BIOL 3130 LEC 8
1) What is the primary difference in "positioning" between an Enhancer and a Proximal Promoter Element?
Proximal Promoter Elements: Are position-dependent (must be adjacent/near the core promoter to work).
Enhancers: Are position-independent (can be located kilobases away or even within introns [non-coding sections of a gene] and still function).
2) What are the three defining physical properties of an Enhancer?
DNA Elements: Usually found upstream, but sometimes within introns.
Position-independent: Can be moved or located far away and still work.
Orientation-independent: Function correctly even if the DNA sequence is flipped to the opposite strand (5′ to 3′).
3) What is a UAS in yeast, and what is its human equivalent?
Upstream Activation Sequence. In yeast, these function similarly to proximal promoter elements in higher eukaryotes.
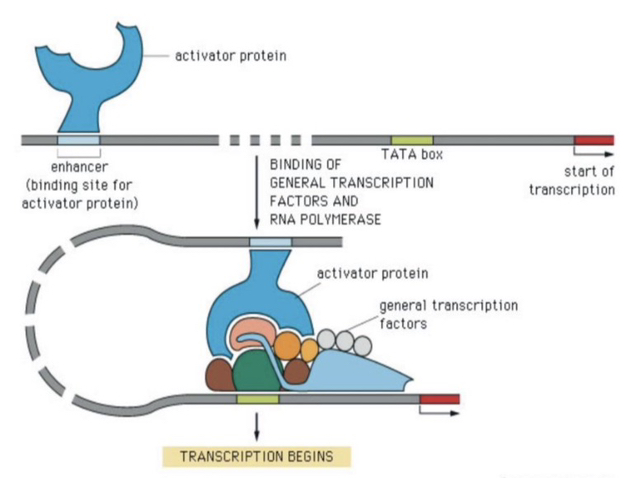
4) How does an enhancer physically communicate with a promoter?
Enhancers bind sequence-specific transcription factors (ssTFs) called activators. These activators recruit factors that promote an interaction between the enhancer and promoter, resulting in a DNA loop.
5) What are the two ways silencers are thought to work?
By binding transcription factors that act as repressors.
By promoting a chromatin structure (the way DNA is packaged) that is unfavorable for transcription.
6) What are General Transcription Factors (GTFs) and which specific ones associate with RNAP II?
They are a specific set of proteins that associate with RNAPs at the core promoters of all active genes. RNAP II requires: TFIIA, TFIIB, TFIID, TFIIE, TFIIF, and TFIIH.
7) Define Sequence specific transcription factors (ssTFs (also known as TFs)) and estimate how many exist in humans.
They are DNA-binding proteins that regulate the transcription of specific genes or groups of related genes. There are an estimated 1400 human TFs.
8) Within the category of ssTFs, what is the difference between an Activator and a Repressor?
Activators: Bind to proximal promoters or enhancers to help assemble RNAPs and GTFs at the core promoter.
Repressors: TFs that inhibit/repress transcription (sometimes binding at silencers).
9) What are Coregulators (Coactivators and Corepressors)?
Proteins or molecules that promote or inhibit transcription by working with activators or repressors. Unlike TFs, they usually function by interacting directly with the activators/repressors rather than the DNA.
10) Compare the primary binding targets of GTFs versus ssTFs.
GTFs: Bind at the core promoter.
ssTFs: Bind at proximal promoters, enhancers, or silencers.
11) Why are GTFs found at the promoters of almost all active genes?
Because they are part of the general transcription machinery; RNAP II absolutely needs these proteins to function and initiate transcription.
