D1.1 DNA Replication Notes
Guiding Questions:
How is new DNA produced?
How has knowledge of DNA replication enabled applications in biotechnology?
How is biotechnology unique in terms of human ingenuity?
What biotechnology processes have been developed for human benefit?
How old is biotechnology?
D1.1.1:
“the structure of DNA is ideally suited to its function.”
Each DNA molecule replicated consists of one new strand and one original strand from the parent molecule due to semi-conservative replication. DNA replication is needed for 3 body functions: Reproduction, Growth, and Repair. (THINK: GRR)
D1.1.2:
When DNA was being discovered, there were 3 leading theories on how DNA was replicated. They were:
semiconservative: the parent DNA strand gets split into two single strands that are the templates.
conservative: an exact copy of the parent strand is made, and the original stays intact.
dispersive: the parent strand is mixed throughout both new strands.
Meselson and Stahl proved that semi-conservative was the right way through their experiment that observed the path of the original, parent strand through multiple rounds of replication.
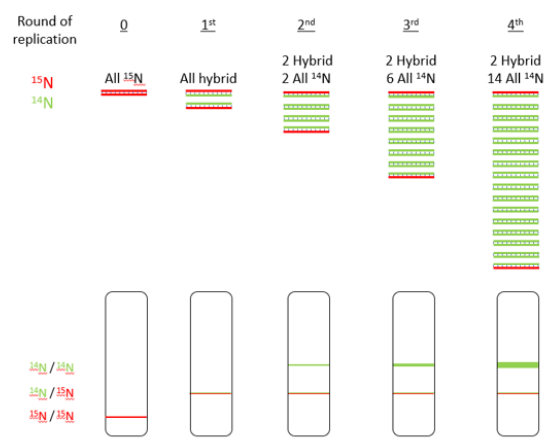
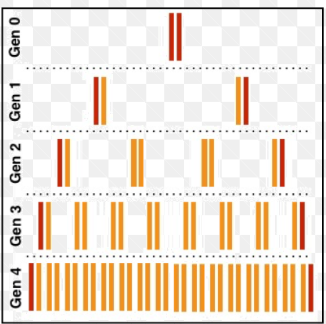 Ultimately, they proved semi-conservative replication throgh the obsersion of two DNA molecules, both with a whole strand from the parent molecule.
Ultimately, they proved semi-conservative replication throgh the obsersion of two DNA molecules, both with a whole strand from the parent molecule.
D1.1.3:
Helicase:
Breaks hydrogen bonds in order to unwind DNA helix
Composed of 6 polypeptides (globular protein) and is arranged in a donut shape.
Uses ATP to move along DNA helix and break bonds
DNA polymerase:
adds complimentary bases to new strand
adds in 5’—>3’ direction
D1.1.4:
Polymerase chain reaction (PCR) is a method used to amplify any piece of DNA. In english this means that a piece of DNA goes through multiple rounds of replication, with the reaction starting with a polymerase (usually artificial). Taq polymerase is often times used in labs due to its ability to work at high efficiency under high temperatures. It also is easily accessible as it can be extracted from Thermus aquaticus, which can be found in hot springs.
Gel electrophoresis is a way to see DNA by seperatin them. It can seperate DNA, RNA, or proteins based on their size. To perform gel electrophoresis, the molecules are placed at the end of a jello-like matrix, called agarose, and an electric current is applied. The molecules move towards the positive pole of the gel, and their distance traveled is based on their size.
D1.1.5:
STRs are short tandem repeats of sections of DNA. For example, CATTAT could be repeated 20 times in someone’s DNA. VNTRs are similar, but they are variable number tandem repeats. This means that there is a larger range in how many repeats of the sequence there are. One person could have 2 repeats, but another could have 7,000 repeats, it varies a lot. STRs are commonly used for DNA profiling as forensic scientists are able to distinguish people’s DNA by their genetic repeats.
VNTRs are inherited from both parents. Therefore, the children will never have the same number of VNTRs as the parents, it will always be a number inbetween the two. VNTRs can show variations between individuals because of the individuality in how many times each sequence is repeated.
