Variation
Phenotypes, Chromosomes, Proteins, DNA
Variation is essential for adaptive evolutionary change
Natural selection cant operate without phenotypic differences
Most studies either measure and compare variation or try to link variation to fitness (reproductive success and survivorship)
Ways variation has been measured
Experimental matings (phenotype) and Chromosomes - 1900-1970
Protein electrophoresis (allozymes) - 1970s
Mitochondrial DNA - 1980s
Nuclear DNA - 1990s
Genomics (sequencing whole genomes) - 2000s
Phenotypes
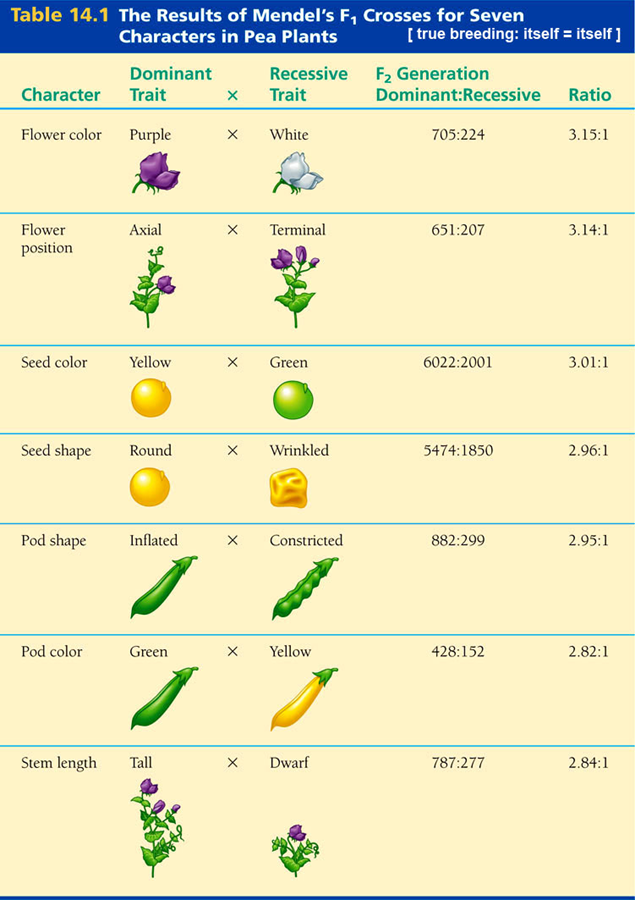
Mendel: 7 traits with discrete variation (yellow vs. green) rather than continuous (height)
Do phenotypic differences have a genetic basis?
Use crosses and frequencies of phenotypes to determine genetic basis
Doesn’t really account for the environment - can be attributed to genetics when you control the environment
Behavior (considered a phenotypic trait)
Genetically based differences in behavior
local adaptation
adaptation to captivity
Ex. migratory behavior in Blackcaps
tendency to migrate and direction to migrate have a genetic basis
Morphology
individuals vary in size, shape, number of body parts
problem using morphological traits to study patterns of genetic variation
variation caused by genetic and environmental differences
need to partition total phenotypic variation caused by genetic and environmental differences
Differences among populations
is difference due to genetic variation or environment?
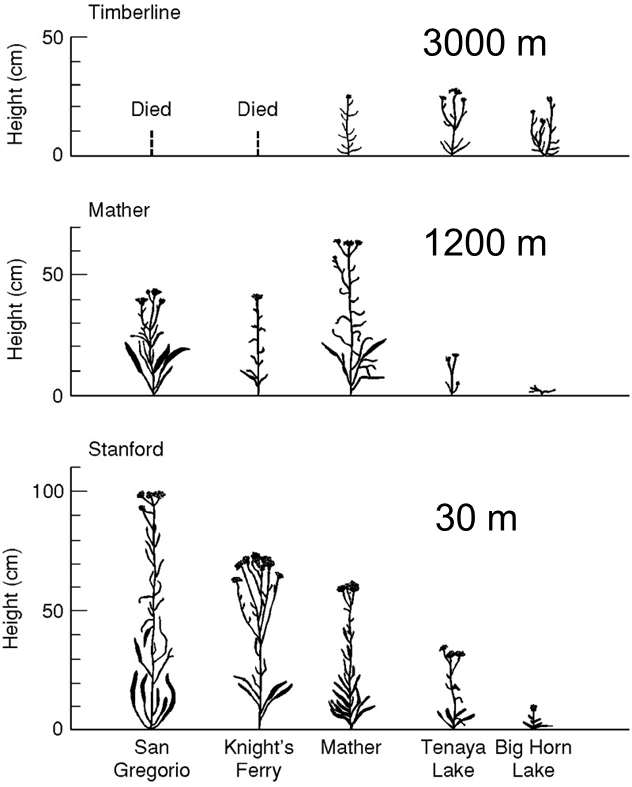
common garden experiment: eliminate the effect of environment by raising individuals in identical environments
ex. altitudinal forms of yarrow
genetic basis to differences
Chromosomes and Proteins
genetic basis to phenotypic variation
examine differences among individuals using chromosomes and proteins
reflects historical sequence of study
these are old tools
Chromosomes
little emphasis on chromosome variation
can reduce fertility/survivorship is you cross individuals with a different complement of chromosomes
some captive breeding programs hybridized individuals that were morphologically similar but had a different number of chromosomes
e.g. orangutans, gazelles, dik-dik
may also be a problem in translocations and reintroductions if individuals are translocated among chromosomally distinct groups
e.g. graceful tarplant
morphologically similar and live in similar habitat
size and shap of 4 chromosomes differenced among pops
experimentals crossings: failed to produce F1 or F1 was sterile
Quanitfying chromosome variation
number
size: largest to smallest
centromere location (middle or towards the end)
banding pattern - some regions of DNA condense more than others forming dark bands
Number of chromosomes
evolves slowly in some taxa and quickly in others
~100 cetacean species all with 2n = 42 or 44
Equus: 2n varies between 32 and 66
polyploidy
most animals are 2n (except eggs and sperm = 1n)
some spp polyploid: more than 2 sets of chromosomes, usually fish, amphibs, lizards, plants (3n to 6n+)
Gray Tree frogs
diploid and tetraploid forms (considered different species) occus in sympatry
reproductively isolated by call recognition
hybrids are triploid and sterile