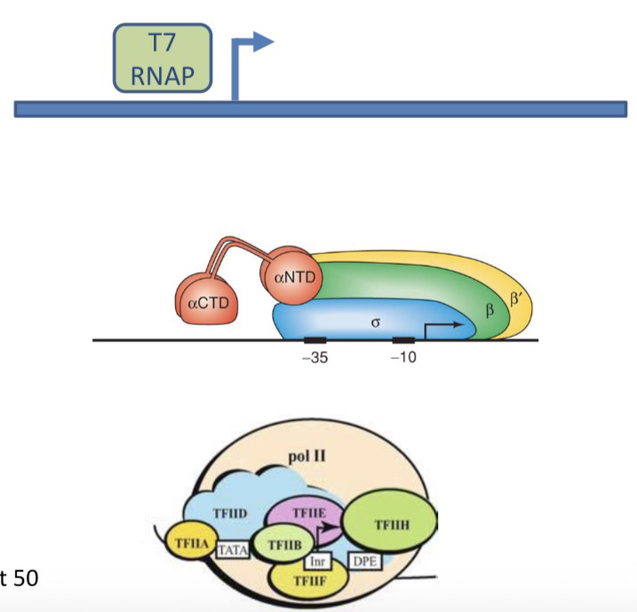
12) Compare the minimal protein requirements for promoter-dependent transcription in T7 Phage, E. coli, and Eukaryotes.
Phage T7: Single polypeptide (T7 RNAP).
E. coli: 6 polypeptides (α2ββ′ωσ).
Eukaryotes: RNAP (I, II, or III) plus GTFs (about 50 polypeptides for RNAP II).
13) What are the four key roles of GTFs at a promoter?
Recruitment of RNAPs to promoters.
Template melting (opening the DNA).
TSS (Transcription Start Site) selection.
Promoter clearance/transcription initiation.
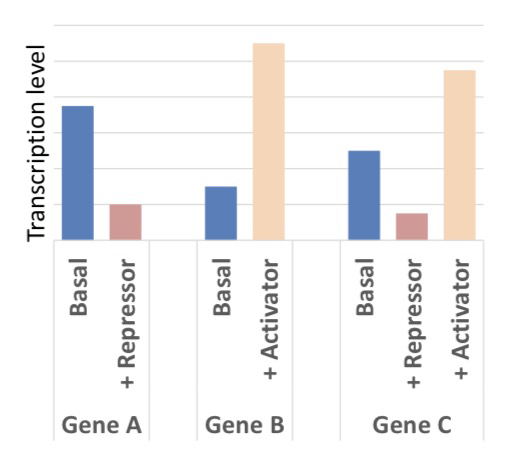
14) Define Basal Transcription and how it is modified.
Basal transcription is the "default" level of activity produced by RNAP + GTFs (the basal/general transcription machinery). It can be elevated by activators or reduced by repressors
15) What is the PIC, and how was its structure determined?
The PIC is the complete assembly of GTFs plus RNAP II on promoter DNA. Its structure was determined by X-ray crystallography.
16) Based on the Greenblatt/Reinberg gel-shift (EMSA) experiments, which factor is critical for any complex to form?
TFIID. It is the first factor to bind the promoter DNA, and no other factors can bind without it.
