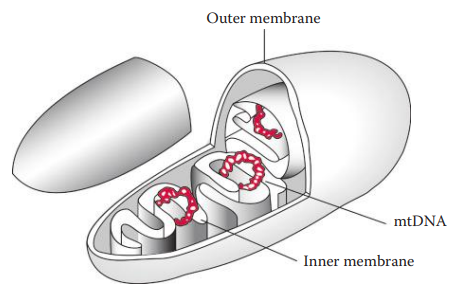
Genetic Contents of Mitochondrial Organelle Genomes
- Cambridge reference sequence (CRS): The sequence was largely derived from a placental sample from an individual of European descent and also partially from HeLa cells and bovine cells.
- It was later discovered, by resequencing the original mtDNA sample, that the CRS contains substitution errors at 10 nucleotide positions.
- The revised Cambridge reference sequence (rCRS) was published in 1999 and presented corrections to these substitution errors.
- Control Region: also known as Displacement Loop; it contains the origin of replication for one of the mtDNA strands but does not code for any gene products.
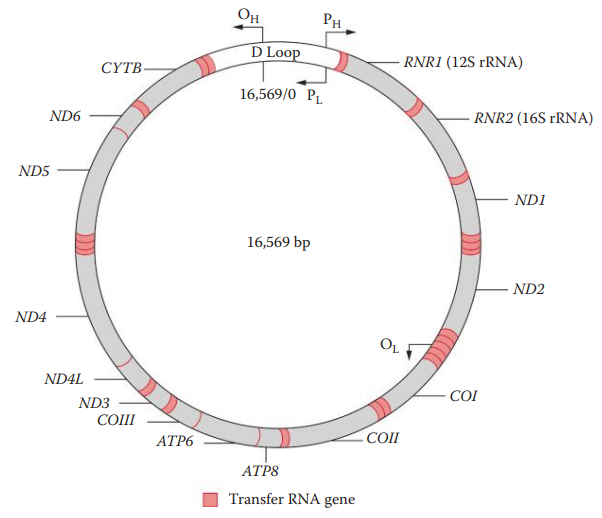
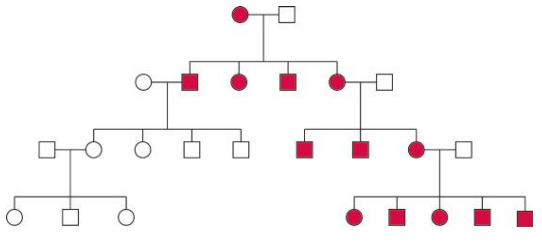
Maternal Inheritance of mtDNA
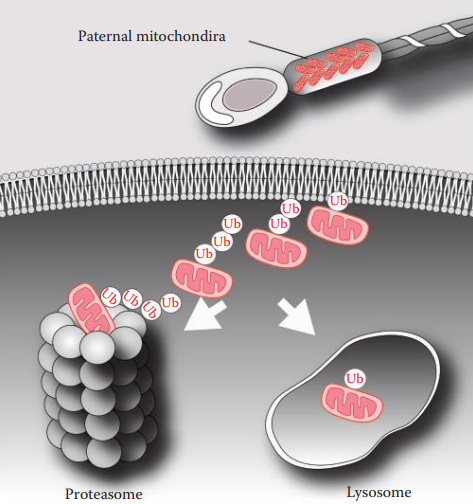
- Maternal inheritance is typically observed for the mtDNA genome, which is inherited differently from nuclear genes.
- The inheritance of the mtDNA genome does not obey the rules of Mendelian inheritance and is thus called non-Mendelian inheritance.
- Mitotype: Considered a a haplotype treated as a single locus.
23.2: mtDNA Polymorphic Regions
Hypervariable Regions
- The most polymorphic region of mtDNA is located within the D-loop.
- The three hypervariable regions in the D-loop are designated: Hypervariable Region I, Hypervariable Region II and, Hypervariable Region III
- The most common polymorphic regions of the human mtDNA genome analyzed for forensic purposes are HV1 and HV2.
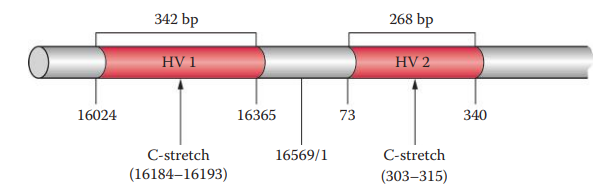
Heteroplasmy
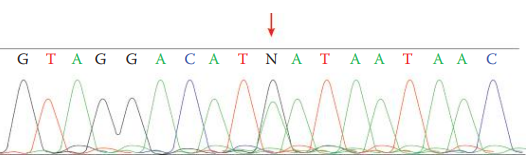
- Heteroplasmy occurs when an individual carries more than one mtDNA haplotype.
- Sequence heteroplasmy: The presence of two different nucleotides at a single position shown as overlapping peaks in a sequence electropherogram.
- Light heteroplasmy: Are often observed at the uninterrupted C stretches in sequencing, in which sequencing products with various lengths of polymeric cytosine residues are present.
23.3: Forensic mtDNA Testing
General Considerations
- mtDNA analysis is often used on samples derived from skeletal or decomposed remains.
- The surface of the sample should be cleaned to remove any adhering debris or contaminants.