D1.1.6: (HL Content)
Franklin made conclusions about the structure of DNA, such as:
Helical shape
Repeating structure
Linus Pauling:
concept of molecular modelling
Rosalind Franklin:
Helical structure
Chargaff:
Base composition
Watson and Crick:
Antiparralell
stability
tight packing of DNA
REMEMBER:
AT —> 2 hydrogen bonds
CG —> 3 hydrogen bonds
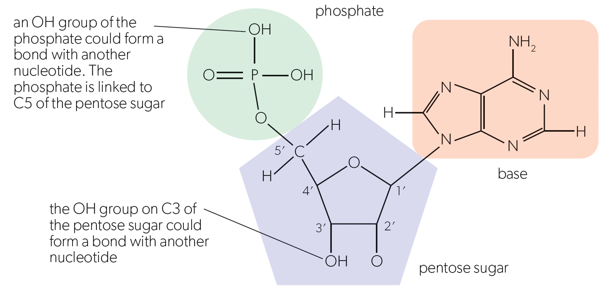 Phosphodiester bonds are between nucleotides (in the backbone of the DNA helix). The energy to make a posphodiester bond comes from the hydrolysis of the first 2 phosphates from the nucleotide. This reaction takes off the other two phosphates, to leave only the one that we want. The removal of these two phosphates releases energy, that is used to make the phosphodiester bond.
Phosphodiester bonds are between nucleotides (in the backbone of the DNA helix). The energy to make a posphodiester bond comes from the hydrolysis of the first 2 phosphates from the nucleotide. This reaction takes off the other two phosphates, to leave only the one that we want. The removal of these two phosphates releases energy, that is used to make the phosphodiester bond.
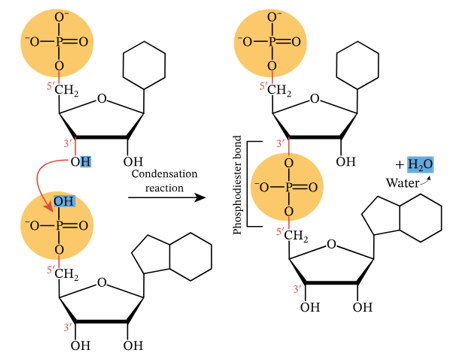 Remember that the addition of new nucleotides is a CONDENSATION reaction, because one water molecule is removed for each phosphodiester bond made.
Remember that the addition of new nucleotides is a CONDENSATION reaction, because one water molecule is removed for each phosphodiester bond made.
D1.1.7:
Because the two strands run antiparallel, a leading and lagging strand are created. The leading strand continuously follows the replication fork as new bases are continiously added. Since it is continuous, it only needs one primer. The lagging strand is made in fragments as its moving away from the replication fork. The fragments are called Okazaki fragments, as nucleotides are discontinuously added to the new strand. The lagging strand needs multiple primers because it starts at multiple different points. Ligase is the enzyme used to connect the Okazaki fragments to turn the lagging strand into a continuous strand. They both add nucleotides in the 5’—> 3’ direction.
D1.1.8:
Topoisomerase:
Releases the supercoiling strain on the helix. Happens before helicase unzips the helix to make it easier.
Helicase:
unzips the DNA helix, but it requires ATP for movement along the DNA strand to unzip it.
DNA Primase:
Creates RNA primer to initiate DNA polymerase adding nucleotides. One on leading strand, but multiple on lagging strand.
DNA Polymerase III:
Adds complimentary base pairs to nucleotide chain, proofreads for errors and makes corrections.
DNA Polymerase I:
Removes RNA primers, and breaks phosphodiester bonds
Ligase:
Connects the gap between Okazaki fragments in the lagging strand.
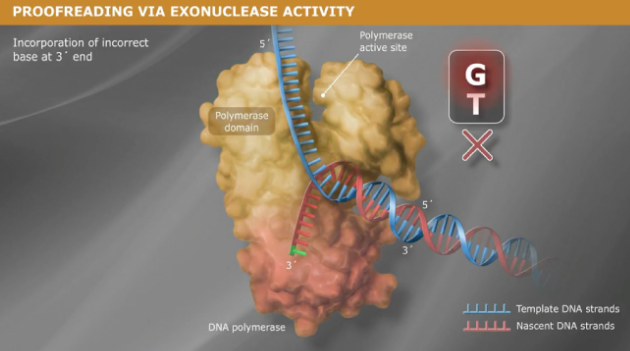
D1.1.9:
Along with adding base pairs, DNA Polymerase III also does proofreading on the new DNA strands immediately after replication. When a disturbance in the helix is discovered, DNA Polymerase III removes the mistake and replaces it with the correct base. By correcting the mistake the helix is fixed, preventing mutations.