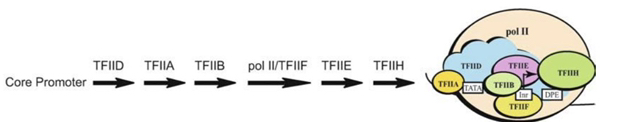
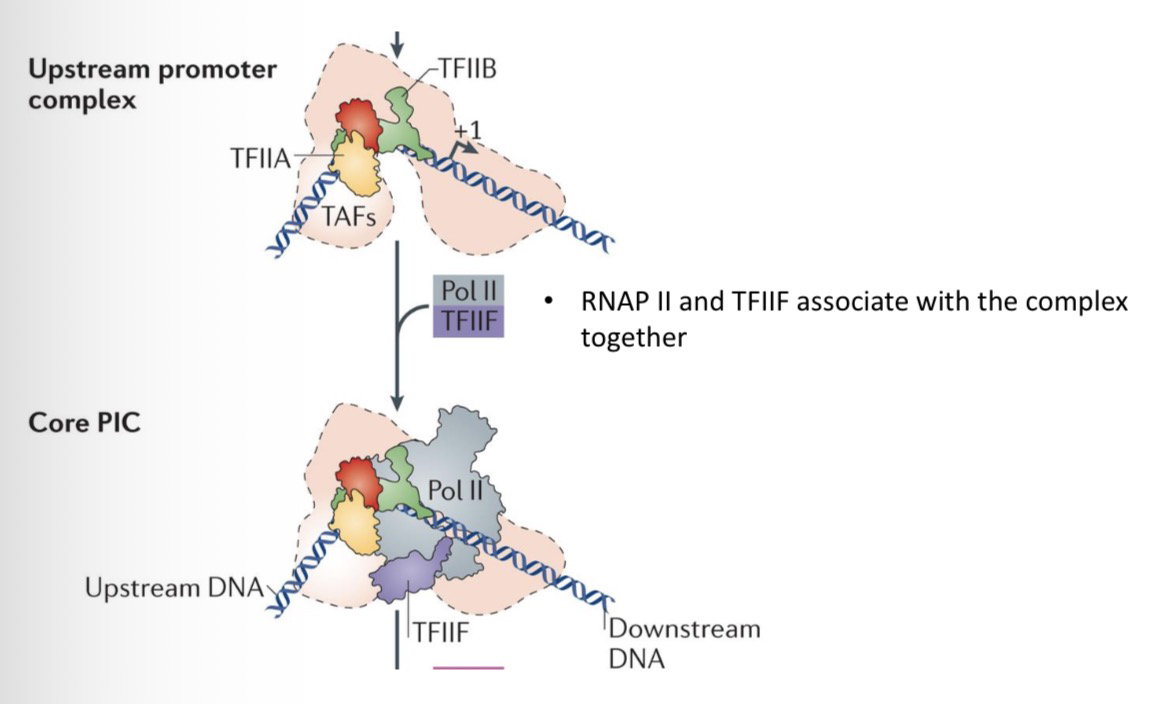
17) How are RNAP II and TFIIF recruited to the promoter?
They bind together as a unit. RNAP II cannot bind the promoter complex unless TFIIF is present, and TFIIF requires RNAP II to join the complex
18) What is the generally accepted step-wise order of PIC assembly?
TFIID→TFIIA→TFIIB→RNAP II + TFIIF→TFIIE→TFIIH.
Note: TFIIA is dispensable (not essential) in vitro but is found in living cells.
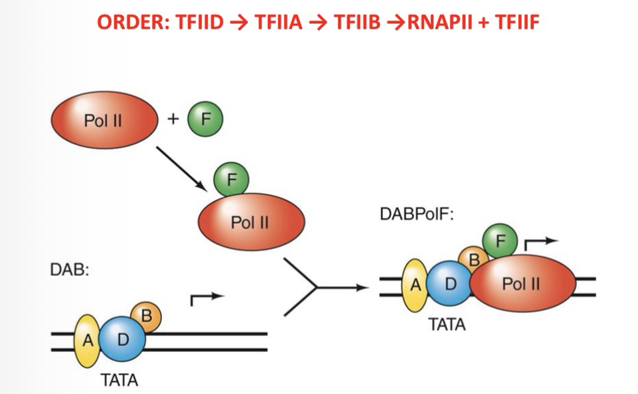
19) Which two researchers/groups were instrumental in determining the steps of PIC assembly?
Danny Reinberg and Jack Greenblatt (University of Toronto). They used the adenovirus major late promoter for their experiments.
20) Once the full PIC is assembled (including TFIIH), what is required for RNAP II to actually start making RNA?
The presence of NTPs (nucleoside triphosphates), which are the building blocks of RNA.