- Bones and teeth are pulverized to facilitate extraction of the mtDNA Assay.
- mtDNA-specific quantization methods using real-time PCR can also be used to directly obtain measurements of mtDNA extracted.
mtDNA Screen Assay
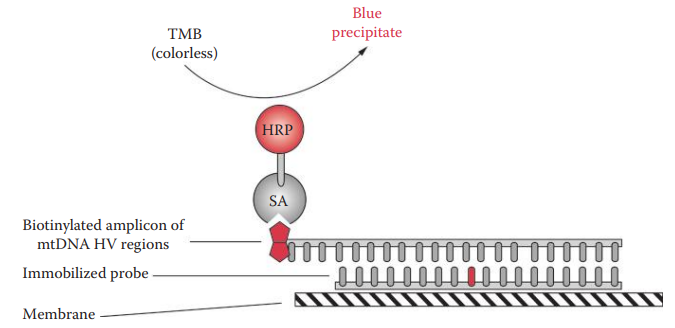
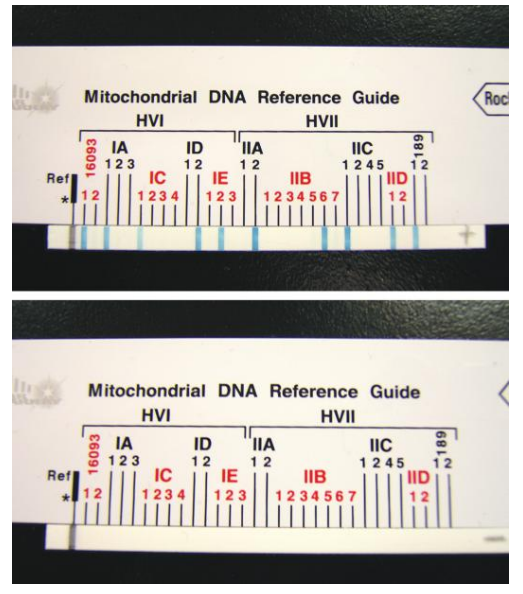
- ASO Assay: It allows initial screening of mtDNA sequence polymorphisms and has the potential to reduce the number of samples required for mtDNA sequencing.
- Linear Array™ mtDNA HV1/HV2 region sequence typing kit: It utilizes reverse ASO configuration with a panel of immobilized ASO probes that detect common polymorphic sites.
mtDNA Sequencing
- A combination of PCR amplification and DNA sequencing techniques is utilized to reduce the time and labor needed to obtain DNA sequences from genomic DNA templates.
- mtDNA sequencing usually consists of:
- PCR amplification;
- DNA sequencing reactions;
- Separation using electrophoresis; and
- Data collection of Sequence analysis.
- DNA Sequencing Reactions
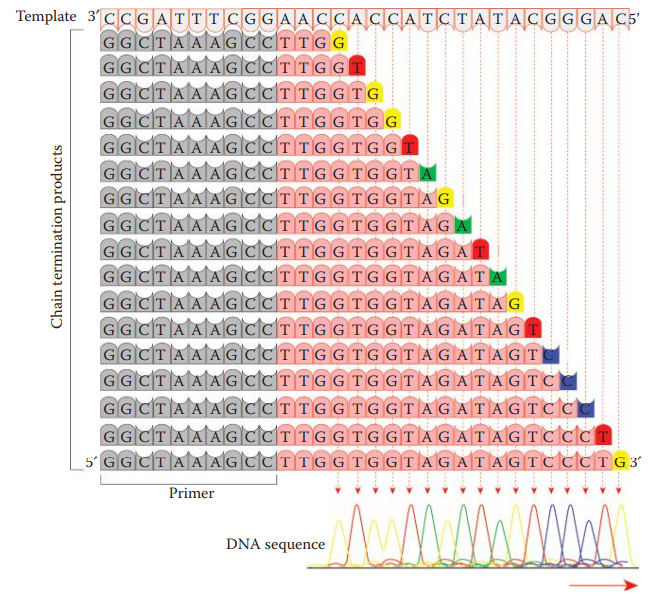
- Chain-Termination or Sanger Method: An oligonucleotide primer that can anneal to a single stranded DNA template is utilized.
- Cycle Sequencing: It utilizes thermal cycling to generate a single-stranded template for chain-termination sequencing reactions.
- Electrophoresis, Sequence Analysis, and Mitotype Designations
- Reporting Format: Sequence differences relative to the rCRS are listed in data format.
- Insertions: Described by noting the position followed by a decimal point and a number.
- Deletions: These site designation is followed by letter d.
- Hetero-plasmic Sites: The IUPAC codes for base calling can be applied to hetero-plasmic sites.
Interpretation of mtDNA Profiling Results
- Interpretation guidelines are used when comparing sequencing results between evidence and reference samples.
- Exclusion: If the sequences of questioned and known samples are different, then the samples can be excluded as originating from the same source.
- It should be taken into account that higher mutation rates are found with the mtDNA genome than are found with the nuclear genome.
- Cannot Exclude: If the sequences are the same, the reference sample and evidence cannot be excluded as arising from the same source.
- When an mtDNA profile cannot be excluded, it is desirable to evaluate the weight of the evidence.
- Inconclusive Result: If the questioned and known samples differ by a single nucleotide, and no heteroplasmy is present, the results are considered to be inconclusive.