Proteins
another method to measure variation
major advance
direction relationship between genes (DNA base pair sequence) and proteins (amino acid sequence)
Allozymes = variant enzymes produced by different alleles at a locus (enzymes are proteins)
Variation in proteins
electrophoresis: separates proteins (enzymes) based on net change and molecular weight
proteins negatively charged
Gel: bands are revealed by straining the gel with the substrate of the protein being screened. if the protein reacts with the substrate, the band appears on the gel
disadvantages:
proteins can be specific to tissue
proteins degrade is not stored properly
needs a lot of tissue
most tissue taken invasively
often reveals little variation
Advantages:
not technically difficult - no markers needed
only game in town before DNA techniques
Measuring Variation
number of alleles
number of polymorphic loci
observed heterozygosity
Types of DNA
Nuclear - biparental inheritance (one copy/cell)
Organelle - uniparental inheritance
Mitochondrial (mtDNA) - 1000s of copies/cell
Chloroplast (cpDNA) - 1000s of copies/cell
mtDNA
First studies of DNA genetic variation used mtDNA in animals
characteristics:
small size, conserved arrangement of genes
variable: less stringent repair of errors during replication so mutations accumulate
no recombination and maternal inheritance in animals so inherited as a single haplotype: lineages can be tracked over time and space easily - often used in phylogenetic studies
b/c haploid and uniparentally inherited it represents ¼ the pop size of diploid nuclear DNA - sensitive to bottlenecks
Chloroplast DNA
mtDNA not very variable in plants so cpDNA generally used instead when haploid markers are required
inherited uniparentally: maternal in flowering plants, paternal in conifers/cycads
slower mutation rate compared to plant nuclear genes
Uniparental DNA: pros
mtDNA, cpDNA, and Y chromosome genes
can follow transition of maternal/paternal genotypes thru time
can look at differences in disperal
distribution of mtDNA (maternally inherited) reflects patterns of seed dispersal but not pollen dispersal
can identify hybrids
Dispersal in black spruce
grow in areas covered by ice 6000 years ago
new pops have little mtDNA variation, lots of nuclear variation
mtDNA dispersed by seeds, dont travel far: only a few established new pops
nuclear genes can disperse widely bc pollen wind-dispersed
pollen that blew to the new pops likely originated from several places so high nuclear genetic variation
if you looked at just mtDNA or just nuclear DNA, would get an incomplete or misleading picture of dispersal
Hybrids
contain a mix of alleles from both parental species
hybrids often have cytoplasmic markers (mt or cp) from one species or pop and nuclear markers from both
known as cytonuclear disequilibrium
Uniparental DNA: cons
behave as single inherited units so effectively a single locus: may not reflect the true history of a species
not representative of pops as a whole, e.g. dispersal and mtDNA
if males disperse and females dont, the mtDNA haplotypes would be pop specific and you may conclude that individuals never move among pops
better to use a mix of markers