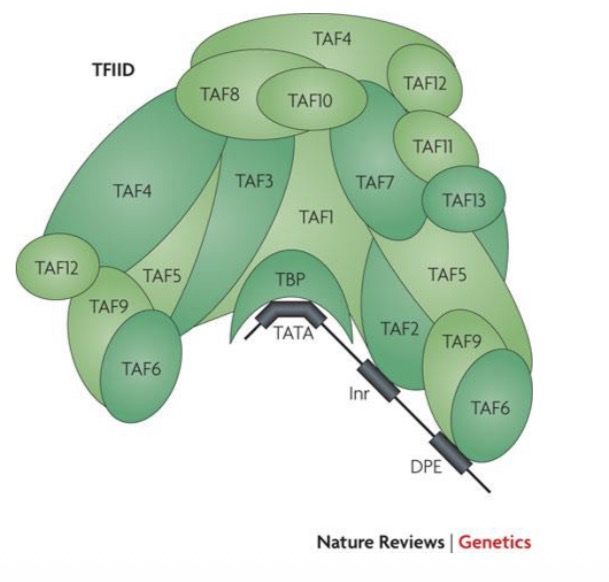
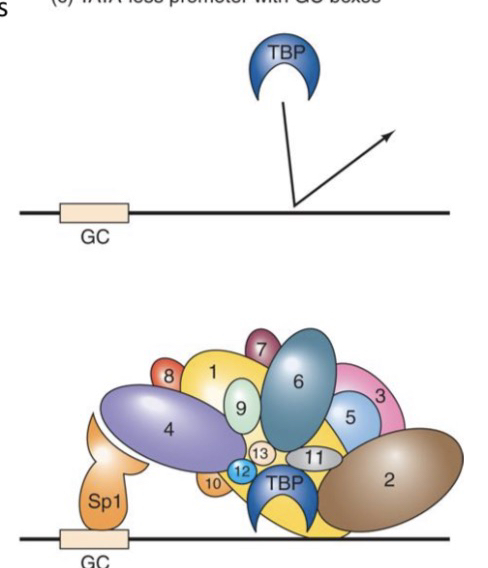
21) What are the two main components of TFIID and their general roles?
TBP (TATA-binding protein): Binds the TATA box.
TAFs (TBP-associated factors): 13 core proteins that help bind DNA (Inr and DPE) and contact activators (ssTFs) to start transcription.
22) Why is TBP unique among transcription factors?
It is believed to be the only component involved in transcription by all three eukaryotic RNAPs (I, II, and III). It is required even for promoters that lack a TATA box.
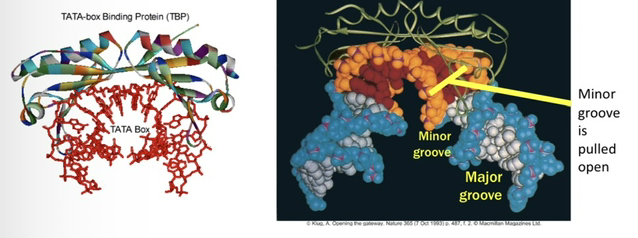
23) Describe the physical interaction between TBP and DNA.
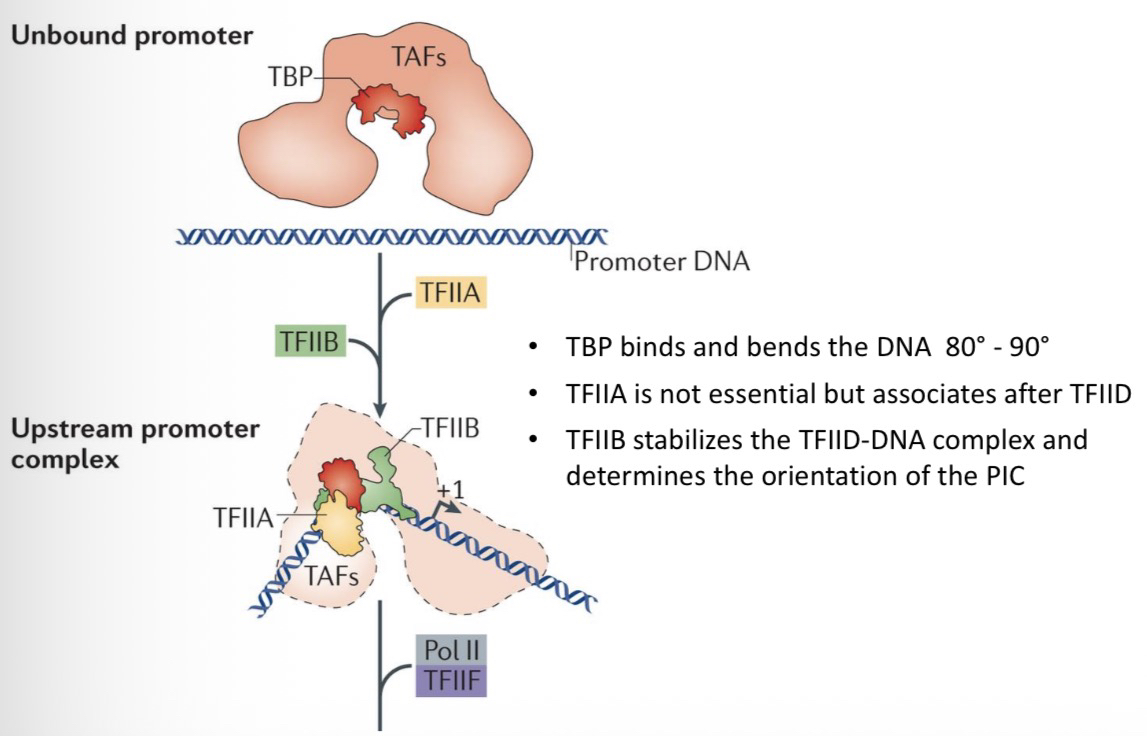
TBP is symmetrical and binds the minor groove of the TATA box like a saddle. It forces the minor groove open and bends the DNA 80° to 90°.
24) Name two specific DNA elements TAFs bind to and the subunits involved.
Initiator (Inr): Contacted by TAF1 and TAF2.
Downstream Promoter Element (DPE): Contacted by TAF6 and TAF9.
Note: This allows TBP to be recruited to TATA-less promoters.
25) How does the activator Sp1 recruit TFIID?
Sp1 binds to the GC box on DNA and makes direct contact with TAF4. This "tethers" TFIID to the promoter, even if a TATA box is missing.
26) Contrast the roles of TFIIA and TFIIB.
TFIIA: Stabilizes the TBP-DNA complex; increases basal transcription.
TFIIB: A single polypeptide that determines PIC orientation(direction of transcription) and recruits RNAP II to the complex.
27) What are the primary roles of TFIIF and TFIIE?
TFIIF: Associated with RNAP II before recruitment; required to bring RNAP II to the promoter.
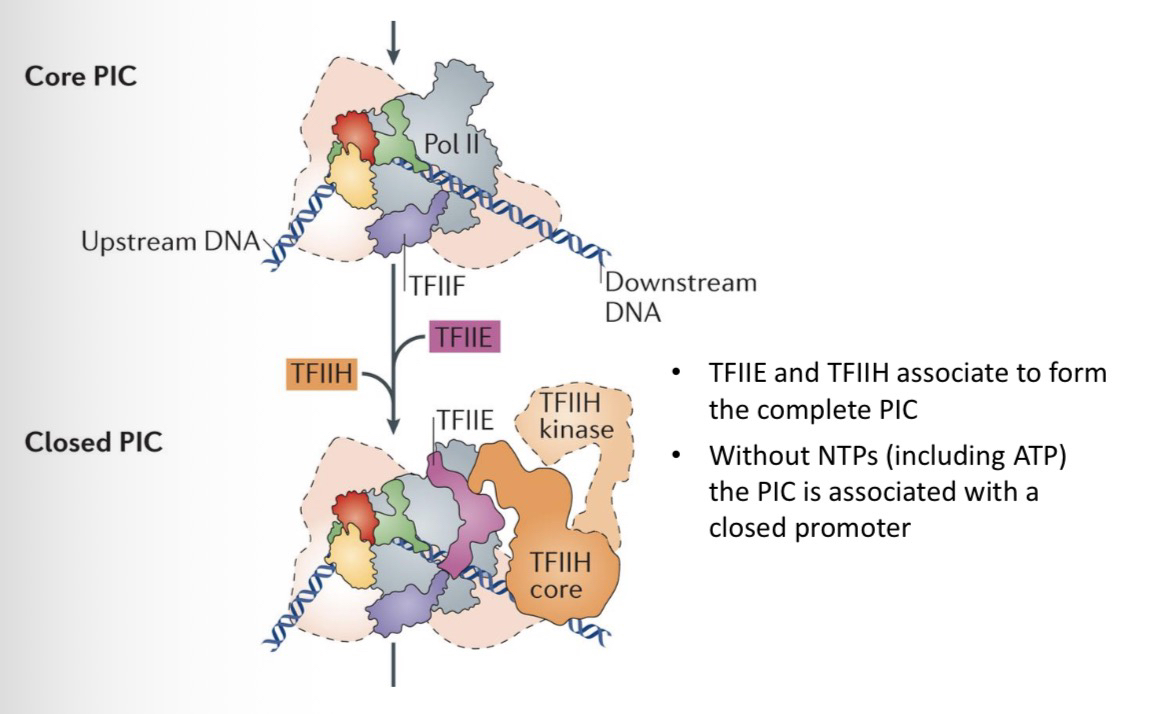
TFIIE: A heterodimer that recruits TFIIH to the complex.
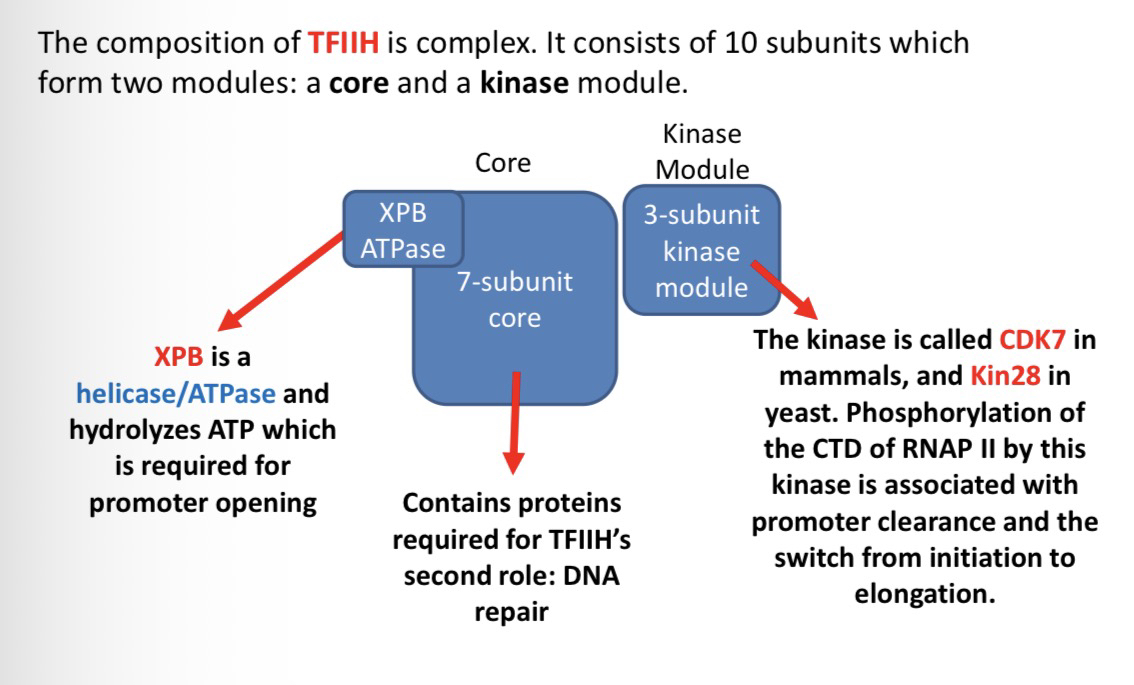
28) Describe the two modules of TFIIH.
Core (7 subunits): Includes XPB (an ATPase/helicase for DNA opening) and proteins for DNA repair.
Kinase Module (3 subunits): Includes CDK7 (mammals) or Kin28(yeast), which phosphorylates the RNAP II CTD.
29) What specific residues does the TFIIH kinase phosphorylate, and why?
It primarily phosphorylates Ser5 (and some Ser7) of the CTD repeats. This triggers promoter clearance (the switch from initiation to elongation).
30) How did in vitro kinase assays prove TFIIH targets the CTD?
RNAP IIa (with CTD) was phosphorylated by TFIIH.
RNAP IIb (lacking CTD) was not phosphorylated.
When phosphorylated Rpb1 was treated with chymotrypsin, the radioactive signal remained on the CTD fragments.
31) What major physical change occurs to the DNA when TBP binds, and which factor determines the direction of the complex?
TBP bends the DNA 80° to 90°.
TFIIB determines the orientation (direction) of the PIC.
32) Contrast the roles of TFIIA and TFIIB in PIC stability.
TFIIA: Not essential, but associates after TFIID.
TFIIB: Essential; stabilizes the TFIID-DNA complex so the other factors can join.
33) How do RNAP II and TFIIF join the growing PIC?
They associate with the complex together (as a pre-formed unit).
34) What is the status of the PIC if all GTFs are present but NTPs (including ATP) are missing?
The PIC is associated with a closed promoter (the DNA strands are not yet separated).
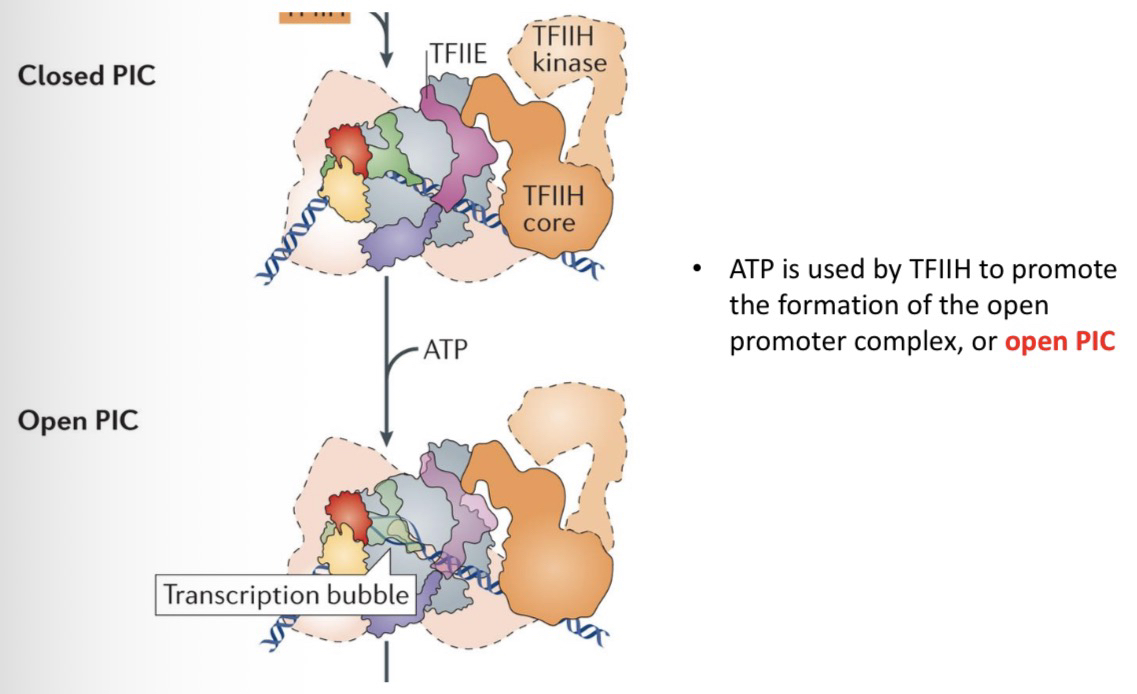
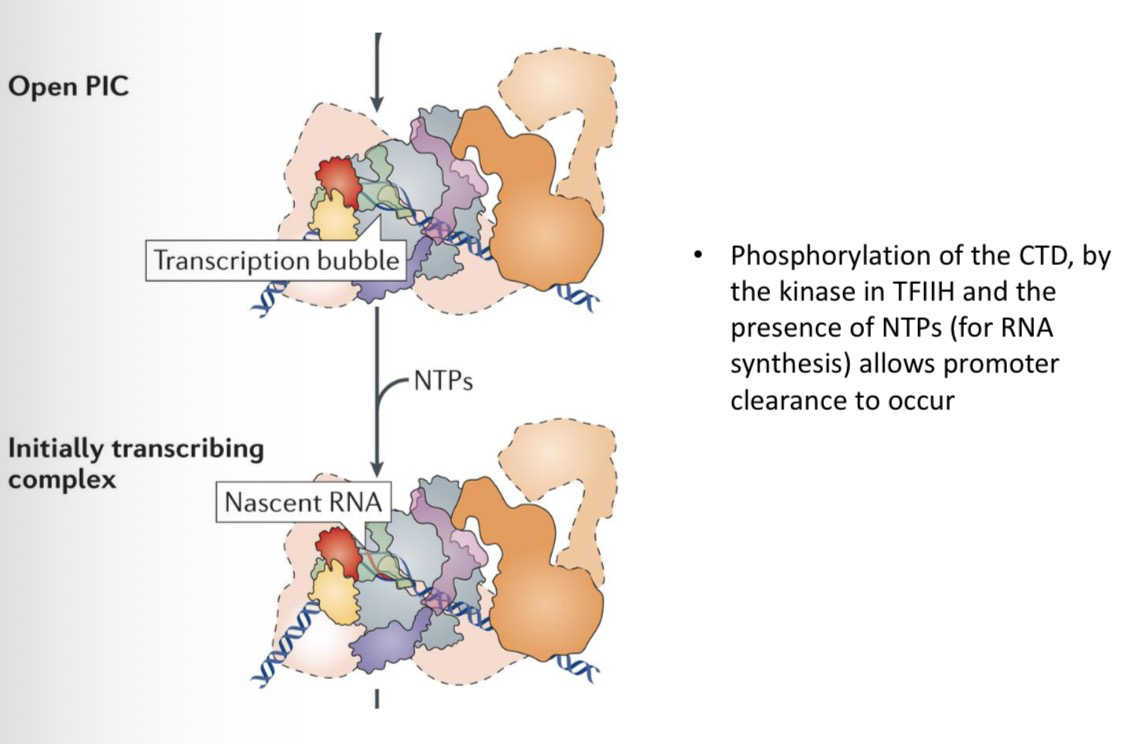
35) What is required to transform a Closed PIC into an Open PIC?
ATP must be hydrolyzed (used) by TFIIH to promote the formation of the open promoter complex (melting/opening the DNA strands).
36) What two events are required for promoter clearance(RNAP II leaving the promoter to start elongation)?
Phosphorylation of the CTD by the kinase module of TFIIH.
The presence of NTPs (nucleoside triphosphates) to begin actual RNA synthesis.
37) Summarize the PIC journey from TBP binding to Elongation.
Assembly: D→A→B→RNAP II/F→E→H (Closed Complex).
Opening: TFIIH uses ATP to open DNA (Open Complex).
Start: TFIIH phosphorylates CTD + NTPs are added→Promoter Clearance.